

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144721.3 - phase: 0 /pseudo
(750 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
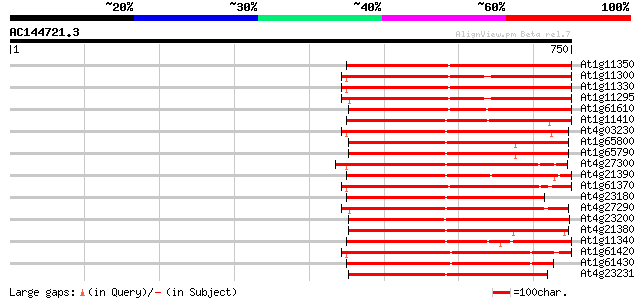
Score E
Sequences producing significant alignments: (bits) Value
At1g11350 putative receptor-like protein kinase gi|4008010; simi... 382 e-106
At1g11300 putative s-locus protein kinase (PPC:1.7.2) 382 e-106
At1g11330 receptor-like protein kinase, putative 382 e-106
At1g11295 putative s-locus protein kinase (PPC:1.7.2) 357 1e-98
At1g61610 348 5e-96
At1g11410 putative brassinosteroid insensitive protein; similar ... 343 2e-94
At4g03230 putative receptor kinase 337 1e-92
At1g65800 receptor kinase, putative 337 1e-92
At1g65790 receptor kinase, putative 336 3e-92
At4g27300 putative receptor protein kinase 335 4e-92
At4g21390 serine/threonine kinase - like protein 335 7e-92
At1g61370 S-like receptor protein kinase; member of gene cluster... 334 1e-91
At4g23180 receptor-like protein kinase 4 (RLK4) 332 4e-91
At4g27290 putative receptor like kinase 326 3e-89
At4g23200 serine /threonine kinase - like protein 325 6e-89
At4g21380 receptor-like serine/threonine protein kinase ARK3 325 6e-89
At1g11340 receptor kinase, putative 325 6e-89
At1g61420 S-like receptor protein kinase; member of gene cluster... 325 7e-89
At1g61430 S-like receptor protein kinase; member of gene cluster... 325 7e-89
At4g23231 unknown protein 323 2e-88
>At1g11350 putative receptor-like protein kinase gi|4008010; similar
to EST gb|AI999419.1
Length = 830
Score = 382 bits (981), Expect = e-106
Identities = 188/300 (62%), Positives = 239/300 (79%), Gaps = 1/300 (0%)
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+Q+G +IAVKRLS+ SGQG+EEF+NEVVVISKLQHRNLVRLLG C+E E+MLVYEFMP
Sbjct: 531 LQEGLDIAVKRLSRTSGQGVEEFVNEVVVISKLQHRNLVRLLGFCIEGEERMLVYEFMPE 590
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
LDA+LFDP++++ LDW+ R NI++GI RG+MYLHRDSRLKIIHRDLKASNILLD+ +
Sbjct: 591 NCLDAYLFDPVKQRLLDWKTRFNIIDGICRGLMYLHRDSRLKIIHRDLKASNILLDENLN 650
Query: 571 PKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVS 630
PKISDFGLARI +G E DE +T RVVGTYGYM PEYAMGGLFSEKSDV+S GV+LLEIVS
Sbjct: 651 PKISDFGLARIFQGNE-DEVSTVRVVGTYGYMAPEYAMGGLFSEKSDVFSLGVILLEIVS 709
Query: 631 GRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQEL 690
GRRN+SFY + + +L +AWKLW I+L+D +++ FE+ + RC+H+GLLCVQ+
Sbjct: 710 GRRNSSFYNDGQNPNLSAYAWKLWNTGEDIALVDPVIFEECFENEIRRCVHVGLLCVQDH 769
Query: 691 PKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSNSNNNVTLSDVIG 750
+RPS++TV+ ML SE ++LP P + AF+ + + ESS QS S NNV+L+ + G
Sbjct: 770 ANDRPSVATVIWMLSSENSNLPEPKQPAFIPRRGTSEVESSGQSDPRASINNVSLTKITG 829
>At1g11300 putative s-locus protein kinase (PPC:1.7.2)
Length = 820
Score = 382 bits (981), Expect = e-106
Identities = 195/311 (62%), Positives = 237/311 (75%), Gaps = 12/311 (3%)
Query: 444 QGRFWP----GIQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERG 499
QG F P +Q+GQEIAVKRLS+ASGQG+EE +NEVVVISKLQHRNLV+LLGCC+
Sbjct: 517 QGGFGPVYKGKLQEGQEIAVKRLSRASGQGLEELVNEVVVISKLQHRNLVKLLGCCIAGE 576
Query: 500 EKMLVYEFMPNKSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLK 559
E+MLVYEFMP KSLD +LFD + K LDW+ R NI+ GI RG++YLHRDSRL+IIHRDLK
Sbjct: 577 ERMLVYEFMPKKSLDYYLFDSRRAKLLDWKTRFNIINGICRGLLYLHRDSRLRIIHRDLK 636
Query: 560 ASNILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVY 619
ASNILLD+ +IPKISDFGLARI G E DEANT+RVVGTYGYM PEYAMGGLFSEKSDV+
