

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144726.11 + phase: 0
(515 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
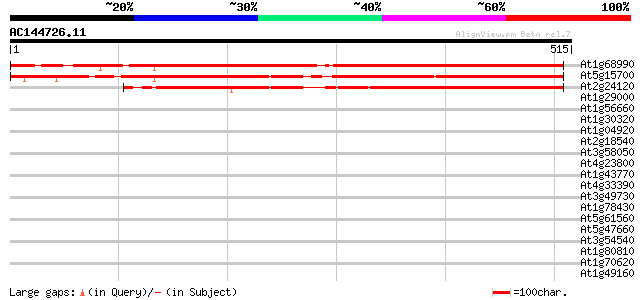
Score E
Sequences producing significant alignments: (bits) Value
At1g68990 DNA-directed RNA polymerase 476 e-134
At5g15700 DNA-directed RNA polymerase (mitochondrial) 432 e-121
At2g24120 chloroplast single subunit DNA-dependent RNA polymerase 362 e-100
At1g29000 hypothetical protein 37 0.034
At1g56660 hypothetical protein 36 0.058
At1g30320 unknown protein 35 0.099
At1g04920 sucrose-phosphate synthase, putative 35 0.099
At2g18540 putative vicilin storage protein (globulin-like) 34 0.17
At3g58050 putative protein 33 0.38
At4g23800 98b like protein 33 0.49
At1g43770 hypothetical protein 33 0.49
At4g33390 putative protein 32 0.84
At3g49730 putative protein 32 0.84
At1g78430 hypothetical protein 32 0.84
At5g61560 putative protein 32 1.1
At5g47660 unknown protein 32 1.1
At3g54540 ABC transporter - like protein 32 1.1
At1g80810 unknown protein 32 1.1
At1g70620 unknown protein 32 1.1
At1g49160 putative kinase 32 1.1
>At1g68990 DNA-directed RNA polymerase
Length = 976
Score = 476 bits (1225), Expect = e-134
Identities = 252/522 (48%), Positives = 347/522 (66%), Gaps = 42/522 (8%)
Query: 1 MWSKLAKRASSRRFILPSQSSTSLRFSHQHNFPEKILRHPFSRFSQVGIFTSEYKLGRSN 60
MW + RAS R+ S SS+S ++P +R S + G+ R+
Sbjct: 1 MWRNILGRASLRKVKFLSDSSSS-----GTHYPVNRVRGILSSVNLSGV--------RNG 47
Query: 61 FSFSEHPFSFSSISLNHFRTYG----SVAAEAIESTDTEDDCSGSEEVVQKLLDQMVIEE 116
S + S H + Y + AA+AI+STD ED+ SGS+EV ++++ E
Sbjct: 48 LSINPVNEMGGLSSFRHGQCYVFEGYATAAQAIDSTDPEDESSGSDEV-----NELITEM 102
Query: 117 EKKNPQLEEKKKNKN----------NSYKYKMLRRRQIKIETEAWEEAAREYQELLEDMR 166
EK+ ++ +K + + K+ ML++RQ+K+ETE WE AARE +E+L DM
Sbjct: 103 EKETERIRKKARLAAIPPKRVIAGMGAQKFYMLKQRQVKMETEEWERAARECREILADMC 162
Query: 167 EQKLAPNLPYMKSLFLGWFEPLKNAIAADQEICKDTKTRLSHAPYFNELPPDMMAVITMH 226
EQKLAPNLPYMKSLFLGWFEP++NAI D + K K ++ +AP+ +LP D MAVITMH
Sbjct: 163 EQKLAPNLPYMKSLFLGWFEPVRNAIQDDLDTFKIKKGKIPYAPFMEQLPADKMAVITMH 222
Query: 227 KLMGLLMTNSNGVGSARVIQAACQIGEAIEHEGRIYRFMEKTKVKKATSHKSDSDSVPAP 286
K+MGLLMTN+ GVG +++ AA QIGEA+E E RI F++K K AT ++++
Sbjct: 223 KMMGLLMTNAEGVGIVKLVNAATQIGEAVEQEVRINSFLQKKNKKNATDKTINTEA---- 278
Query: 287 VKGENLTAEENEKLAKEEKRLRNKVASLIKKQKKQQALGIVKGRDQAKPWGQEAQVKVGS 346
