

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144721.14 - phase: 0
(234 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
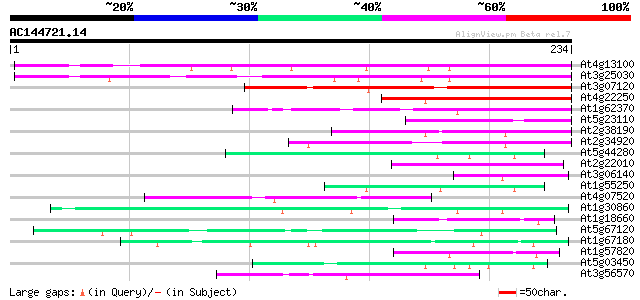
Score E
Sequences producing significant alignments: (bits) Value
At4g13100 putative protein 127 7e-30
At3g25030 unknown protein 125 2e-29
At3g07120 putative RING zinc finger protein 116 1e-26
At4g22250 unknown protein 99 1e-21
At1g62370 hypothetical protein 94 8e-20
At5g23110 putative protein 48 5e-06
At2g38190 unknown protein 46 1e-05
At2g34920 hypothetical protein 46 2e-05
At5g44280 putative protein 45 3e-05
At2g22010 unknown protein 45 3e-05
At3g06140 putative RING zinc finger protein 45 4e-05
At1g55250 unknown protein 44 6e-05
At4g07520 hypothetical protein 44 1e-04
At1g30860 hypothetical protein 43 1e-04
At1g18660 unknown protein 43 1e-04
At5g67120 putative protein 43 2e-04
At1g67180 hypothetical protein 42 2e-04
At1g57820 putative transcription factor protein (At1g57820) 42 3e-04
At5g03450 unknown protein 42 4e-04
At3g56570 putative protein 42 4e-04
>At4g13100 putative protein
Length = 304
Score = 127 bits (318), Expect = 7e-30
Identities = 89/267 (33%), Positives = 135/267 (50%), Gaps = 50/267 (18%)
Query: 3 NRQRITLYDQMTSTSNTRSSLASLIQNDAVLSNQKNTNQTLLDIIRENEPNNVVVNVSVT 62
+R R+TL D+M++ N RSS+ + +L + ++ ++L V
Sbjct: 53 DRLRVTLLDRMSTVENGRSSVTL----EDILMAETSSFRSL-----------TTPTTPVR 97
Query: 63 NSSSSSNNNVVK-----DRKSWKAFKELLLFKRR------------------TSDESTHQ 99
N SSSS +V++ D+ +WK+ ++ L KR T D +H+
Sbjct: 98 NHSSSSLLDVMRRERRRDKTAWKSLRDKLRLKRTATGWISSNPIPTLDNHILTPDNDSHR 157
Query: 100 QQQNEDIPTNSDPSEFL-----PGGEFSDETAQVSMSLMDLLEESEIELERIS--DVVDE 152
+ + TNS+ + E ++ ++ + + E E++ R+S +++++
Sbjct: 158 FNRLGFLLTNSETNRSSRDVSDAAEEAAEREGRLRLGTVLAAEREEMQPPRMSLMELLED 217
Query: 153 GDDVEENNEKREEEEEREEEG-EGEGSNVKV----EHNCCVCMVRDKGAAFIPCGHTFCR 207
D R+E E +G +G G V V E CCVCMVR KGAAFIPCGHTFCR
Sbjct: 218 NDGHMYELSARDEVEVEGRDGCDGGGEAVAVTGAAELGCCVCMVRSKGAAFIPCGHTFCR 277
Query: 208 MCSRELWVSRGNCPLCNHFILEVLDIF 234
+CSRELWV RGNCPLCN IL+VLDIF
Sbjct: 278 LCSRELWVQRGNCPLCNTAILQVLDIF 304
>At3g25030 unknown protein
Length = 250
Score = 125 bits (314), Expect = 2e-29
Identities = 96/263 (36%), Positives = 127/263 (47%), Gaps = 51/263 (19%)
Query: 3 NRQRITLYDQMTSTSNTRSSLASLIQNDAVLSNQKNTN--QTLLDIIRENEPNNVVVNVS 60
+R R+TL D+M++ ++RS L +A+L KN Q L ++ N +V V
Sbjct: 8 DRLRVTLLDRMSTVESSRSCLTL----EAILMADKNRTSPQILSPPPSRSQSNRSLVEVM 63
