

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144722.2 + phase: 0
(624 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
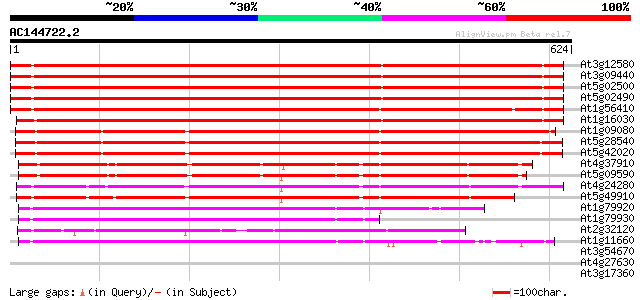
Score E
Sequences producing significant alignments: (bits) Value
At3g12580 putative protein 735 0.0
At3g09440 heat-shock protein (At-hsc70-3) 728 0.0
At5g02500 dnaK-type molecular chaperone hsc70.1 720 0.0
At5g02490 dnaK-type molecular chaperone hsc70.1 - like 716 0.0
At1g56410 hypothetical protein 714 0.0
At1g16030 unknown protein 706 0.0
At1g09080 putative luminal binding protein 604 e-173
At5g28540 luminal binding protein 593 e-170
At5g42020 luminal binding protein (dbj|BAA13948.1) 590 e-169
At4g37910 heat shock protein 70 like protein 439 e-123
At5g09590 heat shock protein 70 (Hsc70-5) 435 e-122
At4g24280 hsp 70-like protein 413 e-115
At5g49910 heat shock protein 70 (Hsc70-7) 409 e-114
At1g79920 putative heat-shock protein (At1g79920) 251 9e-67
At1g79930 putative heat-shock protein 251 9e-67
At2g32120 70kD heat shock protein 241 9e-64
At1g11660 heat-shock protein like 238 6e-63
At3g54670 structural maintenance of chromosomes (SMC) - like pro... 33 0.36
At4g27630 unknown protein 31 2.4
At3g17360 kinesin-like protein 30 3.1
>At3g12580 putative protein
Length = 650
Score = 735 bits (1897), Expect = 0.0
Identities = 388/619 (62%), Positives = 473/619 (75%), Gaps = 7/619 (1%)
Query: 1 MAKEYEGRAVGIDLGTTYSCVAVWLDQHQRVEIIHNDQGNRITPSFVAFNNEQRLIGDAA 60
MA + EG A+GIDLGTTYSCV VW QH RVEII NDQGNR TPS+VAF + +RLIGDAA
Sbjct: 1 MAGKGEGPAIGIDLGTTYSCVGVW--QHDRVEIIANDQGNRTTPSYVAFTDSERLIGDAA 58
Query: 61 KNQSATNPENTIFDAKRLIGRKFSDPVVQRDILLWPFKVISGVNDKPMITVKYKGQEKHF 120
KNQ A NP NT+FDAKRLIGR++SDP VQ D WPFKV+SG +KPMI V +KG+EK F
Sbjct: 59 KNQVAMNPTNTVFDAKRLIGRRYSDPSVQADKSHWPFKVVSGPGEKPMIVVNHKGEEKQF 118
Query: 121 CAEEISSMVLTKMREVAEAYLGSPVKNAVVTVPAYFNDSQRKATIDAGTIAGLNVIRIIN 180
AEEISSMVL KMRE+AEA+LGSPVKNAVVTVPAYFNDSQR+AT DAG I+GLNV+RIIN
Sbjct: 119 SAEEISSMVLIKMREIAEAFLGSPVKNAVVTVPAYFNDSQRQATKDAGVISGLNVMRIIN 178
Query: 181 EPTAAAIAYGLDKRNDCDGSRNIFVFDLGGGTFDVSILTIKGDVFEVKATAGNTHLGGED 240
EPTAAAIAYGLDK+ G +N+ +FDLGGGTFDVS+LTI+ +FEVKATAG+THLGGED
Sbjct: 179 EPTAAAIAYGLDKKASSVGEKNVLIFDLGGGTFDVSLLTIEEGIFEVKATAGDTHLGGED 238
Query: 241 FDNRMVNYFVEELKKKNKVDISQNPKSLRRLRTACERAKRILSFAFVTTVEVDALFTGID 300
FDNRMVN+FV+E K+KNK DI+ NP++LRRLRTACERAKR LS TT+E+D+LF GID
Sbjct: 239 FDNRMVNHFVQEFKRKNKKDITGNPRALRRLRTACERAKRTLSSTAQTTIEIDSLFEGID 298
Query: 301 FCSSITRAKFEELNMDFFNECMNIVDTCLRDSKIYKNDIDDVVLVGGSSRIPKVQEQLLE 360
F ++ITRA+FEELNMD F +CM V+ CLRD+K+ K+ + DVVLVGGS+RIPKVQ+ L +
Sbjct: 299 FYTTITRARFEELNMDLFRKCMEPVEKCLRDAKMDKSSVHDVVLVGGSTRIPKVQQLLQD 358
Query: 361 YFKGKELFMGINPDEAVAYGAAVQAAILS-EGCKNVPNLVLRDVTPLSLGISVSIEHIMS 419
+F GKEL INPDEAVAYGAAVQAAILS EG + V +L+L DVTPLSLG+ + +M+
Sbjct: 359 FFNGKELCKSINPDEAVAYGAAVQAAILSGEGNEKVQDLLLLDVTPLSLGLETA-GGVMT 417
Query: 420 VVIPRNTSIPVKKTSRYSTLYDNQCLVNFLVYEGERPRAADNNLLGSFGLRCL-PGPRGQ 478
V+IPRNT+IP KK +ST DNQ V VYEGER R DNNLLG F L + P PRG
Sbjct: 418 VLIPRNTTIPTKKEQIFSTYSDNQPGVLIQVYEGERARTKDNNLLGKFELSGIPPAPRGV 477
Query: 479 P-LEVCFSIDENGILTVTAREISTGNMNMITITNDKERLSMFDIEKMIKEAEKYHVEDMQ 537
P + VCF ID NGIL V+A + +TG N ITITNDK RLS +IEKM++EAEKY ED +
Sbjct: 478 PQITVCFDIDANGILNVSAEDKTTGQKNKITITNDKGRLSKEEIEKMVQEAEKYKAEDEE 537
Query: 538 FLRRAKVMCELDSCVYNMKKALKKKDVNLILSPQEIAKINNAITVAMDLLDKNKKEKEID 597
++ L++ YNM+ +K + + L + KI +AI A++ LD N+ E D
Sbjct: 538 HKKKVDAKNALENYAYNMRNTIKDEKIASKLDAADKKKIEDAIDQAIEWLDGNQL-AEAD 596
Query: 598 VLEGYLEELERMSKHLISK 616
E ++ELE + +I++
Sbjct: 597 EFEDKMKELESLCNPIIAR 615
>At3g09440 heat-shock protein (At-hsc70-3)
Length = 649
Score = 728 bits (1879), Expect = 0.0
Identities = 385/619 (62%), Positives = 472/619 (76%), Gaps = 7/619 (1%)
Query: 1 MAKEYEGRAVGIDLGTTYSCVAVWLDQHQRVEIIHNDQGNRITPSFVAFNNEQRLIGDAA 60
MA + EG A+GIDLGTTYSCV VW QH RVEII NDQGNR TPS+VAF + +RLIGDAA
Sbjct: 1 MAGKGEGPAIGIDLGTTYSCVGVW--QHDRVEIIANDQGNRTTPSYVAFTDSERLIGDAA 58
Query: 61 KNQSATNPENTIFDAKRLIGRKFSDPVVQRDILLWPFKVISGVNDKPMITVKYKGQEKHF 120
KNQ A NP NT+FDAKRLIGR+F+D VQ DI LWPF + SG +KPMI V YKG++K F
Sbjct: 59 KNQVAMNPINTVFDAKRLIGRRFTDSSVQSDIKLWPFTLKSGPAEKPMIVVNYKGEDKEF 118
Query: 121 CAEEISSMVLTKMREVAEAYLGSPVKNAVVTVPAYFNDSQRKATIDAGTIAGLNVIRIIN 180
AEEISSM+L KMRE+AEAYLG+ +KNAVVTVPAYFNDSQR+AT DAG IAGLNV+RIIN
Sbjct: 119 SAEEISSMILIKMREIAEAYLGTTIKNAVVTVPAYFNDSQRQATKDAGVIAGLNVMRIIN 178
Query: 181 EPTAAAIAYGLDKRNDCDGSRNIFVFDLGGGTFDVSILTIKGDVFEVKATAGNTHLGGED 240
EPTAAAIAYGLDK+ G +N+ +FDLGGGTFDVS+LTI+ +FEVKATAG+THLGGED
Sbjct: 179 EPTAAAIAYGLDKKATSVGEKNVLIFDLGGGTFDVSLLTIEEGIFEVKATAGDTHLGGED 238
Query: 241 FDNRMVNYFVEELKKKNKVDISQNPKSLRRLRTACERAKRILSFAFVTTVEVDALFTGID 300
FDNRMVN+FV+E K+KNK DIS NP++LRRLRTACERAKR LS TT+E+D+LF GID
Sbjct: 239 FDNRMVNHFVQEFKRKNKKDISGNPRALRRLRTACERAKRTLSSTAQTTIEIDSLFDGID 298
Query: 301 FCSSITRAKFEELNMDFFNECMNIVDTCLRDSKIYKNDIDDVVLVGGSSRIPKVQEQLLE 360
F + ITRA+FEELN+D F +CM V+ CLRD+K+ KN IDDVVLVGGS+RIPKVQ+ L++
Sbjct: 299 FYAPITRARFEELNIDLFRKCMEPVEKCLRDAKMDKNSIDDVVLVGGSTRIPKVQQLLVD 358
Query: 361 YFKGKELFMGINPDEAVAYGAAVQAAILS-EGCKNVPNLVLRDVTPLSLGISVSIEHIMS 419
+F GKEL INPDEAVAYGAAVQAAILS EG + V +L+L DVTPLSLG+ + +M+
