

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144617.6 - phase: 0
(178 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
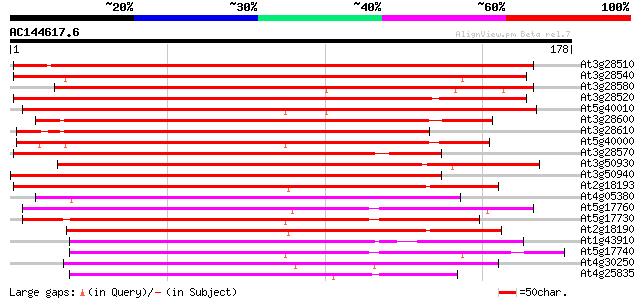
Score E
Sequences producing significant alignments: (bits) Value
At3g28510 hypothetical protein 154 2e-38
At3g28540 hypothetical protein 153 4e-38
At3g28580 putative protein 150 3e-37
At3g28520 hypothetical protein 148 1e-36
At5g40010 putative protein 139 6e-34
At3g28600 unknown protein 139 8e-34
At3g28610 unknown protein 135 1e-32
At5g40000 putative protein 134 3e-32
At3g28570 unknown protein 132 1e-31
At3g50930 BCS1 protein-like protein 124 3e-29
At3g50940 BCS1 protein-like protein 116 6e-27
At2g18193 AAA-type ATPase like protein 112 8e-26
At4g05380 putative protein 112 1e-25
At5g17760 BCS1 - like protein 110 4e-25
At5g17730 BCS1 - like protein 108 2e-24
At2g18190 AAA-type ATPase like protein 107 3e-24
At1g43910 unknown protein 107 4e-24
At5g17740 BCS1 - like protein 103 6e-23
At4g30250 putative protein 96 1e-20
At4g25835 BCS1 like mitochondrial protein 95 2e-20
>At3g28510 hypothetical protein
Length = 530
Score = 154 bits (389), Expect = 2e-38
Identities = 80/165 (48%), Positives = 113/165 (68%), Gaps = 1/165 (0%)
Query: 2 EKKESQAENATKNNKMSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGR 61
EKKE + + ++K S++TL GLLN IDG+WSA +GE++I+FTTN+ +KLD ALI RGR
Sbjct: 327 EKKEGEKKPKV-DDKQSKVTLSGLLNSIDGLWSACSGEKIIVFTTNFVDKLDPALIRRGR 385
Query: 62 MDMLIELPYCCFDGFKMLATKYLSLESHFLFDKIACLLVETNMTPADVAENLMPKVDNED 121
MD IE+ YC F+ FK+LA YL +E+H L+ +I L ET+M+PADVAE LMPK D ED
Sbjct: 386 MDNHIEMSYCKFEAFKVLAKNYLEIETHDLYGEIERKLEETDMSPADVAETLMPKSDEED 445
Query: 122 VATPLLRLIQALRSIEEEAEKEEGTSAKQESDGEDSSAEKKEDAE 166
+ RL++ L +E+A K K++++ E +K E+AE
Sbjct: 446 ADICIKRLVKTLEEEKEKARKLAEEEEKKKAEKEAKKMKKAEEAE 490
>At3g28540 hypothetical protein
Length = 510
Score = 153 bits (387), Expect = 4e-38
Identities = 83/168 (49%), Positives = 113/168 (66%), Gaps = 5/168 (2%)
Query: 2 EKKESQAENATKNNK---MSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALIC 58
E+K+ +AE K + S++TL GLLN IDG+WSA +GE++I+FTTNY +KLD ALI
Sbjct: 322 EEKKKEAEKLLKRERGERESKVTLSGLLNAIDGLWSACSGEKIIVFTTNYLDKLDPALIR 381
Query: 59 RGRMDMLIELPYCCFDGFKMLATKYLSLESHFLFDKIACLLVETNMTPADVAENLMPKVD 118
RGRMD IE+ YC F+ FK+LA YL +ESH LF +I L+ ET+M+PADVAENLMPK D
Sbjct: 382 RGRMDNHIEMSYCRFEAFKVLAKNYLEIESHDLFGEIKRLVEETDMSPADVAENLMPKSD 441
Query: 119 NEDVATPLLRLIQALRSIEEEAEK--EEGTSAKQESDGEDSSAEKKED 164
+D L RL+++L +E+A+K EE K D + +E+
Sbjct: 442 EDDADICLTRLVKSLEEEKEKAKKLAEEEKMKKAARDARRIKKKAEEE 489
>At3g28580 putative protein
Length = 500
Score = 150 bits (379), Expect = 3e-37