Query: 61 VTNSSSSSNNNVVKDRKSWKAFKELLLFKRRTSDESTHQQQQNEDIPTNSDPSEFLPGGE 120
S +D+ +WK+ +E L KR S I +NS PS P
Sbjct: 64 QREHRHS------RDKTAWKSLREKLRLKRNGSVW----------ISSNSIPSLDTPIPN 107
Query: 121 FSDETAQVSMSLMD----LLEESEIELE-RISDVVDEGDDVEENNEKREEEEERE----- 170
+ + Q+ L + E S +E R+ V+ E + E+ E E E
Sbjct: 108 RDNVSHQLGFLLSNSGNVTEEASSVEGRIRLGAVLAEERALSAREEETPAEREVEPARMS 167
Query: 171 -----EEGEGEGSNVKV--------------EHNCCVCMVRDKGAAFIPCGHTFCRMCSR 211
EE EG+ S V V E +CCVCMVR KGAAFIPCGHTFCR+CSR
Sbjct: 168 LMELLEENEGQMSFVSVDGEAEEEVAAVTAAEISCCVCMVRSKGAAFIPCGHTFCRLCSR 227
Query: 212 ELWVSRGNCPLCNHFILEVLDIF 234
ELWV RGNCPLCN ILEVLD+F
Sbjct: 228 ELWVQRGNCPLCNTTILEVLDLF 250
>At3g07120 putative RING zinc finger protein
Length = 360
Score = 116 bits (290), Expect = 1e-26
Identities = 57/138 (41%), Positives = 88/138 (63%), Gaps = 9/138 (6%)
Query: 99 QQQQNEDIPTNSDPSEFLPGGEFSDETAQVSMSLMDLLEESEIELERISD--VVDEGDDV 156
+ + ++D +S +E + S+ + +MSLMDLLEE++ ++ +DE ++
Sbjct: 230 EDEDDDDEEDDSGETEEMKSSSASEP--KQTMSLMDLLEETDRQMGLTGSRYAMDEDEEY 287
Query: 157 EENNEKREEEEEREEEGEGEGSNVKVEHNCCVCMVRDKGAAFIPCGHTFCRMCSRELWVS 216
EE+ E EEE + G GEG E +CCVCMV+ KGA+F PCGHTFC++CS+EL
Sbjct: 288 EEDEEDENNEEEGDSHGGGEG-----ELSCCVCMVKIKGASFTPCGHTFCKLCSKELMAQ 342
Query: 217 RGNCPLCNHFILEVLDIF 234
+G+CP+C+ F+LE L+IF
Sbjct: 343 KGHCPVCSSFVLEFLEIF 360
Score = 26.9 bits (58), Expect = 9.3
Identities = 19/55 (34%), Positives = 23/55 (41%), Gaps = 17/55 (30%)
Query: 39 TNQTLLDIIRENEPNNVVVNVSVTNSSSSSNNNVVKDRKSWKAFKELLLFKRRTS 93
T+QTL DIIRE KDR +W+ F+E L KR S
Sbjct: 48 TSQTLFDIIREEYAKEGH-----------------KDRTTWQIFREKLRLKRTGS 85
>At4g22250 unknown protein
Length = 214
Score = 99.4 bits (246), Expect = 1e-21
Identities = 46/81 (56%), Positives = 59/81 (72%), Gaps = 2/81 (2%)
Query: 156 VEENNEKREEEEEREEE--GEGEGSNVKVEHNCCVCMVRDKGAAFIPCGHTFCRMCSREL 213
+EE EK +++ +E E G+ + CCVCM R KGAAFIPCGHTFCR+CSREL
Sbjct: 134 LEETAEKIVDDDGKETEILTASIGTLTGNDSVCCVCMGRKKGAAFIPCGHTFCRVCSREL 193
Query: 214 WVSRGNCPLCNHFILEVLDIF 234
W++RG+CPLCN I+E+LDIF
Sbjct: 194 WLNRGSCPLCNRPIIEILDIF 214
>At1g62370 hypothetical protein
Length = 204
Score = 93.6 bits (231), Expect = 8e-20
Identities = 57/144 (39%), Positives = 79/144 (54%), Gaps = 17/144 (11%)
Query: 94 DESTHQQQQNEDIPTNSDPSEFLPGGEFSDETAQVSMSLMDLLEESEIELERISDVVDEG 153
D+ ++ QN+ + SDP G E T + +L + L E ER++ V +E
Sbjct: 75 DDDDEEESQNQVVDF-SDPGT---GTELDCLTRGSTRNLAEALAE-----ERLAHVTEEE 125
Query: 154 DDVEENNEKREEEEEREEEGEGEGSNVKVEHN---CCVCMVRDKGAAFIPCGHTFCRMCS 210
