

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
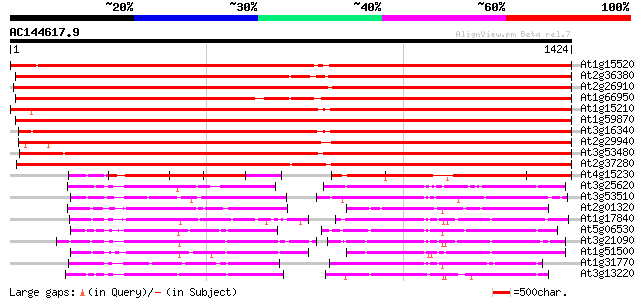
Query= AC144617.9 + phase: 0
(1424 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At1g15520 hypothetical protein 1872 0.0
At2g36380 putative ABC transporter 1669 0.0
At2g26910 putative ABC transporter 1651 0.0
At1g66950 ABC transporter, putative 1641 0.0
At1g15210 putative ABC transporter 1613 0.0
At1g59870 putative ABC transporter (At1g59870) 1608 0.0
At3g16340 putative ABC transporter 1599 0.0
At2g29940 putative ABC transporter 1488 0.0
At3g53480 ABC transporter - like protein 1423 0.0
At2g37280 putative ABC transporter 1414 0.0
At4g15230 ABC transporter like protein 571 e-162
At3g25620 membrane transporter, putative 201 2e-51
At3g53510 ABC transporter -like protein 200 4e-51
At2g01320 putative ABC transporter 199 7e-51
At1g17840 putative ABC transporter (At1g17840) 193 7e-49
At5g06530 ABC transporter like protein 191 3e-48
At3g21090 ABC transporter, putative 188 2e-47
At1g51500 ATP-dependent transmembrane transporter, putative 187 5e-47
At1g31770 unknown protein 187 5e-47
At3g13220 ABC transporter 177 3e-44
>At1g15520 hypothetical protein
Length = 1423
Score = 1872 bits (4850), Expect = 0.0
Identities = 912/1429 (63%), Positives = 1135/1429 (78%), Gaps = 26/1429 (1%)
Query: 3 RSDTKTWKNHC-MDVFSKSERE-DDEEALKCVAIKRILTSSCIRKNVESKGEGKGK---- 56
R ++ WK ++FS+S RE DDEEAL+ A++++ T +RK + + G
Sbjct: 14 RRNSSVWKKDSGREIFSRSSREEDDEEALRWAALEKLPTFDRLRKGILTASHAGGPINEI 73
Query: 57 DVETIQLESTEKRALLARLVKIAEEDNEKFLLKLKERMDRVGLELPTIEVRFEDINVEAQ 116
D++ + + T+K LL RL+K+ ++++EK L KLK+R+DRVG++LPTIEVRF+ + VEA+
Sbjct: 74 DIQKLGFQDTKK--LLERLIKVGDDEHEKLLWKLKKRIDRVGIDLPTIEVRFDHLKVEAE 131
Query: 117 VYVGRRALPTLFNFFVNVIEGCLNNLQIIPSPKKQLHILQNVSGILKPRRMTLLLGPPGS 176
V+VG RALPT NF N + LN L ++P+ KK+ IL +VSGI+KP RM LLLGPP S
Sbjct: 132 VHVGGRALPTFVNFISNFADKFLNTLHLVPNRKKKFTILNDVSGIVKPGRMALLLGPPSS 191
Query: 177 GKTTLLLALAGILGKDLKQSGRVTYNGKGLEEFVPQRTSAYVSQYDNHIGEMTVRETLAF 236
GKTTLLLALAG L ++LKQ+GRVTYNG G+ EFVPQRT+AY+ Q D HIGEMTVRET A+
Sbjct: 192 GKTTLLLALAGKLDQELKQTGRVTYNGHGMNEFVPQRTAAYIGQNDVHIGEMTVRETFAY 251
Query: 237 SARCQGVGQNYEMLTELLRKEKESKIEPDPDINAYMKEAAIEGHQNSVVIDYILKILGLD 296
+AR QGVG Y+MLTEL R+EKE+ I+PDPDI+ +MK + G + +V+ DYILKILGL+
Sbjct: 252 AARFQGVGSRYDMLTELARREKEANIKPDPDIDIFMKAMSTAGEKTNVMTDYILKILGLE 311
Query: 297 VCADTMVGDQMIRGISGGEKKRLTTGEMLVGPIKVLFMDEISNGLDSSTTFQIINSIKQS 356
VCADTMVGD M+RGISGG+KKR+TTGEMLVGP + LFMDEIS GLDSSTT+QI+NS++
Sbjct: 312 VCADTMVGDDMLRGISGGQKKRVTTGEMLVGPSRALFMDEISTGLDSSTTYQIVNSLRNY 371
Query: 357 IHILNGTALVSLLQPAPETYELFDDIILLTDGQIVYQGPREYVLEFFESTGFKCPERKGV 416
+HI NGTAL+SLLQPAPET+ LFDDIIL+ +G+I+Y+GPR++V+EFFE+ GFKCP RKGV
Sbjct: 372 VHIFNGTALISLLQPAPETFNLFDDIILIAEGEIIYEGPRDHVVEFFETMGFKCPPRKGV 431
Query: 417 ADFLQEVTSRKDQWQYWAREDEPYNFVTVKDFARAFELFHIGKQLGEELADPFDKSKFHS 476
ADFLQEVTS+KDQ QYWAR DEPY F+ V++FA AF+ FH+G+++G+ELA PFDK+K H
Sbjct: 432 ADFLQEVTSKKDQMQYWARRDEPYRFIRVREFAEAFQSFHVGRRIGDELALPFDKTKSHP 491
Query: 477 NVLITKKYGINKKELLRACASRELLLMKRNSFVYIFKATQLTYLATLTTTLFLRTKMYHS 536
L TKKYG+ KEL++ SRE LLMKRNSFVY FK QL +A LT TLF RT+M
Sbjct: 492 AALTTKKYGVGIKELVKTSFSREYLLMKRNSFVYYFKFGQLLVMAFLTMTLFFRTEMQKK 551
Query: 537 TIEDAQTYMGALFFTVTVAMFNGISELNMTIMKLPIFYKQRDLLFYPSWAYSLPPWILKI 596
T D Y GALFF + + MFNG+SEL+MTI KLP+FYKQRDLLFYP+W YSLPPW+LKI
Sbjct: 552 TEVDGSLYTGALFFILMMLMFNGMSELSMTIAKLPVFYKQRDLLFYPAWVYSLPPWLLKI 611
Query: 597 PITIIEVAIWECISYYAIGFDPNIGRFFKQSLVVLCINQMASALFRFMAALGRDIVVANT 656
PI+ +E A+ I+YY IGFDPN+GR FKQ ++++ +NQMASALF+ +AALGR+++VANT
Sbjct: 612 PISFMEAALTTFITYYVIGFDPNVGRLFKQYILLVLMNQMASALFKMVAALGRNMIVANT 671
Query: 657 FGTFSLLAVTVLGGFVISREDVHKWFLWGYWSSPLMYGQNAIAVNEFLGHGWRKVAPNSN 716
FG F++L LGG V+SR+D+ KW++WGYW SP+MYGQNAI NEF GH W + NS+
Sbjct: 672 FGAFAMLVFFALGGVVLSRDDIKKWWIWGYWISPIMYGQNAILANEFFGHSWSRAVENSS 731
Query: 717 ETLGVSILKSRGFFPQAYWYWIGVGALIGYVFLFNFLFALALHFLSPFRKDQAGLSQEKL 776
ETLGV+ LKSRGF P AYWYWIG GAL+G+V LFNF F LAL FL+ K QA +++E
Sbjct: 732 ETLGVTFLKSRGFLPHAYWYWIGTGALLGFVVLFNFGFTLALTFLNSLGKPQAVIAEE-- 789
Query: 777 QERNASTDEEFIQSQQQENSSNTKMDEEVSENKASSSGRKGMVLPFQPLSLTFDDITYSV 836
++DE +QS + E +A ++ ++GMVLPF+P S+TFD++ YSV
Sbjct: 790 ----PASDETELQSARSEGVV-----------EAGANKKRGMVLPFEPHSITFDNVVYSV 834
Query: 837 DMPQGMKNQGVTEDRLELLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGIKTSGYIEG 896
DMPQ M QG EDRL LLKGV+GAFRPGVLTALMGVSGAGKTTLMDVLAG KT GYI+G
Sbjct: 835 DMPQEMIEQGTQEDRLVLLKGVNGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDG 894
Query: 897 NIKVSGYQKNQKSFARISGYCEQFDIHSPNVTVYESLLYSAWLRLSPEVDHATRKMFIEE 956
NI +SGY KNQ++FARISGYCEQ DIHSP+VTVYESL+YSAWLRL EVD RK+FIEE
Sbjct: 895 NITISGYPKNQQTFARISGYCEQTDIHSPHVTVYESLVYSAWLRLPKEVDKNKRKIFIEE 954
Query: 957 VMELVELNSLREALVGLPGENGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAI 1016
VMELVEL LR+ALVGLPGE+GLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAI
Sbjct: 955 VMELVELTPLRQALVGLPGESGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAI 1014
Query: 1017 VMRTVRNTVDTGRTVVCTIHQPSIDIFDSFDELLLLKLGGEQIYAGPIGNQCSDLIQYFE 1076
VMRTVRNTVDTGRTVVCTIHQPSIDIF++FDEL LLK GGE+IY GP+G++ + LI YFE
Sbjct: 1015 VMRTVRNTVDTGRTVVCTIHQPSIDIFEAFDELFLLKRGGEEIYVGPLGHESTHLINYFE 1074
Query: 1077 AIQGVPTIKDGYNPATWMLEITSAGKEANLKVNFTDVYKNSELHRRNKQLIQELSVPSQS 1136
+IQG+ I +GYNPATWMLE+++ +EA L V+F VYKNSEL++RNK+LI+ELS P+
Sbjct: 1075 SIQGINKITEGYNPATWMLEVSTTSQEAALGVDFAQVYKNSELYKRNKELIKELSQPAPG 1134
Query: 1137 SKDLHFDAQYSQTFLAQCTYCLWKQHLSYWRNTSYTAVRLLFTIMTGILFGLIFWGVGAK 1196
SKDL+F QYSQ+FL QC LWKQH SYWRN YTAVR LFTI ++FG +FW +G K
Sbjct: 1135 SKDLYFPTQYSQSFLTQCMASLWKQHWSYWRNPPYTAVRFLFTIGIALMFGTMFWDLGGK 1194
Query: 1197 SKKEQDLFNAMGSMYAAVTFIGVVNGASVQPIVAIERTVFYRERAAGMYSAMPYALAQVI 1256
+K QDL NAMGSMY AV F+G+ N ASVQP+V +ERTVFYRE+AAGMYSAMPYA AQV
Sbjct: 1195 TKTRQDLSNAMGSMYTAVLFLGLQNAASVQPVVNVERTVFYREQAAGMYSAMPYAFAQVF 1254
Query: 1257 IELPHILVQAVVYGIIVYAMMGFEWTASKVLWNLFFTYFSFLYYTYYGMMTMAITPNPHV 1316
IE+P++LVQA+VYG+IVYAM+GFEWTA K W LFF Y SFL +T+YGMM +A+TPN H+
Sbjct: 1255 IEIPYVLVQAIVYGLIVYAMIGFEWTAVKFFWYLFFMYGSFLTFTFYGMMAVAMTPNHHI 1314
Query: 1317 AGILSTSFYAIWCLFSGFIIPLSRIPIWWKWYYWICPVAWTLNGLVTSQYGHNMDTL-DN 1375
A ++S++FY IW LFSGF+IP +P+WW+WYYW+CPVAWTL GL+ SQ+G + + D+
Sbjct: 1315 ASVVSSAFYGIWNLFSGFLIPRPSMPVWWEWYYWLCPVAWTLYGLIASQFGDITEPMADS 1374
Query: 1376 GQSVEEFVRNYFGFEYDFLGVVAIVVVSFSVLFALIFTFGIKAFNFQKR 1424
SV++F+R ++G+ FLGVVA + V F +LFA+IF GIK+FNFQKR
Sbjct: 1375 NMSVKQFIREFYGYREGFLGVVAAMNVIFPLLFAVIFAIGIKSFNFQKR 1423
>At2g36380 putative ABC transporter
Length = 1450
Score = 1669 bits (4322), Expect = 0.0
Identities = 816/1424 (57%), Positives = 1065/1424 (74%), Gaps = 31/1424 (2%)
Query: 15 DVFSKSER-EDDEEALKCVAIKRILTSSCIRKNVESKGEGKGK----DVETIQLESTEKR 69
DVF +S+R E+D+ L+ A++R+ T +RK + + GK DV+ L EK+
Sbjct: 44 DVFGRSDRREEDDVELRWAALERLPTYDRLRKGMLPQTMVNGKIGLEDVDVTNLAPKEKK 103
Query: 70 ALLARLVKIAEEDNEKFLLKLKERMDRVGLELPTIEVRFEDINVEAQVYVGRRALPTLFN 129
L+ ++K EEDNEKFL +L+ER DRVG+E+P IEVR+E+++VE V RALPTLFN
Sbjct: 104 HLMEMILKFVEEDNEKFLRRLRERTDRVGIEVPKIEVRYENLSVEGDVRSASRALPTLFN 163
Query: 130 FFVNVIEGCLNNLQIIPSPKKQLHILQNVSGILKPRRMTLLLGPPGSGKTTLLLALAGIL 189
+N IE L ++PS K+++ IL+++SGI+KP RMTLLLGPP SGKTTLL ALAG L
Sbjct: 164 VTLNTIESILGLFHLLPSKKRKIEILKDISGIIKPSRMTLLLGPPSSGKTTLLQALAGKL 223
Query: 190 GKDLKQSGRVTYNGKGLEEFVPQRTSAYVSQYDNHIGEMTVRETLAFSARCQGVGQNYEM 249
L+ SGR+TY G EFVPQ+T AY+SQ+D H GEMTVRE+L FS RC GVG Y++
Sbjct: 224 DDTLQMSGRITYCGHEFREFVPQKTCAYISQHDLHFGEMTVRESLDFSGRCLGVGTRYQL 283
Query: 250 LTELLRKEKESKIEPDPDINAYMKEAAIEGHQNSVVIDYILKILGLDVCADTMVGDQMIR 309
LTEL R+E+E+ I+PDP+I+A+MK AI G + S+V DY+LK+LGLD+CADT+VGD M R
Sbjct: 284 LTELSRREREAGIKPDPEIDAFMKSIAISGQETSLVTDYVLKLLGLDICADTLVGDVMRR 343
Query: 310 GISGGEKKRLTTGEMLVGPIKVLFMDEISNGLDSSTTFQIINSIKQSIHILNGTALVSLL 369
GISGG++KRLTTGEMLVGP LFMDEIS GLDSSTTFQI ++Q +HI + T ++SLL
Sbjct: 344 GISGGQRKRLTTGEMLVGPATALFMDEISTGLDSSTTFQICKFMRQLVHIADVTMVISLL 403
Query: 370 QPAPETYELFDDIILLTDGQIVYQGPREYVLEFFESTGFKCPERKGVADFLQEVTSRKDQ 429
QPAPET+ELFDDIILL++GQIVYQG R+ VLEFFE GFKCPERKG+ADFLQEVTS+KDQ
Sbjct: 404 QPAPETFELFDDIILLSEGQIVYQGSRDNVLEFFEYMGFKCPERKGIADFLQEVTSKKDQ 463
Query: 430 WQYWAREDEPYNFVTVKDFARAFELFHIGKQLGEELADPFDKSKFHSNVLITKKYGINKK 489
QYW R + PY++V+V DF+ F FH G+QL E P+DK+K H L+T+KYGI+ K
Sbjct: 464 EQYWNRREHPYSYVSVHDFSSGFNSFHAGQQLASEFRVPYDKAKTHPAALVTQKYGISNK 523
Query: 490 ELLRACASRELLLMKRNSFVYIFKATQLTYLATLTTTLFLRTKMYHSTIEDAQTYMGALF 549
+L +AC RE LLMKRNSFVY+FK Q+T ++ + T++ RT+M+ T++D Q + GALF
Sbjct: 524 DLFKACFDREWLLMKRNSFVYVFKTVQITIMSLIAMTVYFRTEMHVGTVQDGQKFYGALF 583
Query: 550 FTVTVAMFNGISELNMTIMKLPIFYKQRDLLFYPSWAYSLPPWILKIPITIIEVAIWECI 609
F++ MFNG++EL T+M+LP+F+KQRD LFYP WA++LP ++LKIP+++IE IW +
Sbjct: 584 FSLINLMFNGMAELAFTVMRLPVFFKQRDFLFYPPWAFALPGFLLKIPLSLIESVIWIAL 643
Query: 610 SYYAIGFDPNIGRFFKQSLVVLCINQMASALFRFMAALGRDIVVANTFGTFSLLAVTVLG 669
+YY IGF P+ RFF+Q L C+NQMA +LFRF+ ALGR V+AN+ GT +LL V VLG
Sbjct: 644 TYYTIGFAPSAARFFRQLLAYFCVNQMALSLFRFLGALGRTEVIANSGGTLALLVVFVLG 703
Query: 670 GFVISREDVHKWFLWGYWSSPLMYGQNAIAVNEFLGHGWRKVAPNSN-----ETLGVSIL 724
GF+IS++D+ W W Y++SP+MYGQ A+ +NEFL W +PN++ +T+G +L
Sbjct: 704 GFIISKDDIPSWLTWCYYTSPMMYGQTALVINEFLDERWG--SPNNDTRINAKTVGEVLL 761
Query: 725 KSRGFFPQAYWYWIGVGALIGYVFLFNFLFALALHFLSPFRKDQAGLSQEKLQERNASTD 784
KSRGFF + YW+WI +GAL+G+ LFNF + +AL +L+P L A+T
Sbjct: 762 KSRGFFTEPYWFWICIGALLGFTVLFNFCYIIALMYLNP------------LGNSKATTV 809
Query: 785 EEFIQSQQQENSSNTKMDEEVSENKASSSGRKGMVLPFQPLSLTFDDITYSVDMPQGMKN 844
E + + + + S T ++ + +S +KGMVLPFQPLSL F+++ Y VDMP MK
Sbjct: 810 VEEGKDKHKGSHSGTGVE---LTSTSSHGPKKGMVLPFQPLSLAFNNVNYYVDMPAEMKA 866
Query: 845 QGVTEDRLELLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGIKTSGYIEGNIKVSGYQ 904
QGV DRL+LL+ V GAFRPGVLTAL+GVSGAGKTTLMDVLAG KT GY+EG+I +SGY
Sbjct: 867 QGVEGDRLQLLRDVGGAFRPGVLTALVGVSGAGKTTLMDVLAGRKTGGYVEGSINISGYP 926
Query: 905 KNQKSFARISGYCEQFDIHSPNVTVYESLLYSAWLRLSPEVDHATRKMFIEEVMELVELN 964
KNQ +FAR+SGYCEQ DIHSP+VTVYESL+YSAWLRLS ++D TR+MF+EEVMELVEL
Sbjct: 927 KNQATFARVSGYCEQNDIHSPHVTVYESLIYSAWLRLSADIDTKTREMFVEEVMELVELK 986
Query: 965 SLREALVGLPGENGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAIVMRTVRNT 1024
LR ++VGLPG +GLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAIVMRTVRNT
Sbjct: 987 PLRNSIVGLPGVDGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAIVMRTVRNT 1046
Query: 1025 VDTGRTVVCTIHQPSIDIFDSFDELLLLKLGGEQIYAGPIGNQCSDLIQYFEAIQGVPTI 1084
VDTGRTVVCTIHQPSIDIF+SFDELLL+K GG+ IYAG +G+ L++YFEAI+GVP I
Sbjct: 1047 VDTGRTVVCTIHQPSIDIFESFDELLLMKRGGQVIYAGTLGHHSQKLVEYFEAIEGVPKI 1106
Query: 1085 KDGYNPATWMLEITSAGKEANLKVNFTDVYKNSELHRRNKQLIQELSVPSQSSKDLHFDA 1144
KDGYNPATWML++T+ E+ + V+F ++ NS ++RRN++LI+ELS P S DL+F
Sbjct: 1107 KDGYNPATWMLDVTTPSMESQMSVDFAQIFVNSSVNRRNQELIKELSTPPPGSNDLYFRT 1166
Query: 1145 QYSQTFLAQCTYCLWKQHLSYWRNTSYTAVRLLFTIMTGILFGLIFWGVGAKSKKEQDLF 1204
+Y+Q F Q C WK + S WR Y A+R L T++ G+LFGL+FW G K +KEQDL
Sbjct: 1167 KYAQPFSTQTKACFWKMYWSNWRYPQYNAIRFLMTVVIGVLFGLLFWQTGTKIEKEQDLN 1226
Query: 1205 NAMGSMYAAVTFIGVVNGASVQPIVAIERTVFYRERAAGMYSAMPYALAQVIIELPHILV 1264
N G+MYAAV F+G N A+VQP VAIERTVFYRE+AAGMYSA+PYA++QV +E+ + +
Sbjct: 1227 NFFGAMYAAVLFLGATNAATVQPAVAIERTVFYREKAAGMYSAIPYAISQVAVEIMYNTI 1286
Query: 1265 QAVVYGIIVYAMMGFEWTASKVLWNLFFTYFSFLYYTYYGMMTMAITPNPHVAGILSTSF 1324
Q VY +I+Y+M+G++WT K W ++ F+Y+T YGMM +A+TPN +AGI + F
Sbjct: 1287 QTGVYTLILYSMIGYDWTVVKFFWFYYYMLTCFVYFTLYGMMLVALTPNYQIAGICLSFF 1346
Query: 1325 YAIWCLFSGFIIPLSRIPIWWKWYYWICPVAWTLNGLVTSQYGHNMDTLD----NGQSVE 1380
+ W LFSGF+IP +IPIWW+WYYW PVAWTL G++TSQ G + S++
Sbjct: 1347 LSFWNLFSGFLIPRPQIPIWWRWYYWASPVAWTLYGIITSQVGDRDSIVHITGVGDMSLK 1406
Query: 1381 EFVRNYFGFEYDFLGVVAIVVVSFSVLFALIFTFGIKAFNFQKR 1424
++N FGF+YDFL VVA+V +++ ++F F +GIK NFQ+R
Sbjct: 1407 TLLKNGFGFDYDFLPVVAVVHIAWILIFLFAFAYGIKFLNFQRR 1450
>At2g26910 putative ABC transporter
Length = 1420
Score = 1651 bits (4276), Expect = 0.0
Identities = 807/1425 (56%), Positives = 1066/1425 (74%), Gaps = 18/1425 (1%)
Query: 11 NHCMDVFSKS----EREDDEEALKCVAIKRILTSSCIRKNVESKGEGKGKDVETIQLEST 66
N + FS+S + +DEE L+ A++R+ T S IR+ + G+ K+++ LE++
Sbjct: 3 NSAENAFSRSTSFKDEIEDEEELRWAALQRLPTYSRIRRGIFRDMVGEPKEIQIGNLEAS 62
Query: 67 EKRALLARLVKIAEEDNEKFLLKLKERMDRVGLELPTIEVRFEDINVEAQVYVGRRALPT 126
E+R LL RLV E D E+F ++++R D V L+ P IEVRF+++ VE+ V+VG RALPT
Sbjct: 63 EQRLLLDRLVNSVENDPEQFFARVRKRFDAVDLKFPKIEVRFQNLMVESFVHVGSRALPT 122
Query: 127 LFNFFVNVIEGCLNNLQIIPSPKKQLHILQNVSGILKPRRMTLLLGPPGSGKTTLLLALA 186
+ NF +N+ EG L N+ +I + +L IL +SG+++P R+TLLLGPP SGKTTLLLALA