EN++ E +AKE ++ R +V L++K K +Q +V+ D KPWGQEAQVKVG+
Sbjct: 279 ---ENVS---EEIVAKETEKARKQVTVLMEKNKLRQVKALVRKHDSFKPWGQEAQVKVGA 332
Query: 347 RLIQLLIETAYLEPPANKFGDGGPDIFPAFKHTLKTISNDSNNGSRRYGVIECDPMIQKG 406
RLIQLL+E AY++PPA +F DG PDI PAFK +T++ ++ SRRYG IECDP++ KG
Sbjct: 333 RLIQLLMENAYIQPPAEQFDDGPPDIRPAFKQNFRTVTLENTKTSRRYGCIECDPLVLKG 392
Query: 407 LEKTARNMVIPYMPMLVPPNNWIGYDKGAYLFLPSYVMRIHGAKQQRDALKWAPKNQLDS 466
L+K+AR+MVIPY+PML+PP NW GYD+GA+ FLPSYVMR HGAKQQR +K PK QL+
Sbjct: 393 LDKSARHMVIPYLPMLIPPQNWTGYDQGAHFFLPSYVMRTHGAKQQRTVMKRTPKEQLEP 452
Query: 467 IFEALNTLGNTKWRVNKSVLGVIDQIWANGGRVADLVDRDDV 508
++EAL+TLGNTKW++NK VL ++D+IWANGGR+ LVDR+DV
Sbjct: 453 VYEALDTLGNTKWKINKKVLSLVDRIWANGGRIGGLVDREDV 494
>At5g15700 DNA-directed RNA polymerase (mitochondrial)
Length = 1011
Score = 432 bits (1112), Expect = e-121
Identities = 253/517 (48%), Positives = 327/517 (62%), Gaps = 36/517 (6%)
Query: 1 MWSKLAKRASSR---RFILPSQSSTSLRFSHQHNFPEKI-LRHPF--SRFSQVGIFTSEY 54
MW +AK+A SR R + SQ+ L S + F + + R P S G
Sbjct: 40 MWRNIAKQAISRSAARLNVSSQTRGLLVSSPESIFSKNLSFRFPVLGSPCHGKGFRCLSG 99
Query: 55 KLGRSNFSFSEHPFSFSSISLNHFRTYGSVAAEAIESTDTEDDCSGSEEVVQKLLDQMVI 114
R FS SE S + R Y SVA E + STD E+ E V +LL +M
Sbjct: 100 ITRREEFSKSERCLSGTLA-----RGYTSVAEEEVLSTDVEE-----EPEVDELLKEMKK 149
Query: 115 EEEKKNPQLEEKKKNKN---NSYKYKMLRRRQIKIETEAWEEAAREYQELLEDMREQKLA 171
E+++++ + KK K++ L RRQ+KIETE WE AA EY ELL DM EQKLA
Sbjct: 150 EKKRESHRSWRMKKQDQFGMGRTKFQNLWRRQVKIETEEWERAAAEYMELLTDMCEQKLA 209
Query: 172 PNLPYMKSLFLGWFEPLKNAIAADQEICKDTKTRLSHAPYFNELPPDMMAVITMHKLMGL 231
PNLPY+KSLFLGWFEPL++AIA DQE+ + K++ ++A Y ++LP D ++VITMHKLMG
Sbjct: 210 PNLPYVKSLFLGWFEPLRDAIAKDQELYRLGKSKATYAHYLDQLPADKISVITMHKLMGH 269
Query: 232 LMTNSNGVGSARVIQAACQIGEAIEHEGRIYRFMEKTKVKKATSHKSDSDSVPAPVKGEN 291
LMT + G +V+ AAC +G+AIE E RI F++K K K D + V
Sbjct: 270 LMTGGDN-GCVKVVHAACTVGDAIEQEIRICTFLDKKK-------KGDDNEESGGV---- 317
Query: 292 LTAEENEKLAKEEKRLRNKVASLIKKQKKQQALGIVKGRDQAKPWGQEAQVKVGSRLIQL 351
ENE KE+ +LR KV LIKKQK I++ D KPW + + KVGSRLI+L
Sbjct: 318 ----ENETSMKEQDKLRKKVNELIKKQKLSAVRKILQSHDYTKPWIADVRAKVGSRLIEL 373
Query: 352 LIETAYLEPPANKFGDGGPDIFPAFKHTLKTISNDSNNGSRRYGVIECDPMIQKGLEKTA 411
L+ TAY++ PA++ + PD+ PAF HT K N+G R+YGVIECDP+++KGLEK+
Sbjct: 374 LVRTAYIQSPADQQDNDLPDVRPAFVHTFKVAKGSMNSG-RKYGVIECDPLVRKGLEKSG 432