Sbjct: 637 ASNILLDENLIPKISDFGLARIFPGNE-DEANTRRVVGTYGYMAPEYAMGGLFSEKSDVF 695
Query: 620 SFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRC 679
S GV+LLEI+SGRRN+ + +L+ + W +W E SL+D E++D FE + +C
Sbjct: 696 SLGVILLEIISGRRNS-------NSTLLAYVWSIWNEGEINSLVDPEIFDLLFEKEIHKC 748
Query: 680 MHIGLLCVQELPKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSNS 739
+HIGLLCVQE +RPS+STV ML SEI +P P + AF+ N ESS+ S +S
Sbjct: 749 IHIGLLCVQEAANDRPSVSTVCSMLSSEIADIPEPKQPAFISRNNVPEAESSENSDLKDS 808
Query: 740 NNNVTLSDVIG 750
NNVT++DV G
Sbjct: 809 INNVTITDVTG 819
>At1g11330 receptor-like protein kinase, putative
Length = 840
Score = 382 bits (980), Expect = e-106
Identities = 192/311 (61%), Positives = 242/311 (77%), Gaps = 5/311 (1%)
Query: 444 QGRFWP----GIQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERG 499
QG F P + +GQEIAVKRLS+ SGQG+EE MNEVVVISKLQHRNLV+LLGCC+E
Sbjct: 530 QGGFGPVYKGKLPEGQEIAVKRLSRKSGQGLEELMNEVVVISKLQHRNLVKLLGCCIEGE 589
Query: 500 EKMLVYEFMPNKSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLK 559
E+MLVYE+MP KSLDA+LFDP+++K LDW+ R NI+EGI RG++YLHRDSRLKIIHRDLK
Sbjct: 590 ERMLVYEYMPKKSLDAYLFDPMKQKILDWKTRFNIMEGICRGLLYLHRDSRLKIIHRDLK 649
Query: 560 ASNILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVY 619
ASNILLD+ + PKISDFGLARI + E DEANT+RVVGTYGYM PEYAM G FSEKSDV+
Sbjct: 650 ASNILLDENLNPKISDFGLARIFRANE-DEANTRRVVGTYGYMSPEYAMEGFFSEKSDVF 708
Query: 620 SFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRC 679
S GV+ LEI+SGRRN+S ++ E++L+L+ +AWKLW + SL D V+D FE + +C
Sbjct: 709 SLGVIFLEIISGRRNSSSHKEENNLNLLAYAWKLWNDGEAASLADPAVFDKCFEKEIEKC 768
Query: 680 MHIGLLCVQELPKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSNS 739
+HIGLLCVQE+ +RP++S V+ ML +E L P + AF+ + + ESS QS + S
Sbjct: 769 VHIGLLCVQEVANDRPNVSNVIWMLTTENMSLADPKQPAFIVRRGASEAESSDQSSQKVS 828
Query: 740 NNNVTLSDVIG 750
N+V+L+ V G
Sbjct: 829 INDVSLTAVTG 839
>At1g11295 putative s-locus protein kinase (PPC:1.7.2)
Length = 820
Score = 357 bits (917), Expect = 1e-98
Identities = 183/311 (58%), Positives = 226/311 (71%), Gaps = 12/311 (3%)
Query: 444 QGRFWPGIQ----DGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERG 499
QG F P + +GQEIAVKRLS+ASGQG+EE + EVVVISKLQHRNLV+L GCC+
Sbjct: 517 QGGFGPVYKGMLLEGQEIAVKRLSQASGQGLEELVTEVVVISKLQHRNLVKLFGCCIAGE 576
Query: 500 EKMLVYEFMPNKSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLK 559
E+MLVYEFMP KSLD ++FDP + K LDW R I+ GI RG++YLHRDSRL+IIHRDLK
Sbjct: 577 ERMLVYEFMPKKSLDFYIFDPREAKLLDWNTRFEIINGICRGLLYLHRDSRLRIIHRDLK 636
Query: 560 ASNILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVY 619
ASNILLD+ +IPKISDFGLARI G E DEANT+RVVGTYGYM PEYAMGGLFSEKSDV+
Sbjct: 637 ASNILLDENLIPKISDFGLARIFPGNE-DEANTRRVVGTYGYMAPEYAMGGLFSEKSDVF 695
Query: 620 SFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRC 679
S GV+LLEI+SGRRN+ +L+ W +W E ++D E++D FE + +C
Sbjct: 696 SLGVILLEIISGRRNS-------HSTLLAHVWSIWNEGEINGMVDPEIFDQLFEKEIRKC 748
Query: 680 MHIGLLCVQELPKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSNS 739
+HI LLCVQ+ +RPS+STV +ML SE+ +P P + AF+ E S+ S
Sbjct: 749 VHIALLCVQDAANDRPSVSTVCMMLSSEVADIPEPKQPAFMPRNVGLEAEFSESIALKAS 808
Query: 740 NNNVTLSDVIG 750
NNVT++DV G
Sbjct: 809 INNVTITDVSG 819
>At1g61610
Length = 842
Score = 348 bits (894), Expect = 5e-96
Identities = 179/298 (60%), Positives = 223/298 (74%), Gaps = 2/298 (0%)
Query: 453 DGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKS 512