Sbjct: 123 IPNFIINMAEGLLRNIHVIGGKRNKLTILDGISGVIRPSRLTLLLGPPSSGKTTLLLALA 182
Query: 187 GILGKDLKQSGRVTYNGKGLEEFVPQRTSAYVSQYDNHIGEMTVRETLAFSARCQGVGQN 246
G LG +L+ SG++TYNG L+E + RTSAYVSQ D H+ EMTVR+TL F+ RCQGVG
Sbjct: 183 GRLGTNLQTSGKITYNGYDLKEIIAPRTSAYVSQQDWHVAEMTVRQTLEFAGRCQGVGFK 242
Query: 247 YEMLTELLRKEKESKIEPDPDINAYMKEAAIEGHQNSVVIDYILKILGLDVCADTMVGDQ 306
Y+ML EL R+EK + I PD D++ +MK A+ G + S+V++Y++KILGLD CADT+VGD+
Sbjct: 243 YDMLLELARREKLAGIVPDEDLDIFMKSLALGGMETSLVVEYVMKILGLDTCADTLVGDE 302
Query: 307 MIRGISGGEKKRLTTGEMLVGPIKVLFMDEISNGLDSSTTFQIINSIKQSIHILNGTALV 366
MI+GISGG+KKRLTTGE+LVGP +VLFMDEISNGLDSSTT QII ++ S H L GT ++
Sbjct: 303 MIKGISGGQKKRLTTGELLVGPARVLFMDEISNGLDSSTTHQIIMYMRHSTHALEGTTVI 362
Query: 367 SLLQPAPETYELFDDIILLTDGQIVYQGPREYVLEFFESTGFKCPERKGVADFLQEVTSR 426
SLLQP+PETYELFDD+IL+++GQI+YQGPR+ VL+FF S GF CP+RK VADFLQEVTS+
Sbjct: 363 SLLQPSPETYELFDDVILMSEGQIIYQGPRDEVLDFFSSLGFTCPDRKNVADFLQEVTSK 422
Query: 427 KDQWQYWAREDEPYNFVTVKDFARAFELFHIGKQLGEELADPFDKSKFHSNVLITKKYGI 486
KDQ QYW+ PY +V FA AF + GK+L ++L PFDK HS L T +YG+
Sbjct: 423 KDQQQYWSVPFRPYRYVPPGKFAEAFRSYPTGKKLAKKLEVPFDKRFNHSAALSTSQYGV 482
Query: 487 NKKELLRACASRELLLMKRNSFVYIFKATQLTYLATLTTTLFLRTKMYHSTIEDAQTYMG 546
K ELL+ + + LMK+N+F+Y+FK QL +A +T T+F RT M+H+TI+D Y+G
Sbjct: 483 KKSELLKINFAWQKQLMKQNAFIYVFKFVQLLLVALITMTVFCRTTMHHNTIDDGNIYLG 542
Query: 547 ALFFTVTVAMFNGISELNMTIMKLPIFYKQRDLLFYPSWAYSLPPWILKIPITIIEVAIW 606
+L+F++ + +FNG +E+ M + KLP+ YK RDL FYPSWAY+LP W+L IP +IIE A W
Sbjct: 543 SLYFSMVIILFNGFTEVPMLVAKLPVLYKHRDLHFYPSWAYTLPSWLLSIPTSIIESATW 602
Query: 607 ECISYYAIGFDPNIGRFFKQSLVVLCINQMASALFRFMAALGRDIVVANTFGTFSLLAVT 666
++YY IG+DP RF +Q L+ ++QM+ LFR M +LGR ++VANTFG+F++L V
Sbjct: 603 VAVTYYTIGYDPLFSRFLQQFLLYFSLHQMSLGLFRVMGSLGRHMIVANTFGSFAMLVVM 662
Query: 667 VLGGFVISREDVHKWFLWGYWSSPLMYGQNAIAVNEFLGHGWRKVAPN-SNETLGVSILK 725
LGGF+ISR+ + W++WGYW SPLMY QNA +VNEFLGH W+K A N ++++LG+++LK
Sbjct: 663 TLGGFIISRDSIPSWWIWGYWISPLMYAQNAASVNEFLGHNWQKTAGNHTSDSLGLALLK 722
Query: 726 SRGFFPQAYWYWIGVGALIGYVFLFNFLFALALHFLSPFRKDQAGLSQEKLQERNAS-TD 784
R F YWYWIGV AL+GY LFN LF L L L+P+ K QA +S+E+L ER
Sbjct: 723 ERSLFSGNYWYWIGVAALLGYTVLFNILFTLFLAHLNPWGKFQAVVSREELDEREKKRKG 782
Query: 785 EEFIQSQQQENSSNTKMDEEVSENKASSSGRKGMVLPFQPLSLTFDDITYSVDMPQGMKN 844
+EF+ ++ + + + +N +GMVLPFQPLSL+F +I Y VD+P G+K
Sbjct: 783 DEFVVELREYLQHSGSIHGKYFKN-------RGMVLPFQPLSLSFSNINYYVDVPLGLKE 835
Query: 845 QGVTEDRLELLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGIKTSGYIEGNIKVSGYQ 904
QG+ EDRL+LL ++GAFRPGVLTAL+GVSGAGKTTLMDVLAG KT G IEG++ +SG+
Sbjct: 836 QGILEDRLQLLVNITGAFRPGVLTALVGVSGAGKTTLMDVLAGRKTGGTIEGDVYISGFP 895
Query: 905 KNQKSFARISGYCEQFDIHSPNVTVYESLLYSAWLRLSPEVDHATRKMFIEEVMELVELN 964
K Q++FARISGYCEQ D+HSP +TV ESLL+SA LRL ++D T++ F+ EVMELVEL
Sbjct: 896 KRQETFARISGYCEQNDVHSPCLTVVESLLFSACLRLPADIDSETQRAFVHEVMELVELT 955
Query: 965 SLREALVGLPGENGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAIVMRTVRNT 1024
SL ALVGLPG +GLSTEQRKRLTIAVELVANPSI+FMDEPTSGLDARAAAIVMRTVRN
Sbjct: 956 SLSGALVGLPGVDGLSTEQRKRLTIAVELVANPSIVFMDEPTSGLDARAAAIVMRTVRNI 1015
Query: 1025 VDTGRTVVCTIHQPSIDIFDSFDELLLLKLGGEQIYAGPIGNQCSDLIQYFEAIQGVPTI 1084
V+TGRT+VCTIHQPSIDIF+SFDELL +K GGE IYAGP+G + +LI+YFE+I+GV I
Sbjct: 1016 VNTGRTIVCTIHQPSIDIFESFDELLFMKRGGELIYAGPLGQKSCELIKYFESIEGVQKI 1075
Query: 1085 KDGYNPATWMLEITSAGKEANLKVNFTDVYKNSELHRRNKQLIQELSVPSQSSKDLHFDA 1144
K G+NPA WML++T++ +E L V+F ++Y+NS L +RNK+LI+ LS PS +K++ F
Sbjct: 1076 KPGHNPAAWMLDVTASTEEHRLGVDFAEIYRNSNLCQRNKELIEVLSKPSNIAKEIEFPT 1135
Query: 1145 QYSQTFLAQCTYCLWKQHLSYWRNTSYTAVRLLFTIMTGILFGLIFWGVGAKSKKEQDLF 1204
+YSQ+ +Q CLWKQ+LSYWRN YTAVR +T++ ++ G I W G+K +Q LF
Sbjct: 1136 RYSQSLYSQFVACLWKQNLSYWRNPQYTAVRFFYTVVISLMLGTICWKFGSKRDTQQQLF 1195
Query: 1205 NAMGSMYAAVTFIGVVNGASVQPIVAIERTVFYRERAAGMYSAMPYALAQVIIELPHILV 1264
NAMGSMYAAV FIG+ N + QP+V+IER V YRERAAGMYSA+P+A AQV IE P++L
Sbjct: 1196 NAMGSMYAAVLFIGITNATAAQPVVSIERFVSYRERAAGMYSALPFAFAQVFIEFPYVLA 1255
Query: 1265 QAVVYGIIVYAMMGFEWTASKVLWNLFFTYFSFLYYTYYGMMTMAITPNPHVAGILSTSF 1324
Q+ +Y I YAM FEW+A K LW LFF YFS +Y+T+YGMMT AITPN +VA I++ F
Sbjct: 1256 QSTIYSTIFYAMAAFEWSAVKFLWYLFFMYFSIMYFTFYGMMTTAITPNHNVASIIAAPF 1315
Query: 1325 YAIWCLFSGFIIPLSRIPIWWKWYYWICPVAWTLNGLVTSQYGHNMDT--LDNG---QSV 1379
Y +W LFSGF+IP RIP+WW+WYYW PVAWTL GL+ SQYG + + L +G V
Sbjct: 1316 YMLWNLFSGFMIPYKRIPLWWRWYYWANPVAWTLYGLLVSQYGDDERSVKLSDGIHQVMV 1375
Query: 1380 EEFVRNYFGFEYDFLGVVAIVVVSFSVLFALIFTFGIKAFNFQKR 1424
++ + + G+++DFLGV AI+VV+F V F+L+F F IKAFNFQ+R
Sbjct: 1376 KQLLEDVMGYKHDFLGVSAIMVVAFCVFFSLVFAFAIKAFNFQRR 1420
>At1g66950 ABC transporter, putative
Length = 1434
Score = 1641 bits (4250), Expect = 0.0
Identities = 804/1427 (56%), Positives = 1057/1427 (73%), Gaps = 55/1427 (3%)
Query: 15 DVFSKSER-EDDEEALKCVAIKRILTSSCIRKNVESKGEGKGK----DVETIQLESTEKR 69
+VF +SER E+D+ L+ AI+R+ T +RK + + GK D++ +LE +K+
Sbjct: 46 EVFGRSERREEDDMELRWAAIERLPTFDRLRKGMLPQTSANGKIELEDIDLTRLEPKDKK 105
Query: 70 ALLARLVKIAEEDNEKFLLKLKERMDRVGLELPTIEVRFEDINVEAQVYVGRRALPTLFN 129
L+ ++ EEDNEKFL L+ER DRVG+E+P IEVR+E+I+VE V RALPTLFN
Sbjct: 106 HLMEMILSFVEEDNEKFLRDLRERTDRVGIEVPKIEVRYENISVEGDVRSASRALPTLFN 165
Query: 130 FFVNVIEGCLNNLQIIPSPKKQLHILQNVSGILKPRRMTLLLGPPGSGKTTLLLALAGIL 189
+N +E L ++PS +K++ IL+++SGI+KP RMTLLLGPP SGKTTLL ALAG L
Sbjct: 166 VTLNTLESILGFFHLLPSKRKKIQILKDISGIVKPSRMTLLLGPPSSGKTTLLQALAGKL 225
Query: 190 GKDLKQSGRVTYNGKGLEEFVPQRTSAYVSQYDNHIGEMTVRETLAFSARCQGVGQNYEM 249
L+ SGR+TY G EFVPQ+T AY+SQ+D H GEMTVRE L FS RC GVG Y++
Sbjct: 226 DDTLQMSGRITYCGHEFREFVPQKTCAYISQHDLHFGEMTVREILDFSGRCLGVGSRYQL 285
Query: 250 LTELLRKEKESKIEPDPDINAYMKEAAIEGHQNSVVIDYILKILGLDVCADTMVGDQMIR 309
++EL R+EKE I+PDP I+A+MK AI G + S+V DY+LKILGLD+CAD + GD M R
Sbjct: 286 MSELSRREKEEGIKPDPKIDAFMKSIAISGQETSLVTDYVLKILGLDICADILAGDVMRR 345
Query: 310 GISGGEKKRLTTGEMLVGPIKVLFMDEISNGLDSSTTFQIINSIKQSIHILNGTALVSLL 369
GISGG+KKRLTTGEMLVGP + LFMDEIS GLDSSTTFQI ++Q +HI + T ++SLL
Sbjct: 346 GISGGQKKRLTTGEMLVGPARALFMDEISTGLDSSTTFQICKFMRQLVHISDVTMIISLL 405
Query: 370 QPAPETYELFDDIILLTDGQIVYQGPREYVLEFFESTGFKCPERKGVADFLQEVTSRKDQ 429
QPAPET+ELFDDIILL++GQIVYQGPR+ VLEFFE GF+CPERKGVADFLQEVTS+KDQ
Sbjct: 406 QPAPETFELFDDIILLSEGQIVYQGPRDNVLEFFEYFGFQCPERKGVADFLQEVTSKKDQ 465
Query: 430 WQYWAREDEPYNFVTVKDFARAFELFHIGKQLGEELADPFDKSKFHSNVLITKKYGINKK 489
QYW + ++PYN+V+V DF+ F FH G++L E P+DK+K HS L+T+KYGI+
Sbjct: 466 EQYWNKREQPYNYVSVSDFSSGFSTFHTGQKLTSEFRVPYDKAKTHSAALVTQKYGISNW 525
Query: 490 ELLRACASRELLLMKRNSFVYIFKATQLTYLATLTTTLFLRTKMYHSTIEDAQTYMGALF 549
EL +AC RE LLMKRNSFVY+FK Q+T ++ +T T++LRT+M+ T+ D Q + GA+F
Sbjct: 526 ELFKACFDREWLLMKRNSFVYVFKTVQITIMSLITMTVYLRTEMHVGTVRDGQKFYGAMF 585
Query: 550 FTVTVAMFNGISELNMTIMKLPIFYKQRDLLFYPSWAYSLPPWILKIPITIIEVAIWECI 609
F++ MFNG++EL T+M+LP+FYKQRD LFYP WA++LP W+LKIP+++IE IW +
Sbjct: 586 FSLINVMFNGLAELAFTVMRLPVFYKQRDFLFYPPWAFALPAWLLKIPLSLIESGIWIGL 645
Query: 610 SYYAIGFDPNIGRFFKQSLVVLCINQMASALFRFMAALGRDIVVANTFGTFSLLAVTVLG 669
+YY IGF P+ RF + A+GR V++N+ GTF+LL V LG
Sbjct: 646 TYYTIGFAPSAARF--------------------LGAIGRTEVISNSIGTFTLLIVFTLG 685
Query: 670 GFVISREDVHKWFLWGYWSSPLMYGQNAIAVNEFLGHGWRKVAPNSN-----ETLGVSIL 724
GF+I+++D+ W W Y+ SP+MYGQ AI +NEFL W +PN + +T+G +L
Sbjct: 686 GFIIAKDDIRPWMTWAYYMSPMMYGQTAIVMNEFLDERWS--SPNYDTRINAKTVGEVLL 743
Query: 725 KSRGFFPQAYWYWIGVGALIGYVFLFNFLFALALHFLSPFRKDQAGLSQEKLQERNASTD 784
KSRGFF + YW+WI + AL+G+ LFN + LAL +L+P +A + +E
Sbjct: 744 KSRGFFTEPYWFWICIVALLGFSLLFNLFYILALMYLNPLGNSKATVVEE---------- 793
Query: 785 EEFIQSQQQENSSNTKMDEEVSE-NKASSSG-RKGMVLPFQPLSLTFDDITYSVDMPQGM 842
+ ++ N + V E N +S+ G ++GMVLPFQPLSL F+++ Y VDMP M
Sbjct: 794 -----GKDKQKGENRGTEGSVVELNSSSNKGPKRGMVLPFQPLSLAFNNVNYYVDMPSEM 848
Query: 843 KNQGVTEDRLELLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGIKTSGYIEGNIKVSG 902
K QGV DRL+LL+ V GAFRPG+LTAL+GVSGAGKTTLMDVLAG KT GYIEG+I +SG
Sbjct: 849 KAQGVEGDRLQLLRDVGGAFRPGILTALVGVSGAGKTTLMDVLAGRKTGGYIEGSISISG 908
Query: 903 YQKNQKSFARISGYCEQFDIHSPNVTVYESLLYSAWLRLSPEVDHATRKMFIEEVMELVE 962
Y KNQ +FAR+SGYCEQ DIHSP+VTVYESL+YSAWLRLS ++D TR++F+EEVMELVE
Sbjct: 909 YPKNQTTFARVSGYCEQNDIHSPHVTVYESLIYSAWLRLSTDIDIKTRELFVEEVMELVE 968
Query: 963 LNSLREALVGLPGENGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAIVMRTVR 1022
L LR ++VGLPG +GLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAIVMRTVR
Sbjct: 969 LKPLRNSIVGLPGVDGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAIVMRTVR 1028
Query: 1023 NTVDTGRTVVCTIHQPSIDIFDSFDELLLLKLGGEQIYAGPIGNQCSDLIQYFEAIQGVP 1082
NTVDTGRTVVCTIHQPSIDIF+SFDELLL+K GG+ IYAG +G+ L++YFEA++GVP
Sbjct: 1029 NTVDTGRTVVCTIHQPSIDIFESFDELLLMKRGGQVIYAGSLGHHSQKLVEYFEAVEGVP 1088
Query: 1083 TIKDGYNPATWMLEITSAGKEANLKVNFTDVYKNSELHRRNKQLIQELSVPSQSSKDLHF 1142
I DGYNPATWML++T+ E+ + ++F ++ NS L+RRN++LI++LS P SKD++F
Sbjct: 1089 KINDGYNPATWMLDVTTPSMESQMSLDFAQIFSNSSLYRRNQELIKDLSTPPPGSKDVYF 1148
Query: 1143 DAQYSQTFLAQCTYCLWKQHLSYWRNTSYTAVRLLFTIMTGILFGLIFWGVGAKSKKEQD 1202
+Y+Q+F Q C WKQ+ SYWR+ Y A+R L T++ G+LFGLIFW +G K++ EQD
Sbjct: 1149 KTKYAQSFSTQTKACFWKQYWSYWRHPQYNAIRFLMTVVIGVLFGLIFWQIGTKTENEQD 1208
Query: 1203 LFNAMGSMYAAVTFIGVVNGASVQPIVAIERTVFYRERAAGMYSAMPYALAQVIIELPHI 1262
L N G+MYAAV F+G +N A+VQP +AIERTVFYRE+AAGMYSA+PYA++QV +E+ +
Sbjct: 1209 LNNFFGAMYAAVLFLGALNAATVQPAIAIERTVFYREKAAGMYSAIPYAISQVAVEIMYN 1268
Query: 1263 LVQAVVYGIIVYAMMGFEWTASKVLWNLFFTYFSFLYYTYYGMMTMAITPNPHVAGILST 1322
+Q VY +I+Y+M+G WT +K LW ++ SF+Y+T YGMM MA+TPN +AGI +
Sbjct: 1269 TIQTGVYTLILYSMIGCNWTMAKFLWFYYYMLTSFIYFTLYGMMLMALTPNYQIAGICMS 1328
Query: 1323 SFYAIWCLFSGFIIPLSRIPIWWKWYYWICPVAWTLNGLVTSQYGHNMDTLDNGQSV--- 1379
F ++W LFSGF+IP +IPIWW+WYYW PVAWTL GL+TSQ G + D++ + +
Sbjct: 1329 FFLSLWNLFSGFLIPRPQIPIWWRWYYWATPVAWTLYGLITSQVG-DKDSMVHISGIGDI 1387
Query: 1380 --EEFVRNYFGFEYDFLGVVAIVVVSFSVLFALIFTFGIKAFNFQKR 1424
+ ++ FGFE+DFL VVA+V +++ +LF +F +GIK NFQ+R
Sbjct: 1388 DLKTLLKEGFGFEHDFLPVVAVVHIAWILLFLFVFAYGIKFLNFQRR 1434
>At1g15210 putative ABC transporter
Length = 1442
Score = 1613 bits (4177), Expect = 0.0
Identities = 803/1443 (55%), Positives = 1049/1443 (72%), Gaps = 38/1443 (2%)
Query: 1 MERSDTKTWKNHCMDVFSKSERE-----DDEEALKCVAIKRILTSSCIRKNVESK-GEGK 54
+ RS +K +N D+F+ S R +DEEALK +I+++ T + +R ++ + GE
Sbjct: 19 ISRSVSKASRN-MEDIFNTSSRRTKSVNEDEEALKWASIEKLPTYNRLRTSLMPELGEDD 77
Query: 55 -------GKDVETIQLESTEKRALLARLVKIAEEDNEKFLLKLKERMDRVGLELPTIEVR 107
K V+ +L+ E++ + + K+AE+DNE+ L KL+ R+DRVG++LPT+EVR
Sbjct: 78 VYGNQILNKAVDVTKLDGEERQKFIDMVFKVAEQDNERILTKLRNRIDRVGIQLPTVEVR 137
Query: 108 FEDINVEAQVYVGRRALPTLFNFFVNVIEGCLNNLQIIPSPKKQLHILQNVSGILKPRRM 167
++ + V+A Y G R+LP+L N N+ E L + I + K QL IL++VSGI+KP RM
Sbjct: 138 YDHLTVKADCYTGDRSLPSLLNAVRNMGEAALGMIGIRLAKKAQLTILKDVSGIVKPSRM 197
Query: 168 TLLLGPPGSGKTTLLLALAGILGKDLKQSGRVTYNGKGLEEFVPQRTSAYVSQYDNHIGE 227
TLLLGPP SGKTTLLLALAG L K L SG VTYNG L EFVP +TSAY+SQ D H+G
Sbjct: 198 TLLLGPPSSGKTTLLLALAGKLDKSLDVSGEVTYNGYRLNEFVPIKTSAYISQNDLHVGI 257
Query: 228 MTVRETLAFSARCQGVGQNYEMLTELLRKEKESKIEPDPDINAYMKEAAIEGHQNSVVID 287
MTV+ETL FSARCQGVG Y++L EL R+EK++ I P+ D++ +MK +A +G ++S++ D
Sbjct: 258 MTVKETLDFSARCQGVGTRYDLLNELARREKDAGIFPEADVDLFMKASAAQGVKSSLITD 317
Query: 288 YILKILGLDVCADTMVGDQMIRGISGGEKKRLTTGEMLVGPIKVLFMDEISNGLDSSTTF 347
Y LKILGLD+C DT+VGD M+RGISGG+KKR+TTGEM+VGP K LFMDEIS GLDSSTTF
Sbjct: 318 YTLKILGLDICKDTIVGDDMMRGISGGQKKRVTTGEMIVGPTKTLFMDEISTGLDSSTTF 377
Query: 348 QIINSIKQSIHILNGTALVSLLQPAPETYELFDDIILLTDGQIVYQGPREYVLEFFESTG 407
QI+ ++Q +H+ T L+SLLQPAPET++LFDDIILL++GQIVYQGPR+++LEFFES G
Sbjct: 378 QIVKCLQQIVHLTEATVLISLLQPAPETFDLFDDIILLSEGQIVYQGPRDHILEFFESFG 437
Query: 408 FKCPERKGVADFLQEVTSRKDQWQYWAREDEPYNFVTVKDFARAFELFHIGKQLGEELAD 467
FKCPERKG ADFLQEVTS+KDQ QYW + PY ++ V +FA +F+ FH+G +L EL+
Sbjct: 438 FKCPERKGTADFLQEVTSKKDQEQYWVDPNRPYRYIPVSEFASSFKKFHVGSKLSNELSV 497
Query: 468 PFDKSKFHSNVLITKKYGINKKELLRACASRELLLMKRNSFVYIFKATQLTYLATLTTTL 527
P+DKSK H L+ KY I K ELL++C +E +LMKRNSF Y+FK Q+ +A +T+TL
Sbjct: 498 PYDKSKSHKAALMFDKYSIKKTELLKSCWDKEWMLMKRNSFFYVFKTVQIIIIAAITSTL 557
Query: 528 FLRTKMYHSTIEDAQTYMGALFFTVTVAMFNGISELNMTIMKLPIFYKQRDLLFYPSWAY 587
+LRT+M+ DA Y+G+L F + V MFNG++E+ MTI +LP+FYKQRDLLF+P W Y
Sbjct: 558 YLRTEMHTRNEIDANIYVGSLLFAMIVNMFNGLAEMAMTIQRLPVFYKQRDLLFHPPWTY 617
Query: 588 SLPPWILKIPITIIEVAIWECISYYAIGFDPNIGRFFKQSLVVLCINQMASALFRFMAAL 647
+LP ++L IPI+I E W ++YY+IG+ P+ RFFKQ L++ I QMA+ +FRF+A+
Sbjct: 618 TLPTFLLGIPISIFESTAWMVVTYYSIGYAPDAERFFKQFLIIFLIQQMAAGIFRFIAST 677
Query: 648 GRDIVVANTFGTFSLLAVTVLGGFVISREDVHKWFLWGYWSSPLMYGQNAIAVNEFLGHG 707
R + +ANT G LL V + GGF++ R ++ W+ W YW SPL Y NAI VNE
Sbjct: 678 CRTMTIANTGGVLVLLVVFLTGGFLLPRSEIPVWWRWAYWISPLSYAFNAITVNELFAPR 737