Query: 412 RNMVIPYMPMLVPPNNWIGYDKGAYLFLPSYVMRIHGAKQQRDALKWAPKNQLDSIFEAL 471
R V+PYMPMLVPP W GYDKGAYLFL SY+M+ HGAKQQR+ALK APK QL +FEAL
Sbjct: 433 RYAVMPYMPMLVPPLKWSGYDKGAYLFLTSYIMKTHGAKQQREALKSAPKGQLQPVFEAL 492
Query: 472 NTLGNTKWRVNKSVLGVIDQIWANGGRVADLVDRDDV 508
+TLG+TKWRVNK VL V+D+IW++GG VAD+VDR DV
Sbjct: 493 DTLGSTKWRVNKRVLTVVDRIWSSGGCVADMVDRSDV 529
>At2g24120 chloroplast single subunit DNA-dependent RNA polymerase
Length = 993
Score = 362 bits (929), Expect = e-100
Identities = 190/407 (46%), Positives = 274/407 (66%), Gaps = 37/407 (9%)
Query: 105 VQKLLDQMVIEEEKKNPQLEEKKKNKNNSYKYKMLRRRQIKIETEAWEEAAREYQELLED 164
+ K++DQ + ++E K +K K+ LRRRQ+K ETEAWE EY++L ++
Sbjct: 139 LSKMVDQTL--------KIERKDIDKR---KFDSLRRRQVKEETEAWERMVDEYRDLEKE 187
Query: 165 MREQKLAPNLPYMKSLFLGWFEPLKNAIAADQEICKDT--KTRLSHAPYFNELPPDMMAV 222
M E+ LAPNLPY+K +FLGWF+PLK+ I +Q++ K+ K R ++AP+ LP D MAV
Sbjct: 188 MCEKNLAPNLPYVKHMFLGWFQPLKDVIEREQKLQKNKSKKVRAAYAPHIELLPADKMAV 247
Query: 223 ITMHKLMGLLMTNSNGVGSARVIQAACQIGEAIEHEGRIYRFMEKTKVKKATSHKSDSDS 282
I MHK+MGL+M+ G +V+QAA IG AIE E RI+ F+++T+
Sbjct: 248 IVMHKMMGLVMSGHED-GCIQVVQAAVSIGIAIEQEVRIHNFLKRTR------------- 293
Query: 283 VPAPVKGENLTAEENEKLAKEEKRLRNKVASLIKKQKKQQALGIVKGRDQAKPWGQEAQV 342
+N + E+L KE++ LR +V SLI++++ AL +VK + KPWG+ Q
Sbjct: 294 -------KNNAGDSQEEL-KEKQLLRKRVNSLIRRKRIIDALKVVKS-EGTKPWGRATQA 344
Query: 343 KVGSRLIQLLIETAYLEPPANKFGDGGPDIFPAFKHTLKTISN-DSNNGSRRYGVIECDP 401
K+GSRL++LLIE AY++PP + GD P+ PAF+H KT++ + RRYGVIECD
Sbjct: 345 KLGSRLLELLIEAAYVQPPLTQSGDSIPEFRPAFRHRFKTVTKYPGSKLVRRYGVIECDS 404
Query: 402 MIQKGLEKTARNMVIPYMPMLVPPNNWIGYDKGAYLFLPSYVMRIHGAKQQRDALKWAPK 461
++ GL+K+A++M+IPY+PMLVPP W GYDKG YLFLPSY+MR HG+K+Q+DALK
Sbjct: 405 LLLAGLDKSAKHMLIPYVPMLVPPKRWKGYDKGGYLFLPSYIMRTHGSKKQQDALKDISH 464
Query: 462 NQLDSIFEALNTLGNTKWRVNKSVLGVIDQIWANGGRVADLVDRDDV 508
+FEAL+TLGNTKWRVN+++L V++++WA+GG +A LV+R+DV
Sbjct: 465 KTAHRVFEALDTLGNTKWRVNRNILDVVERLWADGGNIAGLVNREDV 511
>At1g29000 hypothetical protein
Length = 287
Score = 36.6 bits (83), Expect = 0.034
Identities = 24/89 (26%), Positives = 47/89 (51%), Gaps = 1/89 (1%)
Query: 89 IESTDTEDDCSGSEEVVQKLLDQMVIEEEKKNPQLEEKKKNKNNSYKYKMLRRRQIKIE- 147
I S+ TE++ EE +K ++ ++E + + EEKKK + N K ++ ++K+E
Sbjct: 177 IISSKTEEEKKKEEEDKKKKEEEDKKKKEDEKKKEEEKKKEEENKKKEGEKKKEEVKVEV 236