V + R E +G S N CCVCM R+KGAAFIPCGHT+CR+CS
Sbjct: 126 TAVTKVPLMR-----LLAESDGCDSTTTWLGNDPLCCVCMGREKGAAFIPCGHTYCRVCS 180
Query: 211 RELWVSRGNCPLCNHFILEVLDIF 234
RE+W++RG CPLCN I +VLD++
Sbjct: 181 REIWMNRGTCPLCNRSIFDVLDLY 204
>At5g23110 putative protein
Length = 4706
Score = 47.8 bits (112), Expect = 5e-06
Identities = 22/69 (31%), Positives = 30/69 (42%), Gaps = 4/69 (5%)
Query: 166 EEEREEEGEGEGSNVKVEHNCCVCMVRDKGAAFIPCGHTFCRMCSRELWVSRGNCPLCNH 225
E+ER E E K + C +C ++ +PCGH CR CS S CP C
Sbjct: 4640 EQERAEASMKEAETAKSQWLCQICQTKEVEVTIVPCGHVLCRHCS----TSVSRCPFCRL 4695
Query: 226 FILEVLDIF 234
+ + IF
Sbjct: 4696 QVNRTIRIF 4704
>At2g38190 unknown protein
Length = 124
Score = 46.2 bits (108), Expect = 1e-05
Identities = 26/104 (25%), Positives = 48/104 (46%), Gaps = 5/104 (4%)
Query: 135 LLEESEIELERISDVVDEGDDVEENNEKREEEEEREEE---GEGEGSNVKVEHNCCVCMV 191
L E+ ++D D+G + ++ ++ EE +GE SN + C +C
Sbjct: 20 LTEDDSARTRLLADKDDDGSSMGSCDDSYANDDADLEEFMGNDGEASN-RSRRLCAICFD 78
Query: 192 RDKGAAFIPCGHTF-CRMCSRELWVSRGNCPLCNHFILEVLDIF 234
+ F+PCGH+ C C + + G+CP+C + +V I+
Sbjct: 79 VPRDCFFLPCGHSVSCYECGTTMQEADGSCPICRRKMKKVKRIY 122
>At2g34920 hypothetical protein
Length = 785
Score = 45.8 bits (107), Expect = 2e-05
Identities = 32/120 (26%), Positives = 49/120 (40%), Gaps = 14/120 (11%)
Query: 117 PGGEFSD-ETAQVSMSLMDLLEESEIELERISDVVDEGDDVEENNEKREEEEEREEEGEG 175
P G +S +T S +L ++E+ + D V DV + +K E
Sbjct: 673 PAGSWSSLDTGVTSTPTHNLHSTLQLEMSELRDSVKTCLDVNASLQKSVHLEN------- 725
Query: 176 EGSNVKVEHNCCVCMVRDKGAAFIPCGHTF-CRMCSRELWVSRGNCPLCNHFILEVLDIF 234
+ CCVC CGH C C+ EL + G CP+C+ IL+V+ +F
Sbjct: 726 -----PFKRKCCVCNETQVETLLYRCGHMCTCLRCANELQYNGGKCPICHAKILDVVRVF 780
>At5g44280 putative protein
Length = 486
Score = 45.4 bits (106), Expect = 3e-05
Identities = 32/153 (20%), Positives = 62/153 (39%), Gaps = 20/153 (13%)
Query: 91 RTSDESTHQQQQNEDIPTNSDPSEFLPGGEFSDETAQVSMSLMDLLEESEIELERISDVV 150
R + E+T Q+ ++ + E + E ++ V + + ++ + E + V
Sbjct: 23 RFNPEATQDLQEKDETKEEKEGDEEVKHDEAEEDQEVVKPNDAEEDDDGDDAEEDEEEEV 82
Query: 151 DEGDDVEENNEKREEEEEREEEGEGEG------------------SNVKVEHNCCVCM-V 191
+ +D E E+ EEEEE EEE + + ++ + C +C+ +
Sbjct: 83 EAEEDEEAEEEEEEEEEEEEEEEDSKERSPSSISGDQSEFMEIDLGEIRKDVQCPICLGI 142
Query: 192 RDKGAAFIPCGHTFCRMC-SRELWVSRGNCPLC 223
K + C H FCR C + + + CP C
Sbjct: 143 IKKTRTVMECLHRFCRECIDKSMRLGNNECPAC 175
>At2g22010 unknown protein
Length = 343
Score = 45.4 bits (106), Expect = 3e-05
Identities = 19/72 (26%), Positives = 33/72 (45%)