Query: 708 W-RKVAPNSNETLGVSILKSRGFFPQAYWYWIGVGALIGYVFLFNFLFALALHFLSPFRK 766
W K++ NS LG S+L F WYWIGVG L+G+ +FN F LAL +L P K
Sbjct: 738 WMNKMSGNSTTRLGTSVLNIWDVFDDKNWYWIGVGGLLGFTVIFNGFFTLALTYLDPLGK 797
Query: 767 DQAGLSQEKLQERNASTDEEFIQSQQQENSSNTKMDEEVSENKASSSGRKGMVLPFQPLS 826
QA L +E+ +E ++ T+M+ S S +KGMVLPF PL+
Sbjct: 798 AQAILPKEEDEEAKGKAG----------SNKETEME--------SVSAKKGMVLPFTPLA 839
Query: 827 LTFDDITYSVDMPQGMKNQGVTEDRLELLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLA 886
++FDD+ Y VDMP M+ QGV E RL+LLKGV+ AFRPGVLTALMGVSGAGKTTLMDVLA
Sbjct: 840 MSFDDVKYFVDMPAEMREQGVQETRLQLLKGVTSAFRPGVLTALMGVSGAGKTTLMDVLA 899
Query: 887 GIKTSGYIEGNIKVSGYQKNQKSFARISGYCEQFDIHSPNVTVYESLLYSAWLRLSPEVD 946
G KT GYIEG+++VSG+ K Q++FARISGYCEQ DIHSP VTV ESL++SA+LRL+ EV
Sbjct: 900 GRKTGGYIEGDVRVSGFPKKQETFARISGYCEQTDIHSPQVTVRESLIFSAFLRLAKEVS 959
Query: 947 HATRKMFIEEVMELVELNSLREALVGLPGENGLSTEQRKRLTIAVELVANPSIIFMDEPT 1006
+ MF+++VMELVEL LR+A+VGLPG GLSTEQRKRLTIAVELVANPSIIFMDEPT
Sbjct: 960 KEDKLMFVDQVMELVELVDLRDAIVGLPGVTGLSTEQRKRLTIAVELVANPSIIFMDEPT 1019
Query: 1007 SGLDARAAAIVMRTVRNTVDTGRTVVCTIHQPSIDIFDSFDELLLLKLGGEQIYAGPIGN 1066
SGLDARAAAIVMR VRNTVDTGRTVVCTIHQPSIDIF++FDELLL+K GG IY+GP+G
Sbjct: 1020 SGLDARAAAIVMRAVRNTVDTGRTVVCTIHQPSIDIFEAFDELLLMKRGGHVIYSGPLGR 1079
Query: 1067 QCSDLIQYFEAIQGVPTIKDGYNPATWMLEITSAGKEANLKVNFTDVYKNSELHRRNKQL 1126
+++YFE+ GVP I + YNPATWMLE +S E L V+F ++YK S L +RNK L
Sbjct: 1080 NSHKVVEYFESFPGVPKIPEKYNPATWMLEASSLAAELKLGVDFAELYKASALCQRNKAL 1139
Query: 1127 IQELSVPSQSSKDLHFDAQYSQTFLAQCTYCLWKQHLSYWRNTSYTAVRLLFTIMTGILF 1186
+QELSVP Q + DL+F Q+SQ Q CLWKQ +YWR+ Y VR +FT+ T ++
Sbjct: 1140 VQELSVPPQGATDLYFATQFSQNTWGQFKSCLWKQWWTYWRSPDYNLVRFIFTLATSLMI 1199
Query: 1187 GLIFWGVGAKSKKEQDLFNAMGSMYAAVTFIGVVNGASVQPIVAIERTVFYRERAAGMYS 1246
G +FW +G K QDL +G++YAAV F+G+ N ++VQP+VA+ERTVFYRE+AAGMYS
Sbjct: 1200 GSVFWQIGGKRSNVQDLTMVIGAIYAAVVFVGINNCSTVQPMVAVERTVFYREKAAGMYS 1259
Query: 1247 AMPYALAQVIIELPHILVQAVVYGIIVYAMMGFEWTASKVLWNLFFTYFSFLYYTYYGMM 1306
A+PYA++QV ELP++L+Q Y +I+Y+M+GFEW ASK LW +F YFSFLY+TYYGMM
Sbjct: 1260 AIPYAISQVTCELPYVLIQTTYYSLIIYSMVGFEWKASKFLWFIFINYFSFLYWTYYGMM 1319
Query: 1307 TMAITPNPHVAGILSTSFYAIWCLFSGFIIPLSRIPIWWKWYYWICPVAWTLNGLVTSQY 1366
T+++TPN VA I +++FY I+ LFSGF IP +IP WW WYYWICPVAWT+ GL+TSQY
Sbjct: 1320 TVSLTPNQQVASIFASAFYGIFNLFSGFFIPRPKIPKWWVWYYWICPVAWTIYGLITSQY 1379
Query: 1367 GHNMDTL-----DNGQSVEEFVRNYFGFEYDFLGVVAIVVVSFSVLFALIFTFGIKAFNF 1421
G + G +V++++++ +GFE D++G VA V+V F+V FA IF F IK NF
Sbjct: 1380 GDVETPIALLGGAPGLTVKQYIKDQYGFESDYMGPVAGVLVGFTVFFAFIFAFCIKTLNF 1439
Query: 1422 QKR 1424
Q R
Sbjct: 1440 QSR 1442
>At1g59870 putative ABC transporter (At1g59870)
Length = 1469
Score = 1608 bits (4165), Expect = 0.0
Identities = 792/1436 (55%), Positives = 1047/1436 (72%), Gaps = 26/1436 (1%)
Query: 15 DVFSKSERE-----DDEEALKCVAIKRILTSSCIRKN-----VESKGEGK---GKDVETI 61
D+FS R DDEEALK AI+++ T S +R VE G K+V+
Sbjct: 34 DIFSSGSRRTQSVNDDEEALKWAAIEKLPTYSRLRTTLMNAVVEDDVYGNQLMSKEVDVT 93
Query: 62 QLESTEKRALLARLVKIAEEDNEKFLLKLKERMDRVGLELPTIEVRFEDINVEAQVYVGR 121
+L+ +++ + + K+AE+DNE+ L KL+ R+DRVG++LPT+EVR+E + ++A Y G
Sbjct: 94 KLDGEDRQKFIDMVFKVAEQDNERILTKLRNRIDRVGIKLPTVEVRYEHLTIKADCYTGN 153
Query: 122 RALPTLFNFFVNVIEGCLNNLQIIPSPKKQLHILQNVSGILKPRRMTLLLGPPGSGKTTL 181
R+LPTL N N+ E L + I + K QL IL+++SG++KP RMTLLLGPP SGKTTL
Sbjct: 154 RSLPTLLNVVRNMGESALGMIGIQFAKKAQLTILKDISGVIKPGRMTLLLGPPSSGKTTL 213
Query: 182 LLALAGILGKDLKQSGRVTYNGKGLEEFVPQRTSAYVSQYDNHIGEMTVRETLAFSARCQ 241
LLALAG L K L+ SG +TYNG L+EFVP++TSAY+SQ D H+G MTV+ETL FSARCQ
Sbjct: 214 LLALAGKLDKSLQVSGDITYNGYQLDEFVPRKTSAYISQNDLHVGIMTVKETLDFSARCQ 273
Query: 242 GVGQNYEMLTELLRKEKESKIEPDPDINAYMKEAAIEGHQNSVVIDYILKILGLDVCADT 301
GVG Y++L EL R+EK++ I P+ D++ +MK +A +G +NS+V DY LKILGLD+C DT
Sbjct: 274 GVGTRYDLLNELARREKDAGIFPEADVDLFMKASAAQGVKNSLVTDYTLKILGLDICKDT 333
Query: 302 MVGDQMIRGISGGEKKRLTTGEMLVGPIKVLFMDEISNGLDSSTTFQIINSIKQSIHILN 361
+VGD M+RGISGG+KKR+TTGEM+VGP K LFMDEIS GLDSSTTFQI+ ++Q +H+
Sbjct: 334 IVGDDMMRGISGGQKKRVTTGEMIVGPTKTLFMDEISTGLDSSTTFQIVKCLQQIVHLNE 393
Query: 362 GTALVSLLQPAPETYELFDDIILLTDGQIVYQGPREYVLEFFESTGFKCPERKGVADFLQ 421
T L+SLLQPAPET++LFDDIIL+++GQIVYQGPR+ +LEFFES GFKCPERKG ADFLQ
Sbjct: 394 ATVLMSLLQPAPETFDLFDDIILVSEGQIVYQGPRDNILEFFESFGFKCPERKGTADFLQ 453
Query: 422 EVTSRKDQWQYWAREDEPYNFVTVKDFARAFELFHIGKQLGEELADPFDKSKFHSNVLIT 481
EVTS+KDQ QYW + PY+++ V +FA ++ FH+G ++ ELA PFDKS+ H L+
Sbjct: 454 EVTSKKDQEQYWVNPNRPYHYIPVSEFASRYKSFHVGTKMSNELAVPFDKSRGHKAALVF 513
Query: 482 KKYGINKKELLRACASRELLLMKRNSFVYIFKATQLTYLATLTTTLFLRTKMYHSTIEDA 541
KY ++K+ELL++C +E LLM+RN+F Y+FK Q+ +A +T+TLFLRT+M DA
Sbjct: 514 DKYSVSKRELLKSCWDKEWLLMQRNAFFYVFKTVQIVIIAAITSTLFLRTEMNTRNEGDA 573
Query: 542 QTYMGALFFTVTVAMFNGISELNMTIMKLPIFYKQRDLLFYPSWAYSLPPWILKIPITII 601
Y+GAL F + + MFNG +E+ M + +LP+FYKQRDLLFYPSW +SLP ++L IP +I+
Sbjct: 574 NLYIGALLFGMIINMFNGFAEMAMMVSRLPVFYKQRDLLFYPSWTFSLPTFLLGIPSSIL 633
Query: 602 EVAIWECISYYAIGFDPNIGRFFKQSLVVLCINQMASALFRFMAALGRDIVVANTFGTFS 661
E W ++YY+IGF P+ RFFKQ L+V I QMA++LFR +A++ R +++ANT G +
Sbjct: 634 ESTAWMVVTYYSIGFAPDASRFFKQFLLVFLIQQMAASLFRLIASVCRTMMIANTGGALT 693
Query: 662 LLAVTVLGGFVISREDVHKWFLWGYWSSPLMYGQNAIAVNEFLGHGWRKVAPNSNET--L 719
LL V +LGGF++ + + W+ W YW SPL Y N + VNE W +SN T L
Sbjct: 694 LLLVFLLGGFLLPKGKIPDWWGWAYWVSPLTYAFNGLVVNEMFAPRWMNKMASSNSTIKL 753
Query: 720 GVSILKSRGFFPQAYWYWIGVGALIGYVFLFNFLFALALHFLSPFRKDQAGLSQEKLQER 779
G +L + + Q WYWI VGAL+ + LFN LF LAL +L+P K L +E+ ++
Sbjct: 754 GTMVLNTWDVYHQKNWYWISVGALLCFTALFNILFTLALTYLNPLGKKAGLLPEEENEDA 813
Query: 780 NASTD--EEFIQSQQQENSSNTKMDEEVSENKASSSG----RKGMVLPFQPLSLTFDDIT 833
+ D + + M ++ A +SG +KGMVLPF PL+++FDD+
Sbjct: 814 DQGKDPMRRSLSTADGNRRGEVAMGRMSRDSAAEASGGAGNKKGMVLPFTPLAMSFDDVK 873
Query: 834 YSVDMPQGMKNQGVTEDRLELLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGIKTSGY 893
Y VDMP M++QGVTE RL+LLKGV+GAFRPGVLTALMGVSGAGKTTLMDVLAG KT GY
Sbjct: 874 YFVDMPGEMRDQGVTETRLQLLKGVTGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGY 933
Query: 894 IEGNIKVSGYQKNQKSFARISGYCEQFDIHSPNVTVYESLLYSAWLRLSPEVDHATRKMF 953
IEG++++SG+ K Q++FARISGYCEQ DIHSP VTV ESL++SA+LRL EV + MF
Sbjct: 934 IEGDVRISGFPKVQETFARISGYCEQTDIHSPQVTVRESLIFSAFLRLPKEVGKDEKMMF 993
Query: 954 IEEVMELVELNSLREALVGLPGENGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARA 1013
+++VMELVEL+SLR+++VGLPG GLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARA
Sbjct: 994 VDQVMELVELDSLRDSIVGLPGVTGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARA 1053
Query: 1014 AAIVMRTVRNTVDTGRTVVCTIHQPSIDIFDSFDELLLLKLGGEQIYAGPIGNQCSDLIQ 1073
AAIVMR VRNTVDTGRTVVCTIHQPSIDIF++FDEL+L+K GG+ IYAGP+G +++
Sbjct: 1054 AAIVMRAVRNTVDTGRTVVCTIHQPSIDIFEAFDELMLMKRGGQVIYAGPLGQNSHKVVE 1113
Query: 1074 YFEAIQGVPTIKDGYNPATWMLEITSAGKEANLKVNFTDVYKNSELHRRNKQLIQELSVP 1133
YFE+ GV I + YNPATWMLE +S E L V+F ++Y S LH+RNK L++ELSVP
Sbjct: 1114 YFESFPGVSKIPEKYNPATWMLEASSLAAELKLSVDFAELYNQSALHQRNKALVKELSVP 1173
Query: 1134 SQSSKDLHFDAQYSQTFLAQCTYCLWKQHLSYWRNTSYTAVRLLFTIMTGILFGLIFWGV 1193
+ DL+F Q+SQ Q CLWKQ +YWR+ Y VR +FT+ T +L G +FW +
Sbjct: 1174 PAGASDLYFATQFSQNTWGQFKSCLWKQWWTYWRSPDYNLVRFIFTLATSLLIGTVFWQI 1233
Query: 1194 GAKSKKEQDLFNAMGSMYAAVTFIGVVNGASVQPIVAIERTVFYRERAAGMYSAMPYALA 1253
G DL +G++YAA+ F+G+ N ++VQP+VA+ERTVFYRERAAGMYSAMPYA++
Sbjct: 1234 GGNRSNAGDLTMVIGALYAAIIFVGINNCSTVQPMVAVERTVFYRERAAGMYSAMPYAIS 1293
Query: 1254 QVIIELPHILVQAVVYGIIVYAMMGFEWTASKVLWNLFFTYFSFLYYTYYGMMTMAITPN 1313
QV ELP++L+Q V Y +IVYAM+GFEW A K W +F +YFSFLY+TYYGMMT+++TPN
Sbjct: 1294 QVTCELPYVLIQTVYYSLIVYAMVGFEWKAEKFFWFVFVSYFSFLYWTYYGMMTVSLTPN 1353
Query: 1314 PHVAGILSTSFYAIWCLFSGFIIPLSRIPIWWKWYYWICPVAWTLNGLVTSQYGH---NM 1370
VA I +++FY I+ LFSGF IP +IP WW WYYWICPVAWT+ GL+ SQYG +
Sbjct: 1354 QQVASIFASAFYGIFNLFSGFFIPRPKIPKWWIWYYWICPVAWTVYGLIVSQYGDVETRI 1413
Query: 1371 DTLDNGQ--SVEEFVRNYFGFEYDFLGVVAIVVVSFSVLFALIFTFGIKAFNFQKR 1424
L +V++++ +++GF+ DF+G VA V+++F+V FA IF F I+ NFQ R
Sbjct: 1414 QVLGGAPDLTVKQYIEDHYGFQSDFMGPVAAVLIAFTVFFAFIFAFCIRTLNFQTR 1469
>At3g16340 putative ABC transporter
Length = 1416
Score = 1599 bits (4140), Expect = 0.0
Identities = 779/1408 (55%), Positives = 1034/1408 (73%), Gaps = 31/1408 (2%)
Query: 23 EDDEEALKCVAIKRILTSSCIRKNVESKGEGKGKDVETIQLESTEKRALLARLVKIAEED 82
+ DEEALK A++++ T + +R + E V+ +L +++ + + K+ EED
Sbjct: 34 DHDEEALKWAALEKLPTFARLRTTIIHPHEDL---VDVTKLGVDDRQKFIDSIFKVTEED 90
Query: 83 NEKFLLKLKERMDRVGLELPTIEVRFEDINVEAQVYVGRRALPTLFNFFVNVIEGCLNNL 142
NEKFL K + R+DRV ++LPT+EVRFE + +EA ++G+RALPTL N +N+ E L L
Sbjct: 91 NEKFLKKFRNRIDRVRIKLPTVEVRFEKVTIEANCHIGKRALPTLPNAALNIAERGLRLL 150
Query: 143 QIIPSPKKQLHILQNVSGILKPRRMTLLLGPPGSGKTTLLLALAGILGKDLKQSGRVTYN 202
+ ++ IL++VSGI+KP RMTLLLGPP SGKTTLLLALAG L + LK +GRVTYN
Sbjct: 151 GFNFTKTTKVTILRDVSGIIKPSRMTLLLGPPSSGKTTLLLALAGKLDQSLKVTGRVTYN 210
Query: 203 GKGLEEFVPQRTSAYVSQYDNHIGEMTVRETLAFSARCQGVGQNYEMLTELLRKEKESKI 262
G GLEEFVPQ+TSAY+SQ D H+G MTV+ETL FSARCQGVG Y++L+EL+R+EK++ I
Sbjct: 211 GHGLEEFVPQKTSAYISQNDVHVGVMTVQETLDFSARCQGVGTRYDLLSELVRREKDAGI 270
Query: 263 EPDPDINAYMKEAAIEGHQNSVVIDYILKILGLDVCADTMVGDQMIRGISGGEKKRLTTG 322
P+P+++ +MK A ++S++ DY L+ILGLD+C DT+VGD+MIRGISGG+KKR+TTG
Sbjct: 271 LPEPEVDLFMKSIAAGNVKSSLITDYTLRILGLDICKDTVVGDEMIRGISGGQKKRVTTG 330
Query: 323 EMLVGPIKVLFMDEISNGLDSSTTFQIINSIKQSIHILNGTALVSLLQPAPETYELFDDI 382
EM+VGP K LFMDEIS GLDSSTT+QI+ +++ + + T L+SLLQPAPET+ELFDDI
Sbjct: 331 EMIVGPTKTLFMDEISTGLDSSTTYQIVKCLQEIVRFTDATVLMSLLQPAPETFELFDDI 390
Query: 383 ILLTDGQIVYQGPREYVLEFFESTGFKCPERKGVADFLQEVTSRKDQWQYWAREDEPYNF 442
ILL++GQIVYQGPR++VL FFE+ GFKCP+RKG ADFLQEVTSRKDQ QYWA +PY++
Sbjct: 391 ILLSEGQIVYQGPRDHVLTFFETCGFKCPDRKGTADFLQEVTSRKDQEQYWADSKKPYSY 450
Query: 443 VTVKDFARAFELFHIGKQLGEELADPFDKSKFHSNVLITKKYGINKKELLRACASRELLL 502
++V +F++ F FH+G L ++L+ P+D+ K H L+ KK+ + K +L + C RELLL
Sbjct: 451 ISVSEFSKRFRTFHVGANLEKDLSVPYDRFKSHPASLVFKKHSVPKSQLFKVCWDRELLL 510
Query: 503 MKRNSFVYIFKATQLTYLATLTTTLFLRTKMYHSTIEDAQTYMGALFFTVTVAMFNGISE 562
MKRN+F YI K Q+ +A + +T++LRT+M D Y+GAL F++ V MFNG +E
Sbjct: 511 MKRNAFFYITKTVQIIIMALIASTVYLRTEMGTKNESDGAVYIGALMFSMIVNMFNGFAE 570
Query: 563 LNMTIMKLPIFYKQRDLLFYPSWAYSLPPWILKIPITIIEVAIWECISYYAIGFDPNIGR 622
L + I +LP+FYKQRDLLF+P W +SLP ++L IPI+I E +W I+YY IGF P + R
Sbjct: 571 LALMIQRLPVFYKQRDLLFHPPWTFSLPTFLLGIPISIFESVVWVTITYYMIGFAPELSR 630
Query: 623 FFKQSLVVLCINQMASALFRFMAALGRDIVVANTFGTFSLLAVTVLGGFVISREDVHKWF 682
F K LV+ QMA +FRF+AA R +++ANT G +L + +LGGF++ R ++ KW+
Sbjct: 631 FLKHLLVIFLTQQMAGGIFRFIAATCRSMILANTGGALVILLLFLLGGFIVPRGEIPKWW 690
Query: 683 LWGYWSSPLMYGQNAIAVNEFLGHGWRKVAPNSNET-LGVSILKSRGFFPQAYWYWIGVG 741
W YW SP+ Y +A+ VNE L W + N T LG+++L+ F WYWIGVG
Sbjct: 691 KWAYWVSPMAYTYDALTVNEMLAPRWINQPSSDNSTSLGLAVLEIFDIFTDPNWYWIGVG 750
Query: 742 ALIGYVFLFNFLFALALHFLSPFRKDQAGLSQEKLQERNASTDEEFIQSQQQENSSNTKM 801
++G+ LFN L LAL FL+P K QA +S+E +E A EN S +K
Sbjct: 751 GILGFTVLFNILVTLALTFLNPLEKQQAVVSKENTEENRA------------ENGSKSK- 797
Query: 802 DEEVSENKASSSGRKGMVLPFQPLSLTFDDITYSVDMPQGMKNQGVTEDRLELLKGVSGA 861
S ++GMVLPF PL+++FD++ Y VDMP+ MK QGV++D+L+LLK V+G
Sbjct: 798 ---------SIDVKRGMVLPFTPLTMSFDNVNYYVDMPKEMKEQGVSKDKLQLLKEVTGV 848
Query: 862 FRPGVLTALMGVSGAGKTTLMDVLAGIKTSGYIEGNIKVSGYQKNQKSFARISGYCEQFD 921
FRPGVLTALMGVSGAGKTTLMDVLAG KT GYIEG+I++SG+ K Q++FARISGYCEQ D
Sbjct: 849 FRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIEGDIRISGFPKRQETFARISGYCEQND 908
Query: 922 IHSPNVTVYESLLYSAWLRLSPEVDHATRKMFIEEVMELVELNSLREALVGLPGENGLST 981
IHSP VTV ESL+YSA+LRL EV + F++EVMELVEL SL++A+VGLPG GLST
Sbjct: 909 IHSPQVTVKESLIYSAFLRLPKEVTKYEKMRFVDEVMELVELESLKDAVVGLPGITGLST 968
Query: 982 EQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAIVMRTVRNTVDTGRTVVCTIHQPSID 1041
EQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAIVMRTVRNTVDTGRTVVCTIHQPSID
Sbjct: 969 EQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAIVMRTVRNTVDTGRTVVCTIHQPSID 1028
Query: 1042 IFDSFDELLLLKLGGEQIYAGPIGNQCSDLIQYFEAIQGVPTIKDGYNPATWMLEITSAG 1101
IF++FDELLLLK GG+ IYAGP+G +I+YF+AI GVP IK+ YNPATWMLE++S
Sbjct: 1029 IFEAFDELLLLKRGGQVIYAGPLGQNSHKIIEYFQAIHGVPKIKEKYNPATWMLEVSSMA 1088
Query: 1102 KEANLKVNFTDVYKNSELHRRNKQLIQELSVPSQSSKDLHFDAQYSQTFLAQCTYCLWKQ 1161
EA L+++F + YK S L+++NK L++ELS P Q + DL+F ++SQ+ L Q CLWKQ
Sbjct: 1089 AEAKLEIDFAEHYKTSSLYQQNKNLVKELSTPPQGASDLYFSTRFSQSLLGQFKSCLWKQ 1148
Query: 1162 HLSYWRNTSYTAVRLLFTIMTGILFGLIFWGVGAKSKKEQDLFNAMGSMYAAVTFIGVVN 1221
++YWR Y R FT+ ++ G IFW VG K + DL +G+MYAAV F+GV N
Sbjct: 1149 WITYWRTPDYNLARFFFTLAAAVMLGSIFWKVGTKRENANDLTKVIGAMYAAVLFVGVNN 1208
Query: 1222 GASVQPIVAIERTVFYRERAAGMYSAMPYALAQVIIELPHILVQAVVYGIIVYAMMGFEW 1281