Query: 148 TEAWEEAAREYQELLEDMREQKLAPNLPY 176
T EY+E+++ ++ N+PY
Sbjct: 237 TTKTITQVVEYKEIVKVEGQKDKDGNIPY 265
>At1g56660 hypothetical protein
Length = 522
Score = 35.8 bits (81), Expect = 0.058
Identities = 23/81 (28%), Positives = 44/81 (53%), Gaps = 2/81 (2%)
Query: 90 ESTDTEDDCSGSEEVVQKLLDQMVIEEEKKNPQLEEKKKNKNNSYKYKMLRRRQIKIETE 149
+ TD E S++ +K D+ EE+KK P E+K+K+++ + K L+ ++ K E
Sbjct: 236 DETDQEMKEKDSKKNKKKEKDESCAEEKKKKPDKEKKEKDESTEKEDKKLKGKKGKGEKP 295
Query: 150 AWEEAAREYQELLEDMREQKL 170
E+ ++ +E D EQ++
Sbjct: 296 EKEDEGKKTKE--HDATEQEM 314
Score = 33.9 bits (76), Expect = 0.22
Identities = 21/72 (29%), Positives = 41/72 (56%), Gaps = 4/72 (5%)
Query: 106 QKLLDQMVIEEEKKNPQLEEKKKNKNNSYKYKMLRRRQIKIE----TEAWEEAAREYQEL 161
+K D+ EE+KK P+ E+K+K ++ S + K ++ ++ K E + EE +E+ E
Sbjct: 179 KKEKDESGTEEKKKKPKKEKKQKEESKSNEDKKVKGKKEKGEKGDLEKEDEEKKKEHDET 238
Query: 162 LEDMREQKLAPN 173
++M+E+ N
Sbjct: 239 DQEMKEKDSKKN 250
Score = 29.3 bits (64), Expect = 5.5
Identities = 17/67 (25%), Positives = 29/67 (42%)
Query: 256 EHEGRIYRFMEKTKVKKATSHKSDSDSVPAPVKGENLTAEENEKLAKEEKRLRNKVASLI 315
EH+ EK K K +S + K + E++E KE+K+L+ K
Sbjct: 234 EHDETDQEMKEKDSKKNKKKEKDESCAEEKKKKPDKEKKEKDESTEKEDKKLKGKKGKGE 293
Query: 316 KKQKKQQ 322
K +K+ +
Sbjct: 294 KPEKEDE 300
>At1g30320 unknown protein
Length = 509
Score = 35.0 bits (79), Expect = 0.099
Identities = 34/137 (24%), Positives = 60/137 (42%), Gaps = 17/137 (12%)
Query: 83 SVAAEAIESTDTEDDCSGSEEVVQKLLDQMVIEEEKKNPQLEEKKKNKNNSYKYKMLRRR 142
++AA A + + +G E QK IE EK+ EE +K+K+N+ +YK R
Sbjct: 368 NIAAWASKEEEENKKNNGDAEEAQK------IEFEKRATAWEEAEKSKHNA-RYK---RE 417
Query: 143 QIKIETEAWEEAAR-----EYQELLEDMREQKLAPNLPYMKSLFLGWFEPLKNAIAADQE 197
+I+I +AWE + E + + + + K MK + L + A+
Sbjct: 418 EIRI--QAWESQEKAKLEAEMRRIEAKVEQMKAEAEAKIMKKIALAKQRSEEKRALAEAR 475
Query: 198 ICKDTKTRLSHAPYFNE 214
+D + ++ A Y E
Sbjct: 476 KTRDAEKAVAEAQYIRE 492
>At1g04920 sucrose-phosphate synthase, putative
Length = 1062
Score = 35.0 bits (79), Expect = 0.099
Identities = 16/43 (37%), Positives = 28/43 (64%)
Query: 101 SEEVVQKLLDQMVIEEEKKNPQLEEKKKNKNNSYKYKMLRRRQ 143
S + V++++ +M E K P+L+ KK++ N KY +LRRR+
Sbjct: 730 SADPVKQIMSRMRTPEIKSKPELQGKKQSDNLGSKYPVLRRRE 772
>At2g18540 putative vicilin storage protein (globulin-like)
Length = 699
Score = 34.3 bits (77), Expect = 0.17