Query: 160 NEKREEEEEREEEGEGEGSNVKVEHNCCVCMVRDKGAAFIPCGHTFCRMCSRELWVSRGN 219
N +E ER+EE + + ++ CC+C + A PC H C C ++
Sbjct: 254 NRASSQEPERKEESFNKDTTDIEDNTCCICYAGEANAMIAPCSHRSCYGCITRHLLNCQR 313
Query: 220 CPLCNHFILEVL 231
C CN +++V+
Sbjct: 314 CFFCNATVIDVI 325
>At3g06140 putative RING zinc finger protein
Length = 546
Score = 44.7 bits (104), Expect = 4e-05
Identities = 19/49 (38%), Positives = 27/49 (54%), Gaps = 1/49 (2%)
Query: 186 CCVCMVRDKGAAFIPCGHT-FCRMCSRELWVSRGNCPLCNHFILEVLDI 233
C +CM K A +PC H C C++EL + CP+C I E+L+I
Sbjct: 489 CVICMTEAKDTAVLPCRHLCMCSDCAKELRLQSNKCPICRQPIEELLEI 537
>At1g55250 unknown protein
Length = 899
Score = 44.3 bits (103), Expect = 6e-05
Identities = 36/112 (32%), Positives = 44/112 (39%), Gaps = 20/112 (17%)
Query: 132 LMDLLEESEIELERIS----DVVDEGDDVEENNEKREEEEEREEEGEGEGS--------- 178
L + SE E E+IS D+ E DD E + EE E +E E GS
Sbjct: 774 LKSAVSSSEKEYEQISRRTDDIKLELDDEREKKKLEEELMELNKELEELGSESVEAAIVR 833
Query: 179 ------NVKVEHNCCVCMVRDKGAAFIPCGHTFCRMC-SRELWVSRGNCPLC 223
N K C VC R K + C H FC+ C R L + CP C
Sbjct: 834 LQEEVKNCKNILKCGVCFDRPKEVVIVKCYHLFCQQCIQRSLEIRHRKCPGC 885
>At4g07520 hypothetical protein
Length = 734
Score = 43.5 bits (101), Expect = 1e-04
Identities = 37/121 (30%), Positives = 55/121 (44%), Gaps = 7/121 (5%)
Query: 57 VNVSVTNSSSSSNNNVVKDRKSWKAFKELLLFKRRTSDESTHQQQQNEDIPTN-SDPSEF 115
V + +SS S + V R++ +E+ + + DE+ Q+ P N +P E
Sbjct: 4 VRLVTPSSSESEDERVTIIREADMNREEVAAEENKFEDENCEQEP-----PKNLHEPEEE 58
Query: 116 LPGGEFSDETAQVSMSLMDLLEESEIELERISDVVDEGDDVEENNEKREEEEEREEEGEG 175
E DE S + + EE E E E + +EGDDVE + EE +EEE EG
Sbjct: 59 KISEEVDDEEPMQSQGMEENPEEEEKEGEE-EEESEEGDDVEPMQSQGMEENPKEEEKEG 117
Query: 176 E 176
E
Sbjct: 118 E 118
Score = 33.1 bits (74), Expect = 0.13
Identities = 21/47 (44%), Positives = 29/47 (61%), Gaps = 6/47 (12%)
Query: 140 EIELERISDVVD-----EGDDVEENNEKREEEEEREEEGEGEGSNVK 181
E E E+IS+ VD + +EEN E+ E+E E EEE E EG +V+
Sbjct: 54 EPEEEKISEEVDDEEPMQSQGMEENPEEEEKEGEEEEESE-EGDDVE 99
>At1g30860 hypothetical protein
Length = 739
Score = 43.1 bits (100), Expect = 1e-04
Identities = 48/230 (20%), Positives = 93/230 (39%), Gaps = 24/230 (10%)
Query: 18 NTRSSLASLIQNDAVLSNQKNTNQTLLDIIRENEPNNVVVNVSVTNSSSSSNNNVVKDRK 77
NTRS ++D ++ T LD + +N++++ T+S S ++ +
Sbjct: 514 NTRSE-----KDDICRLLERRTVTDFLDSGLREKIDNLMMSRVQTHSDKHSKKWELQQEE 568
Query: 78 SWKAFKELLLFKRRTSDESTHQQQQNEDIPTNSDP--SEFLPGGEFSDETAQVSMSLMDL 135
+ E+ + +Q +D+ +S S P G +S + V+ + +