Sbjct: 359 FFNGKELCKSINPDEAVAYGAAVQAAILSGEGNEKVQDLLLLDVTPLSLGLETA-GGVMT 417
Query: 420 VVIPRNTSIPVKKTSRYSTLYDNQCLVNFLVYEGERPRAADNNLLGSFGLRCL-PGPRGQ 478
V+I RNT+IP KK +ST DNQ V VYEGER R DNNLLG F L + P PRG
Sbjct: 418 VLIQRNTTIPTKKEQVFSTYSDNQPGVLIQVYEGERARTKDNNLLGKFELSGIPPAPRGV 477
Query: 479 P-LEVCFSIDENGILTVTAREISTGNMNMITITNDKERLSMFDIEKMIKEAEKYHVEDMQ 537
P + VCF ID NGIL V+A + +TG N ITITNDK RLS +IEKM++EAEKY ED +
Sbjct: 478 PQITVCFDIDANGILNVSAEDKTTGQKNKITITNDKGRLSKDEIEKMVQEAEKYKSEDEE 537
Query: 538 FLRRAKVMCELDSCVYNMKKALKKKDVNLILSPQEIAKINNAITVAMDLLDKNKKEKEID 597
++ L++ YNM+ ++ + + L+ + KI ++I A++ L+ N+ E D
Sbjct: 538 HKKKVDAKNALENYAYNMRNTIRDEKIGEKLAGDDKKKIEDSIEAAIEWLEANQL-AECD 596
Query: 598 VLEGYLEELERMSKHLISK 616
E ++ELE + +I+K
Sbjct: 597 EFEDKMKELESICNPIIAK 615
>At5g02500 dnaK-type molecular chaperone hsc70.1
Length = 651
Score = 720 bits (1859), Expect = 0.0
Identities = 381/619 (61%), Positives = 470/619 (75%), Gaps = 7/619 (1%)
Query: 1 MAKEYEGRAVGIDLGTTYSCVAVWLDQHQRVEIIHNDQGNRITPSFVAFNNEQRLIGDAA 60
M+ + EG A+GIDLGTTYSCV VW QH RVEII NDQGNR TPS+VAF + +RLIGDAA
Sbjct: 1 MSGKGEGPAIGIDLGTTYSCVGVW--QHDRVEIIANDQGNRTTPSYVAFTDSERLIGDAA 58
Query: 61 KNQSATNPENTIFDAKRLIGRKFSDPVVQRDILLWPFKVISGVNDKPMITVKYKGQEKHF 120
KNQ A NP NT+FDAKRLIGR+FSD VQ D+ LWPFK+ +G DKPMI V+YKG+EK F
Sbjct: 59 KNQVAMNPVNTVFDAKRLIGRRFSDSSVQSDMKLWPFKIQAGPADKPMIYVEYKGEEKEF 118
Query: 121 CAEEISSMVLTKMREVAEAYLGSPVKNAVVTVPAYFNDSQRKATIDAGTIAGLNVIRIIN 180
AEEISSMVL KMRE+AEAYLG +KNAVVTVPAYFNDSQR+AT DAG IAGLNV+RIIN
Sbjct: 119 AAEEISSMVLIKMREIAEAYLGVTIKNAVVTVPAYFNDSQRQATKDAGVIAGLNVMRIIN 178
Query: 181 EPTAAAIAYGLDKRNDCDGSRNIFVFDLGGGTFDVSILTIKGDVFEVKATAGNTHLGGED 240
EPTAAAIAYGLDK+ G +N+ +FDLGGGTFDVS+LTI+ +FEVKATAG+THLGGED
Sbjct: 179 EPTAAAIAYGLDKKATSVGEKNVLIFDLGGGTFDVSLLTIEEGIFEVKATAGDTHLGGED 238
Query: 241 FDNRMVNYFVEELKKKNKVDISQNPKSLRRLRTACERAKRILSFAFVTTVEVDALFTGID 300
FDNRMVN+FV+E K+K+K DI+ NP++LRRLRT+CERAKR LS TT+E+D+L+ GID
Sbjct: 239 FDNRMVNHFVQEFKRKSKKDITGNPRALRRLRTSCERAKRTLSSTAQTTIEIDSLYEGID 298
Query: 301 FCSSITRAKFEELNMDFFNECMNIVDTCLRDSKIYKNDIDDVVLVGGSSRIPKVQEQLLE 360
F S+ITRA+FEELNMD F +CM V+ CLRD+K+ K+ + DVVLVGGS+RIPKVQ+ L +
Sbjct: 299 FYSTITRARFEELNMDLFRKCMEPVEKCLRDAKMDKSTVHDVVLVGGSTRIPKVQQLLQD 358
Query: 361 YFKGKELFMGINPDEAVAYGAAVQAAILS-EGCKNVPNLVLRDVTPLSLGISVSIEHIMS 419
+F GKEL INPDEAVAYGAAVQ AILS EG + V +L+L DVTPLSLG+ + +M+
Sbjct: 359 FFNGKELCKSINPDEAVAYGAAVQGAILSGEGNEKVQDLLLLDVTPLSLGLETA-GGVMT 417
Query: 420 VVIPRNTSIPVKKTSRYSTLYDNQCLVNFLVYEGERPRAADNNLLGSFGLRCL-PGPRGQ 478
+IPRNT+IP KK +ST DNQ V VYEGER R DNNLLG F L + P PRG
Sbjct: 418 TLIPRNTTIPTKKEQVFSTYSDNQPGVLIQVYEGERARTKDNNLLGKFELSGIPPAPRGV 477
Query: 479 P-LEVCFSIDENGILTVTAREISTGNMNMITITNDKERLSMFDIEKMIKEAEKYHVEDMQ 537
P + VCF ID NGIL V+A + +TG N ITITNDK RLS +IEKM++EAEKY ED +
Sbjct: 478 PQITVCFDIDANGILNVSAEDKTTGQKNKITITNDKGRLSKDEIEKMVQEAEKYKSEDEE 537
Query: 538 FLRRAKVMCELDSCVYNMKKALKKKDVNLILSPQEIAKINNAITVAMDLLDKNKKEKEID 597
++ + L++ YNM+ ++ + + L + KI ++I A+ L+ N+ E D
Sbjct: 538 HKKKVEAKNALENYAYNMRNTIQDEKIGEKLPAADKKKIEDSIEQAIQWLEGNQL-AEAD 596
Query: 598 VLEGYLEELERMSKHLISK 616
E ++ELE + +I+K
Sbjct: 597 EFEDKMKELESICNPIIAK 615
>At5g02490 dnaK-type molecular chaperone hsc70.1 - like
Length = 653
Score = 716 bits (1849), Expect = 0.0
Identities = 379/619 (61%), Positives = 467/619 (75%), Gaps = 7/619 (1%)
Query: 1 MAKEYEGRAVGIDLGTTYSCVAVWLDQHQRVEIIHNDQGNRITPSFVAFNNEQRLIGDAA 60
MA + EG A+GIDLGTTYSCV VW QH RVEII NDQGNR TPS+VAF + +RLIGDAA
Sbjct: 1 MAGKGEGPAIGIDLGTTYSCVGVW--QHDRVEIIANDQGNRTTPSYVAFTDSERLIGDAA 58
Query: 61 KNQSATNPENTIFDAKRLIGRKFSDPVVQRDILLWPFKVISGVNDKPMITVKYKGQEKHF 120
KNQ A NP NT+FDAKRLIGR+FSD VQ D LWPF +ISG +KPMI V+YKG+EK F
Sbjct: 59 KNQVAMNPVNTVFDAKRLIGRRFSDASVQSDRQLWPFTIISGTAEKPMIVVEYKGEEKQF 118
Query: 121 CAEEISSMVLTKMREVAEAYLGSPVKNAVVTVPAYFNDSQRKATIDAGTIAGLNVIRIIN 180
AEEISSMVL KMRE+AEA+LG+ VKNAVVTVPAYFNDSQR+AT DAG IAGLNV+RIIN
Sbjct: 119 AAEEISSMVLIKMREIAEAFLGTTVKNAVVTVPAYFNDSQRQATKDAGVIAGLNVLRIIN 178
Query: 181 EPTAAAIAYGLDKRNDCDGSRNIFVFDLGGGTFDVSILTIKGDVFEVKATAGNTHLGGED 240
EPTAAAIAYGLDK+ G +N+ +FDLGGGTFDVS+LTI+ +FEVKATAG+THLGGED
Sbjct: 179 EPTAAAIAYGLDKKATSVGEKNVLIFDLGGGTFDVSLLTIEEGIFEVKATAGDTHLGGED 238
Query: 241 FDNRMVNYFVEELKKKNKVDISQNPKSLRRLRTACERAKRILSFAFVTTVEVDALFTGID 300
FDNRMVN+FV+E K+KNK DI+ P++LRRLRTACERAKR LS TT+E+D+L+ G D
Sbjct: 239 FDNRMVNHFVQEFKRKNKQDITGQPRALRRLRTACERAKRTLSSTAQTTIEIDSLYGGAD 298
Query: 301 FCSSITRAKFEELNMDFFNECMNIVDTCLRDSKIYKNDIDDVVLVGGSSRIPKVQEQLLE 360
F S ITRA+FEE+NMD F +CM V+ CLRD+K+ K+ + ++VLVGGS+RIPKVQ+ L +
Sbjct: 299 FYSPITRARFEEMNMDLFRKCMEPVEKCLRDAKMDKSTVHEIVLVGGSTRIPKVQQLLQD 358
Query: 361 YFKGKELFMGINPDEAVAYGAAVQAAILS-EGCKNVPNLVLRDVTPLSLGISVSIEHIMS 419
+F GKEL INPDEAVAYGAAVQAAILS EG + V +L+L DVTPLSLG+ + +M+
Sbjct: 359 FFNGKELCKSINPDEAVAYGAAVQAAILSGEGNEKVQDLLLLDVTPLSLGLETA-GGVMT 417
Query: 420 VVIPRNTSIPVKKTSRYSTLYDNQCLVNFLVYEGERPRAADNNLLGSFGLRCL-PGPRGQ 478
+I RNT+IP KK +ST DNQ V V+EGER R DNNLLG F L + P PRG
Sbjct: 418 TLIQRNTTIPTKKEQVFSTYSDNQPGVLIQVFEGERARTKDNNLLGKFELSGIPPAPRGV 477
Query: 479 P-LEVCFSIDENGILTVTAREISTGNMNMITITNDKERLSMFDIEKMIKEAEKYHVEDMQ 537