Identities = 22/85 (25%), Positives = 44/85 (50%), Gaps = 4/85 (4%)
Query: 87 EAIESTDTEDDCSGSEEVVQKLLDQMVIEEEKKNPQLEEKKKNKNNSYKYKMLRRRQIKI 146
E + + E EEV +K + E+E+K + E +K+ + + +M +RR+ +
Sbjct: 520 EMAKKREEERQRKEREEVERKRRE----EQERKRREEEARKREEERKREEEMAKRREQER 575
Query: 147 ETEAWEEAAREYQELLEDMREQKLA 171
+ + EE R+ +E E RE+++A
Sbjct: 576 QRKEREEVERKIREEQERKREEEMA 600
>At3g58050 putative protein
Length = 1209
Score = 33.1 bits (74), Expect = 0.38
Identities = 20/68 (29%), Positives = 37/68 (54%), Gaps = 4/68 (5%)
Query: 111 QMVIEEEKKNPQLEEKKKNKNNSYKYKMLRRRQIKIETEAWEE----AAREYQELLEDMR 166
+++ EEEK+ + EE+K+ K + + K LR+++ E + +E + LL R
Sbjct: 530 KLLEEEEKEKREEEERKEKKRSKEREKKLRKKERLKEKDKGKEKKNPECSDKDMLLNSSR 589
Query: 167 EQKLAPNL 174
E++ PNL
Sbjct: 590 EEEDLPNL 597
>At4g23800 98b like protein
Length = 456
Score = 32.7 bits (73), Expect = 0.49
Identities = 56/249 (22%), Positives = 103/249 (40%), Gaps = 27/249 (10%)
Query: 115 EEEKKNPQLEEKKKNKNNSYKYKMLR-------RRQIKIETEAW------EEAAREYQEL 161
E +K+NP+ + K+ + K+K L + ++E EA+ E+ +E +L
Sbjct: 155 EVKKENPEADFKETSNILGAKWKSLSAEDKKPYEERYQVEKEAYLQVIAKEKREKEAMKL 214
Query: 162 LEDMREQKLAPNLPYMKSLFLGWFE-PLKNAIAADQEICKDTKTRLSHAPYFNE----LP 216
LED ++Q+ A L F+ E K +++ K + Y NE L
Sbjct: 215 LEDDQKQRTAMELLDQYLNFVQEAEQDNKKKNKKEKDPLKPKHPVSAFLVYANERRAALR 274
Query: 217 PDMMAVITMHKLMGLLMTNSNGVGSARVIQAACQIGEAIEHEGRIYRFMEKTKVKKATSH 276
+ +V+ + K+ G N + A + A + E Y+ +TK ++A S
Sbjct: 275 EENKSVVEVAKITGEEWKNLSDKKKAPYEKVAKKNKETYLQAMEEYK---RTKEEEALSQ 331
Query: 277 KSDSDSVPAPVKGENL----TAEENEKLAKEEKRLRNKVASLI--KKQKKQQALGIVKGR 330
K + + + K E L E+ + L K+EK + K + K KK + + +
Sbjct: 332 KKEEEELLKLHKQEALQMLKKKEKTDNLIKKEKATKKKKNENVDPNKPKKPASSYFLFSK 391
Query: 331 DQAKPWGQE 339
D+ K +E
Sbjct: 392 DERKKLTEE 400
>At1g43770 hypothetical protein
Length = 431
Score = 32.7 bits (73), Expect = 0.49
Identities = 38/189 (20%), Positives = 75/189 (39%), Gaps = 16/189 (8%)
Query: 58 RSNFSFSEHPFSFSSISLNHFRTYGSVAAEAIESTDTEDDCSGSEEVVQKLLDQM----- 112
R S S P + + N G EA +S+ D S + + +K +DQ
Sbjct: 101 REESSCSRKPHELTCLDGN-----GESVLEAADSSSVPDHSSCTSK--RKEVDQTANLGH 153
Query: 113 VIEEEKKNPQLEEKKKNKNNSYKYKMLRRRQIKIETEAWE----EAAREYQELLEDMREQ 168
++E +K + ++KKK+ N+S + +++ T E ++ +E +E R++
Sbjct: 154 ILERSEKKKKKKKKKKSINHSPPVLAVEDHELQGTTNVVEPVEVSSSSPTKETMESKRQE 213
Query: 169 KLAPNLPYMKSLFLGWFEPLKNAIAADQEICKDTKTRLSHAPYFNELPPDMMAVITMHKL 228