+G+EIAVKRLS S QG+EEF NE+++I+KLQHRNLVRLLGCC+E EKML+YE+MPNKS
Sbjct: 546 EGREIAVKRLSGKSKQGLEEFKNEILLIAKLQHRNLVRLLGCCIEDNEKMLLYEYMPNKS 605
Query: 513 LDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPK 572
LD FLFD ++ LDWRKR ++ GIARG++YLHRDSRLKIIHRDLKASNILLD EM PK
Sbjct: 606 LDRFLFDESKQGSLDWRKRWEVIGGIARGLLYLHRDSRLKIIHRDLKASNILLDTEMNPK 665
Query: 573 ISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGR 632
ISDFG+ARI + D ANT RVVGTYGYM PEYAM G+FSEKSDVYSFGVL+LEIVSGR
Sbjct: 666 ISDFGMARIFNYRQ-DHANTIRVVGTYGYMAPEYAMEGIFSEKSDVYSFGVLILEIVSGR 724
Query: 633 RNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQELPK 692
+N SF + D SL+G+AW LW + T +ID V D + +RC+H+G+LC Q+
Sbjct: 725 KNVSF-RGTDHGSLIGYAWHLWSQGKTKEMIDPIVKDTRDVTEAMRCIHVGMLCTQDSVI 783
Query: 693 ERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSNSNNNVTLSDVIG 750
RP++ +V+LML S+ + LPPP + F NS E + H S N+VT + ++G
Sbjct: 784 HRPNMGSVLLMLESQTSQLPPPRQPTFHSFLNSGDIELNFDGHDVASVNDVTFTTIVG 841
>At1g11410 putative brassinosteroid insensitive protein; similar to
EST gb|W43102
Length = 840
Score = 343 bits (880), Expect = 2e-94
Identities = 179/306 (58%), Positives = 228/306 (74%), Gaps = 9/306 (2%)
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+Q+G EIAVKRLSK+SGQG+EEF NEV +ISKLQHRNLVR+LGCCVE EKMLVYE++PN
Sbjct: 537 LQNGMEIAVKRLSKSSGQGMEEFKNEVKLISKLQHRNLVRILGCCVEFEEKMLVYEYLPN 596
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
KSLD F+F Q+ +LDW KR I+ GI RGI+YLH+DSRL+IIHRDLKASN+LLD+EMI
Sbjct: 597 KSLDYFIFHEEQRAELDWPKRMGIIRGIGRGILYLHQDSRLRIIHRDLKASNVLLDNEMI 656
Query: 571 PKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVS 630
PKI+DFGLARI GG E +T RVVGTYGYM PEYAM G FS KSDVYSFGVL+LEI++
Sbjct: 657 PKIADFGLARIF-GGNQIEGSTNRVVGTYGYMSPEYAMDGQFSIKSDVYSFGVLILEIIT 715
Query: 631 GRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASF-ESSMLRCMHIGLLCVQE 689
G+RN++FY E+SL+LV W W I +ID+ + + ++ E +++C+HIGLLCVQE
Sbjct: 716 GKRNSAFY--EESLNLVKHIWDRWENGEAIEIIDKLMGEETYDEGEVMKCLHIGLLCVQE 773
Query: 690 LPKERPSISTVVLMLISEITHLPPPGKVAFV-----HNQNSRSTESSQQSHRSNSNNNVT 744
+RP +S+VV ML LP P AF + + S+++ S++ N+VT
Sbjct: 774 NSSDRPDMSSVVFMLGHNAIDLPSPKHPAFTAGRRRNTKTGGSSDNWPSGETSSTINDVT 833
Query: 745 LSDVIG 750
L+DV G
Sbjct: 834 LTDVQG 839
>At4g03230 putative receptor kinase
Length = 852
Score = 337 bits (865), Expect = 1e-92
Identities = 180/312 (57%), Positives = 231/312 (73%), Gaps = 9/312 (2%)
Query: 444 QGRFWP---GIQDG-QEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERG 499
QG F P G+ G QEIAVKRLS+ SGQG+EEF NEVV+I+KLQHRNLVRLLG CV
Sbjct: 540 QGGFGPVYKGMFPGDQEIAVKRLSRCSGQGLEEFKNEVVLIAKLQHRNLVRLLGYCVAGE 599
Query: 500 EKMLVYEFMPNKSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLK 559
EK+L+YE+MP+KSLD F+FD ++LDW+ R NI+ GIARG++YLH+DSRL+IIHRDLK
Sbjct: 600 EKLLLYEYMPHKSLDFFIFDRKLCQRLDWKMRCNIILGIARGLLYLHQDSRLRIIHRDLK 659
Query: 560 ASNILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVY 619
SNILLD+EM PKISDFGLARI GG ANT RVVGTYGYM PEYA+ GLFS KSDV+
Sbjct: 660 TSNILLDEEMNPKISDFGLARIF-GGSETSANTNRVVGTYGYMSPEYALEGLFSFKSDVF 718
Query: 620 SFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRC 679
SFGV+++E +SG+RN F++ E SLSL+G AW LW E I L+D+ + ++ L+C
Sbjct: 719 SFGVVVIETISGKRNTGFHEPEKSLSLLGHAWDLWKAERGIELLDQALQESCETEGFLKC 778
Query: 680 MHIGLLCVQELPKERPSISTVVLML-ISEITHLPPPGKVAFVHNQ---NSRSTESSQQSH 735
+++GLLCVQE P +RP++S VV ML SE LP P + AFV + +S+++ S++