P + VCF ID NGIL V+A + +TG N ITITNDK RLS DIEKM++EAEKY ED +
Sbjct: 478 PQITVCFDIDANGILNVSAEDKTTGKKNKITITNDKGRLSKEDIEKMVQEAEKYKSEDEE 537
Query: 538 FLRRAKVMCELDSCVYNMKKALKKKDVNLILSPQEIAKINNAITVAMDLLDKNKKEKEID 597
++ + L++ YNM+ ++ + + L + K+ ++I A+ LD N+ E D
Sbjct: 538 HKKKVEAKNALENYAYNMRNTIRDEKIGEKLPAADKKKVEDSIEEAIQWLDGNQL-GEAD 596
Query: 598 VLEGYLEELERMSKHLISK 616
E ++ELE + +I+K
Sbjct: 597 EFEDKMKELESVCNPIIAK 615
>At1g56410 hypothetical protein
Length = 617
Score = 714 bits (1843), Expect = 0.0
Identities = 389/619 (62%), Positives = 462/619 (73%), Gaps = 9/619 (1%)
Query: 1 MAKEYEGRAVGIDLGTTYSCVAVWLDQHQRVEIIHNDQGNRITPSFVAFNNEQRLIGDAA 60
MA + EG A+GIDLGTTYSCV VW QH RVEII NDQGNR TPS+VAF + +RLIGDAA
Sbjct: 1 MAGKGEGPAIGIDLGTTYSCVGVW--QHDRVEIIANDQGNRTTPSYVAFTDSERLIGDAA 58
Query: 61 KNQSATNPENTIFDAKRLIGRKFSDPVVQRDILLWPFKVISGVNDKPMITVKYKGQEKHF 120
KNQ A NP NT+FDAKRLIGR+FSD VQ D+ WPFKV G DKPMI V YKG+EK F
Sbjct: 59 KNQVAMNPVNTVFDAKRLIGRRFSDASVQSDMKFWPFKVTPGQADKPMIFVNYKGEEKQF 118
Query: 121 CAEEISSMVLTKMREVAEAYLGSPVKNAVVTVPAYFNDSQRKATIDAGTIAGLNVIRIIN 180
AEEISSMVL KMRE+AEAYLGS +KNAVVTVPAYFNDSQR+AT DAG IAGLNV+RIIN
Sbjct: 119 AAEEISSMVLIKMREIAEAYLGSSIKNAVVTVPAYFNDSQRQATKDAGVIAGLNVLRIIN 178
Query: 181 EPTAAAIAYGLDKRNDCDGSRNIFVFDLGGGTFDVSILTIKGDVFEVKATAGNTHLGGED 240
EPTAAAIAYGLDK+ G +N+ +FDLGGGTFDVS+LTI+ +FEVKATAG+THLGGED
Sbjct: 179 EPTAAAIAYGLDKKATSVGIKNVLIFDLGGGTFDVSLLTIEEGIFEVKATAGDTHLGGED 238
Query: 241 FDNRMVNYFVEELKKKNKVDISQNPKSLRRLRTACERAKRILSFAFVTTVEVDALFTGID 300
FDNRMVN+FV+E K+KNK DIS + ++LRRLRTACERAKR LS TTVEVD+LF GID
Sbjct: 239 FDNRMVNHFVQEFKRKNKKDISGDARALRRLRTACERAKRTLSSTAQTTVEVDSLFEGID 298
Query: 301 FCSSITRAKFEELNMDFFNECMNIVDTCLRDSKIYKNDIDDVVLVGGSSRIPKVQEQLLE 360
F S ITRAKFEE+NMD F +CM V CLRDSK+ K+ + DVVLVGGS+RIPKVQ+ L +
Sbjct: 299 FYSPITRAKFEEMNMDLFRKCMEPVMKCLRDSKMDKSMVHDVVLVGGSTRIPKVQQLLQD 358
Query: 361 YFKGKELFMGINPDEAVAYGAAVQAAILS-EGCKNVPNLVLRDVTPLSLGISVSIEHIMS 419
+F GKEL INPDEAVAYGAAVQAAILS EG + V +L+L DVTPLSLGI +I +M+
Sbjct: 359 FFNGKELCKSINPDEAVAYGAAVQAAILSGEGNEKVQDLLLLDVTPLSLGIE-TIGGVMT 417
Query: 420 VVIPRNTSIPVKKTSRYSTLYDNQCLVNFLVYEGERPRAADNNLLGSFGLRCL-PGPRGQ 478
+I RNT+IP KK ++T DNQ V VYEGER R DNN+LG F L + P PRG
Sbjct: 418 TLIQRNTTIPAKKEQEFTTTVDNQPDVLIQVYEGERARTIDNNILGQFVLSGIPPAPRGI 477
Query: 479 P-LEVCFSIDENGILTVTAREISTGNMNMITITNDKERLSMFDIEKMIKEAEKYHVEDMQ 537
P VCF ID NGIL V+A + +TG N ITITNDK RLS DIEKM++EAEKY ED +
Sbjct: 478 PQFTVCFDIDSNGILNVSAEDKATGKKNKITITNDKGRLSKDDIEKMVQEAEKYKSEDEE 537
Query: 538 FLRRAKVMCELDSCVYNMKKALKKKDVNLILSPQEIAKINNAITVAMDLLDKNKKEKEID 597
++ + L++ YN+ L +D+ L + K ++I + LD N+ E D
Sbjct: 538 HKKKVEAKNGLENYAYNVGNTL--RDMGEKLPAADKKKFEDSIEEVIQWLDDNQL-AEAD 594
Query: 598 VLEGYLEELERMSKHLISK 616
E ++ELE + +I+K
Sbjct: 595 EFEHKMKELESVWSTIITK 613
>At1g16030 unknown protein
Length = 646
Score = 706 bits (1823), Expect = 0.0
Identities = 370/612 (60%), Positives = 467/612 (75%), Gaps = 7/612 (1%)
Query: 8 RAVGIDLGTTYSCVAVWLDQHQRVEIIHNDQGNRITPSFVAFNNEQRLIGDAAKNQSATN 67
+A+GIDLGTTYSCV VW++ RVEII NDQGNR TPS+VAF + +RLIGDAAKNQ A N
Sbjct: 7 KAIGIDLGTTYSCVGVWMND--RVEIIPNDQGNRTTPSYVAFTDTERLIGDAAKNQVALN 64
Query: 68 PENTIFDAKRLIGRKFSDPVVQRDILLWPFKVISGVNDKPMITVKYKGQEKHFCAEEISS 127
P+NT+FDAKRLIGRKFSDP VQ DIL WPFKV+SG +KPMI V YK +EK F EEISS
Sbjct: 65 PQNTVFDAKRLIGRKFSDPSVQSDILHWPFKVVSGPGEKPMIVVSYKNEEKQFSPEEISS 124
Query: 128 MVLTKMREVAEAYLGSPVKNAVVTVPAYFNDSQRKATIDAGTIAGLNVIRIINEPTAAAI 187
MVL KM+EVAEA+LG VKNAVVTVPAYFNDSQR+AT DAG I+GLNV+RIINEPTAAAI
Sbjct: 125 MVLVKMKEVAEAFLGRTVKNAVVTVPAYFNDSQRQATKDAGAISGLNVLRIINEPTAAAI 184
Query: 188 AYGLDKRNDCDGSRNIFVFDLGGGTFDVSILTIKGDVFEVKATAGNTHLGGEDFDNRMVN 247
AYGLDK+ G +N+ +FDLGGGTFDVS+LTI+ VFEVKATAG+THLGGEDFDNR+VN
Sbjct: 185 AYGLDKKGTKAGEKNVLIFDLGGGTFDVSLLTIEEGVFEVKATAGDTHLGGEDFDNRLVN 244
Query: 248 YFVEELKKKNKVDISQNPKSLRRLRTACERAKRILSFAFVTTVEVDALFTGIDFCSSITR 307
+FV E ++K+K DI+ N ++LRRLRTACERAKR LS TT+E+D+L GIDF ++I+R
Sbjct: 245 HFVAEFRRKHKKDIAGNARALRRLRTACERAKRTLSSTAQTTIEIDSLHEGIDFYATISR 304
Query: 308 AKFEELNMDFFNECMNIVDTCLRDSKIYKNDIDDVVLVGGSSRIPKVQEQLLEYFKGKEL 367
A+FEE+NMD F +CM+ V+ L+D+K+ K+ + DVVLVGGS+RIPK+Q+ L ++F GKEL
Sbjct: 305 ARFEEMNMDLFRKCMDPVEKVLKDAKLDKSSVHDVVLVGGSTRIPKIQQLLQDFFNGKEL 364
Query: 368 FMGINPDEAVAYGAAVQAAILS-EGCKNVPNLVLRDVTPLSLGISVSIEHIMSVVIPRNT 426
INPDEAVAYGAAVQAAIL+ EG + V +L+L DV PLSLG+ + +M+V+IPRNT
Sbjct: 365 CKSINPDEAVAYGAAVQAAILTGEGSEKVQDLLLLDVAPLSLGLETA-GGVMTVLIPRNT 423
Query: 427 SIPVKKTSRYSTLYDNQCLVNFLVYEGERPRAADNNLLGSFGLRCL-PGPRGQP-LEVCF 484
++P KK +ST DNQ V VYEGER R DNNLLG+F L+ + P PRG P + VCF
Sbjct: 424 TVPCKKEQVFSTYADNQPGVLIQVYEGERARTRDNNLLGTFELKGIPPAPRGVPQINVCF 483
Query: 485 SIDENGILTVTAREISTGNMNMITITNDKERLSMFDIEKMIKEAEKYHVEDMQFLRRAKV 544
ID NGIL V+A + + G N ITITNDK RLS +IEKM+++AEKY ED Q ++ +
Sbjct: 484 DIDANGILNVSAEDKTAGVKNQITITNDKGRLSKEEIEKMVQDAEKYKAEDEQVKKKVEA 543
Query: 545 MCELDSCVYNMKKALKKKDVNLILSPQEIAKINNAITVAMDLLDKNKKEKEIDVLEGYLE 604
L++ YNM+ +K + + L+ ++ KI AI ++ ++ N+ E+D E L+
Sbjct: 544 KNSLENYAYNMRNTIKDEKLAQKLTQEDKQKIEKAIDETIEWIEGNQL-AEVDEFEYKLK 602
Query: 605 ELERMSKHLISK 616
ELE + +ISK
Sbjct: 603 ELEGICNPIISK 614
>At1g09080 putative luminal binding protein
Length = 678