Identities = 84/160 (52%), Positives = 109/160 (67%), Gaps = 8/160 (5%)
Query: 15 NKMSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLIELPYCCFD 74
NK S++TL GLLNFIDG+WSA GER+I+FTTN+ +KLD ALI +GRMD IE+ YCCF+
Sbjct: 338 NKESKVTLSGLLNFIDGLWSACGGERIIVFTTNFVDKLDPALIRKGRMDKHIEMSYCCFE 397
Query: 75 GFKMLATKYLSLESHFLFDKIACLL--VETNMTPADVAENLMPKVDNEDVATPLLRLIQA 132
FK+LA YL +E +F++I LL E MTPADV ENL+PK + E T L RLI+A
Sbjct: 398 AFKVLAKNYLDVEESEMFEEIKRLLEVEEIKMTPADVGENLLPKSEKEGGETCLKRLIEA 457
Query: 133 LRSIEEEA-----EKEEGTSAKQESDGE-DSSAEKKEDAE 166
L+ +EEA E+EE K+E E ++ EKK+ E
Sbjct: 458 LKEEKEEAKKKVEEEEEEKQRKKEKVKEIEAEKEKKKKIE 497
>At3g28520 hypothetical protein
Length = 478
Score = 148 bits (374), Expect = 1e-36
Identities = 78/163 (47%), Positives = 108/163 (65%), Gaps = 2/163 (1%)
Query: 2 EKKESQAENATKNNKMSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGR 61
EKKE++ N S +TL GLLN IDG+WSA + E++IIFTTN+ + LD ALI RGR
Sbjct: 310 EKKEAENLKRVSGNNESNVTLSGLLNAIDGLWSACSDEKIIIFTTNFVDNLDPALIRRGR 369
Query: 62 MDMLIELPYCCFDGFKMLATKYLSLESHFLFDKIACLLVETNMTPADVAENLMPKVDNED 121
MD IE+ YC F+ FK+LA YL ESH L+ +I LL E +++PADVAENLMPK D +D
Sbjct: 370 MDYHIEMSYCRFEAFKVLAKNYLENESHDLYGEIGRLLEEVDVSPADVAENLMPKSDEDD 429
Query: 122 VATPLLRLIQALRSIEEEAEKEEGTSAKQESDGEDSSAEKKED 164
RL+++L EE+ +K E + K + ED+ ++K++
Sbjct: 430 ADICFRRLVKSLE--EEKKKKIEKEARKNKKKAEDNVKQEKQN 470
>At5g40010 putative protein
Length = 514
Score = 139 bits (351), Expect = 6e-34
Identities = 78/168 (46%), Positives = 109/168 (64%), Gaps = 5/168 (2%)
Query: 5 ESQAENATKNNKMSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDM 64
E Q + NK S++TL GLLNFIDG+WSA GER+I+FTTN+ +KLD ALI +GRMD
Sbjct: 330 EKQMKKDQGENKGSKVTLSGLLNFIDGLWSACGGERIIVFTTNFIDKLDPALIRKGRMDK 389
Query: 65 LIELPYCCFDGFKMLATKYLSL---ESHFLFDKIACLL--VETNMTPADVAENLMPKVDN 119
IE+ YC F+ FK+LA YL + + LFD+I LL E MTPADV ENL+ K +
Sbjct: 390 HIEMSYCGFEAFKVLANNYLDAKEEDDNELFDEIKRLLEVEEIKMTPADVGENLLKKSEV 449
Query: 120 EDVATPLLRLIQALRSIEEEAEKEEGTSAKQESDGEDSSAEKKEDAEM 167
E L RLI+AL+ +EEA++ K++ + E+ +K+E+ ++
Sbjct: 450 ETKEICLKRLIEALKEEKEEAKRRIEDEEKKKKEEEEIKRKKREEKKI 497
>At3g28600 unknown protein
Length = 475
Score = 139 bits (350), Expect = 8e-34
Identities = 82/145 (56%), Positives = 94/145 (64%), Gaps = 5/145 (3%)
Query: 9 ENATKNNKMSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLIEL 68
E T+ +K S +TL GLLNFIDGIWSA ER+IIFTTN+ EKLD ALI RGRMDM IEL
Sbjct: 320 EQGTEEDK-SFVTLSGLLNFIDGIWSACGQERIIIFTTNHFEKLDPALIRRGRMDMHIEL 378
Query: 69 PYCCFDGFKMLATKYLSLESHFLFDKIACLLVETNMTPADVAENLMPKVDNEDVATPLLR 128
YC F+ FK+LA YL L++H LF KI LL ET + PADVAENLM K D L