Sbjct: 779 LNVGLLCVQEDPNDRPTMSNVVFMLGSSEAATLPTPKQPAFVLRRCPSSSKASSSTKPET 838
Query: 736 RSNSNNNVTLSD 747
S + +TL D
Sbjct: 839 CSENELTITLED 850
>At1g65800 receptor kinase, putative
Length = 847
Score = 337 bits (864), Expect = 1e-92
Identities = 173/300 (57%), Positives = 222/300 (73%), Gaps = 7/300 (2%)
Query: 453 DGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKS 512
DG+EIAVKRLSK S QG +EFMNEV +I+KLQH NLVRLLGCCV++GEKML+YE++ N S
Sbjct: 544 DGKEIAVKRLSKMSSQGTDEFMNEVRLIAKLQHINLVRLLGCCVDKGEKMLIYEYLENLS 603
Query: 513 LDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPK 572
LD+ LFD + L+W+KR +I+ GIARG++YLH+DSR +IIHRDLKASN+LLD M PK
Sbjct: 604 LDSHLFDQTRSSNLNWQKRFDIINGIARGLLYLHQDSRCRIIHRDLKASNVLLDKNMTPK 663
Query: 573 ISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGR 632
ISDFG+ARI G E EANT+RVVGTYGYM PEYAM G+FS KSDV+SFGVLLLEI+SG+
Sbjct: 664 ISDFGMARIF-GREETEANTRRVVGTYGYMSPEYAMDGIFSMKSDVFSFGVLLLEIISGK 722
Query: 633 RNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFES----SMLRCMHIGLLCVQ 688
RN FY + L+L+GF W+ W E + ++D DA +LRC+ IGLLCVQ
Sbjct: 723 RNKGFYNSNRDLNLLGFVWRHWKEGKELEIVDPINIDALSSEFPTHEILRCIQIGLLCVQ 782
Query: 689 ELPKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSN--SNNNVTLS 746
E ++RP +S+V++ML SE T +P P + F ++S +SS + R + + N VTLS
Sbjct: 783 ERAEDRPVMSSVMVMLGSETTAIPQPKRPGFCVGRSSLEVDSSSSTQRDDECTVNQVTLS 842
>At1g65790 receptor kinase, putative
Length = 843
Score = 336 bits (861), Expect = 3e-92
Identities = 171/300 (57%), Positives = 222/300 (74%), Gaps = 7/300 (2%)
Query: 453 DGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKS 512
DG+EIAVKRLSK S QG +EFMNEV +I+KLQH NLVRLLGCCV++GEKML+YE++ N S
Sbjct: 540 DGKEIAVKRLSKMSSQGTDEFMNEVRLIAKLQHINLVRLLGCCVDKGEKMLIYEYLENLS 599
Query: 513 LDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPK 572
LD+ LFD + L+W+KR +I+ GIARG++YLH+DSR +IIHRDLKASN+LLD M PK
Sbjct: 600 LDSHLFDQTRSSNLNWQKRFDIINGIARGLLYLHQDSRCRIIHRDLKASNVLLDKNMTPK 659
Query: 573 ISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGR 632
ISDFG+ARI G E EANT+RVVGTYGYM PEYAM G+FS KSDV+SFGVLLLEI+SG+
Sbjct: 660 ISDFGMARIF-GREETEANTRRVVGTYGYMSPEYAMDGIFSMKSDVFSFGVLLLEIISGK 718
Query: 633 RNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFES----SMLRCMHIGLLCVQ 688
RN FY + L+L+GF W+ W E N + ++D D+ +LRC+ IGLLCVQ
Sbjct: 719 RNKGFYNSNRDLNLLGFVWRHWKEGNELEIVDPINIDSLSSKFPTHEILRCIQIGLLCVQ 778
Query: 689 ELPKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSN--SNNNVTLS 746
E ++RP +S+V++ML SE T +P P + F ++ +SS + R + + N +TLS
Sbjct: 779 ERAEDRPVMSSVMVMLGSETTAIPQPKRPGFCIGRSPLEADSSSSTQRDDECTVNQITLS 838
>At4g27300 putative receptor protein kinase
Length = 815
Score = 335 bits (860), Expect = 4e-92
Identities = 172/315 (54%), Positives = 234/315 (73%), Gaps = 8/315 (2%)
Query: 436 FSF**YAWQGRFWP----GIQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRL 491
FS+ + +G F P ++DGQEIAVKRLS SGQG+EEF NEV +I+KLQHRNLVRL
Sbjct: 500 FSYVNFLGRGGFGPVYKGKLEDGQEIAVKRLSANSGQGVEEFKNEVKLIAKLQHRNLVRL 559
Query: 492 LGCCVERGEKMLVYEFMPNKSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRL 551
LGCC++ E ML+YE+MPNKSLD F+FD + +LDW+KR NI+ G+ARGI+YLH+DSRL
Sbjct: 560 LGCCIQGEECMLIYEYMPNKSLDFFIFDERRSTELDWKKRMNIINGVARGILYLHQDSRL 619
Query: 552 KIIHRDLKASNILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGL 611
+IIHRDLKA N+LLD++M PKISDFGLA+ GG+ E++T RVVGTYGYMPPEYA+ G
Sbjct: 620 RIIHRDLKAGNVLLDNDMNPKISDFGLAKSF-GGDQSESSTNRVVGTYGYMPPEYAIDGH 678