+SVQP++A+ER+VFYRERAA MYSA+PYALAQV+ E+P++L+Q Y +I+YAMM FEW
Sbjct: 1209 SSSVQPLIAVERSVFYRERAAEMYSALPYALAQVVCEIPYVLIQTTYYTLIIYAMMCFEW 1268
Query: 1282 TASKVLWNLFFTYFSFLYYTYYGMMTMAITPNPHVAGILSTSFYAIWCLFSGFIIPLSRI 1341
T +K W F ++ SFLY+TYYGMMT+A+TPN VA + + +FY ++ LFSGF+IP RI
Sbjct: 1269 TLAKFFWFYFVSFMSFLYFTYYGMMTVALTPNQQVAAVFAGAFYGLFNLFSGFVIPRPRI 1328
Query: 1342 PIWWKWYYWICPVAWTLNGLVTSQYGHNMDTLD-----NGQSVEEFVRNYFGFEYDFLGV 1396
P WW WYYWICPVAWT+ GL+ SQYG DT+ N +++ ++ N++G++ DF+
Sbjct: 1329 PKWWIWYYWICPVAWTVYGLIVSQYGDVEDTIKVPGMANDPTIKWYIENHYGYDADFMIP 1388
Query: 1397 VAIVVVSFSVLFALIFTFGIKAFNFQKR 1424
+A V+V F++ FA +F FGI+ NFQ+R
Sbjct: 1389 IATVLVGFTLFFAFMFAFGIRTLNFQQR 1416
>At2g29940 putative ABC transporter
Length = 1443
Score = 1488 bits (3851), Expect = 0.0
Identities = 740/1436 (51%), Positives = 1010/1436 (69%), Gaps = 58/1436 (4%)
Query: 23 EDDEEALKCVAIKRILT------SSCIRKN---VESKGEGKGKDVETI---QLESTEKRA 70
E DEE L+ AI R+ + ++ +R++ ++ G G V+TI +L+ ++
Sbjct: 32 EQDEEDLRWAAIGRLPSQRQGTHNAILRRSQTQTQTSGYADGNVVQTIDVKKLDRADREM 91
Query: 71 LLARLVKIAEEDNEKFLLKLKERMDR-----------------VGLELPTIEVRFEDINV 113
L+ + + +++DN K L +KER+DR VG+E+P IEVRFE++N+
Sbjct: 92 LVRQALATSDQDNFKLLSAIKERLDRFVTTLRILSVSNFREKKVGMEVPKIEVRFENLNI 151
Query: 114 EAQVYVGRRALPTLFNFFVNVIEGCLNNLQIIPSPKKQLHILQNVSGILKPRRMTLLLGP 173
EA V G RALPTL N + E CL++L+II K +L+IL+++SGI+KP RMTLLLGP
Sbjct: 152 EADVQAGTRALPTLVNVSRDFFERCLSSLRIIKPRKHKLNILKDISGIIKPGRMTLLLGP 211
Query: 174 PGSGKTTLLLALAGILGKDLKQSGRVTYNGKGLEEFVPQRTSAYVSQYDNHIGEMTVRET 233
PGSGK+TLLLALAG L K LK++G +TYNG+ L +F +RTSAY+SQ DNHI E+TVRET
Sbjct: 212 PGSGKSTLLLALAGKLDKSLKKTGNITYNGENLNKFHVKRTSAYISQTDNHIAELTVRET 271
Query: 234 LAFSARCQGVGQNYE-MLTELLRKEKESKIEPDPDINAYMKEAAIEGHQNSVVIDYILKI 292
L F+ARCQG + + + +L R EKE I P +I+A+MK A+++G ++SV DY+LK+
Sbjct: 272 LDFAARCQGASEGFAGYMKDLTRLEKERGIRPSSEIDAFMKAASVKGEKHSVSTDYVLKV 331
Query: 293 LGLDVCADTMVGDQMIRGISGGEKKRLTTGEMLVGPIKVLFMDEISNGLDSSTTFQIINS 352
LGLDVC+DTMVG+ M+RG+SGG++KR+TTGEM VGP K LFMDEIS GLDSSTTFQI+
Sbjct: 332 LGLDVCSDTMVGNDMMRGVSGGQRKRVTTGEMTVGPRKTLFMDEISTGLDSSTTFQIVKC 391
Query: 353 IKQSIHILNGTALVSLLQPAPETYELFDDIILLTDGQIVYQGPREYVLEFFESTGFKCPE 412
I+ +H+++ T L++LLQPAPET++LFDD+ILL++G +VYQGPRE V+ FFES GF+ P
Sbjct: 392 IRNFVHLMDATVLMALLQPAPETFDLFDDLILLSEGYMVYQGPREDVIAFFESLGFRLPP 451
Query: 413 RKGVADFLQEVTSRKDQWQYWAREDEPYNFVTVKDFARAFELFHIGKQLGEELADPFDKS 472
RKGVADFLQEVTS+KDQ QYWA +PY F+ V D A AF G +LA PFDK
Sbjct: 452 RKGVADFLQEVTSKKDQAQYWADPSKPYQFIPVSDIAAAFRNSKYGHAADSKLAAPFDKK 511
Query: 473 KFHSNVLITKKYGINKKELLRACASRELLLMKRNSFVYIFKATQLTYLATLTTTLFLRTK 532
+ L K+ I+ E L+ C RELLL+KR+ F+Y F+ Q+ ++ +T T+FL+T+
Sbjct: 512 SADPSALCRTKFAISGWENLKVCFVRELLLIKRHKFLYTFRTCQVGFVGLVTATVFLKTR 571
Query: 533 MYHSTIEDAQTYMGALFFTVTVAMFNGISELNMTIMKLPIFYKQRDLLFYPSWAYSLPPW 592
++ ++ + Y+ LFF + MFNG SEL + I +LP+FYKQRD F+P+W++S+ W
Sbjct: 572 LHPTSEQFGNEYLSCLFFGLVHMMFNGFSELPLMISRLPVFYKQRDNSFHPAWSWSIASW 631
Query: 593 ILKIPITIIEVAIWECISYYAIGFDPNIGRFFKQSLVVLCINQMASALFRFMAALGRDIV 652
+L++P +++E +W + Y+ +G P+ GRFF+ L++ ++QMA LFR MA+L RD+V
Sbjct: 632 LLRVPYSVLEAVVWSGVVYFTVGLAPSAGRFFRYMLLLFSVHQMALGLFRMMASLARDMV 691
Query: 653 VANTFGTFSLLAVTVLGGFVISREDVHKWFLWGYWSSPLMYGQNAIAVNEFLGHGWRKVA 712
+ANTFG+ ++L V +LGGFVI + D+ W++WG+W SPL YGQ AIAVNEF W +
Sbjct: 692 IANTFGSAAILIVFLLGGFVIPKADIKPWWVWGFWVSPLSYGQRAIAVNEFTATRWMTPS 751
Query: 713 PNSNETLGVSILKSRGFFPQAYWYWIGVGALIGYVFLFNFLFALALHFLSPFRKDQAGLS 772
S+ T+G+++LK R F YWYWIG+ LIGY LFN + LAL +L+P RK +A +
Sbjct: 752 AISDTTIGLNLLKLRSFPTNDYWYWIGIAVLIGYAILFNNVVTLALAYLNPLRKARAVVL 811
Query: 773 QEKLQERNASTDEEFIQSQQQENSSNTKMDEEVSENKASSSGRKGMVLPFQPLSLTFDDI 832
+ +E D + S+ +KGM+LPF+PL++TF ++
Sbjct: 812 DDPNEETALVADANQVISE-----------------------KKGMILPFKPLTMTFHNV 848
Query: 833 TYSVDMPQGMKNQGVTEDRLELLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGIKTSG 892
Y VDMP+ M++QGV E RL+LL VSG F PGVLTAL+G SGAGKTTLMDVLAG KT G
Sbjct: 849 NYYVDMPKEMRSQGVPETRLQLLSNVSGVFSPGVLTALVGSSGAGKTTLMDVLAGRKTGG 908
Query: 893 YIEGNIKVSGYQKNQKSFARISGYCEQFDIHSPNVTVYESLLYSAWLRLSPEVDHATRKM 952
Y EG+I++SG+ K Q++FARISGY EQ DIHSP VTV ESL +SA LRL E+ +K
Sbjct: 909 YTEGDIRISGHPKEQQTFARISGYVEQNDIHSPQVTVEESLWFSASLRLPKEITKEQKKE 968
Query: 953 FIEEVMELVELNSLREALVGLPGENGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDAR 1012
F+E+VM LVEL++LR ALVGLPG GLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDAR
Sbjct: 969 FVEQVMRLVELDTLRYALVGLPGTTGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDAR 1028
Query: 1013 AAAIVMRTVRNTVDTGRTVVCTIHQPSIDIFDSFDELLLLKLGGEQIYAGPIGNQCSDLI 1072
AAAIVMRTVRNTVDTGRTVVCTIHQPSIDIF++FDELLL+K GG+ IY G +G L+
Sbjct: 1029 AAAIVMRTVRNTVDTGRTVVCTIHQPSIDIFEAFDELLLMKRGGQVIYGGKLGTHSQVLV 1088
Query: 1073 QYFEAIQGVPTIKDGYNPATWMLEITSAGKEANLKVNFTDVYKNSELHRRNKQLIQELSV 1132
YF+ I GVP I GYNPATWMLE+T+ E + F D+YK S+ R + I++LSV
Sbjct: 1089 DYFQGINGVPPISSGYNPATWMLEVTTPALEEKYNMEFADLYKKSDQFREVEANIKQLSV 1148
Query: 1133 PSQSSKDLHFDAQYSQTFLAQCTYCLWKQHLSYWRNTSYTAVRLLFTIMTGILFGLIFWG 1192
P + S+ + F ++YSQ L+Q CLWKQ+L YWR+ Y VRL+FT + + G +FW
Sbjct: 1149 PPEGSEPISFTSRYSQNQLSQFLLCLWKQNLVYWRSPEYNLVRLVFTTIAAFILGTVFWD 1208
Query: 1193 VGAKSKKEQDLFNAMGSMYAAVTFIGVVNGASVQPIVAIERTVFYRERAAGMYSAMPYAL 1252
+G+K QDL MG++Y+A F+GV N +SVQPIV+IERTVFYRE+AAGMY+ +PYA
Sbjct: 1209 IGSKRTSSQDLITVMGALYSACLFLGVSNASSVQPIVSIERTVFYREKAAGMYAPIPYAA 1268
Query: 1253 AQVIIELPHILVQAVVYGIIVYAMMGFEWTASKVLWNLFFTYFSFLYYTYYGMMTMAITP 1312
AQ ++E+P+IL Q ++YG+I Y +GFE T SK + L F + +F Y+T+YGMM + +TP
Sbjct: 1269 AQGLVEIPYILTQTILYGVITYFTIGFERTFSKFVLYLVFMFLTFTYFTFYGMMAVGLTP 1328
Query: 1313 NPHVAGILSTSFYAIWCLFSGFIIPLSRIPIWWKWYYWICPVAWTLNGLVTSQYGHNMDT 1372
N H+A ++S++FY++W L SGF++ IP+WW W+Y+ICPVAWTL G++ SQ G ++++
Sbjct: 1329 NQHLAAVISSAFYSLWNLLSGFLVQKPLIPVWWIWFYYICPVAWTLQGVILSQLG-DVES 1387
Query: 1373 LDNGQ----SVEEFVRNYFGFEYDFLGVVAIVVVSFSVLFALIFTFGIKAFNFQKR 1424
+ N +V+EF+ YFG++ + +GV A V+V F LF F +K NFQ+R
Sbjct: 1388 MINEPLFHGTVKEFIEYYFGYKPNMIGVSAAVLVGFCALFFSAFALSVKYLNFQRR 1443
>At3g53480 ABC transporter - like protein
Length = 1450
Score = 1423 bits (3683), Expect = 0.0
Identities = 702/1410 (49%), Positives = 984/1410 (69%), Gaps = 22/1410 (1%)
Query: 24 DDEEALKCVAIKRILTSSCIRKNV-----ESKGEGKGKDVETIQLESTEKRALLARLVKI 78
D E AL+ I+R+ T +R + ES E + V+ +L + E+ ++ +L+K
Sbjct: 54 DAEYALQWAEIERLPTVKRMRSTLLDDGDESMTEKGRRVVDVTKLGAVERHLMIEKLIKH 113
Query: 79 AEEDNEKFLLKLKERMDRVGLELPTIEVRFEDINVEAQVYVGR-RALPTLFNFFVNVIEG 137
E DN K L K++ R+DRVG+ELPTIEVR+E + V A+ V +ALPTL+N V+
Sbjct: 114 IENDNLKLLKKIRRRIDRVGMELPTIEVRYESLKVVAECEVVEGKALPTLWNTAKRVLSE 173
Query: 138 CLNNLQIIPSPKKQLHILQNVSGILKPRRMTLLLGPPGSGKTTLLLALAGILGKDLKQSG 197
L L + + +++I+ +V+GI+KP R+TLLLGPP GKTTLL AL+G L +LK SG
Sbjct: 174 -LVKLTGAKTHEAKINIINDVNGIIKPGRLTLLLGPPSCGKTTLLKALSGNLENNLKCSG 232
Query: 198 RVTYNGKGLEEFVPQRTSAYVSQYDNHIGEMTVRETLAFSARCQGVGQNYEMLTELLRKE 257
++YNG L+EFVPQ+TSAY+SQYD HI EMTVRET+ FSARCQGVG +++ E+ ++E
Sbjct: 233 EISYNGHRLDEFVPQKTSAYISQYDLHIAEMTVRETVDFSARCQGVGSRTDIMMEVSKRE 292
Query: 258 KESKIEPDPDINAYMKEAAIEGHQNSVVIDYILKILGLDVCADTMVGDQMIRGISGGEKK 317
KE I PD +++AYMK ++EG Q S+ DYILKILGLD+CA+ ++GD M RGISGG+KK
Sbjct: 293 KEKGIIPDTEVDAYMKAISVEGLQRSLQTDYILKILGLDICAEILIGDVMRRGISGGQKK 352
Query: 318 RLTTGEMLVGPIKVLFMDEISNGLDSSTTFQIINSIKQSIHILNGTALVSLLQPAPETYE 377
RLTT EM+VGP K LFMDEI+NGLDSST FQI+ S++Q HI + T LVSLLQPAPE+Y+
Sbjct: 353 RLTTAEMIVGPTKALFMDEITNGLDSSTAFQIVKSLQQFAHISSATVLVSLLQPAPESYD 412
Query: 378 LFDDIILLTDGQIVYQGPREYVLEFFESTGFKCPERKGVADFLQEVTSRKDQWQYWARED 437
LFDDI+L+ G+IVY GPR VL FFE GF+CPERKGVADFLQEV S+KDQ QYW ED
Sbjct: 413 LFDDIMLMAKGRIVYHGPRGEVLNFFEDCGFRCPERKGVADFLQEVISKKDQAQYWWHED 472
Query: 438 EPYNFVTVKDFARAFELFHIGKQLGEELADPFDKSKFHSNVLITKKYGINKKELLRACAS 497
PY+FV+V+ ++ F+ IGK++ + L+ P+D+SK H + L Y + EL AC S
Sbjct: 473 LPYSFVSVEMLSKKFKDLSIGKKIEDTLSKPYDRSKSHKDALSFSVYSLPNWELFIACIS 532
Query: 498 RELLLMKRNSFVYIFKATQLTYLATLTTTLFLRTKMYHSTIEDAQTYMGALFFTVTVAMF 557
RE LLMKRN FVYIFK QL A +T T+F+RT+M I +YM ALFF + + +
Sbjct: 533 REYLLMKRNYFVYIFKTAQLVMAAFITMTVFIRTRMGIDIIH-GNSYMSALFFALIILLV 591
Query: 558 NGISELNMTIMKLPIFYKQRDLLFYPSWAYSLPPWILKIPITIIEVAIWECISYYAIGFD 617
+G EL+MT +L +FYKQ+ L FYP+WAY++P +LK+P++ E +W C+SYY IG+
Sbjct: 592 DGFPELSMTAQRLAVFYKQKQLCFYPAWAYAIPATVLKVPLSFFESLVWTCLSYYVIGYT 651
Query: 618 PNIGRFFKQSLVVLCINQMASALFRFMAALGRDIVVANTFGTFSLLAVTVLGGFVISRED 677
P RFFKQ +++ ++ + ++FR +AA+ + +V + T G+F +L V GFVI
Sbjct: 652 PEASRFFKQFILLFAVHFTSISMFRCLAAIFQTVVASITAGSFGILFTFVFAGFVIPPPS 711
Query: 678 VHKWFLWGYWSSPLMYGQNAIAVNEFLGHGWRKVAPNSNETLGVSILKSRGFFPQAYWYW 737
+ W WG+W++PL YG+ ++VNEFL W ++ PN N TLG +IL++RG Y YW
Sbjct: 712 MPAWLKWGFWANPLSYGEIGLSVNEFLAPRWNQMQPN-NFTLGRTILQTRGMDYNGYMYW 770
Query: 738 IGVGALIGYVFLFNFLFALALHFLSPFRKDQAGLSQEKLQERNASTDEEFIQSQQQENSS 797
+ + AL+G+ LFN +F LAL FL +A +SQ+KL E + ++++
Sbjct: 771 VSLCALLGFTVLFNIIFTLALTFLKSPTSSRAMISQDKLSELQGT----------EKSTE 820
Query: 798 NTKMDEEVSENKASSSGRKGMVLPFQPLSLTFDDITYSVDMPQGMKNQGVTEDRLELLKG 857
++ + ++ +++ + MVLPF+PL++TF D+ Y VDMP M++QG + +L+LL
Sbjct: 821 DSSVRKKTTDSPVKTEEEDKMVLPFKPLTVTFQDLNYFVDMPVEMRDQGYDQKKLQLLSD 880
Query: 858 VSGAFRPGVLTALMGVSGAGKTTLMDVLAGIKTSGYIEGNIKVSGYQKNQKSFARISGYC 917
++GAFRPG+LTALMGVSGAGKTTL+DVLAG KTSGYIEG+I++SG+ K Q++FAR+SGYC
Sbjct: 881 ITGAFRPGILTALMGVSGAGKTTLLDVLAGRKTSGYIEGDIRISGFPKVQETFARVSGYC 940
Query: 918 EQFDIHSPNVTVYESLLYSAWLRLSPEVDHATRKMFIEEVMELVELNSLREALVGLPGEN 977
EQ DIHSPN+TV ES++YSAWLRL+PE+D T+ F+++V+E +EL+ ++++LVG+ G +
Sbjct: 941 EQTDIHSPNITVEESVIYSAWLRLAPEIDATTKTKFVKQVLETIELDEIKDSLVGVTGVS 1000
Query: 978 GLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAIVMRTVRNTVDTGRTVVCTIHQ 1037
GLSTEQRKRLTIAVELVANPSIIFMDEPT+GLDARAAAIVMR V+N DTGRT+VCTIHQ
Sbjct: 1001 GLSTEQRKRLTIAVELVANPSIIFMDEPTTGLDARAAAIVMRAVKNVADTGRTIVCTIHQ 1060
Query: 1038 PSIDIFDSFDELLLLKLGGEQIYAGPIGNQCSDLIQYFEAIQGVPTIKDGYNPATWMLEI 1097
PSIDIF++FDEL+LLK GG IY GP+G +I+YFE++ +P IKD +NPATWML++
Sbjct: 1061 PSIDIFEAFDELVLLKRGGRMIYTGPLGQHSRHIIEYFESVPEIPKIKDNHNPATWMLDV 1120
Query: 1098 TSAGKEANLKVNFTDVYKNSELHRRNKQLIQELSVPSQSSKDLHFDAQYSQTFLAQCTYC 1157
+S E L V+F +Y +S L++RN +L+++LS P S D+ F ++Q++ Q
Sbjct: 1121 SSQSVEIELGVDFAKIYHDSALYKRNSELVKQLSQPDSGSSDIQFKRTFAQSWWGQFKSI 1180
Query: 1158 LWKQHLSYWRNTSYTAVRLLFTIMTGILFGLIFWGVGAKSKKEQDLFNAMGSMYAAVTFI 1217
LWK +LSYWR+ SY +R++ T+++ ++FG +FW G +Q +F G++Y V F+
Sbjct: 1181 LWKMNLSYWRSPSYNLMRMMHTLVSSLIFGALFWKQGQNLDTQQSMFTVFGAIYGLVLFL 1240
Query: 1218 GVVNGASVQPIVAIERTVFYRERAAGMYSAMPYALAQVIIELPHILVQAVVYGIIVYAMM 1277
G+ N AS ER V YRER AGMYSA YAL QV+ E+P+I +QA + I+ Y M+
Sbjct: 1241 GINNCASALQYFETERNVMYRERFAGMYSATAYALGQVVTEIPYIFIQAAEFVIVTYPMI 1300
Query: 1278 GFEWTASKVLWNLFFTYFSFLYYTYYGMMTMAITPNPHVAGILSTSFYAIWCLFSGFIIP 1337
GF +A KV W+L+ + S L + Y M ++ITPN VA IL + FY + LFSGF+IP
Sbjct: 1301 GFYPSAYKVFWSLYSMFCSLLTFNYLAMFLVSITPNFMVAAILQSLFYVGFNLFSGFLIP 1360
Query: 1338 LSRIPIWWKWYYWICPVAWTLNGLVTSQYG---HNMDTLDNGQSVEEFVRNYFGFEYDFL 1394
+++P WW W Y++ P +WTLNG ++SQYG ++ +V F+++YFGF +D L
Sbjct: 1361 QTQVPGWWIWLYYLTPTSWTLNGFISSQYGDIHEEINVFGQSTTVARFLKDYFGFHHDLL 1420
Query: 1395 GVVAIVVVSFSVLFALIFTFGIKAFNFQKR 1424
V A+V ++F + A +F F + NFQ+R
Sbjct: 1421 AVTAVVQIAFPIALASMFAFFVGKLNFQRR 1450
>At2g37280 putative ABC transporter
Length = 1413
Score = 1414 bits (3660), Expect = 0.0
Identities = 710/1419 (50%), Positives = 982/1419 (69%), Gaps = 27/1419 (1%)
Query: 18 SKSEREDD----EEALKCVAIKRILTSSCIRKNVESK-GEG--KGKDV-ETIQLESTEKR 69
S++E ED E AL+ I+R+ T +R ++ K GEG KGK V + +L + E+
Sbjct: 10 SRNEHEDGGDEAEHALQWAEIQRLPTFKRLRSSLVDKYGEGTEKGKKVVDVTKLGAMERH 69
Query: 70 ALLARLVKIAEEDNEKFLLKLKERMDRVGLELPTIEVRFEDINVEAQVYVGR-RALPTLF 128
++ +L+K E DN K L K++ RM+RVG+E P+IEVR+E + VEA V +ALPTL+
Sbjct: 70 LMIEKLIKHIENDNLKLLKKIRRRMERVGVEFPSIEVRYEHLGVEAACEVVEGKALPTLW 129
Query: 129 NFFVNVIEGCLNNLQIIPSPKKQLHILQNVSGILKPRRMTLLLGPPGSGKTTLLLALAGI 188
N +V L L + + + + IL +VSGI+ P R+TLLLGPPG GKTTLL AL+G
Sbjct: 130 NSLKHVFLDLLK-LSGVRTNEANIKILTDVSGIISPGRLTLLLGPPGCGKTTLLKALSGN 188
Query: 189 LGKDLKQSGRVTYNGKGLEEFVPQRTSAYVSQYDNHIGEMTVRETLAFSARCQGVGQNYE 248
L +LK G ++YNG GL E VPQ+TSAY+SQ+D HI EMT RET+ FSARCQGVG +
Sbjct: 189 LENNLKCYGEISYNGHGLNEVVPQKTSAYISQHDLHIAEMTTRETIDFSARCQGVGSRTD 248
Query: 249 MLTELLRKEKESKIEPDPDINAYMKEAAIEGHQNSVVIDYILKILGLDVCADTMVGDQMI 308
++ E+ ++EK+ I PDP+I+AYMK +++G + S+ DYILKILGLD+CA+T+VG+ M
Sbjct: 249 IMMEVSKREKDGGIIPDPEIDAYMKAISVKGLKRSLQTDYILKILGLDICAETLVGNAMK 308
Query: 309 RGISGGEKKRLTTGEMLVGPIKVLFMDEISNGLDSSTTFQIINSIKQSIHILNGTALVSL 368
RGISGG+KKRLTT EM+VGP K LFMDEI+NGLDSST FQII S++Q HI N T VSL