P+ + +G E ++ + C K +LS +L +
Sbjct: 214 SSDSRKPHELTCLVGDSETANSSSVPEHNSCAVKKRKLSSGNKQIQLADGSSSCRVAESN 273
Query: 229 MGLLMTNSN 237
M L +T+ N
Sbjct: 274 MPLTLTSRN 282
>At4g33390 putative protein
Length = 779
Score = 32.0 bits (71), Expect = 0.84
Identities = 53/283 (18%), Positives = 111/283 (38%), Gaps = 21/283 (7%)
Query: 74 SLNHFRTYGSVAAEAIESTDTEDDCSGSEEVVQKLLDQMVIEEEKKNP-QLEEKKKNKNN 132
+L+ + +A+ + S + E D + E + K ++ EE + P QL++ + +
Sbjct: 495 ALDSLKQREGMASVTVASLEAEIDITRCEIALVKSKEKETREEMVELPKQLQQASQEADE 554
Query: 133 SYKYKMLRRRQIKIETEAWEEAAREYQELLEDM-REQKLAPNLPYMKSLFLGWFEPLKNA 191
+ + L R +++ E E+A + + QK + + L L + L+ +
Sbjct: 555 AKSFAELAREELRKSQEEAEQAKAGASTMESRLFAAQKEIEAIKASERLALAAIKALQES 614
Query: 192 IAADQEICKDTKTRLSHAPYFNELPPDMMAVITMHKLMGLLMTNSNGVGSARVIQAACQI 251
++ +E D+ P + I + + + +ARV A ++
Sbjct: 615 ESSSKENAVDS-------------PRTVTLTIEEYYELSKRAHEAEEAANARVAAAVSEV 661
Query: 252 GEAIEHEGRIYRFME---KTKVKKATSHKSDSDSVPAPVKGENLTAEENEKLAKEEKRLR 308
GEA E E R +E K V++ + + +G+ +E K + ++ R
Sbjct: 662 GEAKETEKRSLEKLEEVNKEMVERKATLAGAMEKAEKAKEGKLGVEQELRKWREVSEKKR 721
Query: 309 NKVAS---LIKKQKKQQALGIVKGRDQAKPWGQEAQVKVGSRL 348
+S I+ K+++A V + P Q VK +L
Sbjct: 722 KNGSSHGKSIQGSKEKEAETSVSNETETNPIPQVNPVKKKKKL 764
>At3g49730 putative protein
Length = 1184
Score = 32.0 bits (71), Expect = 0.84
Identities = 15/31 (48%), Positives = 19/31 (60%), Gaps = 1/31 (3%)
Query: 154 AAREYQELLEDMREQKLAPNLPYMKSLFLGW 184
+ +E ++ EDMRE K PNL Y SL GW
Sbjct: 217 SVKEASKVFEDMRE-KFPPNLRYFTSLLYGW 246
>At1g78430 hypothetical protein
Length = 326
Score = 32.0 bits (71), Expect = 0.84
Identities = 18/73 (24%), Positives = 40/73 (54%), Gaps = 4/73 (5%)
Query: 87 EAIESTDTEDDCS-GSEEVVQKLLDQMVIEEEKKNP---QLEEKKKNKNNSYKYKMLRRR 142
+ ++ TDTE C+ E+ + + Q+ E E+ N +L++K ++ + + +
Sbjct: 187 DQLKKTDTEMSCAKAKEDEIASKVSQIGEELEESNETTAKLKKKLESVEEAKETLEAEMK 246
Query: 143 QIKIETEAWEEAA 155
++K++TE W +AA
Sbjct: 247 KLKVQTEQWRKAA 259
>At5g61560 putative protein
Length = 815
Score = 31.6 bits (70), Expect = 1.1
Identities = 28/113 (24%), Positives = 50/113 (43%), Gaps = 7/113 (6%)
Query: 73 ISLNHFRTYGSVA-AEAIESTDTEDDCSGSEEVVQKLLDQMVIEEEKKNPQLEEKKKNKN 131
I L H + +VA +E I+++ D + L + I EE+ + +E +++ +
Sbjct: 323 IELRHIKGMYAVAQSEVIDASKKMQDLNQRRSEEATRLKNLTIREEEADEVVEMERERQE 382