Score = 604 bits (1558), Expect = e-173
Identities = 332/605 (54%), Positives = 430/605 (70%), Gaps = 12/605 (1%)
Query: 7 GRAVGIDLGTTYSCVAVWLDQHQRVEIIHNDQGNRITPSFVAFNNEQRLIGDAAKNQSAT 66
G +GIDLGTTYSCV V+ ++H VEII NDQGNRITPS+VAF + +RLIG+AAKNQ+A
Sbjct: 50 GTVIGIDLGTTYSCVGVYHNKH--VEIIANDQGNRITPSWVAFTDTERLIGEAAKNQAAK 107
Query: 67 NPENTIFDAKRLIGRKFSDPVVQRDILLWPFKVISGVNDKPMITVKYKGQEKHFCAEEIS 126
NPE TIFD KRLIGRKF DP VQRDI P+KV++ + KP I VK KG+EK F EEIS
Sbjct: 108 NPERTIFDPKRLIGRKFDDPDVQRDIKFLPYKVVNK-DGKPYIQVKVKGEEKLFSPEEIS 166
Query: 127 SMVLTKMREVAEAYLGSPVKNAVVTVPAYFNDSQRKATIDAGTIAGLNVIRIINEPTAAA 186
+M+LTKM+E AEA+LG +K+AV+TVPAYFND+QR+AT DAG IAGLNV+RIINEPT AA
Sbjct: 167 AMILTKMKETAEAFLGKKIKDAVITVPAYFNDAQRQATKDAGAIAGLNVVRIINEPTGAA 226
Query: 187 IAYGLDKRNDCDGSRNIFVFDLGGGTFDVSILTIKGDVFEVKATAGNTHLGGEDFDNRMV 246
IAYGLDK+ G NI V+DLGGGTFDVSILTI VFEV +T+G+THLGGEDFD+R++
Sbjct: 227 IAYGLDKKG---GESNILVYDLGGGTFDVSILTIDNGVFEVLSTSGDTHLGGEDFDHRVM 283
Query: 247 NYFVEELKKKNKVDISQNPKSLRRLRTACERAKRILSFAFVTTVEVDALFTGIDFCSSIT 306
+YF++ +KKK DIS++ K+L +LR CE AKR LS VE+++LF G+DF +T
Sbjct: 284 DYFIKLVKKKYNKDISKDHKALGKLRRECELAKRSLSNQHQVRVEIESLFDGVDFSEPLT 343
Query: 307 RAKFEELNMDFFNECMNIVDTCLRDSKIYKNDIDDVVLVGGSSRIPKVQEQLLEYFKGKE 366
RA+FEELNMD F + M V L+D+ + K+DID++VLVGGS+RIPKVQ+ L ++F GKE
Sbjct: 344 RARFEELNMDLFKKTMEPVKKALKDAGLKKSDIDEIVLVGGSTRIPKVQQMLKDFFDGKE 403
Query: 367 LFMGINPDEAVAYGAAVQAAILS-EGCKNVPNLVLRDVTPLSLGISVSIEHIMSVVIPRN 425
G NPDEAVAYGAAVQ +LS EG + N++L DV PLSLGI ++ +M+ +IPRN
Sbjct: 404 PSKGTNPDEAVAYGAAVQGGVLSGEGGEETQNILLLDVAPLSLGIE-TVGGVMTNIIPRN 462
Query: 426 TSIPVKKTSRYSTLYDNQCLVNFLVYEGERPRAADNNLLGSFGLR-CLPGPRGQP-LEVC 483
T IP KK+ ++T D Q V VYEGER DN LG F L LP PRG P +EV
Sbjct: 463 TVIPTKKSQVFTTYQDQQTTVTINVYEGERSMTKDNRELGKFDLTGILPAPRGVPQIEVT 522
Query: 484 FSIDENGILTVTAREISTGNMNMITITNDKERLSMFDIEKMIKEAEKYHVEDMQFLRRAK 543
F +D NGIL V A + ITITNDK RL+ +IE+MI+EAE++ ED +
Sbjct: 523 FEVDANGILQVKAEDKVAKTSQSITITNDKGRLTEEEIEEMIREAEEFAEEDKIMKEKID 582
Query: 544 VMCELDSCVYNMKKALKKKD-VNLILSPQEIAKINNAITVAMDLLDKNKKEKEIDVLEGY 602
+L++ VYNMK + K+ + +S ++ K+ + A++ L++N ++ D E
Sbjct: 583 ARNKLETYVYNMKSTVADKEKLAKKISDEDKEKMEGVLKEALEWLEENVNAEKEDYDE-K 641
Query: 603 LEELE 607
L+E+E
Sbjct: 642 LKEVE 646
>At5g28540 luminal binding protein
Length = 669
Score = 593 bits (1530), Expect = e-170
Identities = 328/614 (53%), Positives = 432/614 (69%), Gaps = 13/614 (2%)
Query: 7 GRAVGIDLGTTYSCVAVWLDQHQRVEIIHNDQGNRITPSFVAFNNEQRLIGDAAKNQSAT 66
G +GIDLGTTYSCV V+ + H VEII NDQGNRITPS+V F + +RLIG+AAKNQ+A
Sbjct: 35 GSVIGIDLGTTYSCVGVYKNGH--VEIIANDQGNRITPSWVGFTDSERLIGEAAKNQAAV 92
Query: 67 NPENTIFDAKRLIGRKFSDPVVQRDILLWPFKVISGVNDKPMITVKYK-GQEKHFCAEEI 125
NPE T+FD KRLIGRKF D VQ+D L P+++++ + KP I VK K G+ K F EEI
Sbjct: 93 NPERTVFDVKRLIGRKFEDKEVQKDRKLVPYQIVNK-DGKPYIQVKIKDGETKVFSPEEI 151
Query: 126 SSMVLTKMREVAEAYLGSPVKNAVVTVPAYFNDSQRKATIDAGTIAGLNVIRIINEPTAA 185
S+M+LTKM+E AEAYLG +K+AVVTVPAYFND+QR+AT DAG IAGLNV RIINEPTAA
Sbjct: 152 SAMILTKMKETAEAYLGKKIKDAVVTVPAYFNDAQRQATKDAGVIAGLNVARIINEPTAA 211
Query: 186 AIAYGLDKRNDCDGSRNIFVFDLGGGTFDVSILTIKGDVFEVKATAGNTHLGGEDFDNRM 245
AIAYGLDK+ G +NI VFDLGGGTFDVS+LTI VFEV +T G+THLGGEDFD+R+
Sbjct: 212 AIAYGLDKKG---GEKNILVFDLGGGTFDVSVLTIDNGVFEVLSTNGDTHLGGEDFDHRV 268
Query: 246 VNYFVEELKKKNKVDISQNPKSLRRLRTACERAKRILSFAFVTTVEVDALFTGIDFCSSI 305
+ YF++ +KKK++ DIS++ K+L +LR CERAKR LS VE+++LF G+DF +
Sbjct: 269 MEYFIKLIKKKHQKDISKDNKALGKLRRECERAKRALSSQHQVRVEIESLFDGVDFSEPL 328
Query: 306 TRAKFEELNMDFFNECMNIVDTCLRDSKIYKNDIDDVVLVGGSSRIPKVQEQLLEYFKGK 365
TRA+FEELN D F + M V + D+ + K+ ID++VLVGGS+RIPKVQ+ L ++F+GK
Sbjct: 329 TRARFEELNNDLFRKTMGPVKKAMDDAGLQKSQIDEIVLVGGSTRIPKVQQLLKDFFEGK 388
Query: 366 ELFMGINPDEAVAYGAAVQAAILS-EGCKNVPNLVLRDVTPLSLGISVSIEHIMSVVIPR 424
E G+NPDEAVAYGAAVQ ILS EG +++L DV PL+LGI ++ +M+ +IPR
Sbjct: 389 EPNKGVNPDEAVAYGAAVQGGILSGEGGDETKDILLLDVAPLTLGIE-TVGGVMTKLIPR 447
Query: 425 NTSIPVKKTSRYSTLYDNQCLVNFLVYEGERPRAADNNLLGSFGLRCL-PGPRGQP-LEV 482
NT IP KK+ ++T D Q V+ V+EGER D LLG F L + P PRG P +EV
Sbjct: 448 NTVIPTKKSQVFTTYQDQQTTVSIQVFEGERSLTKDCRLLGKFDLNGIPPAPRGTPQIEV 507
Query: 483 CFSIDENGILTVTAREISTGNMNMITITNDKERLSMFDIEKMIKEAEKYHVEDMQFLRRA 542
F +D NGIL V A + ++G ITITN+K RLS +I++M+KEAE++ ED + +
Sbjct: 508 TFEVDANGILNVKAEDKASGKSEKITITNEKGRLSQEEIDRMVKEAEEFAEEDKKVKEKI 567
Query: 543 KVMCELDSCVYNMKKALKKKD-VNLILSPQEIAKINNAITVAMDLLDKNKKEKEIDVLEG 601
L++ VYNMK + KD + L E KI A A++ LD+N + E + +
Sbjct: 568 DARNALETYVYNMKNQVNDKDKLADKLEGDEKEKIEAATKEALEWLDEN-QNSEKEEYDE 626
Query: 602 YLEELERMSKHLIS 615
L+E+E + +I+
Sbjct: 627 KLKEVEAVCNPIIT 640
>At5g42020 luminal binding protein (dbj|BAA13948.1)
Length = 668
Score = 590 bits (1522), Expect = e-169
Identities = 327/614 (53%), Positives = 431/614 (69%), Gaps = 13/614 (2%)
Query: 7 GRAVGIDLGTTYSCVAVWLDQHQRVEIIHNDQGNRITPSFVAFNNEQRLIGDAAKNQSAT 66
G +GIDLGTTYSCV V+ + H VEII NDQGNRITPS+V F + +RLIG+AAKNQ+A
Sbjct: 35 GSVIGIDLGTTYSCVGVYKNGH--VEIIANDQGNRITPSWVGFTDSERLIGEAAKNQAAV 92
Query: 67 NPENTIFDAKRLIGRKFSDPVVQRDILLWPFKVISGVNDKPMITVKYK-GQEKHFCAEEI 125
NPE T+FD KRLIGRKF D VQ+D L P+++++ + KP I VK K G+ K F EEI
Sbjct: 93 NPERTVFDVKRLIGRKFEDKEVQKDRKLVPYQIVNK-DGKPYIQVKIKDGETKVFSPEEI 151