Sbjct: 309 RGISGGQKKRLTTAEMIVGPTKALFMDEITNGLDSSTAFQIIKSLQQVAHITNATVFVSL 368
Query: 369 LQPAPETYELFDDIILLTDGQIVYQGPREYVLEFFESTGFKCPERKGVADFLQEVTSRKD 428
LQPAPE+Y+LFDDI+L+ +G+IVY GPR+ VL+FFE GF+CPERKGVADFLQEV S+KD
Sbjct: 369 LQPAPESYDLFDDIVLMAEGKIVYHGPRDDVLKFFEECGFQCPERKGVADFLQEVISKKD 428
Query: 429 QWQYWAREDEPYNFVTVKDFARAFELFHIGKQLGEELADPFDKSKFHSNVLITKKYGINK 488
Q QYW ++ P++FV+V ++ F+ IG+++ E L+ P+D SK H + L Y + K
Sbjct: 429 QGQYWLHQNLPHSFVSVDTLSKRFKDLEIGRKIEEALSKPYDISKTHKDALSFNVYSLPK 488
Query: 489 KELLRACASRELLLMKRNSFVYIFKATQLTYLATLTTTLFLRTKMYHSTIEDAQTYMGAL 548
EL RAC SRE LLMKRN FVY+FK QL A +T T+F+RT+M I +YM L
Sbjct: 489 WELFRACISREFLLMKRNYFVYLFKTFQLVLAAIITMTVFIRTRMDIDIIH-GNSYMSCL 547
Query: 549 FFTVTVAMFNGISELNMTIMKLPIFYKQRDLLFYPSWAYSLPPWILKIPITIIEVAIWEC 608
FF V + +GI EL+MT+ +L +FYKQ+ L FYP+WAY++P +LKIP++ E +W C
Sbjct: 548 FFATVVLLVDGIPELSMTVQRLSVFYKQKQLCFYPAWAYAIPATVLKIPLSFFESLVWTC 607
Query: 609 ISYYAIGFDPNIGRFFKQSLVVLCINQMASALFRFMAALGRDIVVANTFGTFSLLAVTVL 668
++YY IG+ P RFF+Q +++ ++ + ++FR +AA+ + V A T G+F +L V
Sbjct: 608 LTYYVIGYTPEPYRFFRQFMILFAVHFTSISMFRCIAAIFQTGVAAMTAGSFVMLITFVF 667
Query: 669 GGFVISREDVHKWFLWGYWSSPLMYGQNAIAVNEFLGHGWRKVAPNSNETLGVSILKSRG 728
GF I D+ W WG+W +P+ Y + ++VNEFL W+K+ P +N TLG +IL+SRG
Sbjct: 668 AGFAIPYTDMPGWLKWGFWVNPISYAEIGLSVNEFLAPRWQKMQP-TNVTLGRTILESRG 726
Query: 729 FFPQAYWYWIGVGALIGYVFLFNFLFALALHFLSPFRKDQAGLSQEKLQERNASTDEEFI 788
Y YW+ + AL+G +FN +F LAL FL + +SQ+KL E + D
Sbjct: 727 LNYDDYMYWVSLSALLGLTIIFNTIFTLALSFLKSPTSSRPMISQDKLSELQGTKDSSVK 786
Query: 789 QSQQQENSSNTKMDEEVSENKASSSGRKGMVLPFQPLSLTFDDITYSVDMPQGMKNQGVT 848
+++ ++S T D G+ M+LPF+PL++TF D+ Y VD+P MK QG
Sbjct: 787 KNKPLDSSIKTNEDP----------GK--MILPFKPLTITFQDLNYYVDVPVEMKGQGYN 834
Query: 849 EDRLELLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGIKTSGYIEGNIKVSGYQKNQK 908
E +L+LL ++GAFRPGVLTALMG+SGAGKTTL+DVLAG KTSGYIEG I++SG+ K Q+
Sbjct: 835 EKKLQLLSEITGAFRPGVLTALMGISGAGKTTLLDVLAGRKTSGYIEGEIRISGFLKVQE 894
Query: 909 SFARISGYCEQFDIHSPNVTVYESLLYSAWLRLSPEVDHATRKMFIEEVMELVELNSLRE 968
+FAR+SGYCEQ DIHSP++TV ESL+YSAWLRL PE++ T+ F+++V+E +EL +++
Sbjct: 895 TFARVSGYCEQTDIHSPSITVEESLIYSAWLRLVPEINPQTKIRFVKQVLETIELEEIKD 954
Query: 969 ALVGLPGENGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAIVMRTVRNTVDTG 1028
ALVG+ G +GLSTEQRKRLT+AVELVANPSIIFMDEPT+GLDARAAAIVMR V+N +TG
Sbjct: 955 ALVGVAGVSGLSTEQRKRLTVAVELVANPSIIFMDEPTTGLDARAAAIVMRAVKNVAETG 1014
Query: 1029 RTVVCTIHQPSIDIFDSFDELLLLKLGGEQIYAGPIGNQCSDLIQYFEAIQGVPTIKDGY 1088
RT+VCTIHQPSI IF++FDEL+LLK GG IY+GP+G S +I+YF+ I GV I+D Y
Sbjct: 1015 RTIVCTIHQPSIHIFEAFDELVLLKRGGRMIYSGPLGQHSSCVIEYFQNIPGVAKIRDKY 1074
Query: 1089 NPATWMLEITSAGKEANLKVNFTDVYKNSELHRRNKQLIQELSVPSQSSKDLHFDAQYSQ 1148
NPATWMLE+TS E L ++F +Y S+L++ N +L++ELS P S DLHF ++Q
Sbjct: 1075 NPATWMLEVTSESVETELDMDFAKIYNESDLYKNNSELVKELSKPDHGSSDLHFKRTFAQ 1134
Query: 1149 TFLAQCTYCLWKQHLSYWRNTSYTAVRLLFTIMTGILFGLIFWGVGAKSKKEQDLFNAMG 1208
+ Q CLWK LSYWR+ SY +R+ T ++ +FGL+FW G K +Q+LF +G
Sbjct: 1135 NWWEQFKSCLWKMSLSYWRSPSYNLMRIGHTFISSFIFGLLFWNQGKKIDTQQNLFTVLG 1194
Query: 1209 SMYAAVTFIGVVNGASVQPIVAIERTVFYRERAAGMYSAMPYALAQVIIELPHILVQAVV 1268
++Y V F+G+ N S ER V YRER AGMYSA YALAQV+ E+P+I +Q+
Sbjct: 1195 AIYGLVLFVGINNCTSALQYFETERNVMYRERFAGMYSAFAYALAQVVTEIPYIFIQSAE 1254
Query: 1269 YGIIVYAMMGFEWTASKVLWNLFFTYFSFLYYTYYGMMTMAITPNPHVAGILSTSFYAIW 1328
+ I++Y M+GF + SKV W+L+ + + L + Y M ++ITPN VA IL + F+ +
Sbjct: 1255 FVIVIYPMIGFYASFSKVFWSLYAMFCNLLCFNYLAMFLISITPNFMVAAILQSLFFTTF 1314
Query: 1329 CLFSGFIIPLSRIPIWWKWYYWICPVAWTLNGLVTSQYG---HNMDTLDNGQSVEEFVRN 1385
+F+GF+IP +IP WW W+Y+I P +WTLN +SQYG ++ ++V F+ +
Sbjct: 1315 NIFAGFLIPKPQIPKWWVWFYYITPTSWTLNLFFSSQYGDIHQKINAFGETKTVASFLED 1374
Query: 1386 YFGFEYDFLGVVAIVVVSFSVLFALIFTFGIKAFNFQKR 1424
YFGF +D L + AI++++F + A ++ F + NFQKR
Sbjct: 1375 YFGFHHDRLMITAIILIAFPIALATMYAFFVAKLNFQKR 1413
>At4g15230 ABC transporter like protein
Length = 979
Score = 571 bits (1471), Expect = e-162
Identities = 293/624 (46%), Positives = 408/624 (64%), Gaps = 71/624 (11%)
Query: 818 MVLPFQPLSLTFDDITYSVDMPQGMKNQGVTEDRLELLKGVSGAFRPGVLTALMGVSGAG 877
++LPF+PL++TF ++ Y ++ PQG Q LL ++GA +PGVLT+LMGVSGAG
Sbjct: 410 IILPFKPLTVTFQNVQYYIETPQGKTRQ--------LLSDITGALKPGVLTSLMGVSGAG 461
Query: 878 KTTLMDVLAGIKTSGYIEGNIKVSGYQKNQKSFARISGYCEQFDIHSPNVTVYESLLYSA 937
KTTL+DVL+G KT G I+G IKV GY K Q++FAR+SGYCEQFDIHSPN+TV ESL YSA
Sbjct: 462 KTTLLDVLSGRKTRGIIKGEIKVGGYPKVQETFARVSGYCEQFDIHSPNITVEESLKYSA 521
Query: 938 WLRLSPEVDHATRKM--------------FIEEVMELVELNSLREALVGLPGENGLSTEQ 983
WLRL +D T+ + ++EV+E VEL+ +++++VGLPG +GLS EQ
Sbjct: 522 WLRLPYNIDSKTKNVRNYTLKTNRLKEIELVKEVLETVELDDIKDSVVGLPGISGLSIEQ 581
Query: 984 RKRLTIAVELVANPSIIFMDEPTSGLDARAAAIVMRTVRNTVDTGRTVVCTIHQPSIDIF 1043
RKRLTIAVELVANPSIIFMDEPT+GLDARAAAIVMR V+N +TGRTVVCTIHQPSIDIF
Sbjct: 582 RKRLTIAVELVANPSIIFMDEPTTGLDARAAAIVMRAVKNVAETGRTVVCTIHQPSIDIF 641
Query: 1044 DSFDELLLLKLGGEQIYAGPIGNQCSDLIQYFEAIQGVPTIKDGYNPATWMLEITSAGKE 1103
++FDEL+L+K GG+ +Y GP G S +I+YFE
Sbjct: 642 ETFDELILMKNGGQLVYYGPPGQNSSKVIEYFE--------------------------- 674
Query: 1104 ANLKVNFTDVYKNSELHRRNKQLIQELSVPSQSSKDLHFDAQYSQTFLAQCTYCLWKQHL 1163
NK ++++LS S S+ L F +Q+SQT Q CLWKQH
Sbjct: 675 -------------------NKMVVEQLSSASLGSEALRFPSQFSQTAWVQLKACLWKQHY 715
Query: 1164 SYWRNTSYTAVRLLFTIMTGILFGLIFWGVGAKSKKEQDLFNAMGSMYAAVTFIGVVNGA 1223
SYWRN S+ R++F ++ L GL+FW +QDL + GSMY V F G+ N A
Sbjct: 716 SYWRNPSHNITRIVFILLDSTLCGLLFWQKAEDINNQQDLISIFGSMYTLVVFPGMNNCA 775
Query: 1224 SVQPIVAIERTVFYRERAAGMYSAMPYALAQVIIELPHILVQAVVYGIIVYAMMGFEWTA 1283
+V +A ER VFYRER A MYS+ Y+ +QV+IE+P+ L+Q+++ IIVY +G+ +
Sbjct: 776 AVINFIAAERNVFYRERFARMYSSWAYSFSQVLIEVPYSLLQSLLCTIIVYPTIGYHMSV 835
Query: 1284 SKVLWNLFFTYFSFLYYTYYGMMTMAITPNPHVAGILSTSFYAIWCLFSGFIIPLSRIPI 1343
K+ W+L+ + S L + Y GM+ +A+TPN H+A L +SF+++ LF+GF+IP +IP
Sbjct: 836 YKMFWSLYSIFCSLLIFNYSGMLMVALTPNIHMAVTLRSSFFSMLNLFAGFVIPKQKIPK 895
Query: 1344 WWKWYYWICPVAWTLNGLVTSQYGH-NMDTLDNGQS--VEEFVRNYFGFEYDFLGVVAIV 1400
WW W Y++ P +W L GL++SQYG + + L G+ V F+ +YFG++++ L VVA V
Sbjct: 896 WWIWMYYLSPTSWVLEGLLSSQYGDVDKEILVFGEKKRVSAFLEDYFGYKHESLAVVAFV 955
Query: 1401 VVSFSVLFALIFTFGIKAFNFQKR 1424
++++ ++ A +F F + +FQK+
Sbjct: 956 LIAYPIIVATLFAFFMSKLSFQKK 979
Score = 359 bits (921), Expect = 7e-99
Identities = 180/350 (51%), Positives = 242/350 (68%), Gaps = 19/350 (5%)
Query: 250 LTELLRKEKESKIEPDPDINAYMKEAAIEGHQNSVVIDYILKILGLDVCADTMVGDQMIR 309
+ E+ R EK +I PDP ++AYMK ILGLD+CADT VGD
Sbjct: 1 MKEISRMEKLQEIIPDPAVDAYMK------------------ILGLDICADTRVGDATRP 42
Query: 310 GISGGEKKRLTTGEMLVGPIKVLFMDEISNGLDSSTTFQIINSIKQSIHILNGTALVSLL 369
GISGGEK+RLTTGE++VGP LFMDEISNGLDSSTTFQI++ ++Q HI T L+SLL
Sbjct: 43 GISGGEKRRLTTGELVVGPATTLFMDEISNGLDSSTTFQIVSCLQQLAHIAEATILISLL 102
Query: 370 QPAPETYELFDDIILLTDGQIVYQGPREYVLEFFESTGFKCPERKGVADFLQEVTSRKDQ 429
QPAPET+ELFDD+IL+ +G+I+Y PR + FFE GFKCPERKGVADFLQE+ S+KDQ
Sbjct: 103 QPAPETFELFDDVILMGEGKIIYHAPRADICRFFEEFGFKCPERKGVADFLQEIMSKKDQ 162
Query: 430 WQYWAREDEPYNFVTVKDFARAFELFHIGKQLGEELADPFDKSKFHSNVLITKKYGINKK 489
QYW D+PY++++V F F+ ++G L EEL+ PF+KS+ + L KKY + K
Sbjct: 163 EQYWCHRDKPYSYISVDSFINKFKESNLGLLLKEELSKPFNKSQTRKDGLCYKKYSLGKW 222
Query: 490 ELLRACASRELLLMKRNSFVYIFKATQLTYLATLTTTLFLRTKMYHSTIEDAQTYMGALF 549
E+L+AC+ RE LLMKRNSF+Y+FK+ L + A +T T+FL+ ++ MG+LF
Sbjct: 223 EMLKACSRREFLLMKRNSFIYLFKSALLVFNALVTMTVFLQVGATTDSLH-GNYLMGSLF 281
Query: 550 FTVTVAMFNGISELNMTIMKLPIFYKQRDLLFYPSWAYSLPPWILKIPIT 599
+ + +G+ EL +TI +L +F KQ+DL FYP+WAY++P ILKIP++
Sbjct: 282 TALFRLLADGLPELTLTISRLGVFCKQKDLYFYPAWAYAIPSIILKIPLS 331
Score = 103 bits (256), Expect = 9e-22
Identities = 78/261 (29%), Positives = 138/261 (51%), Gaps = 24/261 (9%)
Query: 150 KQLHILQNVSGILKPRRMTLLLGPPGSGKTTLLLALAGILGKDLKQSGRVTYNGKGLEEF 209
K +L +++G LKP +T L+G G+GKTTLL L+G + + + G + G +
Sbjct: 434 KTRQLLSDITGALKPGVLTSLMGVSGAGKTTLLDVLSGRKTRGIIK-GEIKVGGYPKVQE 492
Query: 210 VPQRTSAYVSQYDNHIGEMTVRETLAFSARCQGVGQNYEMLTELLRKEKESKIEPDPDIN 269
R S Y Q+D H +TV E+L +SA + + N + T+ +R N
Sbjct: 493 TFARVSGYCEQFDIHSPNITVEESLKYSAWLR-LPYNIDSKTKNVR-------------N 538
Query: 270 AYMKEAAIEGHQNSVVIDYILKILGLDVCADTMVGDQMIRGISGGEKKRLTTGEMLVGPI 329
+K ++ + ++ +L+ + LD D++VG I G+S ++KRLT LV
Sbjct: 539 YTLKTNRLKEIE---LVKEVLETVELDDIKDSVVGLPGISGLSIEQRKRLTIAVELVANP 595
Query: 330 KVLFMDEISNGLDSSTTFQIINSIKQSIHILNGTALVSLLQPAPETYELFDDIILLTD-G 388
++FMDE + GLD+ ++ ++K ++ T + ++ QP+ + +E FD++IL+ + G
Sbjct: 596 SIIFMDEPTTGLDARAAAIVMRAVK-NVAETGRTVVCTIHQPSIDIFETFDELILMKNGG 654
Query: 389 QIVYQGP----REYVLEFFES 405
Q+VY GP V+E+FE+
Sbjct: 655 QLVYYGPPGQNSSKVIEYFEN 675
Score = 73.9 bits (180), Expect = 6e-13
Identities = 48/200 (24%), Positives = 94/200 (47%), Gaps = 1/200 (0%)
Query: 492 LRACASRELLLMKRNSFVYIFKATQLTYLATLTTTLFLRTKMYHSTIEDAQTYMGALFFT 551
L+AC ++ RN I + + +TL LF + + +D + G+++
Sbjct: 706 LKACLWKQHYSYWRNPSHNITRIVFILLDSTLCGLLFWQKAEDINNQQDLISIFGSMYTL 765
Query: 552 VTV-AMFNGISELNMTIMKLPIFYKQRDLLFYPSWAYSLPPWILKIPITIIEVAIWECIS 610
V M N + +N + +FY++R Y SWAYS ++++P ++++ + I
Sbjct: 766 VVFPGMNNCAAVINFIAAERNVFYRERFARMYSSWAYSFSQVLIEVPYSLLQSLLCTIIV 825
Query: 611 YYAIGFDPNIGRFFKQSLVVLCINQMASALFRFMAALGRDIVVANTFGTFSLLAVTVLGG 670
Y IG+ ++ + F + C + + M AL +I +A T + + + G
Sbjct: 826 YPTIGYHMSVYKMFWSLYSIFCSLLIFNYSGMLMVALTPNIHMAVTLRSSFFSMLNLFAG 885
Query: 671 FVISREDVHKWFLWGYWSSP 690
FVI ++ + KW++W Y+ SP
Sbjct: 886 FVIPKQKIPKWWIWMYYLSP 905
Score = 67.8 bits (164), Expect = 4e-11
Identities = 79/375 (21%), Positives = 164/375 (43%), Gaps = 29/375 (7%)
Query: 954 IEEVMELVELNSLREALVGLPGENGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARA 1013
++ M+++ L+ + VG G+S +++RLT +V + +FMDE ++GLD+
Sbjct: 19 VDAYMKILGLDICADTRVGDATRPGISGGEKRRLTTGELVVGPATTLFMDEISNGLDSST 78
Query: 1014 AAIVMRTVRNTVDTGR-TVVCTIHQPSIDIFDSFDELLLLKLGGEQIYAGPIGNQCSDLI 1072
++ ++ T++ ++ QP+ + F+ FD+++L+ G+ IY P +D+
Sbjct: 79 TFQIVSCLQQLAHIAEATILISLLQPAPETFELFDDVILMG-EGKIIYHAP----RADIC 133
Query: 1073 QYFEAIQGVPTIKDGYNPATWMLEITSAGKEANLKV------------NFTDVYKNSELH 1120
++FE + G A ++ EI S + +F + +K S L
Sbjct: 134 RFFEEFGFKCPERKGV--ADFLQEIMSKKDQEQYWCHRDKPYSYISVDSFINKFKESNLG 191
Query: 1121 RRNKQLIQELSVPSQSSKDLHFDAQYSQTFLAQCTYCLWKQHLSYWRNTSYTAVRLLFTI 1180
K+ + + SQ+ KD +YS C ++ L RN+ + +
Sbjct: 192 LLLKEELSKPFNKSQTRKDGLCYKKYSLGKWEMLKACSRREFLLMKRNSFIYLFKSALLV 251
Query: 1181 MTGILFGLIFWGVGAKSKKEQDLFNAMGSMYAAVTFIGVVNGASVQPIVAIERTVFYRER 1240
++ +F VGA + + MGS++ A+ F + +G + VF +++
Sbjct: 252 FNALVTMTVFLQVGATTDSLHGNY-LMGSLFTAL-FRLLADGLPELTLTISRLGVFCKQK 309
Query: 1241 AAGMYSAMPYALAQVIIELP-----HILVQAVVYGIIVYAMMGFEWTASKVLWNLFFTYF 1295
Y A YA+ +I+++P ++ AV Y ++Y S+ NL Y
Sbjct: 310 DLYFYPAWAYAIPSIILKIPLSPRGQKVLPAVPY--LIYIQPFLCINVSRYCGNLSHNYC 367
Query: 1296 SFLYYTYYGMMTMAI 1310
+++Y+ + ++ I
Sbjct: 368 FHNHWSYFHIGSLII 382
>At3g25620 membrane transporter, putative
Length = 646
Score = 201 bits (511), Expect = 2e-51
Identities = 176/623 (28%), Positives = 312/623 (49%), Gaps = 34/623 (5%)
Query: 798 NTKMDEEVSENKASSSGRKGMVL--PFQPLSLTFDDITYSVDMPQG-----MKNQGVTED 850
N +D++ + S R+ VL +P+ L F+++TYS+ G +Q +
Sbjct: 36 NPCLDDDNDHDGPSHQSRQSSVLRQSLRPIILKFEELTYSIKSQTGKGSYWFGSQEPKPN 95
Query: 851 RLELLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGIKTSGYIEGNIKVSGYQKNQKSF 910
RL +LK VSG +PG L A++G SG+GKTTL+ LAG + G + G + +G + S
Sbjct: 96 RL-VLKCVSGIVKPGELLAMLGPSGSGKTTLVTALAG-RLQGKLSGTVSYNG-EPFTSSV 152
Query: 911 ARISGYCEQFDIHSPNVTVYESLLYSAWLRLSPEVDHATRKMFIEEVMELVELNSLREAL 970
R +G+ Q D+ P++TV E+L Y+A LRL E+ + +E V+ + L ++
Sbjct: 153 KRKTGFVTQDDVLYPHLTVMETLTYTALLRLPKELTRKEKLEQVEMVVSDLGLTRCCNSV 212
Query: 971 VGLPGENGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAIVMRTVRNTVDTGRT 1030
+G G+S +RKR++I E++ NPS++ +DEPTSGLD+ AA ++ T+R+ GRT
Sbjct: 213 IGGGLIRGISGGERKRVSIGQEMLVNPSLLLLDEPTSGLDSTTAARIVATLRSLARGGRT 272
Query: 1031 VVCTIHQPSIDIFDSFDELLLLKLGGEQIYAGPIGNQCSDLIQYFEAIQGVPTIKDGYNP 1090
VV TIHQPS ++ FD++L+L G IY+G G +++YF +I G NP
Sbjct: 273 VVTTIHQPSSRLYRMFDKVLVLS-EGCPIYSGDSGR----VMEYFGSI-GYQPGSSFVNP 326
Query: 1091 ATWMLEITSAGKEANLKVNFTDVYKNSELHRRNKQLIQELSVPSQSSKDLH--FDAQYSQ 1148
A ++L++ + G ++ K + + N L R +Q + S+ S K+L+ + S+
Sbjct: 327 ADFVLDLAN-GITSDTK-QYDQIETNGRLDRLEEQNSVKQSLISSYKKNLYPPLKEEVSR 384
Query: 1149 TFLAQCTYC-LWKQHLSYWRNTSYTAVRLLFTIMTGILFGLIFWGVGAKSKKEQDLFNAM 1207
TF T L K+ ++ N T+ + F+++ + GL++W + L + +
Sbjct: 385 TFPQDQTNARLRKKAIT---NRWPTSWWMQFSVL--LKRGLLWW-----HSRVAHLQDQV 434
Query: 1208 GSMYAAVTFIGVVNGASVQPIVAIERTVFYRERAAGMYSAMPYALAQVIIELPHILVQAV 1267
G ++ F G + ER + +ER++G+Y Y +A+ + +LP L+
Sbjct: 435 GLLFFFSIFWGFFPLFNAIFTFPQERPMLIKERSSGIYRLSSYYIARTVGDLPMELILPT 494