Query: 132 NSYKYKMLRRRQI------KIETEAWEEAAREYQELLEDMREQKLAPNLPYMK 178
++ L R I ++E EA E R+ ++ LED E YMK
Sbjct: 383 DAENEAELVRECIERETEERLEAEARAEEVRKEKQRLEDALEGGPLQRQQYMK 435
Score = 29.6 bits (65), Expect = 4.2
Identities = 31/164 (18%), Positives = 73/164 (43%), Gaps = 4/164 (2%)
Query: 3 SKLAKRASSRRFILPSQSSTSLRFSHQHNFPEKILRHPFSRFS--QVGIFTSEYKLGRSN 60
S++ + +SS + P+ SS+ + + + LRH ++ Q + + K+ N
Sbjct: 291 SQMEEASSSSTYSDPTSSSSQIHKDFELEKLKIELRHIKGMYAVAQSEVIDASKKMQDLN 350
Query: 61 FSFSEHPFSFSSISLNHFRTYGSVAAEAIESTDTEDDCSGSEEVVQKLLDQMVIEEEKKN 120
SE ++++ V E D E++ E +++ ++ +E E +
Sbjct: 351 QRRSEEATRLKNLTIREEEADEVVEMERERQEDAENEAELVRECIERETEER-LEAEARA 409
Query: 121 PQLEEKKKNKNNSYKYKMLRRRQ-IKIETEAWEEAAREYQELLE 163
++ ++K+ ++ + L+R+Q +K E E EA + + L+
Sbjct: 410 EEVRKEKQRLEDALEGGPLQRQQYMKFEWEEIVEATSSFSDELK 453
>At5g47660 unknown protein
Length = 398
Score = 31.6 bits (70), Expect = 1.1
Identities = 45/201 (22%), Positives = 74/201 (36%), Gaps = 41/201 (20%)
Query: 5 LAKRASSRRFILPSQSSTSLRFSHQHNFPEKILRHPFSRFSQVGIFTSEYKLGRS----- 59
LA AS + + P + L S + P IL R ++G E LG
Sbjct: 51 LADLASPPQKLKPIRCGVKLPSSSEDRHPLDILAGTLDRLPEMGFGCFEAPLGSKIADVE 110
Query: 60 -------NFSFSEH----PFSFSSISLNHFRTYGSVAAEAIESTDTEDD------CSGS- 101
FS E P + N G + +++S+D++ +G
Sbjct: 111 ESGQLTRGFSKEEDDSLPPLQMEFQARNRISWDGLSLSSSVDSSDSDSSPDVRKTVTGKR 170
Query: 102 --------EEVVQKLLDQMVIEEEKKNPQLEEKKKNKNNSYKYKMLRRRQIKIETEAWEE 153
E ++KL+ M+ +EK + QL N + + +RR EAW +
Sbjct: 171 KRETRVKLEHFLEKLVGSMMKRQEKMHNQLI----NVMEKMEVERIRRE------EAWRQ 220
Query: 154 AAREYQELLEDMREQKLAPNL 174
E E+ R+Q++A NL
Sbjct: 221 QETERMTQNEEARKQEMARNL 241
>At3g54540 ABC transporter - like protein
Length = 723
Score = 31.6 bits (70), Expect = 1.1
Identities = 48/258 (18%), Positives = 99/258 (37%), Gaps = 45/258 (17%)
Query: 92 TDTEDDCSGSEEVVQKLLDQMVIEEEKKNPQLEEKKKNKNNSYKYKML-------RRRQI 144
+D EDD EE QK + + E++ K+ K K ++ +R +
Sbjct: 73 SDEEDDGESDEEERQKEARRKLKSEQRHLEISVTDKEQKKREAKERLALQAAESAKREAM 132
Query: 145 KIETEAWEEAAREYQELLE--DMREQKLAPNLPYMKSLFLGWFEPLKNAIAADQEICKDT 202
K + +A+ +LE DM + + K + + F + A +E+ K+
Sbjct: 133 KDDHDAFTVVIGSKTSVLEGDDMADANV-------KDITIESF----SVSARGKELLKNA 181
Query: 203 KTRLSHAPYFNELPPDMMAVITMHKLMG-----------LLMTNSNGVGSARVIQAACQI 251
R+SH + + P+ M T+ KL+ +L+ VG + +
Sbjct: 182 SVRISHGKRYGLIGPNGMGKSTLLKLLAWRKIPVPKNIDVLLVEQEVVGDEK-----SAL 236