Sbjct: 379 SYCSFEAFKILAKNYLDLDTHPLFKKIESLLKETKIAPADVAENLMKKNTEIDADGSLKD 438
Query: 129 LIQALRSIEEEAEKEEGTSAKQESD 153
LIQAL E +K G + D
Sbjct: 439 LIQAL----EGKKKIHGAQVDEPKD 459
>At3g28610 unknown protein
Length = 473
Score = 135 bits (340), Expect = 1e-32
Identities = 76/131 (58%), Positives = 90/131 (68%), Gaps = 3/131 (2%)
Query: 3 KKESQAENATKNNKMSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRM 62
+K+ +N + NK S +TL GLLNFIDGIWSA ER+I+FTTN+ KLD ALI RGRM
Sbjct: 317 RKDGDQDN--EENK-SFVTLSGLLNFIDGIWSACGQERIIVFTTNHLAKLDPALIRRGRM 373
Query: 63 DMLIELPYCCFDGFKMLATKYLSLESHFLFDKIACLLVETNMTPADVAENLMPKVDNEDV 122
DM IEL YC F+ FK LA YL L+SH LF KI L+ ETN+ PADVAENLM K D
Sbjct: 374 DMHIELSYCTFEAFKTLAKNYLDLDSHPLFSKIESLMKETNIAPADVAENLMKKNRETDA 433
Query: 123 ATPLLRLIQAL 133
L LI++L
Sbjct: 434 DGSLNDLIESL 444
>At5g40000 putative protein
Length = 470
Score = 134 bits (337), Expect = 3e-32
Identities = 81/155 (52%), Positives = 102/155 (65%), Gaps = 8/155 (5%)
Query: 3 KKESQA-ENATKNNK-MSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRG 60
KKES + E K +K + +TL GLLNFIDGIWSA ER+++FTTN+ EKLD ALI RG
Sbjct: 313 KKESNSRERYGKEDKDENSVTLSGLLNFIDGIWSACGQERIVVFTTNHLEKLDPALIRRG 372
Query: 61 RMDMLIELPYCCFDGFKMLATKYLSL---ESHFLFDKIACLLVETNMTPADVAENLMPKV 117
RMDM IEL YC ++ FK+LA YL L ++H LF +I LL ET ++PADVAENLM +
Sbjct: 373 RMDMHIELSYCTYEAFKILAKNYLDLDGDDAHPLFSEIKALLEETKISPADVAENLMARN 432
Query: 118 DNEDVATPLLRLIQALRSIEEEAEKEEGTSAKQES 152
DV L LI AL EEE + + K++S
Sbjct: 433 QQIDVDKSLNLLISAL---EEENQYQRSQQEKKKS 464
>At3g28570 unknown protein
Length = 451
Score = 132 bits (332), Expect = 1e-31
Identities = 71/136 (52%), Positives = 92/136 (67%), Gaps = 4/136 (2%)
Query: 2 EKKESQAENATKNNKMSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGR 61
E++ + + K + +TL GLLNFIDGIWSA ER+++FTTN+ KLD ALI RGR
Sbjct: 305 EREVKDLKGDKEGKKSNAVTLSGLLNFIDGIWSACGQERILVFTTNHVGKLDQALIRRGR 364
Query: 62 MDMLIELPYCCFDGFKMLATKYLSLESHFLFDKIACLLVETNMTPADVAENLMPKVDNED 121
MDM IEL YC F FK+LA YL+++SH LF +I LL ET +TPADVAE++M K +
Sbjct: 365 MDMHIELSYCTFGAFKILAKNYLNIDSHHLFGEIESLLKETKITPADVAEHMMAK----E 420
Query: 122 VATPLLRLIQALRSIE 137
V L LI+AL I+
Sbjct: 421 VDGSLKGLIRALERIK 436
>At3g50930 BCS1 protein-like protein
Length = 576
Score = 124 bits (310), Expect = 3e-29
Identities = 70/158 (44%), Positives = 97/158 (61%), Gaps = 6/158 (3%)
Query: 16 KMSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLIELPYCCFDG 75
+ ++TL GLLNFIDG+WS+ ER+IIFTTNY EKLD AL+ GRMDM I + YC
Sbjct: 384 RYKKVTLSGLLNFIDGLWSSCGDERIIIFTTNYKEKLDAALLRPGRMDMHIHMSYCTPST 443
Query: 76 FKMLATKYLSLESHFLFDKIACLLVETNMTPADVAENLMPKVDNEDVATPLLRLIQALRS 135