Query: 1268 VYGIIVYAMMGFEWTASKVLWNLFFTYFSFLYYTYYGMMTMAITPNPHVAGILSTSFYAI 1327
++ I Y M G + + + + L ++ L G+ AI + A LS+ +
Sbjct: 495 IFVTITYWMGGLKPSLTTFIMTLMIVLYNVLVAQGVGLALGAILMDAKKAATLSSVLMLV 554
Query: 1328 WCLFSGFIIPLSRIPIWWKWYYWICPVAWTLNGLVTSQYGHNMDTLDNGQSVEEFVRNYF 1387
+ L G+ I IP + W ++ + LV QY + + + G + V +Y
Sbjct: 555 FLLAGGYYI--QHIPGFIAWLKYVSFSHYCYKLLVGVQYTWD-EVYECGSGLHCSVMDYE 611
Query: 1388 GFEYDFLGVVAIVVVSFSVLFAL 1410
G + +G + V++ +V+ L
Sbjct: 612 GIKNLRIGNMMWDVLALAVMLLL 634
Score = 162 bits (410), Expect = 1e-39
Identities = 150/537 (27%), Positives = 256/537 (46%), Gaps = 76/537 (14%)
Query: 148 PKKQLHILQNVSGILKPRRMTLLLGPPGSGKTTLLLALAGILGKDLKQSGRVTYNGKGLE 207
PK +L+ VSGI+KP + +LGP GSGKTTL+ ALAG L L SG V+YNG+
Sbjct: 92 PKPNRLVLKCVSGIVKPGELLAMLGPSGSGKTTLVTALAGRLQGKL--SGTVSYNGEPFT 149
Query: 208 EFVPQRTSAYVSQYDNHIGEMTVRETLAFSARCQGVGQNYEMLTELLRKEKESKIEPDPD 267
V ++T +V+Q D +TV ETL ++A + + EL RKEK ++E
Sbjct: 150 SSVKRKTG-FVTQDDVLYPHLTVMETLTYTALLR-------LPKELTRKEKLEQVE---- 197
Query: 268 INAYMKEAAIEGHQNSVVIDYILKILGLDVCADTMVGDQMIRGISGGEKKRLTTG-EMLV 326
++ LGL C ++++G +IRGISGGE+KR++ G EMLV
Sbjct: 198 --------------------MVVSDLGLTRCCNSVIGGGLIRGISGGERKRVSIGQEMLV 237
Query: 327 GPIKVLFMDEISNGLDSSTTFQIINSIKQSIHILNGTALVSLLQPAPETYELFDDIILLT 386
P +L +DE ++GLDS+T +I+ +++ S+ T + ++ QP+ Y +FD +++L+
Sbjct: 238 NP-SLLLLDEPTSGLDSTTAARIVATLR-SLARGGRTVVTTIHQPSSRLYRMFDKVLVLS 295
Query: 387 DGQIVYQGPREYVLEFFESTGFKCPERKGV--ADFLQEV-------TSRKDQWQYWARED 437
+G +Y G V+E+F S G++ P V ADF+ ++ T + DQ + R D
Sbjct: 296 EGCPIYSGDSGRVMEYFGSIGYQ-PGSSFVNPADFVLDLANGITSDTKQYDQIETNGRLD 354
Query: 438 EPYNFVTVKDFARAFELFHIGKQLGEELADPFDKSKFHSNVLITKKYGINKKELLRACAS 497
+VK + ++ L EE++ F + + +N + KK N+ +
Sbjct: 355 RLEEQNSVKQSLISSYKKNLYPPLKEEVSRTFPQDQ--TNARLRKKAITNRWP--TSWWM 410
Query: 498 RELLLMKRNSFVYIFKATQLTYLATLTTTLFLRTKMYHSTIEDAQTYMGAL-FFTVTVAM 556
+ +L+KR +HS + Q +G L FF++
Sbjct: 411 QFSVLLKRGLL------------------------WWHSRVAHLQDQVGLLFFFSIFWGF 446
Query: 557 FNGISELNMTIMKLPIFYKQRDLLFYPSWAYSLPPWILKIPITIIEVAIWECISYYAIGF 616
F + + + P+ K+R Y +Y + + +P+ +I I+ I+Y+ G
Sbjct: 447 FPLFNAIFTFPQERPMLIKERSSGIYRLSSYYIARTVGDLPMELILPTIFVTITYWMGGL 506
Query: 617 DPNIGRFFKQSLVVLCINQMASALFRFMAALGRDIVVANTFGTFSLLAVTVLGGFVI 673
P++ F ++VL +A + + A+ D A T + +L + GG+ I
Sbjct: 507 KPSLTTFIMTLMIVLYNVLVAQGVGLALGAILMDAKKAATLSSVLMLVFLLAGGYYI 563
>At3g53510 ABC transporter -like protein
Length = 739
Score = 200 bits (509), Expect = 4e-51
Identities = 169/666 (25%), Positives = 304/666 (45%), Gaps = 50/666 (7%)
Query: 779 RNASTDEEFIQSQQQENSSNTK-MDEEVSENKASSSGRKGMVLPFQPLSLTFDDITYSVD 837
R + T E + S + E ++ MD V+ N SS V P L+F D+TYSV
Sbjct: 40 RVSVTLAELLMSVEDEGDDQSRAMDIAVASNFWSSVP-SSRVPSSSPFVLSFKDLTYSVK 98
Query: 838 MPQGMK------NQGVTEDRLE-----LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLA 886
+ + K N + +E LL G+SG R G + A++G SG+GK+TL+D LA
Sbjct: 99 IKKKFKPFPCCGNSPFDGNDMEMNTKVLLNGISGEAREGEMMAVLGASGSGKSTLIDALA 158
Query: 887 GIKTSGYIEGNIKVSGYQKNQKSFARISGYCEQFDIHSPNVTVYESLLYSAWLRLSPEVD 946
+ + G+I ++G IS Y Q D+ P +TV E+L++SA RL +
Sbjct: 159 NRISKESLRGDITLNGEVLESSLHKVISAYVMQDDLLFPMLTVEETLMFSAEFRLPSSLS 218
Query: 947 HATRKMFIEEVMELVELNSLREALVGLPGENGLSTEQRKRLTIAVELVANPSIIFMDEPT 1006
+K ++ +++ + L + + ++G G G+S +R+R++I +++ +P I+F+DEPT
Sbjct: 219 KKKKKARVQALIDQLGLRNAAKTVIGDEGHRGVSGGERRRVSIGTDIIHDPIILFLDEPT 278
Query: 1007 SGLDARAAAIVMRTVRNTVDTGRTVVCTIHQPSIDIFDSFDELLLLKLGGEQIYAGPIGN 1066
SGLD+ +A +V++ ++ +G V+ +IHQPS I D+L+ L G +Y+G
Sbjct: 279 SGLDSTSAYMVVKVLQRIAQSGSIVIMSIHQPSYRILGLLDKLIFLS-RGNTVYSG---- 333
Query: 1067 QCSDLIQYFEAIQGVPTIKDGYNPATWML----EITSAGKEANLKVNFTDVYK---NSEL 1119
+ L Q+F G P I + N + L E+ + + V F ++ S
Sbjct: 334 SPTHLPQFFSEF-GHP-IPENENKPEFALDLIRELEDSPEGTKSLVEFHKQWRAKQTSSQ 391
Query: 1120 HRRNKQLIQELSVPSQSSK--------DLHFDAQ-YSQTFLAQCTYCLWKQHLSYWRNTS 1170
RRN + + ++ + S+ +L Q ++ F + + L+ R
Sbjct: 392 SRRNTNVSLKDAISASISRGKLVSGATNLRSSFQTFANPFWTEMLVIGKRSILNSRRQPE 451
Query: 1171 YTAVRLLFTIMTGILFGLIFWGVGAKSKKEQDLFNAMGSMYAAVTFIGVVNGASVQPIVA 1230
+RL ++TG++ IFW + + Q+ +A A P+
Sbjct: 452 LFGIRLGAVLVTGMILATIFWKLDNSPRGIQERL----GFFAFAMSTTFYTCAEAIPVFL 507
Query: 1231 IERTVFYRERAAGMYSAMPYALAQVIIELPHILVQAVVYGIIVYAMMGFEWTASKVLWNL 1290
ER +F RE A Y Y LA II +P +++ + + ++ +G + L+
Sbjct: 508 QERYIFMRETAYNAYRRSSYVLAHTIISIPALIILSAAFAASTFSAVGLAGGSEGFLFFF 567
Query: 1291 FFTYFSFLYYTYYGMMTMAITPNPHVAGILSTSFYAIWCLFSGFIIPLSRIPIWWKWYYW 1350
F +F + + + + + + + A + LFSGF I RIP++W W+++
Sbjct: 568 FTILTAFWAGSSFVTFLSGVVSHVMIGFTVVVAILAYFLLFSGFFISRDRIPLYWIWFHY 627
Query: 1351 ICPVAWTLNGLVTSQYGHNMDTLDNGQSVEEFVRNYFGFEYDFLGVVAIVVVSFSVLFAL 1410
+ V + G++ +++ + FVR F+ LG V V S+L ++
Sbjct: 628 LSLVKYPYEGVLQNEF---------EDPTKCFVRGIQMFDNSPLGQVP-TAVKISLLKSM 677
Query: 1411 IFTFGI 1416
GI
Sbjct: 678 SGVLGI 683
Score = 151 bits (382), Expect = 2e-36
Identities = 133/566 (23%), Positives = 250/566 (43%), Gaps = 64/566 (11%)
Query: 154 ILQNVSGILKPRRMTLLLGPPGSGKTTLLLALAGILGKDLKQSGRVTYNGKGLEEFVPQR 213
+L +SG + M +LG GSGK+TL+ ALA + K+ + G +T NG+ LE + +
Sbjct: 126 LLNGISGEAREGEMMAVLGASGSGKSTLIDALANRISKESLR-GDITLNGEVLESSLHKV 184
Query: 214 TSAYVSQYDNHIGEMTVRETLAFSARCQGVGQNYEMLTELLRKEKESKIEPDPDINAYMK 273
SAYV Q D +TV ETL FSA + + + L +K+K+++++
Sbjct: 185 ISAYVMQDDLLFPMLTVEETLMFSAE-------FRLPSSLSKKKKKARVQA--------- 228
Query: 274 EAAIEGHQNSVVIDYILKILGLDVCADTMVGDQMIRGISGGEKKRLTTGEMLVGPIKVLF 333
++ LGL A T++GD+ RG+SGGE++R++ G ++ +LF
Sbjct: 229 ---------------LIDQLGLRNAAKTVIGDEGHRGVSGGERRRVSIGTDIIHDPIILF 273
Query: 334 MDEISNGLDSSTTFQIINSIKQSIHILNGTALVSLLQPAPETYELFDDIILLTDGQIVYQ 393
+DE ++GLDS++ + ++ + Q I ++S+ QP+ L D +I L+ G VY
Sbjct: 274 LDEPTSGLDSTSAYMVV-KVLQRIAQSGSIVIMSIHQPSYRILGLLDKLIFLSRGNTVYS 332
Query: 394 GPREYVLEFFESTGFKCPERKGVADFLQEVTSRKDQWQYWAREDEPYNFVTVKDFARAFE 453
G ++ +FF G PE + +F ++ ED P ++ +F + +
Sbjct: 333 GSPTHLPQFFSEFGHPIPENENKPEFALDLIRE--------LEDSPEGTKSLVEFHKQWR 384
Query: 454 LFHIGKQ--------LGEELADPFDKSKFHSNVLITKKYGINKKELLRACAS---RELLL 502
Q L + ++ + K S N + + A+ E+L+
Sbjct: 385 AKQTSSQSRRNTNVSLKDAISASISRGKLVSG-------ATNLRSSFQTFANPFWTEMLV 437
Query: 503 MKRNSFVYIFKATQLTYL---ATLTTTLFLRTKMY--HSTIEDAQTYMGALFFTVTVAMF 557
+ + S + + +L + A L T + L T + ++ Q +G F ++ +
Sbjct: 438 IGKRSILNSRRQPELFGIRLGAVLVTGMILATIFWKLDNSPRGIQERLGFFAFAMSTTFY 497
Query: 558 NGISELNMTIMKLPIFYKQRDLLFYPSWAYSLPPWILKIPITIIEVAIWECISYYAIGFD 617
+ + + + IF ++ Y +Y L I+ IP II A + ++ A+G
Sbjct: 498 TCAEAIPVFLQERYIFMRETAYNAYRRSSYVLAHTIISIPALIILSAAFAASTFSAVGLA 557
Query: 618 PNIGRFFKQSLVVLCINQMASALFRFMAALGRDIVVANTFGTFSLLAVTVLGGFVISRED 677
F +L S+ F++ + +++ T L + GF ISR+
Sbjct: 558 GGSEGFLFFFFTILTAFWAGSSFVTFLSGVVSHVMIGFTVVVAILAYFLLFSGFFISRDR 617
Query: 678 VHKWFLWGYWSSPLMYGQNAIAVNEF 703
+ +++W ++ S + Y + NEF
Sbjct: 618 IPLYWIWFHYLSLVKYPYEGVLQNEF 643
>At2g01320 putative ABC transporter
Length = 725
Score = 199 bits (507), Expect = 7e-51
Identities = 147/531 (27%), Positives = 255/531 (47%), Gaps = 35/531 (6%)
Query: 854 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAG---IKTSGYIEGNIKVSGYQKNQKSF 910
LLK VSG +PG L A+MG SG+GKTTL++VLAG + ++ G ++V+G + K++
Sbjct: 90 LLKNVSGEAKPGRLLAIMGPSGSGKTTLLNVLAGQLSLSPRLHLSGLLEVNGKPSSSKAY 149
Query: 911 ARISGYCEQFDIHSPNVTVYESLLYSAWLRLSPEVDHAT-RKMFIEEVMELVELNSLREA 969
+ Q D+ +TV E+L ++A L+L PE+ A R ++ ++ + L S ++
Sbjct: 150 KL--AFVRQEDLFFSQLTVRETLSFAAELQL-PEISSAEERDEYVNNLLLKLGLVSCADS 206
Query: 970 LVGLPGENGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAIVMRTVRNTVDTGR 1029
VG G+S ++KRL++A EL+A+PS+IF DEPT+GLDA A VM T++ G
Sbjct: 207 CVGDAKVRGISGGEKKRLSLACELIASPSVIFADEPTTGLDAFQAEKVMETLQKLAQDGH 266
Query: 1030 TVVCTIHQPSIDIFDSFDELLLLKLGGEQIYAGPIGNQCSDLIQYFEAIQGVPTIKDGYN 1089
TV+C+IHQP ++ FD+++LL G +YAGP G + F + + N
Sbjct: 267 TVICSIHQPRGSVYAKFDDIVLL-TEGTLVYAGPAGKEPLTYFGNFGFL-----CPEHVN 320
Query: 1090 PATWMLE-----------ITSAGKEANLKVNFTDVYKNSELHRRNKQLIQELS---VPSQ 1135
PA ++ + + S+ K + V+ +S L+ + +E P +
Sbjct: 321 PAEFLADLISVDYSSSETVYSSQKRVHALVDAFSQRSSSVLYATPLSMKEETKNGMRPRR 380
Query: 1136 SSKDLHFDAQYSQTFLAQCTYCLWKQHLSYWRNTSYTAVRLLFTIMTGILFGLIFWGVGA 1195
+ D + Q FL L + + R+ VR ++ + ++FG +FW +G
Sbjct: 381 KAIVERTDGWWRQFFL-----LLKRAWMQASRDGPTNKVRARMSVASAVIFGSVFWRMG- 434
Query: 1196 KSKKEQDLFNAMGSMYAAVTFIGVVNGASVQPIVAIERTVFYRERAAGMYSAMPYALAQV 1255
K + + + MG + A + + ER + RER+ G YS PY L++
Sbjct: 435 --KSQTSIQDRMGLLQVAAINTAMAALTKTVGVFPKERAIVDRERSKGSYSLGPYLLSKT 492
Query: 1256 IIELPHILVQAVVYGIIVYAMMGFEWTASKVLWNLFFTYFSFLYYTYYGMMTMAITPNPH 1315
I E+P +++G ++Y M T S+ + G+ A+ P+
Sbjct: 493 IAEIPIGAAFPLMFGAVLYPMARLNPTLSRFGKFCGIVTVESFAASAMGLTVGAMVPSTE 552
Query: 1316 VAGILSTSFYAIWCLFSGFIIPLSRIPIWWKWYYWICPVAWTLNGLVTSQY 1366
A + S ++ +F G+ + PI ++W + W GL +++
Sbjct: 553 AAMAVGPSLMTVFIVFGGYYVNADNTPIIFRWIPRASLIRWAFQGLCINEF 603
Score = 165 bits (418), Expect = 2e-40
Identities = 162/572 (28%), Positives = 251/572 (43%), Gaps = 62/572 (10%)
Query: 147 SPKKQLHILQNVSGILKPRRMTLLLGPPGSGKTTLLLALAGILGKD--LKQSGRVTYNGK 204
S K +L+NVSG KP R+ ++GP GSGKTTLL LAG L L SG + NGK
Sbjct: 83 SSKSVRFLLKNVSGEAKPGRLLAIMGPSGSGKTTLLNVLAGQLSLSPRLHLSGLLEVNGK 142
Query: 205 GLEEFVPQRTSAY----VSQYDNHIGEMTVRETLAFSARCQGVGQNYEMLTELLRKEKES 260
P + AY V Q D ++TVRETL+F+A EL E S
Sbjct: 143 ------PSSSKAYKLAFVRQEDLFFSQLTVRETLSFAA-------------ELQLPEISS 183
Query: 261 KIEPDPDINAYMKEAAIEGHQNSVVIDYILKILGLDVCADTMVGDQMIRGISGGEKKRLT 320
E D +N + +LK LGL CAD+ VGD +RGISGGEKKRL+
Sbjct: 184 AEERDEYVN-----------------NLLLK-LGLVSCADSCVGDAKVRGISGGEKKRLS 225
Query: 321 TGEMLVGPIKVLFMDEISNGLDSSTTFQIINSIKQSIHILNGTALVSLLQPAPETYELFD 380
L+ V+F DE + GLD+ +++ ++ Q + T + S+ QP Y FD
Sbjct: 226 LACELIASPSVIFADEPTTGLDAFQAEKVMETL-QKLAQDGHTVICSIHQPRGSVYAKFD 284
Query: 381 DIILLTDGQIVYQGPR-EYVLEFFESTGFKCPERKGVADFLQEVTS--RKDQWQYWARED 437
DI+LLT+G +VY GP + L +F + GF CPE A+FL ++ S ++ +
Sbjct: 285 DIVLLTEGTLVYAGPAGKEPLTYFGNFGFLCPEHVNPAEFLADLISVDYSSSETVYSSQK 344
Query: 438 EPYNFVTVKDFARAFELFHIGKQLGEELADPFDKSKFHSNVLITKKYGINKKELLRACAS 497
+ V + L+ + EE + + ++ + G ++ L
Sbjct: 345 RVHALVDAFSQRSSSVLYATPLSMKEETKNGMRPRR---KAIVERTDGWWRQFFL--LLK 399
Query: 498 RELLLMKRNSFVYIFKATQLTYLATLTTTLFLRTKMYHSTIEDAQTYMGALFFTVTVAMF 557
R + R+ +A A + ++F R ++I+D MG L A+
Sbjct: 400 RAWMQASRDGPTNKVRARMSVASAVIFGSVFWRMGKSQTSIQDR---MGLL---QVAAIN 453
Query: 558 NGISELNMTIMKLP----IFYKQRDLLFYPSWAYSLPPWILKIPITIIEVAIWECISYYA 613
++ L T+ P I ++R Y Y L I +IPI ++ + Y
Sbjct: 454 TAMAALTKTVGVFPKERAIVDRERSKGSYSLGPYLLSKTIAEIPIGAAFPLMFGAVLYPM 513
Query: 614 IGFDPNIGRFFKQSLVVLCINQMASALFRFMAALGRDIVVANTFGTFSLLAVTVLGGFVI 673
+P + RF K +V + ASA+ + A+ A G + V GG+ +
Sbjct: 514 ARLNPTLSRFGKFCGIVTVESFAASAMGLTVGAMVPSTEAAMAVGPSLMTVFIVFGGYYV 573
Query: 674 SREDVHKWFLWGYWSSPLMYGQNAIAVNEFLG 705
+ ++ F W +S + + + +NEF G
Sbjct: 574 NADNTPIIFRWIPRASLIRWAFQGLCINEFSG 605
>At1g17840 putative ABC transporter (At1g17840)
Length = 703
Score = 193 bits (490), Expect = 7e-49
Identities = 172/630 (27%), Positives = 293/630 (46%), Gaps = 69/630 (10%)
Query: 827 LTFDDITYSVDMPQGMKNQGVTEDRLELLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLA 886
LT+ D+T V M G + +L+G++G PG LTALMG SG+GK+T++D LA
Sbjct: 50 LTWQDLTVMVTMGDG--------ETQNVLEGLTGYAEPGSLTALMGPSGSGKSTMLDALA 101
Query: 887 G-IKTSGYIEGNIKVSGYQKNQKSFARISGYCEQFDIHSPNVTVYESLLYSAWLRLSPEV 945
+ + ++ G + ++G +K + SF + Y Q D +TV E++ YSA +RL ++
Sbjct: 102 SRLAANAFLSGTVLLNG-RKTKLSFGT-AAYVTQDDNLIGTLTVRETIWYSARVRLPDKM 159
Query: 946 DHATRKMFIEEVMELVELNSLREALVGLPGENGLSTEQRKRLTIAVELVANPSIIFMDEP 1005
+ ++ +E + + L + ++G G+S +++R++IA+E++ P ++F+DEP
Sbjct: 160 LRSEKRALVERTIIEMGLQDCADTVIGNWHLRGISGGEKRRVSIALEILMRPRLLFLDEP 219
Query: 1006 TSGLDARAAAIVMRTVRNTVDTGRTVVCTIHQPSIDIFDSFDELLLLKLGGEQIYAGPIG 1065
TSGLD+ +A V +T+R GRTV+ +IHQPS ++F+ FD L LL GG+ +Y G
Sbjct: 220 TSGLDSASAFFVTQTLRALSRDGRTVIASIHQPSSEVFELFDRLYLLS-GGKTVYFG--- 275
Query: 1066 NQCSDLIQYF-EAIQGVPTIKDGYNPATWMLEITS-------AGKEANLKVNF------- 1110
Q SD ++F +A P ++ NP+ L + A + ++K+ F
Sbjct: 276 -QASDAYEFFAQAGFPCPALR---NPSDHFLRCINSDFDKVRATLKGSMKLRFEASDDPL 331
Query: 1111 ------------TDVYKNSELHRRNKQLIQELSVPSQSSKDLHFDAQYSQTFLAQCTYCL 1158
D Y S+ + K ++E+S K D+ SQ TY L
Sbjct: 332 EKITTAEAIRLLVDYYHTSDYYYTAKAKVEEIS----QFKGTILDSGGSQASFLLQTYTL 387
Query: 1159 WKQ-HLSYWRNTSYTAVRLLFTIMTGILFGLIFWGVGAKSKKEQDLFNAMGSMYAAVTFI 1217
K+ ++ R+ Y +RLL I+ + G I+ VG + ++ VTF+
Sbjct: 388 TKRSFINMSRDFGYYWLRLLIYILVTVCIGTIYLNVGTSYSAILARGSCASFVFGFVTFM 447
Query: 1218 GVVNGASVQPIVAIERTVFYRERAAGMYSAMPYALAQVIIELPHILVQAVVYGIIVYAMM 1277
+ P + VF RER G Y + +A + P +++ + G I Y M+