Query: 126 SSMVLTKMREVAEAYLGSPVKNAVVTVPAYFNDSQRKATIDAGTIAGLNVIRIINEPTAA 185
S+M+LTKM+E AEAYLG +K+AVVTVPAYFND+QR+AT DAG IAGLNV RIINEPTAA
Sbjct: 152 SAMILTKMKETAEAYLGKKIKDAVVTVPAYFNDAQRQATKDAGVIAGLNVARIINEPTAA 211
Query: 186 AIAYGLDKRNDCDGSRNIFVFDLGGGTFDVSILTIKGDVFEVKATAGNTHLGGEDFDNRM 245
AIAYGLDK+ G +NI VFDLGGGTFDVS+LTI VFEV +T G+THLGGEDFD+R+
Sbjct: 212 AIAYGLDKKG---GEKNILVFDLGGGTFDVSVLTIDNGVFEVLSTNGDTHLGGEDFDHRI 268
Query: 246 VNYFVEELKKKNKVDISQNPKSLRRLRTACERAKRILSFAFVTTVEVDALFTGIDFCSSI 305
+ YF++ +KKK++ DIS++ K+L +LR CERAKR LS VE+++LF G+D +
Sbjct: 269 MEYFIKLIKKKHQKDISKDNKALGKLRRECERAKRALSSQHQVRVEIESLFDGVDLSEPL 328
Query: 306 TRAKFEELNMDFFNECMNIVDTCLRDSKIYKNDIDDVVLVGGSSRIPKVQEQLLEYFKGK 365
TRA+FEELN D F + M V + D+ + K+ ID++VLVGGS+RIPKVQ+ L ++F+GK
Sbjct: 329 TRARFEELNNDLFRKTMGPVKKAMDDAGLQKSQIDEIVLVGGSTRIPKVQQLLKDFFEGK 388
Query: 366 ELFMGINPDEAVAYGAAVQAAILS-EGCKNVPNLVLRDVTPLSLGISVSIEHIMSVVIPR 424
E G+NPDEAVAYGAAVQ ILS EG +++L DV PL+LGI ++ +M+ +IPR
Sbjct: 389 EPNKGVNPDEAVAYGAAVQGGILSGEGGDETKDILLLDVAPLTLGIE-TVGGVMTKLIPR 447
Query: 425 NTSIPVKKTSRYSTLYDNQCLVNFLVYEGERPRAADNNLLGSFGLRCL-PGPRGQP-LEV 482
NT IP KK+ ++T D Q V+ V+EGER D LLG F L + P PRG P +EV
Sbjct: 448 NTVIPTKKSQVFTTYQDQQTTVSIQVFEGERSLTKDCRLLGKFDLTGVPPAPRGTPQIEV 507
Query: 483 CFSIDENGILTVTAREISTGNMNMITITNDKERLSMFDIEKMIKEAEKYHVEDMQFLRRA 542
F +D NGIL V A + ++G ITITN+K RLS +I++M+KEAE++ ED + +
Sbjct: 508 TFEVDANGILNVKAEDKASGKSEKITITNEKGRLSQEEIDRMVKEAEEFAEEDKKVKEKI 567
Query: 543 KVMCELDSCVYNMKKALKKKD-VNLILSPQEIAKINNAITVAMDLLDKNKKEKEIDVLEG 601
L++ VYNMK + KD + L E KI A A++ LD+N + E + +
Sbjct: 568 DARNALETYVYNMKNQVSDKDKLADKLEGDEKEKIEAATKEALEWLDEN-QNSEKEEYDE 626
Query: 602 YLEELERMSKHLIS 615
L+E+E + +I+
Sbjct: 627 KLKEVEAVCNPIIT 640
>At4g37910 heat shock protein 70 like protein
Length = 682
Score = 439 bits (1130), Expect = e-123
Identities = 267/582 (45%), Positives = 369/582 (62%), Gaps = 32/582 (5%)
Query: 10 VGIDLGTTYSCVAVWLDQHQRVEIIHNDQGNRITPSFVAFNNE-QRLIGDAAKNQSATNP 68
+GIDLGTT SCV+V + RV I N +G+R TPS VA N + + L+G AK Q+ TNP
Sbjct: 55 IGIDLGTTNSCVSVMEGKTARV--IENAEGSRTTPSVVAMNQKGELLVGTPAKRQAVTNP 112
Query: 69 ENTIFDAKRLIGRKFSDPVVQRDILLWPFKVISGVNDKPMITVKYKGQEKHFCAEEISSM 128
NTIF +KRLIGR+F DP Q+++ + P+K++ N V+ GQ+ F +I +
Sbjct: 113 TNTIFGSKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAW--VEANGQK--FSPSQIGAN 168
Query: 129 VLTKMREVAEAYLGSPVKNAVVTVPAYFNDSQRKATIDAGTIAGLNVIRIINEPTAAAIA 188
VLTKM+E AEAYLG + AVVTVPAYFND+QR+AT DAG IAGL+V RIINEPTAAA++
Sbjct: 169 VLTKMKETAEAYLGKSINKAVVTVPAYFNDAQRQATKDAGKIAGLDVQRIINEPTAAALS 228
Query: 189 YGLDKRNDCDGSRNIFVFDLGGGTFDVSILTIKGDVFEVKATAGNTHLGGEDFDNRMVNY 248
YG++ + I VFDLGGGTFDVSIL I VFEVKAT G+T LGGEDFDN ++ Y
Sbjct: 229 YGMNNKEGV-----IAVFDLGGGTFDVSILEISSGVFEVKATNGDTFLGGEDFDNTLLEY 283
Query: 249 FVEELKKKNKVDISQNPKSLRRLRTACERAKRILSFAFVTTVEVDALFTGIDFCS----- 303
V E K+ + +D++++ +L+RLR A E+AK + + T E++ F D
Sbjct: 284 LVNEFKRSDNIDLTKDNLALQRLREAAEKAK--IELSSTTQTEINLPFITADASGAKHLN 341
Query: 304 -SITRAKFEELNMDFFNECMNIVDTCLRDSKIYKNDIDDVVLVGGSSRIPKVQEQLLEYF 362
++TR+KFE L + CL+D+ + ++D+V+LVGG +R+PKVQE + E F
Sbjct: 342 ITLTRSKFEGLVGKLIERTRSPCQNCLKDAGVTIKEVDEVLLVGGMTRVPKVQEIVSEIF 401
Query: 363 KGKELFMGINPDEAVAYGAAVQAAILSEGCKNVPNLVLRDVTPLSLGISVSIEHIMSVVI 422
GK G+NPDEAVA GAA+Q IL +V +L+L DV PLSLGI ++ + + +I
Sbjct: 402 -GKSPCKGVNPDEAVAMGAAIQGGILR---GDVKDLLLLDVVPLSLGIE-TLGAVFTKLI 456
Query: 423 PRNTSIPVKKTSRYSTLYDNQCLVNFLVYEGERPRAADNNLLGSFGLRCL-PGPRGQP-L 480
PRNT+IP KK+ +ST DNQ V V +GER AADN +LG F L + P PRG P +
Sbjct: 457 PRNTTIPTKKSQVFSTAADNQMQVGIKVLQGEREMAADNKVLGEFDLVGIPPAPRGMPQI 516
Query: 481 EVCFSIDENGILTVTAREISTGNMNMITITNDKERLSMFDIEKMIKEAEKYHVEDMQFLR 540
EV F ID NGI TV+A++ +TG ITI LS +I +M+KEAE +D + +
Sbjct: 517 EVTFDIDANGITTVSAKDKATGKEQNITI-RSSGGLSDDEINRMVKEAELNAQKDQEKKQ 575
Query: 541 RAKVMCELDSCVYNMKKALKKKDVNLILSPQEIA-KINNAIT 581
+ D+ +Y+++K+L + + P EIA +I A++
Sbjct: 576 LIDLRNSADTTIYSVEKSLSEYREKI---PAEIASEIETAVS 614
>At5g09590 heat shock protein 70 (Hsc70-5)
Length = 682
Score = 435 bits (1118), Expect = e-122
Identities = 263/575 (45%), Positives = 362/575 (62%), Gaps = 31/575 (5%)
Query: 10 VGIDLGTTYSCVAVWLDQHQRVEIIHNDQGNRITPSFVAFNNE-QRLIGDAAKNQSATNP 68
+GIDLGTT SCVAV ++ +V I N +G R TPS VAFN + + L+G AK Q+ TNP
Sbjct: 60 IGIDLGTTNSCVAVMEGKNPKV--IENAEGARTTPSVVAFNTKGELLVGTPAKRQAVTNP 117
Query: 69 ENTIFDAKRLIGRKFSDPVVQRDILLWPFKVISGVNDKPMITVKYKGQEKHFCAEEISSM 128
NT+ KRLIGRKF DP Q+++ + P+K++ N V+ GQ+ + +I +
Sbjct: 118 TNTVSGTKRLIGRKFDDPQTQKEMKMVPYKIVRAPNGDAW--VEANGQQ--YSPSQIGAF 173
Query: 129 VLTKMREVAEAYLGSPVKNAVVTVPAYFNDSQRKATIDAGTIAGLNVIRIINEPTAAAIA 188
+LTKM+E AEAYLG V AVVTVPAYFND+QR+AT DAG IAGL+V RIINEPTAAA++
Sbjct: 174 ILTKMKETAEAYLGKSVTKAVVTVPAYFNDAQRQATKDAGRIAGLDVERIINEPTAAALS 233
Query: 189 YGLDKRNDCDGSRNIFVFDLGGGTFDVSILTIKGDVFEVKATAGNTHLGGEDFDNRMVNY 248
YG+ + I VFDLGGGTFDVS+L I VFEVKAT G+T LGGEDFDN ++++
Sbjct: 234 YGMTNKEGL-----IAVFDLGGGTFDVSVLEISNGVFEVKATNGDTFLGGEDFDNALLDF 288
Query: 249 FVEELKKKNKVDISQNPKSLRRLRTACERAKRILSFAFVTTVEVDALFTGID------FC 302
V E K +D++++ +L+RLR A E+AK + + + E++ F D F
Sbjct: 289 LVNEFKTTEGIDLAKDRLALQRLREAAEKAK--IELSSTSQTEINLPFITADASGAKHFN 346