Query: 612 FSEKSDVYSFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDRE-VWDA 670
FS KSDV+SFGVL+LEI++G+ N F + L+L+G WK+W+E+ I + + E + +
Sbjct: 679 FSVKSDVFSFGVLVLEIITGKTNRGFRHADHDLNLLGHVWKMWVEDREIEVPEEEWLEET 738
Query: 671 SFESSMLRCMHIGLLCVQELPKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTES 730
S +LRC+H+ LLCVQ+ P++RP++++VVLM S+ + LP P + F N+N S
Sbjct: 739 SVIPEVLRCIHVALLCVQQKPEDRPTMASVVLMFGSD-SSLPHPTQPGFFTNRNVPDI-S 796
Query: 731 SQQSHRSNSNNNVTL 745
S S RS + ++T+
Sbjct: 797 SSLSLRSQNEVSITM 811
>At4g21390 serine/threonine kinase - like protein
Length = 849
Score = 335 bits (858), Expect = 7e-92
Identities = 175/305 (57%), Positives = 226/305 (73%), Gaps = 9/305 (2%)
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
++DG+EIAVKRLS SGQG++EF NE+++I+KLQHRNLVRLLGCC E EKMLVYE+MPN
Sbjct: 548 LEDGREIAVKRLSGKSGQGVDEFKNEIILIAKLQHRNLVRLLGCCFEGEEKMLVYEYMPN 607
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
KSLD FLFD ++ +DW+ R +I+EGIARG++YLHRDSRL+IIHRDLK SN+LLD EM
Sbjct: 608 KSLDFFLFDETKQALIDWKLRFSIIEGIARGLLYLHRDSRLRIIHRDLKVSNVLLDAEMN 667
Query: 571 PKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVS 630
PKISDFG+ARI GG +EANT RVVGTYGYM PEYAM GLFS KSDVYSFGVLLLEIVS
Sbjct: 668 PKISDFGMARIF-GGNQNEANTVRVVGTYGYMSPEYAMEGLFSVKSDVYSFGVLLLEIVS 726
Query: 631 GRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQEL 690
G+RN S +E SL+G+AW L+ + L+D ++ + LRC+H+ +LCVQ+
Sbjct: 727 GKRNTSLRSSEHG-SLIGYAWYLYTHGRSEELVDPKIRVTCSKREALRCIHVAMLCVQDS 785
Query: 691 PKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSR-----STESSQQSHRSNSNNNVTL 745
ERP++++V+LML S+ L P + F + + + +SSQQ S+N +T
Sbjct: 786 AAERPNMASVLLMLESDTATLAAPRQPTFTSTRRNSIDVNFALDSSQQ--YIVSSNEITS 843
Query: 746 SDVIG 750
+ V+G
Sbjct: 844 TVVLG 848
>At1g61370 S-like receptor protein kinase; member of gene cluster;
similar to EST gb|T41816
Length = 814
Score = 334 bits (856), Expect = 1e-91
Identities = 168/311 (54%), Positives = 228/311 (73%), Gaps = 10/311 (3%)
Query: 444 QGRFWP----GIQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERG 499
QG F P +QDG+EIA+KRLS SGQG+EEFMNE+++ISKLQHRNLVRLLGCC+E
Sbjct: 509 QGGFGPVYKGNLQDGKEIAIKRLSSTSGQGLEEFMNEIILISKLQHRNLVRLLGCCIEGE 568
Query: 500 EKMLVYEFMPNKSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLK 559
EK+L+YEFM NKSL+ F+FD +K +LDW KR I++GIA G++YLHRDS L+++HRD+K
Sbjct: 569 EKLLIYEFMANKSLNTFIFDSTKKLELDWPKRFEIIQGIACGLLYLHRDSCLRVVHRDMK 628
Query: 560 ASNILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVY 619
SNILLD+EM PKISDFGLAR+ +G + +ANT+RVVGT GYM PEYA G+FSEKSD+Y
Sbjct: 629 VSNILLDEEMNPKISDFGLARMFQGTQ-HQANTRRVVGTLGYMSPEYAWTGMFSEKSDIY 687
Query: 620 SFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRC 679
+FGVLLLEI++G+R +SF E+ +L+ FAW W E L+D+++ + ES + RC
Sbjct: 688 AFGVLLLEIITGKRISSFTIGEEGKTLLEFAWDSWCESGGSDLLDQDISSSGSESEVARC 747
Query: 680 MHIGLLCVQELPKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSNS 739
+ IGLLC+Q+ +RP+I+ V+ ML + + LP P + F + ES +S S
Sbjct: 748 VQIGLLCIQQQAGDRPNIAQVMSMLTTTM-DLPKPKQPVFA----MQVQESDSESKTMYS 802
Query: 740 NNNVTLSDVIG 750
NN+T + ++G
Sbjct: 803 VNNITQTAIVG 813
>At4g23180 receptor-like protein kinase 4 (RLK4)
Length = 669
Score = 332 bits (852), Expect = 4e-91
Identities = 163/264 (61%), Positives = 207/264 (77%), Gaps = 1/264 (0%)
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ DG E+AVKRLSK+SGQG EF NEVV+++KLQHRNLVRLLG C++ E++LVYE++PN