Sbjct: 569 EEEVNFEIDEEIKEEPLRGGEEQDDRDDLSQSSSSHISASSPAGSWSSQDTDVTSTPALV 628
Query: 136 LEESEI-ELERISDVVDEGDDVEENNEKREEEEEREEEGEGEGSNVKVEHN--------- 185
++ + E+E IS + + +++ E R+ +N ++H
Sbjct: 629 VQNPQSPEMELISGMRSQIQQLQQ-----EMSVLRDSVKTCLDANASLQHKAHQENPMKR 683
Query: 186 -CCVCMVRDKGAAFIPCGHT-FCRMCSRELWVSRGNCPLCNHFILEVLDI 233
CCVC A CGH C C+ EL S G CP+C I++V+ +
Sbjct: 684 KCCVCDETQVEAVLYRCGHMCMCLKCANELHWSGGKCPICRAQIVDVVRV 733
>At1g18660 unknown protein
Length = 486
Score = 43.1 bits (100), Expect = 1e-04
Identities = 27/68 (39%), Positives = 34/68 (49%), Gaps = 5/68 (7%)
Query: 161 EKREEEEEREEEGEGEGSNVKVEHNCCVCMVRDKGAAFIPCGHTFCRMCSRELWVSRGN- 219
EK R+ G E S+ + +C VC+ A PCGHTFCR C + + RGN
Sbjct: 174 EKVMPNSMRKTHGMAERSD---DFDCTVCLKLLYEPATTPCGHTFCRSCLFQS-MDRGNK 229
Query: 220 CPLCNHFI 227
CPLC I
Sbjct: 230 CPLCRTVI 237
>At5g67120 putative protein
Length = 272
Score = 42.7 bits (99), Expect = 2e-04
Identities = 49/230 (21%), Positives = 92/230 (39%), Gaps = 42/230 (18%)
Query: 11 DQMTSTSNTRSSLASLIQNDAVLSNQK---NTNQTLLDIIRE------NEPNNVVVNVSV 61
D + S ++ +S S+++ V S+ N +TL R NE +
Sbjct: 73 DSLQSWNDEQSETTSVVEYTDVSSHGNTFTNEEETLERYWRNWLQSSTNEQSETGSQEEY 132
Query: 62 TNSSSSSNNNVVKDRKSWKAFKELLLFKRRTSDESTHQQQQNEDIPTNSDPSEFLPGGEF 121
TN+SS + ++ + L + R ST++Q + E + ++PS G F
Sbjct: 133 TNASSHGGTFIYEE-------ETLEQYWRNWLQSSTNEQFETESLEEYTNPSSH--GDIF 183
Query: 122 SDETAQVSMSLMDLLEESEIELERISDVVDEGDDVEENNEKREEEEEREEEGEGEGSNVK 181
+ E L+ + +E+ E +S+ V + + + EKR ++E +
Sbjct: 184 TYE------ELLSITDETGDERTGLSEEVIDENLIRRKYEKRSDDETKR----------- 226
Query: 182 VEHNCCVCMVRDKG---AAFIPCGHTFCRMCSRELWVSRGNCPLCNHFIL 228
C +C + K + + CGH F C + + CPLCN ++
Sbjct: 227 ----CVICQQKLKDNEEVSKLGCGHDFHFGCIKNWLMVTNKCPLCNREVV 272
>At1g67180 hypothetical protein
Length = 453
Score = 42.4 bits (98), Expect = 2e-04
Identities = 46/209 (22%), Positives = 77/209 (36%), Gaps = 28/209 (13%)
Query: 47 IRENEPNNVVVNVS---VTNSSSSSNNNVVKDRKSWKAFKELLLFKRRTSDESTHQ---- 99
+R P++++ N V SS VVK R +++ L+ + ++ H
Sbjct: 152 LRTKRPSSILENKENSGVAESSRKGKKRVVKQR----SYRNLIDLESDEESDNNHHDNSD 207
Query: 100 QQQNEDIPTNSDPSEFLPGGEFSD-ETA------QVSMSLMDLLEESEIELERISDVVDE 152
+ QNE E + G F ET+ ++ D+ E E E S V
Sbjct: 208 ENQNETQDHREPADENVRGFVFEQGETSALRHPGDLATPNWDVDEIEESENWSHSAVFKR 267
Query: 153 GDDVEENNEKREEEEEREEEGEGEGSNVKVEHNCCVCMVR-DKGAAFIPCGHTFCRMCSR 211
+ ++++E EE E + +C +C +PCGH FC C +
Sbjct: 268 PRSFSPEIKPQDDDESTREETEAT-EKAPAQVSCIICWTEFSSSRGILPCGHRFCYSCIQ 326