Query: 303 SSITRAKFEELNMDFFNECMNIVDTCLRDSKIYKNDIDDVVLVGGSSRIPKVQEQLLEYF 362
++TR++FE L + CL+D+ I ++D+V+LVGG +R+PKVQ + E F
Sbjct: 347 ITLTRSRFETLVNHLIERTRDPCKNCLKDAGISAKEVDEVLLVGGMTRVPKVQSIVAEIF 406
Query: 363 KGKELFMGINPDEAVAYGAAVQAAILSEGCKNVPNLVLRDVTPLSLGISVSIEHIMSVVI 422
GK G+NPDEAVA GAA+Q IL +V L+L DVTPLSLGI ++ + + +I
Sbjct: 407 -GKSPSKGVNPDEAVAMGAALQGGILR---GDVKELLLLDVTPLSLGIE-TLGGVFTRLI 461
Query: 423 PRNTSIPVKKTSRYSTLYDNQCLVNFLVYEGERPRAADNNLLGSFGLRCL-PGPRGQP-L 480
RNT+IP KK+ +ST DNQ V V +GER A DN LLG F L + P PRG P +
Sbjct: 462 TRNTTIPTKKSQVFSTAADNQTQVGIRVLQGEREMATDNKLLGEFDLVGIPPSPRGVPQI 521
Query: 481 EVCFSIDENGILTVTAREISTGNMNMITITNDKERLSMFDIEKMIKEAEKYHVEDMQFLR 540
EV F ID NGI+TV+A++ +TG + ITI LS DI+KM++EAE + +D +
Sbjct: 522 EVTFDIDANGIVTVSAKDKTTGKVQQITI-RSSGGLSEDDIQKMVREAELHAQKDKERKE 580
Query: 541 RAKVMCELDSCVYNMKKALKKKDVNLILSPQEIAK 575
D+ +Y+++K+L + + P EIAK
Sbjct: 581 LIDTKNTADTTIYSIEKSLGEYREKI---PSEIAK 612
>At4g24280 hsp 70-like protein
Length = 718
Score = 413 bits (1062), Expect = e-115
Identities = 257/617 (41%), Positives = 369/617 (59%), Gaps = 27/617 (4%)
Query: 8 RAVGIDLGTTYSCVAVWLDQHQRVEIIHNDQGNRITPSFVAFNNE-QRLIGDAAKNQSAT 66
+ VGIDLGTT S VA + + I+ N +G R TPS VA+ RL+G AK Q+
Sbjct: 79 KVVGIDLGTTNSAVAAM--EGGKPTIVTNAEGQRTTPSVVAYTKSGDRLVGQIAKRQAVV 136
Query: 67 NPENTIFDAKRLIGRKFSDPVVQRDILLWPFKVISGVNDKPMITVKYKGQEKHFCAEEIS 126
NPENT F KR IGRK ++ V + ++V+ N+ + ++ K F AEEIS
Sbjct: 137 NPENTFFSVKRFIGRKMNE--VDEESKQVSYRVVRDENNN--VKLECPAINKQFAAEEIS 192
Query: 127 SMVLTKMREVAEAYLGSPVKNAVVTVPAYFNDSQRKATIDAGTIAGLNVIRIINEPTAAA 186
+ VL K+ + A +L V AV+TVPAYFNDSQR AT DAG IAGL V+RIINEPTAA+
Sbjct: 193 AQVLRKLVDDASRFLNDKVTKAVITVPAYFNDSQRTATKDAGRIAGLEVLRIINEPTAAS 252
Query: 187 IAYGLDKRNDCDGSRNIFVFDLGGGTFDVSILTIKGDVFEVKATAGNTHLGGEDFDNRMV 246
+AYG D++ + I VFDLGGGTFDVS+L + VFEV +T+G+THLGG+DFD R+V
Sbjct: 253 LAYGFDRK----ANETILVFDLGGGTFDVSVLEVGDGVFEVLSTSGDTHLGGDDFDKRVV 308
Query: 247 NYFVEELKKKNKVDISQNPKSLRRLRTACERAKRILSFAFVTTVEVDALFTGID----FC 302
++ E KK +D+ ++ ++L+RL A E+AK LS T + + + D
Sbjct: 309 DWLAAEFKKDEGIDLLKDKQALQRLTEAAEKAKIELSSLTQTNMSLPFITATADGPKHIE 368
Query: 303 SSITRAKFEELNMDFFNECMNIVDTCLRDSKIYKNDIDDVVLVGGSSRIPKVQEQLLEYF 362
+++TRAKFEEL D + V+ LRD+K+ DID+V+LVGGS+RIP VQE L+
Sbjct: 369 TTLTRAKFEELCSDLLDRVRTPVENSLRDAKLSFKDIDEVILVGGSTRIPAVQE-LVRKV 427
Query: 363 KGKELFMGINPDEAVAYGAAVQAAILSEGCKNVPNLVLRDVTPLSLGISVSIEHIMSVVI 422
GKE + +NPDE VA GAAVQA +L+ +V ++VL DVTPLS+G+ ++ +M+ +I
Sbjct: 428 TGKEPNVTVNPDEVVALGAAVQAGVLA---GDVSDIVLLDVTPLSIGLE-TLGGVMTKII 483
Query: 423 PRNTSIPVKKTSRYSTLYDNQCLVNFLVYEGERPRAADNNLLGSFGLRCL-PGPRGQP-L 480
PRNT++P K+ +ST D Q V V +GER DN LGSF L + P PRG P +
Sbjct: 484 PRNTTLPTSKSEVFSTAADGQTSVEINVLQGEREFVRDNKSLGSFRLDGIPPAPRGVPQI 543
Query: 481 EVCFSIDENGILTVTAREISTGNMNMITITNDKERLSMFDIEKMIKEAEKYHVEDMQFLR 540
EV F ID NGIL+V+A + TG ITIT L ++++M++EAE++ +D +
Sbjct: 544 EVKFDIDANGILSVSAVDKGTGKKQDITITG-ASTLPKDEVDQMVQEAERFAKDDKEKRD 602
Query: 541 RAKVMCELDSCVYNMKKALKKKDVNLILSPQEI-AKINNAITVAMDLLDKNKKEKEIDVL 599
+ DS VY +K LK+ + P E+ K+ + D + ++ D +
Sbjct: 603 AIDTKNQADSVVYQTEKQLKELGEKI---PGEVKEKVEAKLQELKDKIGSGSTQEIKDAM 659
Query: 600 EGYLEELERMSKHLISK 616
+E+ ++ + L ++
Sbjct: 660 AALNQEVMQIGQSLYNQ 676
>At5g49910 heat shock protein 70 (Hsc70-7)
Length = 718
Score = 409 bits (1051), Expect = e-114
Identities = 249/561 (44%), Positives = 345/561 (61%), Gaps = 23/561 (4%)
Query: 8 RAVGIDLGTTYSCVAVWLDQHQRVEIIHNDQGNRITPSFVAFN-NEQRLIGDAAKNQSAT 66
+ VGIDLGTT S VA + + I+ N +G R TPS VA+ ++ RL+G AK Q+
Sbjct: 79 KVVGIDLGTTNSAVAAM--EGGKPTIVTNAEGQRTTPSVVAYTKSKDRLVGQIAKRQAVV 136
Query: 67 NPENTIFDAKRLIGRKFSDPVVQRDILLWPFKVISGVNDKPMITVKYKGQEKHFCAEEIS 126
NPENT F KR IGR+ ++ V + ++VI N + G K F AEEIS
Sbjct: 137 NPENTFFSVKRFIGRRMNE--VAEESKQVSYRVIKDENGNVKLDCPAIG--KQFAAEEIS 192
Query: 127 SMVLTKMREVAEAYLGSPVKNAVVTVPAYFNDSQRKATIDAGTIAGLNVIRIINEPTAAA 186
+ VL K+ + A +L V AV+TVPAYFNDSQR AT DAG IAGL V+RIINEPTAA+
Sbjct: 193 AQVLRKLVDDASRFLNDKVTKAVITVPAYFNDSQRTATKDAGRIAGLEVLRIINEPTAAS 252
Query: 187 IAYGLDKRNDCDGSRNIFVFDLGGGTFDVSILTIKGDVFEVKATAGNTHLGGEDFDNRMV 246
+AYG ++++ + I VFDLGGGTFDVS+L + VFEV +T+G+THLGG+DFD R+V
Sbjct: 253 LAYGFERKS----NETILVFDLGGGTFDVSVLEVGDGVFEVLSTSGDTHLGGDDFDKRVV 308
Query: 247 NYFVEELKKKNKVDISQNPKSLRRLRTACERAKRILSFAFVTTVEVDALFTGID----FC 302
++ KK +D+ ++ ++L+RL A E+AK LS T + + + D
Sbjct: 309 DWLASTFKKDEGIDLLKDKQALQRLTEAAEKAKIELSSLTQTNMSLPFITATADGPKHIE 368
Query: 303 SSITRAKFEELNMDFFNECMNIVDTCLRDSKIYKNDIDDVVLVGGSSRIPKVQEQLLEYF 362
+++TR KFEEL D + V+ LRD+K+ DID+V+LVGGS+RIP VQ+ L+
Sbjct: 369 TTLTRGKFEELCSDLLDRVRTPVENSLRDAKLSFKDIDEVILVGGSTRIPAVQD-LVRKL 427
Query: 363 KGKELFMGINPDEAVAYGAAVQAAILSEGCKNVPNLVLRDVTPLSLGISVSIEHIMSVVI 422
GKE + +NPDE VA GAAVQA +LS +V ++VL DVTPLSLG+ ++ +M+ +I
Sbjct: 428 TGKEPNVSVNPDEVVALGAAVQAGVLS---GDVSDIVLLDVTPLSLGLE-TLGGVMTKII 483
Query: 423 PRNTSIPVKKTSRYSTLYDNQCLVNFLVYEGERPRAADNNLLGSFGLRCL-PGPRGQP-L 480
PRNT++P K+ +ST D Q V V +GER DN +GSF L + P PRG P +
Sbjct: 484 PRNTTLPTSKSEVFSTAADGQTSVEINVLQGEREFVRDNKSIGSFRLDGIPPAPRGVPQI 543
Query: 481 EVCFSIDENGILTVTAREISTGNMNMITITNDKERLSMFDIEKMIKEAEKYHVEDMQFLR 540
EV F ID NGIL+V+A + TG ITIT L +++ M++EAE++ ED +