Sbjct: 367 LSDGTEVAVKRLSKSSGQGEVEFKNEVVLVAKLQHRNLVRLLGFCLDGEERVLVYEYVPN 426
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
KSLD FLFDP +K +LDW +R I+ G+ARGI+YLH+DSRL IIHRDLKASNILLD +M
Sbjct: 427 KSLDYFLFDPAKKGQLDWTRRYKIIGGVARGILYLHQDSRLTIIHRDLKASNILLDADMN 486
Query: 571 PKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVS 630
PKI+DFG+ARI G + E NT R+VGTYGYM PEYAM G +S KSDVYSFGVL+LEI+S
Sbjct: 487 PKIADFGMARIF-GLDQTEENTSRIVGTYGYMSPEYAMHGQYSMKSDVYSFGVLVLEIIS 545
Query: 631 GRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQEL 690
G++N+SFYQ + + LV +AW LW + L+D + + + ++RC+HIGLLCVQE
Sbjct: 546 GKKNSSFYQTDGAHDLVSYAWGLWSNGRPLELVDPAIVENCQRNEVVRCVHIGLLCVQED 605
Query: 691 PKERPSISTVVLMLISEITHLPPP 714
P ERP++ST+VLML S LP P
Sbjct: 606 PAERPTLSTIVLMLTSNTVTLPVP 629
>At4g27290 putative receptor like kinase
Length = 772
Score = 326 bits (836), Expect = 3e-89
Identities = 174/310 (56%), Positives = 228/310 (73%), Gaps = 11/310 (3%)
Query: 444 QGRFWPGIQD----GQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERG 499
QG F P + GQE+AVKRLS+ S QG+EEF NE+ +I+KLQHRNLV++LG CV+
Sbjct: 462 QGGFGPVYKGTLACGQEVAVKRLSRTSRQGVEEFKNEIKLIAKLQHRNLVKILGYCVDEE 521
Query: 500 EKMLVYEFMPNKSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLK 559
E+ML+YE+ PNKSLD+F+FD ++++LDW KR I++GIARG++YLH DSRL+IIHRDLK
Sbjct: 522 ERMLIYEYQPNKSLDSFIFDKERRRELDWPKRVEIIKGIARGMLYLHEDSRLRIIHRDLK 581
Query: 560 ASNILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVY 619
ASN+LLD +M KISDFGLAR + GG+ EANT RVVGTYGYM PEY + G FS KSDV+
Sbjct: 582 ASNVLLDSDMNAKISDFGLARTL-GGDETEANTTRVVGTYGYMSPEYQIDGYFSLKSDVF 640
Query: 620 SFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFE-SSMLR 678
SFGVL+LEIVSGRRN F E L+L+G AW+ +LE+ +ID V ++ + S +LR
Sbjct: 641 SFGVLVLEIVSGRRNRGFRNEEHKLNLLGHAWRQFLEDKAYEIIDEAVNESCTDISEVLR 700
Query: 679 CMHIGLLCVQELPKERPSISTVVLMLISEITHLPP--PGKVAFVHNQNSRSTESSQQSHR 736
+HIGLLCVQ+ PK+RP++S VVLML SE+ L P PG F + +N +++ +
Sbjct: 701 VIHIGLLCVQQDPKDRPNMSVVVLMLSSEMLLLDPRQPG---FFNERNLLFSDTVSINLE 757
Query: 737 SNSNNNVTLS 746
SNN T+S
Sbjct: 758 IPSNNFQTMS 767
>At4g23200 serine /threonine kinase - like protein
Length = 648
Score = 325 bits (833), Expect = 6e-89
Identities = 168/301 (55%), Positives = 221/301 (72%), Gaps = 6/301 (1%)
Query: 453 DGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKS 512
+G E+AVKRLSK S QG +EF NEVV+++KLQHRNLV+LLG C+E EK+LVYEF+PNKS
Sbjct: 346 NGTEVAVKRLSKTSEQGAQEFKNEVVLVAKLQHRNLVKLLGYCLEPEEKILVYEFVPNKS 405
Query: 513 LDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPK 572
LD FLFDP ++ +LDW KR NI+ GI RGI+YLH+DSRL IIHRDLKASNILLD +MIPK
Sbjct: 406 LDYFLFDPTKQGQLDWTKRYNIIGGITRGILYLHQDSRLTIIHRDLKASNILLDADMIPK 465
Query: 573 ISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGR 632
I+DFG+ARI G + ANTKR+ GT+GYMPPEY + G FS KSDVYSFGVL+LEI+ G+
Sbjct: 466 IADFGMARI-SGIDQSVANTKRIAGTFGYMPPEYVIHGQFSMKSDVYSFGVLILEIICGK 524
Query: 633 RNNSFYQNEDSL-SLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQELP 691
+N SFYQ + +LV + W+LW + + L+D + + ++RC+HI LLCVQE P
Sbjct: 525 KNRSFYQADTKAENLVTYVWRLWTNGSPLELVDLTISENCQTEEVIRCIHIALLCVQEDP 584
Query: 692 KERPSISTVVLMLI--SEITHLPPPGKVAFVHNQNSRSTESSQ--QSHRSNSNNNVTLSD 747
K+RP++ST+++ML S I +P P N+ S SSQ S + N+VT+++
Sbjct: 585 KDRPNLSTIMMMLTNSSLILSVPQPPGFFVPQNKERDSFLSSQFTMGCTSQTKNDVTITN 644
Query: 748 V 748
+
Sbjct: 645 L 645
>At4g21380 receptor-like serine/threonine protein kinase ARK3