Sbjct: 448 SI----GGFPSFVEDMKVFQRERLNGHYGVAAFVIANTLSATPFLIMITFISGTICYFMV 503
Query: 1278 GFEWTASKVLWNLFFTYFSFLYYTYYGMMTMAITPNPHVAGILSTSFYAIWCLFSGFIIP 1337
G + L+ + Y S M +I PN + I+ I+ L SGF
Sbjct: 504 GLHPGFTHYLFFVLCLYASVTVVESLMMAIASIVPNFLMGIIIGAGIQGIFMLVSGFFRL 563
Query: 1338 LSRI--PIWWKWYYWICPVAWTLNGLVTSQYGHNMD--TLDNGQSV-----EEFVRNYFG 1388
+ I P W +I W L G QY +++ T D+ S E + N F
Sbjct: 564 PNDIPKPFWRYPMSYISFHFWALQG----QYQNDLRGLTFDSQGSAFKIPGEYVLENVFQ 619
Query: 1389 FE-YDFLGVVAIVVVSFSVLFALIFTFGIK 1417
+ + + V++S +++ +IF IK
Sbjct: 620 IDLHRSKWINLSVILSMIIIYRIIFFIMIK 649
Score = 168 bits (425), Expect = 2e-41
Identities = 162/642 (25%), Positives = 277/642 (42%), Gaps = 97/642 (15%)
Query: 153 HILQNVSGILKPRRMTLLLGPPGSGKTTLLLALAGILGKDLKQSGRVTYNGKGLEEFVPQ 212
++L+ ++G +P +T L+GP GSGK+T+L ALA L + SG V NG+ + +
Sbjct: 68 NVLEGLTGYAEPGSLTALMGPSGSGKSTMLDALASRLAANAFLSGTVLLNGRKTK--LSF 125
Query: 213 RTSAYVSQYDNHIGEMTVRETLAFSARCQGVGQNYEMLTELLRKEKESKIEPDPDINAYM 272
T+AYV+Q DN IG +TVRET+ +SAR + + ++LR EK + +E
Sbjct: 126 GTAAYVTQDDNLIGTLTVRETIWYSARVR-------LPDKMLRSEKRALVE--------- 169
Query: 273 KEAAIEGHQNSVVIDYILKILGLDVCADTMVGDQMIRGISGGEKKRLTTG-EMLVGPIKV 331
+I+ +GL CADT++G+ +RGISGGEK+R++ E+L+ P ++
Sbjct: 170 ----------RTIIE-----MGLQDCADTVIGNWHLRGISGGEKRRVSIALEILMRP-RL 213
Query: 332 LFMDEISNGLDSSTTFQIINSIKQSIHILNGTALVSLLQPAPETYELFDDIILLTDGQIV 391
LF+DE ++GLDS++ F + +++ ++ T + S+ QP+ E +ELFD + LL+ G+ V
Sbjct: 214 LFLDEPTSGLDSASAFFVTQTLR-ALSRDGRTVIASIHQPSSEVFELFDRLYLLSGGKTV 272
Query: 392 YQGPREYVLEFFESTGFKCPERKGVAD-FLQEVTSRKDQWQ---------YWAREDEPYN 441
Y G EFF GF CP + +D FL+ + S D+ + + D+P
Sbjct: 273 YFGQASDAYEFFAQAGFPCPALRNPSDHFLRCINSDFDKVRATLKGSMKLRFEASDDPLE 332
Query: 442 FVTVKDFARAF-ELFHIGKQLGEELADPFDKSKFHSNVLITKKYGINKKELLR--ACASR 498
+T + R + +H A + S+F +L G LL+ R
Sbjct: 333 KITTAEAIRLLVDYYHTSDYYYTAKAKVEEISQFKGTIL--DSGGSQASFLLQTYTLTKR 390
Query: 499 ELLLMKRNSFVYIFKATQLTYLATLTTTLFLRTKMYHSTIEDAQTYMGALFFTVTVAMFN 558
+ M R+ Y + + T++L +S I + +F VT
Sbjct: 391 SFINMSRDFGYYWLRLLIYILVTVCIGTIYLNVGTSYSAILARGSCASFVFGFVTFMSIG 450
Query: 559 GISELNMTIMKLPIFYKQRDLLFYPSWAYSLPPWILKIPITIIEVAIWECISYYAIGFDP 618
G + + +F ++R Y A+ + + P I+ I I Y+ +G P
Sbjct: 451 GFPSF---VEDMKVFQRERLNGHYGVAAFVIANTLSATPFLIMITFISGTICYFMVGLHP 507
Query: 619 NIGRFFKQSLVVLCINQMASALFRFMAALGR--------DIVVANTFGTFSLLAVTVLGG 670
+ VLC+ + + M A+ I+ A G F L++ G
Sbjct: 508 GFTHYL---FFVLCLYASVTVVESLMMAIASIVPNFLMGIIIGAGIQGIFMLVS-----G 559
Query: 671 FVISREDVHKWFLWGYWSSPLMYGQNAIAVNEFLGHGWRKVAPNSNETLGVSILKSRGFF 730
F D+ K F W P+ Y H W N+ G++ F
Sbjct: 560 FFRLPNDIPKPF----WRYPMSY---------ISFHFWALQGQYQNDLRGLTFDSQGSAF 606
Query: 731 PQAYWY--------------WIGVGALIGYVFLFNFLFALAL 758
Y WI + ++ + ++ +F + +
Sbjct: 607 KIPGEYVLENVFQIDLHRSKWINLSVILSMIIIYRIIFFIMI 648
>At5g06530 ABC transporter like protein
Length = 627
Score = 191 bits (485), Expect = 3e-48
Identities = 168/636 (26%), Positives = 292/636 (45%), Gaps = 64/636 (10%)
Query: 792 QQENSSNTKMDEEVSENKASSSGRKGMVLPFQPLSLTFDDITYSVDMPQGMKNQGVTEDR 851
QQ N + + D E + K K P P+ L F D+TY V + + +
Sbjct: 3 QQCNVTVSAEDIEAGKKKP-----KFQAEPTLPIFLKFRDVTYKVVI-----KKLTSSVE 52
Query: 852 LELLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGIKTSGYIEGNIKVSG--YQKNQKS 909
E+L G+SG+ PG + ALMG SG+GKTTL+ +LAG + G++ + Y K KS
Sbjct: 53 KEILTGISGSVNPGEVLALMGPSGSGKTTLLSLLAGRISQSSTGGSVTYNDKPYSKYLKS 112
Query: 910 FARISGYCEQFDIHSPNVTVYESLLYSAWLRLSPEVDHATRKMFIEEVMELVELNSLREA 969
G+ Q D+ P++TV E+L Y+A LRL + +K +V++ + L ++
Sbjct: 113 KI---GFVTQDDVLFPHLTVKETLTYAARLRLPKTLTREQKKQRALDVIQELGLERCQDT 169
Query: 970 LVGLPGENGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAIVMRTVRNTVDTGR 1029
++G G+S +RKR++I E++ NPS++ +DEPTSGLD+ A + + + + G+
Sbjct: 170 MIGGAFVRGVSGGERKRVSIGNEIIINPSLLLLDEPTSGLDSTTALRTILMLHDIAEAGK 229
Query: 1030 TVVCTIHQPSIDIFDSFDELLLLKLGGEQIYAGPIGNQCSDLIQYFEAIQGVPTIKDGYN 1089
TV+ TIHQPS +F FD+L+LL G +Y G + S+ + YF +I P I N
Sbjct: 230 TVITTIHQPSSRLFHRFDKLILLG-RGSLLYFG----KSSEALDYFSSIGCSPLI--AMN 282
Query: 1090 PATWMLEI--------------------------TSAGKEANLKVN--FTDVYKNSELHR 1121
PA ++L++ T GK + V+ + Y+ +
Sbjct: 283 PAEFLLDLANGNINDISVPSELDDRVQVGNSGRETQTGKPSPAAVHEYLVEAYETRVAEQ 342
Query: 1122 RNKQLIQELSVPSQS-SKDLHFDAQYSQTFLAQCTYC-LWKQHLSYWRNTSYTAVRLLFT 1179
K+L+ + + ++ +K Q+ + Q YC L+ + L R+ ++ +R+
Sbjct: 343 EKKKLLDPVPLDEEAKAKSTRLKRQWGTCWWEQ--YCILFCRGLKERRHEYFSWLRVTQV 400
Query: 1180 IMTGILFGLIFWGVGAKSKKEQDLFNAMGSMYAAVTFIGVVNGASVQPIVAIERTVFYRE 1239
+ T ++ GL++W + + L + G ++ F G + ER + +E
Sbjct: 401 LSTAVILGLLWW--QSDIRTPMGLQDQAGLLFFIAVFWGFFPVFTAIFAFPQERAMLNKE 458
Query: 1240 RAAGMYSAMPYALAQVIIELPHILVQAVVYGIIVYAMMGFEWTASKVLWNLFFTYFSFLY 1299
RAA MY Y LA+ +LP + ++ ++VY M G + ++ + +
Sbjct: 459 RAADMYRLSAYFLARTTSDLPLDFILPSLFLLVVYFMTGLRISPYPFFLSMLTVFLCIIA 518
Query: 1300 YTYYGMMTMAITPNPHVAGILSTSFYAIWCLFSGFIIPLSRIPIWWKWYYWICPVAWTLN 1359
G+ AI + A L++ + L GF + ++P++ W ++ T
Sbjct: 519 AQGLGLAIGAILMDLKKATTLASVTVMTFMLAGGFFV--KKVPVFISWIRYLSFNYHTYK 576
Query: 1360 GLVTSQY-----GHNMDTLDNG-QSVEEFVRNYFGF 1389
L+ QY N +DNG V V FG+
Sbjct: 577 LLLKVQYQDFAVSINGMRIDNGLTEVAALVVMIFGY 612
Score = 145 bits (367), Expect = 1e-34
Identities = 139/547 (25%), Positives = 244/547 (44%), Gaps = 63/547 (11%)
Query: 154 ILQNVSGILKPRRMTLLLGPPGSGKTTLLLALAGILGKDLKQSGRVTYNGKGLEEFVPQR 213
IL +SG + P + L+GP GSGKTTLL LAG + + G VTYN K +++ +
Sbjct: 55 ILTGISGSVNPGEVLALMGPSGSGKTTLLSLLAGRISQS-STGGSVTYNDKPYSKYLKSK 113
Query: 214 TSAYVSQYDNHIGEMTVRETLAFSARCQGVGQNYEMLTELLRKEKESKIEPDPDINAYMK 273
+V+Q D +TV+ETL ++AR + L + L +E++ K
Sbjct: 114 IG-FVTQDDVLFPHLTVKETLTYAARLR--------LPKTLTREQK-------------K 151
Query: 274 EAAIEGHQNSVVIDYILKILGLDVCADTMVGDQMIRGISGGEKKRLTTGEMLVGPIKVLF 333
+ A++ +++ LGL+ C DTM+G +RG+SGGE+KR++ G ++ +L
Sbjct: 152 QRALD----------VIQELGLERCQDTMIGGAFVRGVSGGERKRVSIGNEIIINPSLLL 201
Query: 334 MDEISNGLDSSTTFQIINSIKQSIHILNGTALVSLLQPAPETYELFDDIILLTDGQIVYQ 393
+DE ++GLDS+T + I + I T + ++ QP+ + FD +ILL G ++Y
Sbjct: 202 LDEPTSGLDSTTALRTILML-HDIAEAGKTVITTIHQPSSRLFHRFDKLILLGRGSLLYF 260
Query: 394 GPREYVLEFFESTGFKCPERKGVADFLQEVTSRK-----------DQWQYW-----ARED 437
G L++F S G A+FL ++ + D+ Q +
Sbjct: 261 GKSSEALDYFSSIGCSPLIAMNPAEFLLDLANGNINDISVPSELDDRVQVGNSGRETQTG 320
Query: 438 EPYNFVTVKDFARAFELFHIGKQLGEELAD--PFDKSKFHSNVLITKKYGINKKELLRAC 495
+P + A+E + +Q ++L D P D+ + + +++G E
Sbjct: 321 KPSPAAVHEYLVEAYET-RVAEQEKKKLLDPVPLDEEAKAKSTRLKRQWGTCWWEQYCIL 379
Query: 496 ASRELLLMKRNSFVYIFKATQLTYLATLTTTLFLRTKMYHSTIEDAQTYMGALFFTVTVA 555
R L + F ++ + TQ+ A + L+ ++ + T Q G LFF +A
Sbjct: 380 FCRGLKERRHEYFSWL-RVTQVLSTAVILGLLWWQSDI--RTPMGLQDQAGLLFF---IA 433
Query: 556 MFNGISELNMTIMKLP----IFYKQRDLLFYPSWAYSLPPWILKIPITIIEVAIWECISY 611
+F G + I P + K+R Y AY L +P+ I +++ + Y
Sbjct: 434 VFWGFFPVFTAIFAFPQERAMLNKERAADMYRLSAYFLARTTSDLPLDFILPSLFLLVVY 493
Query: 612 YAIGFDPNIGRFFKQSLVVLCINQMASALFRFMAALGRDIVVANTFGTFSLLAVTVLGGF 671
+ G + FF L V A L + A+ D+ A T + +++ + GGF
Sbjct: 494 FMTGLRISPYPFFLSMLTVFLCIIAAQGLGLAIGAILMDLKKATTLASVTVMTFMLAGGF 553
Query: 672 VISREDV 678
+ + V
Sbjct: 554 FVKKVPV 560
>At3g21090 ABC transporter, putative
Length = 680
Score = 188 bits (478), Expect = 2e-47
Identities = 173/645 (26%), Positives = 296/645 (45%), Gaps = 71/645 (11%)
Query: 807 ENKASSSGRKGMVLPFQPLS---LTFDDITYSVDMPQGMKNQGVTEDRLELLKGVSGAFR 863
E + SSSGR+ + + L ++D+T V +P + G T LL+ ++G
Sbjct: 2 ELEGSSSGRRQLPSKLEMSRGAYLAWEDLT--VVIPNF--SDGPTR---RLLQRLNGYAE 54
Query: 864 PGVLTALMGVSGAGKTTLMDVLAG-IKTSGYIEGNIKVSGYQKNQKSFARISGYCEQFDI 922
PG + A+MG SG+GK+TL+D LAG + + + GN+ ++G K + + Y Q D+
Sbjct: 55 PGRIMAIMGPSGSGKSTLLDSLAGRLARNVVMTGNLLLNG--KKARLDYGLVAYVTQEDV 112
Query: 923 HSPNVTVYESLLYSAWLRLSPEVDHATRKMFIEEVMELVELNSLREALVGLPGENGLSTE 982
+TV E++ YSA LRL ++ +E + + L + ++G G+S
Sbjct: 113 LLGTLTVRETITYSAHLRLPSDMSKEEVSDIVEGTIMELGLQDCSDRVIGNWHARGVSGG 172
Query: 983 QRKRLTIAVELVANPSIIFMDEPTSGLDARAAAIVMRTVRNTVDTGRTVVCTIHQPSIDI 1042
+RKR++IA+E++ P I+F+DEPTSGLD+ +A V++ +RN GRTV+ ++HQPS ++
Sbjct: 173 ERKRVSIALEILTRPQILFLDEPTSGLDSASAFFVIQALRNIARDGRTVISSVHQPSSEV 232
Query: 1043 FDSFDELLLLKLGGEQIYAGPIGNQCSDLIQYFEAIQGVPTIKDGYNPATWMLE------ 1096
F FD+L LL GE +Y G + +++F A G P K NP+ L
Sbjct: 233 FALFDDLFLLS-SGESVYFG----EAKSAVEFF-AESGFPCPKK-RNPSDHFLRCINSDF 285
Query: 1097 --ITSAGKEAN-------------------LKVNFTDVYKNSELHRRNKQLIQELSVPSQ 1135
+T+ K + +K + YK S+ + K I+ELS +
Sbjct: 286 DTVTATLKGSQRIQETPATSDPLMNLATSVIKARLVENYKRSKYAKSAKSRIRELS--NI 343
Query: 1136 SSKDLHFDAQYSQTFLAQCTYCLWKQHLSYWRNTSYTAVRLLFTIMTGILFGLIFWGVGA 1195
++ T+ Q + ++ R+ Y R++ I+ I G IF+ VG
Sbjct: 344 EGLEMEIRKGSEATWWKQLRTLTARSFINMCRDVGYYWTRIISYIVVSISVGTIFYDVGY 403
Query: 1196 KSKKEQDLFNAMGSMYAAVTFIGVVNGASVQPIVAIERTVFYRERAAGMYSAMPYALAQV 1255
+ G + +TF+ + P E VFY+ER +G Y Y L+
Sbjct: 404 SYTSILARVSCGGFITGFMTFMSI----GGFPSFLEEMKVFYKERLSGYYGVSVYILSNY 459
Query: 1256 IIELPHILVQAVVYGIIVYAMMGFEWTASKVLWNLFFTYFSFLYYTYYGMMTMAITPNPH 1315
I P ++ +V+ G I Y ++ F S + +FS M+ ++ PN
Sbjct: 460 ISSFPFLVAISVITGTITYNLVKFRPGFSHYAFFCLNIFFSVSVIESLMMVVASVVPNFL 519
Query: 1316 VAGILSTSFYAIWCLFSGFIIPLSRIP-IWWKWYYWICPVAWTLNGLVTSQYGHNMDTLD 1374
+ I I + SGF L +P I+W++ PV++ G Q+ + L
Sbjct: 520 MGLITGAGLIGIIMMTSGFFRLLPDLPKIFWRY-----PVSYISYGSWAIQF----EPLF 570
Query: 1375 NGQ---SVEEFVRNYFGFEYDF-----LGVVAIVVVSFSVLFALI 1411
G+ + EE + FG + + L V ++V + +LF ++
Sbjct: 571 PGEPKMTGEEVIEKVFGVKVTYSKWWDLAAVVAILVCYRLLFFVV 615
Score = 172 bits (435), Expect = 2e-42
Identities = 168/677 (24%), Positives = 298/677 (43%), Gaps = 70/677 (10%)
Query: 120 GRRALPTLFNFFVNVIEGCLNNLQIIPS----PKKQLHILQNVSGILKPRRMTLLLGPPG 175
GRR LP+ + +IP+ P ++L LQ ++G +P R+ ++GP G
Sbjct: 9 GRRQLPSKLEMSRGAYLAWEDLTVVIPNFSDGPTRRL--LQRLNGYAEPGRIMAIMGPSG 66
Query: 176 SGKTTLLLALAGILGKDLKQSGRVTYNGKGLEEFVPQRTSAYVSQYDNHIGEMTVRETLA 235
SGK+TLL +LAG L +++ +G + NGK + + AYV+Q D +G +TVRET+
Sbjct: 67 SGKSTLLDSLAGRLARNVVMTGNLLLNGK--KARLDYGLVAYVTQEDVLLGTLTVRETIT 124
Query: 236 FSARCQGVGQNYEMLTELLRKEKESKIEPDPDINAYMKEAAIEGHQNSVVIDYILKILGL 295
+SA + L + KE+ S I +EG + LGL
Sbjct: 125 YSAHLR--------LPSDMSKEEVSDI--------------VEG---------TIMELGL 153
Query: 296 DVCADTMVGDQMIRGISGGEKKRLTTGEMLVGPIKVLFMDEISNGLDSSTTFQIINSIKQ 355
C+D ++G+ RG+SGGE+KR++ ++ ++LF+DE ++GLDS++ F +I +++
Sbjct: 154 QDCSDRVIGNWHARGVSGGERKRVSIALEILTRPQILFLDEPTSGLDSASAFFVIQALR- 212
Query: 356 SIHILNGTALVSLLQPAPETYELFDDIILLTDGQIVYQGPREYVLEFFESTGFKCPERKG 415
+I T + S+ QP+ E + LFDD+ LL+ G+ VY G + +EFF +GF CP+++
Sbjct: 213 NIARDGRTVISSVHQPSSEVFALFDDLFLLSSGESVYFGEAKSAVEFFAESGFPCPKKRN 272
Query: 416 VAD-FLQEVTSRKDQ-----------WQYWAREDEPYNFVTVKDFARAFELFHIGKQLGE 463
+D FL+ + S D + A D N T AR E + K
Sbjct: 273 PSDHFLRCINSDFDTVTATLKGSQRIQETPATSDPLMNLATSVIKARLVENYKRSKYAKS 332
Query: 464 ELADPFDKSKFHS-NVLITKKYGINKKELLRACASRELLLMKRNSFVYIFKATQLTYLAT 522
+ + S + I K + LR +R + M R+ Y + ++
Sbjct: 333 AKSRIRELSNIEGLEMEIRKGSEATWWKQLRTLTARSFINMCRDVGYYWTRIISYIVVSI 392
Query: 523 LTTTLFLRTKMYHSTIEDAQTYMGALFFTVTVAMFNGISELNMTIMKLPIFYKQRDLLFY 582
T+F +++I + G F F I + ++ +FYK+R +Y
Sbjct: 393 SVGTIFYDVGYSYTSILARVSCGG---FITGFMTFMSIGGFPSFLEEMKVFYKERLSGYY 449
Query: 583 PSWAYSLPPWILKIPITIIEVAIWECISYYAIGFDPNIGRFFKQSLVVLCINQMASALFR 642
Y L +I P + I I+Y + F P + L + + +L
Sbjct: 450 GVSVYILSNYISSFPFLVAISVITGTITYNLVKFRPGFSHYAFFCLNIFFSVSVIESLMM 509
Query: 643 FMAALGRDIVVANTFGTFSLLAVTVLGGFVISREDVHKWFLWGYWSSPLMYGQNAIAVNE 702
+A++ + ++ G + + + GF D+ K F W Y S + YG AI
Sbjct: 510 VVASVVPNFLMGLITGAGLIGIIMMTSGFFRLLPDLPKIF-WRYPVSYISYGSWAIQ--- 565
Query: 703 FLGHGWRKVAPNSNETLGVSILKSRGFFPQAYWYWIGVGALIGYVFLFNFLFALALHFLS 762
+ + P + G +++ Y W + A++ + + LF + L
Sbjct: 566 -----FEPLFPGEPKMTGEEVIEKVFGVKVTYSKWWDLAAVVAILVCYRLLFFVVLKL-- 618
Query: 763 PFRKDQAGLSQEKLQER 779
+++AG + + +Q +
Sbjct: 619 ---RERAGPALKAIQAK 632
>At1g51500 ATP-dependent transmembrane transporter, putative
Length = 687
Score = 187 bits (474), Expect = 5e-47
Identities = 159/597 (26%), Positives = 276/597 (45%), Gaps = 52/597 (8%)
Query: 854 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAG-IKTSGYIEGNIKVSGYQKNQKSFAR 912
LL G++G PG + A+MG SG+GK+TL+D LAG + + + GN+ ++G K +
Sbjct: 44 LLDGLNGHAEPGRIMAIMGPSGSGKSTLLDSLAGRLARNVIMTGNLLLNG--KKARLDYG 101
Query: 913 ISGYCEQFDIHSPNVTVYESLLYSAWLRLSPEVDHATRKMFIEEVMELVELNSLREALVG 972
+ Y Q DI +TV E++ YSA LRLS ++ +E + + L + ++G
Sbjct: 102 LVAYVTQEDILMGTLTVRETITYSAHLRLSSDLTKEEVNDIVEGTIIELGLQDCADRVIG 161
Query: 973 LPGENGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAIVMRTVRNTV-DTGRTV 1031
G+S +RKR+++A+E++ P I+F+DEPTSGLD+ +A V++ +RN D GRTV
Sbjct: 162 NWHSRGVSGGERKRVSVALEILTRPQILFLDEPTSGLDSASAFFVIQALRNIARDGGRTV 221