Sbjct: 544 EVKFDIDANGILSVSASDKGTGKKQDITITG-ASTLPKDEVDTMVQEAERFAKEDKEKRD 602
Query: 541 RAKVMCELDSCVYNMKKALKK 561
+ DS VY +K LK+
Sbjct: 603 AIDTKNQADSVVYQTEKQLKE 623
>At1g79920 putative heat-shock protein (At1g79920)
Length = 736
Score = 251 bits (641), Expect = 9e-67
Identities = 167/533 (31%), Positives = 278/533 (51%), Gaps = 21/533 (3%)
Query: 10 VGIDLGTTYSCVAVWLDQHQRVEIIHNDQGNRITPSFVAFNNEQRLIGDAAKNQSATNPE 69
VG D G VAV + + ++++ ND+ NR TP+ V F ++QR IG A + NP+
Sbjct: 4 VGFDFGNENCLVAV--ARQRGIDVVLNDESNRETPAIVCFGDKQRFIGTAGAASTMMNPK 61
Query: 70 NTIFDAKRLIGRKFSDPVVQRDILLWPFKVISGVNDKPMITVKYKGQEKHFCAEEISSMV 129
N+I KRLIGR+FSDP +QRDI PF V G + P+I Y G+ + F ++ M+
Sbjct: 62 NSISQIKRLIGRQFSDPELQRDIKSLPFSVTEGPDGYPLIHANYLGEIRAFTPTQVMGMM 121
Query: 130 LTKMREVAEAYLGSPVKNAVVTVPAYFNDSQRKATIDAGTIAGLNVIRIINEPTAAAIAY 189
L+ ++ +AE L + V + + +P YF D QR+A +DA TIAGL+ + +I+E TA A+AY
Sbjct: 122 LSNLKGIAEKNLNTAVVDCCIGIPVYFTDLQRRAVLDAATIAGLHPLHLIHETTATALAY 181
Query: 190 GLDKRNDCDGSR-NIFVFDLGGGTFDVSILTIKGDVFEVKATAGNTHLGGEDFDNRMVNY 248
G+ K + + + N+ D+G + V I K ++ + A + LGG DFD + N+
Sbjct: 182 GIYKTDLPENDQLNVAFIDIGHASMQVCIAGFKKGQLKILSHAFDRSLGGRDFDEVLFNH 241
Query: 249 FVEELKKKNKVDISQNPKSLRRLRTACERAKRILSFAFVTTVEVDALFTGIDFCSSITRA 308
F + K + K+D+SQN K+ RLR CE+ K++LS + + ++ L D I R
Sbjct: 242 FAAKFKDEYKIDVSQNAKASLRLRATCEKLKKVLSANPMAPLNIECLMAEKDVRGVIKRE 301
Query: 309 KFEELNMDFFNECMNIVDTCLRDSKIYKNDIDDVVLVGGSSRIPKVQEQLLEYFKGKELF 368
+FEE+++ ++ L D+ + D+ V +VG SR+P + + L E+F GKE
Sbjct: 302 EFEEISIPILERVKRPLEKALSDAGLTVEDVHMVEVVGSGSRVPAMIKILTEFF-GKEPR 360
Query: 369 MGINPDEAVAYGAAVQAAILSEGCKNVPNLVLRDVTPLSLGIS-----------VSIEHI 417
+N E V+ G A+Q AILS K V + + P S+ ++ +
Sbjct: 361 RTMNASECVSRGCALQCAILSPTFK-VREFQVHESFPFSISLAWKGAATDAQNGGTENQQ 419
Query: 418 MSVVIPRNTSIP-VKKTSRYSTLYDNQCLVNFLVYEGERPRAADNNLLGSFGLRCLPGPR 476
++V P+ IP VK + Y + + + V + + P +G F + G R
Sbjct: 420 STIVFPKGNPIPSVKALTFYRSGTFSIDVQYSDVNDLQAPPKISTYTIGPF--QSSKGER 477
Query: 477 GQPLEVCFSIDENGILTVTAREISTGNMNMITITNDK-ERLSMFDIEKMIKEA 528
+ L+V ++ +GI++V + + +++T D+ E + D +K EA
Sbjct: 478 AK-LKVKVRLNLHGIVSVESATLLEEEEVEVSVTKDQSEETAKMDTDKASAEA 529
>At1g79930 putative heat-shock protein
Length = 831
Score = 251 bits (641), Expect = 9e-67
Identities = 142/403 (35%), Positives = 228/403 (56%), Gaps = 5/403 (1%)
Query: 10 VGIDLGTTYSCVAVWLDQHQRVEIIHNDQGNRITPSFVAFNNEQRLIGDAAKNQSATNPE 69
VG D G VAV + + ++++ ND+ NR TP+ V F ++QR IG A + NP+
Sbjct: 4 VGFDFGNENCLVAV--ARQRGIDVVLNDESNRETPAIVCFGDKQRFIGTAGAASTMMNPK 61
Query: 70 NTIFDAKRLIGRKFSDPVVQRDILLWPFKVISGVNDKPMITVKYKGQEKHFCAEEISSMV 129
N+I KRLIGR+FSDP +QRDI PF V G + P+I Y G+++ F ++ M+
Sbjct: 62 NSISQIKRLIGRQFSDPELQRDIKSLPFSVTEGPDGYPLIHANYLGEKRAFTPTQVMGMM 121
Query: 130 LTKMREVAEAYLGSPVKNAVVTVPAYFNDSQRKATIDAGTIAGLNVIRIINEPTAAAIAY 189
L+ ++ +AE L + V + + +P YF D QR+A +DA TIAGL+ +R+I+E TA A+AY
Sbjct: 122 LSNLKGIAEKNLNTAVVDCCIGIPVYFTDLQRRAVLDAATIAGLHPLRLIHETTATALAY 181
Query: 190 GLDKRNDCDGSR-NIFVFDLGGGTFDVSILTIKGDVFEVKATAGNTHLGGEDFDNRMVNY 248
G+ K + + + N+ D+G + V I K ++ + A + LGG DFD + N+
Sbjct: 182 GIYKTDLPESDQLNVAFIDIGHASMQVCIAGFKKGQLKILSHAFDRSLGGRDFDEVLFNH 241
Query: 249 FVEELKKKNKVDISQNPKSLRRLRTACERAKRILSFAFVTTVEVDALFTGIDFCSSITRA 308
F + K + K+D+SQN K+ RLR CE+ K++LS + + ++ L D I R
Sbjct: 242 FAAKFKDEYKIDVSQNAKASLRLRATCEKLKKVLSANPLAPLNIECLMDEKDVRGVIKRE 301
Query: 309 KFEELNMDFFNECMNIVDTCLRDSKIYKNDIDDVVLVGGSSRIPKVQEQLLEYFKGKELF 368
+FEE+++ ++ L D+ + D+ V ++G SR+P + + L E+F GKE
Sbjct: 302 EFEEISIPILERVKRPLEKALSDAGLTVEDVHMVEVIGSGSRVPAMIKILTEFF-GKEPR 360
Query: 369 MGINPDEAVAYGAAVQAAILSEGCKNVPNLVLRDVTPLSLGIS 411
+N E V+ G A+Q AILS K V + + P S+ ++
Sbjct: 361 RTMNASECVSRGCALQCAILSPTFK-VREFQVHESFPFSISLA 402
>At2g32120 70kD heat shock protein
Length = 563
Score = 241 bits (615), Expect = 9e-64
Identities = 175/514 (34%), Positives = 267/514 (51%), Gaps = 34/514 (6%)
Query: 9 AVGIDLGTTYSCVAVWLDQHQRVEIIHNDQGNRITPSFVAFNNEQRLIGDAAKNQSATNP 68
A+GID+GT+ +AVW +V I+ N + ++ SFV F +E G NQ A
Sbjct: 30 ALGIDIGTSQCSIAVW--NGSQVHILRNTRNQKLIKSFVTFKDEVPAGG--VSNQLAHEQ 85
Query: 69 EN----TIFDAKRLIGRKFSDPVVQRDILLWPFKVIS-GVNDKPMITVKYKGQEKHFCAE 123
E IF+ KRL+GR +DPVV L PF V + + +P I + E
Sbjct: 86 EMLTGAAIFNMKRLVGRVDTDPVVHASKNL-PFLVQTLDIGVRPFIAALVNNAWRSTTPE 144
Query: 124 EISSMVLTKMREVAEAYLGSPVKNAVVTVPAYFNDSQRKATIDAGTIAGLNVIRIINEPT 183
E+ ++ L ++R +AEA L PV+N V+TVP F+ Q A +AGL+V+R++ EPT
Sbjct: 145 EVLAIFLVELRLMAEAQLKRPVRNVVLTVPVSFSRFQLTRFERACAMAGLHVLRLMPEPT 204
Query: 184 AAAIAYGLDKR-----NDCDGSRNIFV-FDLGGGTFDVSILTIKGDVFEVKATAGNTHLG 237
A A+ Y ++ N GS + V F++G G DV++ G V ++KA AG+ +G
Sbjct: 205 AIALLYAQQQQMTTHDNMGSGSERLAVIFNMGAGYCDVAVTATAGGVSQIKALAGSP-IG 263
Query: 238 GEDFDNRMVNYFVEELKKKNKVDISQNPKSLRRLRTACERAKRILSFAFVTTVEVDALFT 297
GED + + N ++ LR A + A L+ +EVD L
Sbjct: 264 GEDILQNTIRHIAPP-----------NEEASGLLRVAAQDAIHRLTDQENVQIEVD-LGN 311
Query: 298 GIDFCSSITRAKFEELNMDFFNECMNIVDTCLRDSKIYKNDIDDVVLVGGSSRIPKVQEQ 357
G + R +FEE+N F EC +V CLRD+++ DIDD+++VGG S IPKV+
Sbjct: 312 GNKISKVLDRLEFEEVNQKVFEECERLVVQCLRDARVNGGDIDDLIMVGGCSYIPKVRTI 371
Query: 358 LLEYFKGKELFMGINPDEAVAYGAAVQAAILSEGCKNVPNLVLRDV--TPLSLGISVSIE 415