Length = 850
Score = 325 bits (833), Expect = 6e-89
Identities = 166/300 (55%), Positives = 219/300 (72%), Gaps = 7/300 (2%)
Query: 453 DGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKS 512
DGQE+AVKRLSK S QG +EF NEV +I++LQH NLVRLL CCV+ GEKML+YE++ N S
Sbjct: 547 DGQEMAVKRLSKTSVQGTDEFKNEVKLIARLQHINLVRLLACCVDAGEKMLIYEYLENLS 606
Query: 513 LDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPK 572
LD+ LFD + KL+W+ R +I+ GIARG++YLH+DSR +IIHRDLKASNILLD M PK
Sbjct: 607 LDSHLFDKSRNSKLNWQMRFDIINGIARGLLYLHQDSRFRIIHRDLKASNILLDKYMTPK 666
Query: 573 ISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGR 632
ISDFG+ARI G + EANT++VVGTYGYM PEYAM G+FS KSDV+SFGVLLLEI+S +
Sbjct: 667 ISDFGMARIF-GRDETEANTRKVVGTYGYMSPEYAMDGIFSMKSDVFSFGVLLLEIISSK 725
Query: 633 RNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASF---ESSMLRCMHIGLLCVQE 689
RN FY ++ L+L+G W+ W E + +ID + D+S + +LRC+ IGLLCVQE
Sbjct: 726 RNKGFYNSDRDLNLLGCVWRNWKEGKGLEIIDPIITDSSSTFRQHEILRCIQIGLLCVQE 785
Query: 690 LPKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSNSN---NNVTLS 746
++RP++S V+LML SE T +P P + ++ T+SS R + + N +T+S
Sbjct: 786 RAEDRPTMSLVILMLGSESTTIPQPKAPGYCLERSLLDTDSSSSKQRDDESWTVNQITVS 845
>At1g11340 receptor kinase, putative
Length = 901
Score = 325 bits (833), Expect = 6e-89
Identities = 177/305 (58%), Positives = 221/305 (72%), Gaps = 11/305 (3%)
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+Q+ EIAVKRLS+ SGQG+EEF NEV +ISKLQHRNLVR+LGCCVE EKMLVYE++PN
Sbjct: 602 LQNRMEIAVKRLSRNSGQGMEEFKNEVKLISKLQHRNLVRILGCCVELEEKMLVYEYLPN 661
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
KSLD F+F Q+ +LDW KR IV GIARGI+YLH+DSRL+IIHRDLKASNILLD EMI
Sbjct: 662 KSLDYFIFHEEQRAELDWPKRMEIVRGIARGILYLHQDSRLRIIHRDLKASNILLDSEMI 721
Query: 571 PKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVS 630
PKISDFG+ARI GG E T RVVGT+GYM PEYAM G FS KSDVYSFGVL+LEI++
Sbjct: 722 PKISDFGMARIF-GGNQMEGCTSRVVGTFGYMAPEYAMEGQFSIKSDVYSFGVLMLEIIT 780
Query: 631 GRRNNSFYQNEDSLSLVGFAWKLW----LEENTISLIDREVWDASFESSMLRCMHIGLLC 686
G++N++F+ E+S +LVG W LW E +L+D+E +D E +++C+ IGLLC
Sbjct: 781 GKKNSAFH--EESSNLVGHIWDLWENGEATEIIDNLMDQETYD---EREVMKCIQIGLLC 835
Query: 687 VQELPKERPSISTVVLMLISEITHLPPPGKVAFVH-NQNSRSTESSQQSHRSNSNNNVTL 745
VQE +R +S+VV+ML T+LP P AF + + + S N+VT
Sbjct: 836 VQENASDRVDMSSVVIMLGHNATNLPNPKHPAFTSARRRGGENGACLKGQTGISVNDVTF 895
Query: 746 SDVIG 750
SD+ G
Sbjct: 896 SDIQG 900
>At1g61420 S-like receptor protein kinase; member of gene cluster;
similar to EST gb|T41816
Length = 807
Score = 325 bits (832), Expect = 7e-89
Identities = 168/312 (53%), Positives = 228/312 (72%), Gaps = 12/312 (3%)
Query: 444 QGRFWP----GIQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERG 499
QG F P +QDG+EIAVKRLS +SGQG EEFMNE+V+ISKLQH+NLVR+LGCC+E
Sbjct: 502 QGGFGPVYKGKLQDGKEIAVKRLSSSSGQGKEEFMNEIVLISKLQHKNLVRILGCCIEGE 561
Query: 500 EKMLVYEFMPNKSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLK 559
EK+L+YEFM N SLD FLFD ++ ++DW KR +I++GIARGI YLHRDS LK+IHRDLK
Sbjct: 562 EKLLIYEFMLNNSLDTFLFDSRKRLEIDWPKRLDIIQGIARGIHYLHRDSHLKVIHRDLK 621
Query: 560 ASNILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVY 619
SNILLD++M PKISDFGLAR+ +G E + NT+RVVGT GYM PEYA G+FSEKSD+Y
Sbjct: 622 VSNILLDEKMNPKISDFGLARMYQGTEYQD-NTRRVVGTLGYMAPEYAWTGMFSEKSDIY 680
Query: 620 SFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRC 679
SFGVL+LEI+SG + + F ++ +L+ +AW+ W + I L+D++V D+ + RC