Query: 252 GEAIEHEGRIYRFMEKTKVKKATSHKSDSDSVPAPVKGENLTAEENEKLAKEEKRLRNKV 311
+ + + E+ + + +S +D GEN+ E+++ ++ L +++
Sbjct: 237 NAVVSANEELVKLREEAEALQKSSSGAD---------GENVDGEDDDDTGEKLAELYDRL 287
Query: 312 ASLIKKQKKQQALGIVKG 329
L + QA I+ G
Sbjct: 288 QILGSDAAEAQASKILAG 305
>At1g80810 unknown protein
Length = 826
Score = 31.6 bits (70), Expect = 1.1
Identities = 22/80 (27%), Positives = 41/80 (50%), Gaps = 6/80 (7%)
Query: 90 ESTDTEDDCSGSEEVVQ-KLLDQMVIEEEKKNPQLEEKKKNKNNSYKYKMLRRRQIKIET 148
E + EDDCS +E Q K ++ EEEK+ P E + + +++ + + R ET
Sbjct: 716 EEQEYEDDCSDKKEQSQDKGVEAETKEEEKQYPNSEGESEGEDSESEEEPKWR-----ET 770
Query: 149 EAWEEAAREYQELLEDMREQ 168
+ E+ E +E ++ M ++
Sbjct: 771 DDMEDDEEEEEEEIDHMEDE 790
>At1g70620 unknown protein
Length = 897
Score = 31.6 bits (70), Expect = 1.1
Identities = 16/55 (29%), Positives = 25/55 (45%)
Query: 121 PQLEEKKKNKNNSYKYKMLRRRQIKIETEAWEEAAREYQELLEDMREQKLAPNLP 175
PQ ++ N N Y+ + Q E W A+E+ +D + Q+ APN P
Sbjct: 118 PQANQEWGNPNWGYQQQQGHTPQANSNVEDWAVKAKEWAAANKDQQSQQSAPNQP 172
>At1g49160 putative kinase
Length = 539
Score = 31.6 bits (70), Expect = 1.1
Identities = 26/109 (23%), Positives = 46/109 (41%), Gaps = 4/109 (3%)
Query: 66 HPFSFSSISLNHFRTYGSVAAEAIESTDTEDDCSGSEEVVQKLLDQMVIEEEKKNP---- 121
HP S NH S A E+I + + D C S++ + ++E + P
Sbjct: 430 HPQQNQSSKDNHQNGASSQAGESISHSLSSDYCPRSDDEANPTVAATTEDQEAEKPGSLE 489
Query: 122 QLEEKKKNKNNSYKYKMLRRRQIKIETEAWEEAAREYQELLEDMREQKL 170
+ EE ++ K K + R ++K T EEA E + + + Q++
Sbjct: 490 EEEEDERLKEELEKIEERFREEMKEITRKREEATMETKNRFFEKKMQQV 538
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.132 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,804,926
Number of Sequences: 26719
Number of extensions: 525496
Number of successful extensions: 2903
Number of sequences better than 10.0: 82
Number of HSP's better than 10.0 without gapping: 20
Number of HSP's successfully gapped in prelim test: 62
Number of HSP's that attempted gapping in prelim test: 2645
Number of HSP's gapped (non-prelim): 221
length of query: 515
length of database: 11,318,596
effective HSP length: 104
effective length of query: 411
effective length of database: 8,539,820
effective search space: 3509866020
effective search space used: 3509866020
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC144726.11