+ K E++ G+NP EA GAA++ A+ S +L L + TPL++G+ +
Sbjct: 372 IKNVCKKDEIYKGVNPLEAAVRGAALEGAVTSGIHDPFGSLDLLTIQATPLAVGVRANGN 431
Query: 416 HIMSVVIPRNTSIPVKKTSRYSTLYDNQCLVNFLVYEGERPRAADNNLLGSFGLRCL-PG 474
+ VIPRNT +P +K ++T+ DNQ ++YEGE +N+LLG F L + P
Sbjct: 432 KFIP-VIPRNTMVPARKDLFFTTVQDNQKEALIIIYEGEGETVEENHLLGYFKLVGIPPA 490
Query: 475 PRGQP-LEVCFSIDENGILTVTAREISTGNMNMI 507
P+G P + VC ID + L V A + G+ + +
Sbjct: 491 PKGVPEINVCMDIDASNALRVFAAVLMPGSSSPV 524
>At1g11660 heat-shock protein like
Length = 773
Score = 238 bits (608), Expect = 6e-63
Identities = 178/615 (28%), Positives = 313/615 (49%), Gaps = 34/615 (5%)
Query: 10 VGIDLGTTYSCVAVWLDQHQRVEIIHNDQGNRITPSFVAFNNEQRLIGDAAKNQSATNPE 69
VG D+G +AV + + ++++ ND+ NR P+ V+F +QR +G AA + +P+
Sbjct: 4 VGFDVGNENCVIAV--AKQRGIDVLLNDESNRENPAMVSFGEKQRFMGAAAAASATMHPK 61
Query: 70 NTIFDAKRLIGRKFSDPVVQRDILLWPFKVISGVNDKPMITVKYKGQEKHFCAEEISSMV 129
+TI KRLIGRKF +P VQ D+ L+PF+ + I ++Y G+ + F +I M+
Sbjct: 62 STISQLKRLIGRKFREPDVQNDLRLFPFETSEDSDGGIQIRLRYMGEIQSFSPVQILGML 121
Query: 130 LTKMREVAEAYLGSPVKNAVVTVPAYFNDSQRKATIDAGTIAGLNVIRIINEPTAAAIAY 189
L+ ++++AE L +PV + V+ +P+YF +SQR A +DA IAGL +R++++ TA A+ Y
Sbjct: 122 LSHLKQIAEKSLKTPVSDCVIGIPSYFTNSQRLAYLDAAAIAGLRPLRLMHDSTATALGY 181
Query: 190 GLDKRNDCDGSRNIFV--FDLGGGTFDVSILTIKGDVFEVKATAGNTHLGGEDFDNRMVN 247
G+ K + S ++ D+G V + + + V++ A + +LGG DFD + N
Sbjct: 182 GIYKTDLVANSSPTYIVFIDIGHCDTQVCVASFESGSMRVRSHAFDRNLGGRDFDEVLFN 241
Query: 248 YFVEELKKKNKVDISQNPKSLRRLRTACERAKRILSFAFVTTVEVDALFTGIDFCSSITR 307
+F E K+K +D+ N K+ RLR +CE+ K++LS + ++ L D S I R
Sbjct: 242 HFALEFKEKYNIDVYTNTKACVRLRASCEKVKKVLSANAEAQLNIECLMEEKDVRSFIKR 301
Query: 308 AKFEELNMDFFNECMNIVDTCLRDSKIYKNDIDDVVLVGGSSRIPKVQEQLLEYFKGKEL 367
+FE+L+ + L DS + + I V LVG SRIP + + L FK +EL
Sbjct: 302 EEFEQLSAGLLERLIVPCQKALADSGLSLDQIHSVELVGSGSRIPAISKMLSSLFK-REL 360
Query: 368 FMGINPDEAVAYGAAVQAAILSEGCKNVPNLVLRDVTPLSLGISVSIEHIMS----VVIP 423
+N E VA G A+Q A+LS + V + ++D P ++G S I + ++ P
Sbjct: 361 GRTVNASECVARGCALQCAMLSPVFR-VRDYEVQDSYPFAIGFSSDKGPINTPSNELLFP 419
Query: 424 RN---TSIPVKKTSRYSTLYDNQCLVNFLVYEGERPRAADNNLLGSFGLRCLPGPRGQPL 480
+ S+ V R +T N + P + ++G F + R +
Sbjct: 420 KGQIFPSVKVLTLHRENTFQLEAFYANHNELSPDIPTQISSFMIGPFHISHGEAAR---V 476
Query: 481 EVCFSIDENGILTVTAREI-STGNMNMITITNDKERLSMFDIEKMIKEAEKYHVEDMQFL 539
+V ++ +GI+T+ + + S +++ I KE ++ E+MI E E + M
Sbjct: 477 KVRVQLNLHGIVTIDSATLESKLSVSEQLIEYHKENITS---EEMISE-ENHQSSAM--- 529
Query: 540 RRAKVMCELDSCVYNMKKALKKKDVNLI------LSPQEI--AKINNAITVAMDLLDKNK 591
+ + + N KA+K+ ++ ++ L+ E+ AK V DL ++
Sbjct: 530 -KDGSLDPSSGSIGNEPKAIKRMEIPVVANVSGALTKDELSEAKQRENSLVEQDLKMEST 588
Query: 592 KEKEIDVLEGYLEEL 606
K+K+ + LE ++ E+
Sbjct: 589 KDKK-NALESFVYEM 602
>At3g54670 structural maintenance of chromosomes (SMC) - like
protein
Length = 1265
Score = 33.5 bits (75), Expect = 0.36
Identities = 30/101 (29%), Positives = 49/101 (47%), Gaps = 10/101 (9%)
Query: 518 MFDIEKMIKEAEKYHVEDMQFLR----RAKVMCELDSCVYNMKKA-LKKKDVNLILSPQE 572
M ++EK +EA K VE ++L+ R K + E S + + K L+ +E
Sbjct: 263 MRELEKFEREAGKRKVEQAKYLKEIAQREKKIAEKSSKLGKIVSIPWKSVQPELLRFKEE 322
Query: 573 IAKINNAITVAMDLLDKNKKE-----KEIDVLEGYLEELER 608
IA+I I +DK KKE KEI+ ++ ++EL +
Sbjct: 323 IARIKAKIETNRKDVDKRKKEKGKHSKEIEQMQKSIKELNK 363
>At4g27630 unknown protein
Length = 467
Score = 30.8 bits (68), Expect = 2.4
Identities = 23/98 (23%), Positives = 49/98 (49%), Gaps = 9/98 (9%)
Query: 521 IEKMIKEAEKYHVEDMQFLRRAKVMCELDSCVYNMKKALK-KKDVNLILSPQEIAK---- 575
I I+E E+ ++ ++ ++M +++C+ KK L + +V L +E K
Sbjct: 176 ISLFIREIEESEIKSLE----RQLMQSMETCIAKKKKILLCQVEVERSLVSEEHQKGKSF 231
Query: 576 INNAITVAMDLLDKNKKEKEIDVLEGYLEELERMSKHL 613
+ + + ++KE++I ++E +E LE +SK L
Sbjct: 232 FRRFVGTVVRSVQDDQKEQDIKLMEAEVEGLEELSKQL 269
>At3g17360 kinesin-like protein
Length = 2008
Score = 30.4 bits (67), Expect = 3.1
Identities = 39/159 (24%), Positives = 65/159 (40%), Gaps = 31/159 (19%)
Query: 480 LEVCFSIDENGILTVTAREISTGN--MNMITITNDKERLSMFDIEKMIKEAEKYHVEDMQ 537
LE C SID L + ++ + ++ ND+ R + D+ K AE+ E
Sbjct: 1508 LENC-SIDLKTALFTSQSDLEQAKQRIQILAEQNDELRALVSDLCKEKAAAEEGLDEQRD 1566
Query: 538 FLRRAKVMC---------ELDSCVYNMKKALKKKDVNLILSPQEIAKINNAITVAMDLLD 588
+ R + +L S V ++K+ LKK EI +NN + +A + D
Sbjct: 1567 LVNRLEKEILHLTTTAEKQLLSAVKSIKENLKKTSDEKDQIVDEICSLNNKLELAYAIAD 1626
Query: 589 KNK-------------------KEKEIDVLEGYLEELER 608
+ + KE+E+ +LE +EELER
Sbjct: 1627 EKEAIAVEAHQESEASKIYAEQKEEEVKILEISVEELER 1665
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.137 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,627,243
Number of Sequences: 26719
Number of extensions: 593760
Number of successful extensions: 1718
Number of sequences better than 10.0: 31
Number of HSP's better than 10.0 without gapping: 19
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 1602
Number of HSP's gapped (non-prelim): 33
length of query: 624
length of database: 11,318,596
effective HSP length: 105
effective length of query: 519
effective length of database: 8,513,101
effective search space: 4418299419
effective search space used: 4418299419
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 63 (28.9 bits)
Medicago: description of AC144722.2