Query: 1032 VCTIHQPSIDIFDSFDELLLLKLGGEQIYAG----------PIGNQC------------- 1068
V +IHQPS ++F FD+L LL GE +Y G G C
Sbjct: 222 VSSIHQPSSEVFALFDDLFLLS-SGETVYFGESKFAVEFFAEAGFPCPKKRNPSDHFLRC 280
Query: 1069 --SDLIQYFEAIQGVPTIKDGYNPATWMLEITSAGKEANLKVNFTDVYKNSELHRRNKQL 1126
SD ++G I++ PAT + A E +K + Y+ S + K
Sbjct: 281 INSDFDTVTATLKGSQRIRE--TPATSDPLMNLATSE--IKARLVENYRRSVYAKSAKSR 336
Query: 1127 IQEL-SVPSQSSKDLHFDAQYSQTFLAQCTYCLWKQHLSYWRNTSYTAVRLLFTIMTGIL 1185
I+EL S+ ++ ++ T+ Q + ++ R+ Y R++ I+
Sbjct: 337 IRELASIEGHHGMEVRKGSE--ATWFKQLRTLTKRSFVNMCRDIGYYWSRIVIYIVVSFC 394
Query: 1186 FGLIFWGVGAKSKKEQDLFNAMGSMYAAVTFIGVVNGASVQPIVAIERTVFYRERAAGMY 1245
G IF+ VG + G + +TF+ + P E VFY+ER +G Y
Sbjct: 395 VGTIFYDVGHSYTSILARVSCGGFITGFMTFMSI----GGFPSFIEEMKVFYKERLSGYY 450
Query: 1246 SAMPYALAQVIIELPHILVQAVVYGIIVYAMMGFEWTASKVLWNLFFTYFSFLYYTYYGM 1305
Y ++ + P ++ A++ G I Y M+ F S + +FS M
Sbjct: 451 GVSVYIISNYVSSFPFLVAIALITGSITYNMVKFRPGVSHWAFFCLNIFFSVSVIESLMM 510
Query: 1306 MTMAITPNPHVAGILSTSFYAIWCLFSGFIIPLSRIP-IWWKW-YYWICPVAWTLNGLVT 1363
+ ++ PN + I I + SGF L +P ++W++ ++ +W + G
Sbjct: 511 VVASLVPNFLMGLITGAGIIGIIMMTSGFFRLLPDLPKVFWRYPISFMSYGSWAIQGAYK 570
Query: 1364 SQY-GHNMDTLDNGQ---SVEEFVRNYFGFEYDF-----LGVVAIVVVSFSVLFALI 1411
+ + G D + G+ + E+ + FG + L + +++V + +LF ++
Sbjct: 571 NDFLGLEFDPMFAGEPKMTGEQVINKIFGVQVTHSKWWDLSAIVLILVCYRILFFIV 627
Score = 175 bits (443), Expect = 2e-43
Identities = 157/628 (25%), Positives = 279/628 (44%), Gaps = 66/628 (10%)
Query: 154 ILQNVSGILKPRRMTLLLGPPGSGKTTLLLALAGILGKDLKQSGRVTYNGKGLEEFVPQR 213
+L ++G +P R+ ++GP GSGK+TLL +LAG L +++ +G + NGK +
Sbjct: 44 LLDGLNGHAEPGRIMAIMGPSGSGKSTLLDSLAGRLARNVIMTGNLLLNGKKAR--LDYG 101
Query: 214 TSAYVSQYDNHIGEMTVRETLAFSARCQGVGQNYEMLTELLRKEKESKIEPDPDINAYMK 273
AYV+Q D +G +TVRET+ +SA + L+ L KE ++N ++
Sbjct: 102 LVAYVTQEDILMGTLTVRETITYSAHLR--------LSSDLTKE---------EVNDIVE 144
Query: 274 EAAIEGHQNSVVIDYILKILGLDVCADTMVGDQMIRGISGGEKKRLTTGEMLVGPIKVLF 333
IE LGL CAD ++G+ RG+SGGE+KR++ ++ ++LF
Sbjct: 145 GTIIE--------------LGLQDCADRVIGNWHSRGVSGGERKRVSVALEILTRPQILF 190
Query: 334 MDEISNGLDSSTTFQIINSIKQSIHILNGTALVSLLQPAPETYELFDDIILLTDGQIVYQ 393
+DE ++GLDS++ F +I +++ T + S+ QP+ E + LFDD+ LL+ G+ VY
Sbjct: 191 LDEPTSGLDSASAFFVIQALRNIARDGGRTVVSSIHQPSSEVFALFDDLFLLSSGETVYF 250
Query: 394 GPREYVLEFFESTGFKCPERKGVAD-FLQEVTSRKDQ-----------WQYWAREDEPYN 441
G ++ +EFF GF CP+++ +D FL+ + S D + A D N
Sbjct: 251 GESKFAVEFFAEAGFPCPKKRNPSDHFLRCINSDFDTVTATLKGSQRIRETPATSDPLMN 310
Query: 442 FVTVKDFARAFELFHIGKQLGEELADPFDKSKFHSNVLITKKYGINKKELLRACASRELL 501
T + AR E + + KS+ I +G+ ++ A ++L
Sbjct: 311 LATSEIKARLVENYR------RSVYAKSAKSRIRELASIEGHHGMEVRKGSEATWFKQLR 364
Query: 502 LMKRNSFV--------YIFKATQLTYLATLTTTLFLRTKMYHSTIEDAQTYMGALFFTVT 553
+ + SFV Y + ++ T+F +++I + G F
Sbjct: 365 TLTKRSFVNMCRDIGYYWSRIVIYIVVSFCVGTIFYDVGHSYTSILARVSCGG---FITG 421
Query: 554 VAMFNGISELNMTIMKLPIFYKQRDLLFYPSWAYSLPPWILKIPITIIEVAIWECISYYA 613
F I I ++ +FYK+R +Y Y + ++ P + I I+Y
Sbjct: 422 FMTFMSIGGFPSFIEEMKVFYKERLSGYYGVSVYIISNYVSSFPFLVAIALITGSITYNM 481
Query: 614 IGFDPNIGRFFKQSLVVLCINQMASALFRFMAALGRDIVVANTFGTFSLLAVTVLGGFVI 673
+ F P + + L + + +L +A+L + ++ G + + + GF
Sbjct: 482 VKFRPGVSHWAFFCLNIFFSVSVIESLMMVVASLVPNFLMGLITGAGIIGIIMMTSGFFR 541
Query: 674 SREDVHKWFLWGYWSSPLMYGQNAIA---VNEFLGHGWRKVAPNSNETLGVSILKSRGFF 730
D+ K F W Y S + YG AI N+FLG + + + G ++
Sbjct: 542 LLPDLPKVF-WRYPISFMSYGSWAIQGAYKNDFLGLEFDPMFAGEPKMTGEQVINKIFGV 600
Query: 731 PQAYWYWIGVGALIGYVFLFNFLFALAL 758
+ W + A++ + + LF + L
Sbjct: 601 QVTHSKWWDLSAIVLILVCYRILFFIVL 628
>At1g31770 unknown protein
Length = 648
Score = 187 bits (474), Expect = 5e-47
Identities = 147/561 (26%), Positives = 273/561 (48%), Gaps = 39/561 (6%)
Query: 813 SGRKGMVLPFQPLSLTFDDITYSVDMPQGMKNQGVTEDRLE-LLKGVSGAFRPGVLTALM 871
+ + G+ + P++L F+++ Y V + Q + G + + + +L G++G PG A++
Sbjct: 39 TSQPGLQMSMYPITLKFEEVVYKVKIEQTSQCMGSWKSKEKTILNGITGMVCPGEFLAML 98
Query: 872 GVSGAGKTTLMDVLAGIKTSGYIEGNIKVSGYQKNQKSFARISGYCEQFDIHSPNVTVYE 931
G SG+GKTTL+ L G + S G + +G Q R +G+ Q D+ P++TV+E
Sbjct: 99 GPSGSGKTTLLSALGG-RLSKTFSGKVMYNG-QPFSGCIKRRTGFVAQDDVLYPHLTVWE 156
Query: 932 SLLYSAWLRLSPEVDHATRKMFIEEVMELVELNSLREALVGLPGENGLSTEQRKRLTIAV 991
+L ++A LRL + + ++ V+ + LN +++G P G+S ++KR++I
Sbjct: 157 TLFFTALLRLPSSLTRDEKAEHVDRVIAELGLNRCTNSMIGGPLFRGISGGEKKRVSIGQ 216
Query: 992 ELVANPSIIFMDEPTSGLDARAAAIVMRTVRNTVDTGRTVVCTIHQPSIDIFDSFDELLL 1051
E++ NPS++ +DEPTSGLD+ A ++ T++ GRTVV TIHQPS I+ FD+++L
Sbjct: 217 EMLINPSLLLLDEPTSGLDSTTAHRIVTTIKRLASGGRTVVTTIHQPSSRIYHMFDKVVL 276
Query: 1052 LKLGGEQIYAGPIGNQCSDLIQYFEAIQGVPTIKDGYNPATWMLEI-----------TSA 1100
L G IY G S ++YF ++ ++ NPA +L++ TS
Sbjct: 277 LS-EGSPIYYG----AASSAVEYFSSLGFSTSLT--VNPADLLLDLANGIPPDTQKETSE 329
Query: 1101 GKEANLKVNFTDVYKNSELHRRNKQLIQELS----VPSQSSKDLHFDAQYSQTFLAQCTY 1156
++ +K Y+ + + +L S ++K+L + Q+ T+ Q T
Sbjct: 330 QEQKTVKETLVSAYEKNISTKLKAELCNAESHSYEYTKAAAKNLKSE-QWCTTWWYQFTV 388
Query: 1157 CLWKQHLSYWRNTSYTAVRLLFTIMTGILFGLIFWGVGAKSKKEQDLFNAMGSMYAAVTF 1216
L ++ + R S+ +R+ I L GL++W +++ S++ F
Sbjct: 389 LL-QRGVRERRFESFNKLRIFQVISVAFLGGLLWWHTPKSHIQDRTALLFFFSVFWG--F 445
Query: 1217 IGVVNGASVQPIVAIERTVFYRERAAGMYSAMPYALAQVIIELPHILVQAVVYGIIVYAM 1276
+ N P E+ + +ER++GMY Y +A+ + +LP L + I+Y M
Sbjct: 446 YPLYNAVFTFP---QEKRMLIKERSSGMYRLSSYFMARNVGDLPLELALPTAFVFIIYWM 502
Query: 1277 MGFEWTASKVLWNLFFTYFSFLYYTYYGMMTMAITPNPHVAGILSTSFYAIWCLFSGFII 1336
G + + + +L +S L G+ A+ N A L++ ++ + G+ +
Sbjct: 503 GGLKPDPTTFILSLLVVLYSVLVAQGLGLAFGALLMNIKQATTLASVTTLVFLIAGGYYV 562
Query: 1337 PLSRIP---IWWKW--YYWIC 1352
+IP +W K+ Y + C
Sbjct: 563 --QQIPPFIVWLKYLSYSYYC 581
Score = 156 bits (395), Expect = 7e-38
Identities = 147/550 (26%), Positives = 252/550 (45%), Gaps = 70/550 (12%)
Query: 150 KQLHILQNVSGILKPRRMTLLLGPPGSGKTTLLLALAGILGKDLKQSGRVTYNGKGLEEF 209
K+ IL ++G++ P +LGP GSGKTTLL AL G L K SG+V YNG+
Sbjct: 77 KEKTILNGITGMVCPGEFLAMLGPSGSGKTTLLSALGGRLSKTF--SGKVMYNGQPFSGC 134
Query: 210 VPQRTSAYVSQYDNHIGEMTVRETLAFSARCQGVGQNYEMLTELLRKEKESKIEPDPDIN 269
+ +RT +V+Q D +TV ETL F+A + + + L R EK
Sbjct: 135 IKRRT-GFVAQDDVLYPHLTVWETLFFTALLR-------LPSSLTRDEKAEH-------- 178
Query: 270 AYMKEAAIEGHQNSVVIDYILKILGLDVCADTMVGDQMIRGISGGEKKRLTTG-EMLVGP 328
+D ++ LGL+ C ++M+G + RGISGGEKKR++ G EML+ P
Sbjct: 179 ----------------VDRVIAELGLNRCTNSMIGGPLFRGISGGEKKRVSIGQEMLINP 222
Query: 329 IKVLFMDEISNGLDSSTTFQIINSIKQSIHILNG--TALVSLLQPAPETYELFDDIILLT 386
+L +DE ++GLDS+T +I+ +IK+ + +G T + ++ QP+ Y +FD ++LL+
Sbjct: 223 -SLLLLDEPTSGLDSTTAHRIVTTIKR---LASGGRTVVTTIHQPSSRIYHMFDKVVLLS 278
Query: 387 DGQIVYQGPREYVLEFFESTGFKCPERKGVADFLQEVTS--RKDQWQYWAREDEPYNFVT 444
+G +Y G +E+F S GF AD L ++ + D + + +++ T
Sbjct: 279 EGSPIYYGAASSAVEYFSSLGFSTSLTVNPADLLLDLANGIPPDTQKETSEQEQK----T 334
Query: 445 VKDFARAFELFHIGKQLGEELAD----PFDKSKFHSNVLITKKYGINKKELLRACASREL 500
VK+ + +I +L EL + ++ +K + L ++++ R +
Sbjct: 335 VKETLVSAYEKNISTKLKAELCNAESHSYEYTKAAAKNLKSEQWCTTWWYQFTVLLQRGV 394
Query: 501 LLMKRNSF--VYIFKATQLTYLATLTTTLFLRTKMYHSTIEDAQTYMGALFFTVTVAMFN 558
+ SF + IF+ + +L L +H+ Q LFF ++F
Sbjct: 395 RERRFESFNKLRIFQVISVAFLGGLL--------WWHTPKSHIQDRTALLFF---FSVFW 443
Query: 559 GISELNMTIMKLP----IFYKQRDLLFYPSWAYSLPPWILKIPITIIEVAIWECISYYAI 614
G L + P + K+R Y +Y + + +P+ + + I Y+
Sbjct: 444 GFYPLYNAVFTFPQEKRMLIKERSSGMYRLSSYFMARNVGDLPLELALPTAFVFIIYWMG 503
Query: 615 GFDPNIGRFFKQSLVVLCINQMASALFRFMAALGRDIVVANTFGTFSLLAVTVLGGFVIS 674
G P+ F LVVL +A L AL +I A T + + L + GG+ +
Sbjct: 504 GLKPDPTTFILSLLVVLYSVLVAQGLGLAFGALLMNIKQATTLASVTTLVFLIAGGYYV- 562
Query: 675 REDVHKWFLW 684
+ + + +W
Sbjct: 563 -QQIPPFIVW 571
>At3g13220 ABC transporter
Length = 647
Score = 177 bits (450), Expect = 3e-44
Identities = 156/605 (25%), Positives = 276/605 (44%), Gaps = 68/605 (11%)
Query: 803 EEVSENKASS-SGRKGMV--LPFQPLSLTFDDITYSVDMPQGMKNQGVTEDR------LE 853
EEV EN +G G+V + F P + + + +D+ + + ED
Sbjct: 8 EEVEENHVMQITGSNGIVHNMEFMPQAYLRNQYSSEIDIDEEFVSTYPLEDAPLPIFLKH 67
Query: 854 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGIKTSGYIEGNIKVSGYQKNQKSFARI 913
+LKG++G+ PG + ALMG SG+GKTTL+ ++ G T ++G + + + RI
Sbjct: 68 ILKGITGSTGPGEILALMGPSGSGKTTLLKIMGGRLTDN-VKGKLTYNDIPYSPSVKRRI 126
Query: 914 SGYCEQFDIHSPNVTVYESLLYSAWLRLSPEVDHATRKMFIEEVMELVELNSLREALVGL 973
G+ Q D+ P +TV E+L ++A+LRL + + IE +++ + L R VG
Sbjct: 127 -GFVTQDDVLLPQLTVEETLAFAAFLRLPSSMSKEQKYAKIEMIIKELGLERCRRTRVGG 185
Query: 974 PGENGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAIVMRTVRNTVDTGRTVVC 1033
G+S +RKR +IA E++ +PS++ +DEPTSGLD+ +A ++ ++ GRTV+
Sbjct: 186 GFVKGISGGERKRASIAYEILVDPSLLLLDEPTSGLDSTSATKLLHILQGVAKAGRTVIT 245
Query: 1034 TIHQPSIDIFDSFDELLLLKLGGEQIYAGPIGNQCSDLIQYFEAIQGVPTIKDGYNPATW 1093
TIHQPS +F FD+LLL+ G Y + + ++YF +++ +P I NPA +
Sbjct: 246 TIHQPSSRMFHMFDKLLLISEGHPAFY-----GKARESMEYFSSLRILPEI--AMNPAEF 298
Query: 1094 MLEITS-------------AGKEAN----------LKVNF-TDVY-KNSELHRRNKQLIQ 1128
+L++ + A K A LK + TD+ K E + RN++ +
Sbjct: 299 LLDLATGQVSDISLPDELLAAKTAQPDSEEVLLKYLKQRYKTDLEPKEKEENHRNRKAPE 358
Query: 1129 ELSVPSQSSKDLHFDAQYSQTFLAQCTYCLWKQHLSYWRNT-------SYTAVRLLFTIM 1181
L + Q KD T W Q L R T + +RL+ ++
Sbjct: 359 HLQIAIQVKKD--------------WTLSWWDQFLILSRRTFRERRRDYFDKLRLVQSLG 404
Query: 1182 TGILFGLIFWGVGAKSKKEQDLFNAMGSMYAAVTFIGVVNGASVQPIVAIERTVFYRERA 1241
++ GL++W +K+ E L + +G M+ F + + E+ +ER
Sbjct: 405 VAVVLGLLWW--KSKTDTEAHLRDQVGLMFYICIFWTSSSLFGAVYVFPFEKIYLVKERK 462
Query: 1242 AGMYSAMPYALAQVIIELPHILVQAVVYGIIVYAMMGFEWTASKVLWNLFFTYFSFLYYT 1301
A MY Y + + ++ ++ + IIVY M F L+ + +
Sbjct: 463 AEMYRLSVYYVCSTLCDMVAHVLYPTFFMIIVYFMAEFNRNIPCFLFTVLTILLIAITSQ 522
Query: 1302 YYGMMTMAITPNPHVAGILSTSFYAIWCLFSGFIIPLSRIPIWWKWYYWICPVAWTLNGL 1361
G A + AG++++ ++ L G+ + IP + +W ++ + + L
Sbjct: 523 GAGEFLGASVLSIKRAGMIASLVLMLFLLTGGYYV--QHIPKFMQWLKYLSFMHYGFRLL 580
Query: 1362 VTSQY 1366
+ QY
Sbjct: 581 LKVQY 585
Score = 133 bits (334), Expect = 8e-31
Identities = 139/565 (24%), Positives = 249/565 (43%), Gaps = 56/565 (9%)
Query: 142 LQIIPSPKKQLHILQNVSGILKPRRMTLLLGPPGSGKTTLLLALAGILGKDLKQSGRVTY 201
L+ P P HIL+ ++G P + L+GP GSGKTTLL + G L ++K G++TY
Sbjct: 56 LEDAPLPIFLKHILKGITGSTGPGEILALMGPSGSGKTTLLKIMGGRLTDNVK--GKLTY 113
Query: 202 NGKGLEEFVPQRTSAYVSQYDNHIGEMTVRETLAFSARCQGVGQNYEMLTELLRKEKESK 261
N V +R +V+Q D + ++TV ETLAF+A + + + + +++K +K
Sbjct: 114 NDIPYSPSVKRRIG-FVTQDDVLLPQLTVEETLAFAAFLR-------LPSSMSKEQKYAK 165
Query: 262 IEPDPDINAYMKEAAIEGHQNSVVIDYILKILGLDVCADTMVGDQMIRGISGGEKKRLTT 321
IE I+K LGL+ C T VG ++GISGGE+KR +
Sbjct: 166 IE------------------------MIIKELGLERCRRTRVGGGFVKGISGGERKRASI 201
Query: 322 G-EMLVGPIKVLFMDEISNGLDSSTTFQIINSIKQSIHILNGTALVSLLQPAPETYELFD 380
E+LV P +L +DE ++GLDS++ ++++ I Q + T + ++ QP+ + +FD
Sbjct: 202 AYEILVDP-SLLLLDEPTSGLDSTSATKLLH-ILQGVAKAGRTVITTIHQPSSRMFHMFD 259
Query: 381 DIILLTDGQIVYQGPREYVLEFFESTGFKCPERKGVADFLQEVTSRKDQWQYWAREDEPY 440
++L+++G + G +E+F S A+FL ++ + Q + DE
Sbjct: 260 KLLLISEGHPAFYGKARESMEYFSSLRILPEIAMNPAEFLLDLAT--GQVSDISLPDELL 317
Query: 441 NFVTVKDFARAFELFHIGKQLGEELADPFDKSKFHSNVLITK--KYGINKKELLRACASR 498
T + + L ++ ++ +L +P +K + H N + + I K+
Sbjct: 318 AAKTAQPDSEEVLLKYLKQRYKTDL-EPKEKEENHRNRKAPEHLQIAIQVKKDWTLSWWD 376
Query: 499 ELLLMKRNSF-----VYIFKATQLTYLATLTTTLFLRTKMYHSTIEDAQTYMGALFFTVT 553
+ L++ R +F Y K + L L K T + +G +F+
Sbjct: 377 QFLILSRRTFRERRRDYFDKLRLVQSLGVAVVLGLLWWKSKTDTEAHLRDQVGLMFY--- 433
Query: 554 VAMFNGISELNMTIMKLPI----FYKQRDLLFYPSWAYSLPPWILKIPITIIEVAIWECI 609
+ +F S L + P K+R Y Y + + + ++ + I
Sbjct: 434 ICIFWTSSSLFGAVYVFPFEKIYLVKERKAEMYRLSVYYVCSTLCDMVAHVLYPTFFMII 493
Query: 610 SYYAIGFDPNIGRFFKQSLVVLCINQMASALFRFMAALGRDIVVANTFGTFSLLAVTVLG 669
Y+ F+ NI F L +L I + F+ A I A + L+ + G
Sbjct: 494 VYFMAEFNRNIPCFLFTVLTILLIAITSQGAGEFLGASVLSIKRAGMIASLVLMLFLLTG 553
Query: 670 GFVISREDVHKWFLWGYWSSPLMYG 694
G+ + + + K+ W + S + YG
Sbjct: 554 GYYV--QHIPKFMQWLKYLSFMHYG 576
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.138 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 31,013,565
Number of Sequences: 26719
Number of extensions: 1363730
Number of successful extensions: 5297
Number of sequences better than 10.0: 189
Number of HSP's better than 10.0 without gapping: 164
Number of HSP's successfully gapped in prelim test: 26
Number of HSP's that attempted gapping in prelim test: 4328
Number of HSP's gapped (non-prelim): 580
length of query: 1424
length of database: 11,318,596
effective HSP length: 112
effective length of query: 1312
effective length of database: 8,326,068
effective search space: 10923801216
effective search space used: 10923801216
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 66 (30.0 bits)
Medicago: description of AC144617.9