FK LA YL ++ H LF KI + T +TPA+VAE LM + V L+ ++ ++
Sbjct: 444 FKALALNYLEIKEHRLFSKIEEGIEATEVTPAEVAEQLMRNDSVDKVLEGLIEFLK-VKK 502
Query: 136 IEEE-----AEKEEGTSAKQESDGEDSSAEKKEDAEMV 168
IE E EK+E + K+ +G DS +K+ D ++V
Sbjct: 503 IENEQDKAKTEKQELENKKKTKEGTDSVVKKEVDEQLV 540
>At3g50940 BCS1 protein-like protein
Length = 451
Score = 116 bits (291), Expect = 6e-27
Identities = 58/137 (42%), Positives = 85/137 (61%)
Query: 1 MEKKESQAENATKNNKMSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRG 60
+E K+ + + +TL GLLNF+DG+WS+ ER+I+FTTNY EKLD AL+ G
Sbjct: 311 IELKDRSTDQENNDPLHKTVTLSGLLNFVDGLWSSCGNERIIVFTTNYREKLDPALLRPG 370
Query: 61 RMDMLIELPYCCFDGFKMLATKYLSLESHFLFDKIACLLVETNMTPADVAENLMPKVDNE 120
RMDM I + YC FK+LA+ YL ++ H LF++I + E +TPA+VAE LM +
Sbjct: 371 RMDMHIHMSYCTPAAFKVLASNYLEIQDHILFEQIEEFIREIEVTPAEVAEQLMRSDSVD 430
Query: 121 DVATPLLRLIQALRSIE 137
V L+ ++A + I+
Sbjct: 431 KVLQGLVEFLKAKKQID 447
>At2g18193 AAA-type ATPase like protein
Length = 495
Score = 112 bits (281), Expect = 8e-26
Identities = 61/156 (39%), Positives = 94/156 (60%), Gaps = 3/156 (1%)
Query: 2 EKKESQAENATKNNKMSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGR 61
E ++ +AEN ++TL G+LNFIDG+WS+ ER+I+FTTN+ E+LD AL+ GR
Sbjct: 308 EVRDREAENQEDEQIKGKVTLSGILNFIDGLWSSFGDERIIVFTTNHKERLDPALLRPGR 367
Query: 62 MDMLIELPYCCFDGFKMLATKYLSLE--SHFLFDKIACLLVETNMTPADVAENLMPKVDN 119
MD+ I + YC GF+ L + YL L+ +H L ++I L+ T +TPA++AE LM D
Sbjct: 368 MDVHINMSYCTGLGFRTLVSNYLGLDGLNHPLCEEIEALVDSTEVTPAELAEELMQDDDT 427
Query: 120 EDVATPLLRLIQALRSIEEEAEKEEGTSAKQESDGE 155
+ V ++ ++ R +E K+E + K D E
Sbjct: 428 DVVLRGVISFVEK-RKVERSKTKKEVSICKATDDDE 462
>At4g05380 putative protein
Length = 248
Score = 112 bits (280), Expect = 1e-25
Identities = 59/136 (43%), Positives = 82/136 (59%), Gaps = 1/136 (0%)
Query: 9 ENATKNNKMS-QITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLIE 67
EN K K ++TL GLLNF+DG+WS+ ER+IIFTTN+ EKLD AL+ GRMD+ I
Sbjct: 112 ENQNKKKKKDPKVTLSGLLNFVDGLWSSCVEERIIIFTTNHKEKLDPALLRPGRMDVHIL 171
Query: 68 LPYCCFDGFKMLATKYLSLESHFLFDKIACLLVETNMTPADVAENLMPKVDNEDVATPLL 127
+ YC FK LA YL +E H LFD I + +E TPA++ E LM D + L+
Sbjct: 172 MDYCTPIVFKKLAALYLEIEEHELFDPIEKMFLEVKATPAEITEKLMVSKDPDVTLKGLV 231
Query: 128 RLIQALRSIEEEAEKE 143
+++ + +E + E
Sbjct: 232 EFLESKKMTKESVDSE 247
>At5g17760 BCS1 - like protein
Length = 505
Score = 110 bits (275), Expect = 4e-25
Identities = 69/167 (41%), Positives = 90/167 (53%), Gaps = 8/167 (4%)
Query: 5 ESQAENATKNNKMSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDM 64
E E + +TL GLLNFIDG+WS+ ER+IIFTTN+ ++LD AL+ GRMDM
Sbjct: 324 EQPVEGKNRGESQGPLTLSGLLNFIDGLWSSCGDERIIIFTTNHKDRLDPALLRPGRMDM 383