Query: 212 ELWVSR-------GNCPLCNHFILEVLDI 233
+ W R CPLC + + I
Sbjct: 327 K-WADRLVSERKKTTCPLCKSNFITITKI 354
>At1g57820 putative transcription factor protein (At1g57820)
Length = 645
Score = 42.0 bits (97), Expect = 3e-04
Identities = 25/77 (32%), Positives = 37/77 (47%), Gaps = 9/77 (11%)
Query: 161 EKREEEEEREEEGEGEGSNVKV------EHNCCVCMVRDKGAAFIPCGHTFCRMCSRELW 214
E+ +EEE+R+++G+G+ N+ V C CM + PCGH C C E W
Sbjct: 115 EEDDEEEKRKKKGKGKNPNLDVLSALGDNLMCSFCMQLPERPVTKPCGHNACLKCF-EKW 173
Query: 215 VSRG--NCPLCNHFILE 229
+ +G C C I E
Sbjct: 174 MGQGKRTCGKCRSIIPE 190
>At5g03450 unknown protein
Length = 630
Score = 41.6 bits (96), Expect = 4e-04
Identities = 36/141 (25%), Positives = 55/141 (38%), Gaps = 23/141 (16%)
Query: 102 QNEDIPTNSDPSEFLPGGEFSDETAQVSMSLMDLLEESEIELERISDVVDEGDDVEENNE 161
+ ED + ++ GE +E + E+SE + DV+D V +N
Sbjct: 31 EEEDEEEEDEEEAYVDYGEVQEEDEEEEEE-----EDSERQTREYIDVLDSPVRVSQNES 85
Query: 162 KREEEEEREEE--GEGEGSNVKVEH--------NCCVCM-VRDKGAAF----IPCGHTFC 206
+ E + R + GS VE +C +CM V G +PCGH +
Sbjct: 86 EGIENQNRSQGVCSSSSGSQEDVEWKQGDTEGLSCSICMEVWTSGGQHQVCCLPCGHLYG 145
Query: 207 RMCSRELWVSR---GNCPLCN 224
C + + R G CPLCN
Sbjct: 146 YSCINKWFQQRRSGGKCPLCN 166
>At3g56570 putative protein
Length = 531
Score = 41.6 bits (96), Expect = 4e-04
Identities = 33/112 (29%), Positives = 49/112 (43%), Gaps = 5/112 (4%)
Query: 87 LFKRRTSDESTHQQQQNEDIPTNSDPSEFLPGGEFSDETAQVSMSLMDLLEES--EIELE 144
LF +T E H +++ SD + E +DE + S + E+S E+ E
Sbjct: 194 LFNHKTGAEDVHFTHESDSEADESDNDD--AANETTDED-EPSSKISSSPEQSFEEVPGE 250
Query: 145 RISDVVDEGDDVEENNEKREEEEEREEEGEGEGSNVKVEHNCCVCMVRDKGA 196
D E ++ EE E+ EEEEE EEE E + + MV+D A
Sbjct: 251 NTDDEAKEEEEEEEEEEEGEEEEEGEEEEENSSMLQNDQSGLKMIMVKDVSA 302
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.311 0.128 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,614,712
Number of Sequences: 26719
Number of extensions: 267800
Number of successful extensions: 5335
Number of sequences better than 10.0: 629
Number of HSP's better than 10.0 without gapping: 308
Number of HSP's successfully gapped in prelim test: 330
Number of HSP's that attempted gapping in prelim test: 2862
Number of HSP's gapped (non-prelim): 1526
length of query: 234
length of database: 11,318,596
effective HSP length: 96
effective length of query: 138
effective length of database: 8,753,572
effective search space: 1207992936
effective search space used: 1207992936
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC144721.14