Sbjct: 681 SFGVLMLEIISGEKISRFSYGKEEKTLIAYAWESWCDTGGIDLLDKDVADSCRPLEVERC 740
Query: 680 MHIGLLCVQELPKERPSISTVVLMLISEITHLPPPGKVAF-VHNQNSRSTESSQQSHRSN 738
+ IGLLCVQ P +RP+ + +L +++ + LPPP + F VH ++ +S+ S
Sbjct: 741 VQIGLLCVQHQPADRPN-TLELLSMLTTTSDLPPPEQPTFVVHRRDDKSS-----SEDLI 794
Query: 739 SNNNVTLSDVIG 750
+ N +T S ++G
Sbjct: 795 TVNEMTKSVILG 806
>At1g61430 S-like receptor protein kinase; member of gene cluster;
similar to EST gb|T41816
Length = 806
Score = 325 bits (832), Expect = 7e-89
Identities = 162/278 (58%), Positives = 214/278 (76%), Gaps = 3/278 (1%)
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+QDG+EIAVKRLS +SGQG +EFMNE+V+ISKLQHRNLVR+LGCCVE EK+L+Y F+ N
Sbjct: 511 LQDGREIAVKRLSSSSGQGKQEFMNEIVLISKLQHRNLVRVLGCCVEGTEKLLIYGFLKN 570
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
KSLD F+FD +K +LDW KR I+EGIARG++YLHRDSRL++IHRDLK SNILLD++M
Sbjct: 571 KSLDTFVFDARKKLELDWPKRFEIIEGIARGLLYLHRDSRLRVIHRDLKVSNILLDEKMN 630
Query: 571 PKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVS 630
PKISDFGLAR+ +G + E T+RVVGT GYM PEYA G+FSEKSD+YSFGVLLLEI+S
Sbjct: 631 PKISDFGLARMFQGTQYQE-KTRRVVGTLGYMSPEYAWTGVFSEKSDIYSFGVLLLEIIS 689
Query: 631 GRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQEL 690
G++ +SF E+ +L+ +AW+ W E ++ +D+ + D+S S + RC+ IGLLCVQ
Sbjct: 690 GKKISSFSYGEEGKALLAYAWECWCETREVNFLDQALADSSHPSEVGRCVQIGLLCVQHE 749
Query: 691 PKERPSISTVVLMLISEITHLPPPGKVAF-VHNQNSRS 727
P +RP+ + +L +++ + LP P K F VH + S
Sbjct: 750 PADRPN-TLELLSMLTTTSDLPLPKKPTFVVHTRKDES 786
>At4g23231 unknown protein
Length = 627
Score = 323 bits (829), Expect = 2e-88
Identities = 161/267 (60%), Positives = 205/267 (76%), Gaps = 1/267 (0%)
Query: 453 DGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKS 512
+G E+AVKRLSK+SGQG EF NEVVV++KLQHRNLVRLLG + GE++LVYE+MPNKS
Sbjct: 358 NGTEVAVKRLSKSSGQGDTEFKNEVVVVAKLQHRNLVRLLGFSIGGGERILVYEYMPNKS 417
Query: 513 LDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPK 572
LD FLFDP ++ +LDW +R ++ GIARGI+YLH+DSRL IIHRDLKASNILLD +M PK
Sbjct: 418 LDYFLFDPAKQNQLDWTRRYKVIGGIARGILYLHQDSRLTIIHRDLKASNILLDADMNPK 477
Query: 573 ISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGR 632
++DFGLARI G + + NT R+VGT+GYM PEYA+ G FS KSDVYSFGVL+LEI+SG+
Sbjct: 478 LADFGLARIF-GMDQTQENTSRIVGTFGYMAPEYAIHGQFSVKSDVYSFGVLVLEIISGK 536
Query: 633 RNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQELPK 692
+NNSFY+ + + LV AW+LW + L+D + D +S ++RC+HI LLCVQE P
Sbjct: 537 KNNSFYETDGAHDLVTHAWRLWSNGTALDLVDPIIIDNCQKSEVVRCIHICLLCVQEDPA 596
Query: 693 ERPSISTVVLMLISEITHLPPPGKVAF 719
ERP +ST+ +ML S LP P + F
Sbjct: 597 ERPILSTIFMMLTSNTVTLPVPLQPGF 623
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.358 0.159 0.601
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,862,008
Number of Sequences: 26719
Number of extensions: 568105
Number of successful extensions: 6421
Number of sequences better than 10.0: 996
Number of HSP's better than 10.0 without gapping: 843
Number of HSP's successfully gapped in prelim test: 153
Number of HSP's that attempted gapping in prelim test: 3226
Number of HSP's gapped (non-prelim): 1238
length of query: 750
length of database: 11,318,596
effective HSP length: 107
effective length of query: 643
effective length of database: 8,459,663
effective search space: 5439563309
effective search space used: 5439563309
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 64 (29.3 bits)
Medicago: description of AC144721.3