Query: 65 LIELPYCCFDGFKMLATKYLSLES----HFLFDKIACLLVETNMTPADVAENLMPKVDNE 120
I + +C F GFK LA+ YL L H LF +I L+ MTPA VAE LM +E
Sbjct: 384 HIYMGHCSFQGFKTLASNYLGLSDAAMPHRLFPEIERLIDGEVMTPAQVAEELM---KSE 440
Query: 121 DVATPLLRLIQALRSIEEEAEKEEGTSAKQ-ESDGEDSSAEKKEDAE 166
D L L+ L + ++++ KQ ES E K D E
Sbjct: 441 DADVALEGLVNVLEKMRLKSKESNPVMMKQKESRLEMEEMRLKSDTE 487
>At5g17730 BCS1 - like protein
Length = 470
Score = 108 bits (269), Expect = 2e-24
Identities = 63/148 (42%), Positives = 90/148 (60%), Gaps = 8/148 (5%)
Query: 5 ESQAENATKNNKMSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDM 64
+ + ++ TK + M +TL GLL IDG+WS+ ER++IFTT + E+LD AL+ GRMDM
Sbjct: 316 QPKTQDDTKGSSM--LTLSGLLTCIDGLWSSCGDERIVIFTTTHKERLDPALLRPGRMDM 373
Query: 65 LIELPYCCFDGFKMLATKYLSL---ESHFLFDKIACLLVETNMTPADVAENLMPKVDNED 121
I + +CCFD FK LA+ YL L + H L+ +I L+ +TPA VAE LM NED
Sbjct: 374 HIHMGHCCFDVFKTLASNYLGLSHDDPHHLYPEIERLIKGEVLTPAQVAEELM---KNED 430
Query: 122 VATPLLRLIQALRSIEEEAEKEEGTSAK 149
L L++ L+ E EK +G + +
Sbjct: 431 PDVALEGLVKVLKRKRLELEKYDGETGR 458
>At2g18190 AAA-type ATPase like protein
Length = 494
Score = 107 bits (268), Expect = 3e-24
Identities = 59/140 (42%), Positives = 89/140 (63%), Gaps = 3/140 (2%)
Query: 19 QITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLIELPYCCFDGFKM 78
++TL GLLNF+DG+WS+ ER+I+FTTN+ E+LD AL+ GRMDM I + YC GF+
Sbjct: 329 RVTLSGLLNFVDGLWSSFGDERIIVFTTNHKERLDPALLRPGRMDMHINMSYCTGLGFRT 388
Query: 79 LATKYLSLE--SHFLFDKIACLLVETNMTPADVAENLMPKVDNEDVATPLLRLIQALRSI 136
L + YL L +H L ++I L+ T +TPA++AE LM + D + V ++ ++ R +
Sbjct: 389 LVSNYLGLGGLNHPLCEEIEALIDSTEVTPAELAEELMQEDDTDVVLRGVVSFVEN-RKV 447
Query: 137 EEEAEKEEGTSAKQESDGED 156
E KE S ++ DG+D
Sbjct: 448 EISKTKELEGSTCRKLDGDD 467
>At1g43910 unknown protein
Length = 475
Score = 107 bits (266), Expect = 4e-24
Identities = 61/144 (42%), Positives = 85/144 (58%), Gaps = 7/144 (4%)
Query: 20 ITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLIELPYCCFDGFKML 79
I+L GLLNF+DG+WS+ E++IIFTTN+ EKLD AL+ GRMD+ I + C FK L
Sbjct: 336 ISLSGLLNFVDGLWSSCGEEKIIIFTTNHKEKLDPALLRPGRMDVHILMDNCTPFVFKKL 395
Query: 80 ATKYLSLESHFLFDKIACLLVETNMTPADVAENLMPKVDNEDVATPLLRLIQALRSIEEE 139
YL + H LFD I L++E + TPA+V + LM N D+A ++ L E
Sbjct: 396 VALYLKTDEHVLFDPIEKLILEVSSTPAEVTQQLMAS-KNADIA------LKGLAEFLEN 448
Query: 140 AEKEEGTSAKQESDGEDSSAEKKE 163
+ ++G + E +GE AE KE
Sbjct: 449 KKLKKGEDSSVEEEGEIEDAETKE 472
>At5g17740 BCS1 - like protein
Length = 533
Score = 103 bits (256), Expect = 6e-23
Identities = 68/163 (41%), Positives = 96/163 (58%), Gaps = 13/163 (7%)
Query: 20 ITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLIELPYCCFDGFKML 79
+TL GLLN IDG+WS+ ER+IIFTTN EKLD AL+ GRMDM I + +C F GFK L
Sbjct: 334 LTLSGLLNCIDGLWSSCGNERIIIFTTNNKEKLDPALLRPGRMDMHIYMGHCSFQGFKTL 393
Query: 80 ATKYLSL-----ESHFLFDKIACLLVETNMTPADVAENLMPKVDNEDVATPLLRLIQALR 134
A+ YL L ++H L I L+ +TPA VAE LM +ED L L++ L+
Sbjct: 394 ASNYLGLSDENDDTHPLCPDIKHLIDGHVLTPAQVAEELM---KDEDADAALEGLVKVLK 450
Query: 135 SIEEEAEK-EEGTSAKQESDGEDSSAEKKEDAEMVTSSMLRLE 176
E +K ++ + K+ +GE++ A DAE+ + ++E
Sbjct: 451 RKRLEPKKCDDESKMKKLKEGEEAIA----DAELAVLTPAQVE 489
>At4g30250 putative protein
Length = 512
Score = 95.9 bits (237), Expect = 1e-20
Identities = 54/148 (36%), Positives = 85/148 (56%), Gaps = 10/148 (6%)
Query: 18 SQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLIELPYCCFDGFK 77
S +TL GLLNF DG+WS E++ +FTTN+ EKLD AL+ GRMDM + + +C F K
Sbjct: 334 SSVTLSGLLNFTDGLWSCCGSEKIFVFTTNHIEKLDSALMRSGRMDMHVHMGFCKFPALK 393
Query: 78 MLATKYLSLESH----FLFDKIACLLVETNMTPADVAENLM------PKVDNEDVATPLL 127
+L YL LE + ++ + E +TPADV+E L+ K E V+
Sbjct: 394 ILLKNYLRLEEEDMDSVVLKEMEECVEEAEITPADVSEVLIRNRSDAEKAVREIVSVLKE 453
Query: 128 RLIQALRSIEEEAEKEEGTSAKQESDGE 155
R+++ +S+ + +K+EG ++E++ E
Sbjct: 454 RVVKRRKSVGLKKKKQEGQEEEEEAEEE 481
>At4g25835 BCS1 like mitochondrial protein
Length = 506
Score = 94.7 bits (234), Expect = 2e-20
Identities = 57/127 (44%), Positives = 71/127 (55%), Gaps = 6/127 (4%)
Query: 20 ITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLIELPYCCFDGFKML 79
ITL GLLNF DG+WS ER+ +FTTN+ EKLD AL+ GRMDM I + YC F K+L
Sbjct: 336 ITLSGLLNFTDGLWSCCGSERIFVFTTNHIEKLDPALLRSGRMDMHIHMSYCTFSSVKIL 395
Query: 80 ATKYLSLESHFLFDKIACLLVE----TNMTPADVAENLMPKVDNEDVATPLLRLIQALRS 135
YL E L D + L E +TPADV+E L+ + D + L+ LRS
Sbjct: 396 LRNYLGFEEGDLNDVVLKELAEVVDRAEITPADVSEALIK--NRRDKERAVRELLVDLRS 453
Query: 136 IEEEAEK 142
E EK
Sbjct: 454 RVERNEK 460
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.314 0.130 0.357
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,766,553
Number of Sequences: 26719
Number of extensions: 154356
Number of successful extensions: 1507
Number of sequences better than 10.0: 148
Number of HSP's better than 10.0 without gapping: 97
Number of HSP's successfully gapped in prelim test: 52
Number of HSP's that attempted gapping in prelim test: 1077
Number of HSP's gapped (non-prelim): 409
length of query: 178
length of database: 11,318,596
effective HSP length: 93
effective length of query: 85
effective length of database: 8,833,729
effective search space: 750866965
effective search space used: 750866965
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 56 (26.2 bits)
Medicago: description of AC144617.6


