

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144541.14 - phase: 0
(63 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
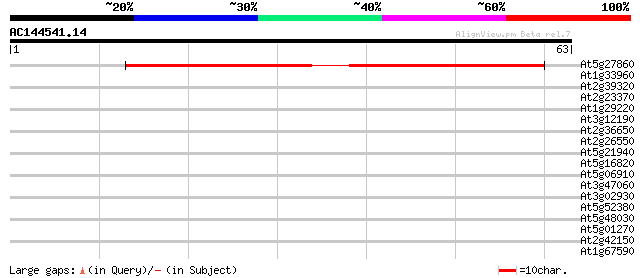
At5g27860 putative protein 69 6e-13
At1g33960 AIG1 29 0.38
At2g39320 hypothetical protein 28 0.85
At2g23370 unknown protein 26 4.2
At1g29220 unknown protein 26 4.2
At3g12190 hypothetical protein 25 5.5
At2g36650 hypothetical protein 25 5.5
At2g26550 heme oxygenase 2 (HO2) 25 5.5
At5g21940 unknown protein 25 7.2
At5g16820 Heat Shock Factor 3 25 7.2
At5g06910 DnaJ homologue (gb|AAB91418.1|) 25 7.2
At3g47060 FtsH metalloprotease - like protein 25 7.2
At3g02930 unknown protein 25 7.2
At5g52380 unknown protein 25 9.4
At5g48030 DnaJ protein-like 25 9.4
At5g01270 hypothetical protein 25 9.4
At2g42150 unknown protein 25 9.4
At1g67590 unknown protein 25 9.4
>At5g27860 putative protein
Length = 224
Score = 68.6 bits (166), Expect = 6e-13
Identities = 36/47 (76%), Positives = 39/47 (82%), Gaps = 4/47 (8%)
Query: 14 KFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNELLNFLNA 60
KFL R+KDDG RRSAVSGKK+ DK+KEDK AESKRNELL FLNA
Sbjct: 113 KFLNRDKDDGERRSAVSGKKV----DKSKEDKAAESKRNELLKFLNA 155
>At1g33960 AIG1
Length = 353
Score = 29.3 bits (64), Expect = 0.38
Identities = 15/49 (30%), Positives = 31/49 (62%), Gaps = 1/49 (2%)
Query: 12 LSKFLGRNKDDGVRRSAV-SGKKILLKLDKTKEDKEAESKRNELLNFLN 59
L +LG N D ++R + G++++L +KTK+D++ + +ELL ++
Sbjct: 180 LEDYLGDNMPDFLKRVLILCGQRMILFDNKTKDDEKKTKQVHELLKLID 228
>At2g39320 hypothetical protein
Length = 189
Score = 28.1 bits (61), Expect = 0.85
Identities = 14/44 (31%), Positives = 24/44 (53%), Gaps = 1/44 (2%)
Query: 10 VQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNE 53
+ + +NK+ G R S+ S + +KL + KE+ EA+ K E
Sbjct: 95 IHFNSIYKKNKEKGSRSSSSSSSAVWMKLQRKKEN-EAKKKEEE 137
>At2g23370 unknown protein
Length = 288
Score = 25.8 bits (55), Expect = 4.2
Identities = 15/46 (32%), Positives = 21/46 (45%)
Query: 2 NGERSSSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEA 47
NG+ S+ V + R G R K+LLKL + E K+A
Sbjct: 143 NGDHVSALVATEFYTKRGNFPGFARPFAFNAKVLLKLGRNLEAKDA 188
>At1g29220 unknown protein
Length = 351
Score = 25.8 bits (55), Expect = 4.2
Identities = 11/27 (40%), Positives = 19/27 (69%)
Query: 34 ILLKLDKTKEDKEAESKRNELLNFLNA 60
I + + +E KE ESK+N+ L+F++A
Sbjct: 213 IEIDMKNERERKEQESKKNQKLDFVSA 239
>At3g12190 hypothetical protein
Length = 269
Score = 25.4 bits (54), Expect = 5.5
Identities = 15/41 (36%), Positives = 22/41 (53%), Gaps = 3/41 (7%)
Query: 19 NKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNELLNFLN 59
N DG RR ++ KLD+ K + E+E K+ L+ LN
Sbjct: 87 NTADGFRRDFEEKQR---KLDRLKREIESEEKKRFLVQKLN 124
>At2g36650 hypothetical protein
Length = 374
Score = 25.4 bits (54), Expect = 5.5
Identities = 14/54 (25%), Positives = 29/54 (52%)
Query: 6 SSSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNELLNFLN 59
S++ + L++F+ RN+D+ V S + + + E++EAES + L+
Sbjct: 27 SATGLILARFVSRNEDNEVTSSTSNPESSSSPSRENDEEEEAESPNQQKQEILS 80
>At2g26550 heme oxygenase 2 (HO2)
Length = 299
Score = 25.4 bits (54), Expect = 5.5
Identities = 12/30 (40%), Positives = 18/30 (60%)
Query: 24 VRRSAVSGKKILLKLDKTKEDKEAESKRNE 53
+R V+GKK+ L DKT +KE E + +
Sbjct: 85 MRLRNVNGKKLDLSEDKTDTEKEEEEEEED 114
>At5g21940 unknown protein
Length = 264
Score = 25.0 bits (53), Expect = 7.2
Identities = 12/31 (38%), Positives = 15/31 (47%)
Query: 1 PNGERSSSPVQLSKFLGRNKDDGVRRSAVSG 31
P+ SS S +GRN DDG + S G
Sbjct: 29 PSDSSSSPSSSASSSIGRNSDDGEKSSEDGG 59
>At5g16820 Heat Shock Factor 3
Length = 447
Score = 25.0 bits (53), Expect = 7.2
Identities = 17/52 (32%), Positives = 28/52 (53%), Gaps = 6/52 (11%)
Query: 8 SPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNELLNFLN 59
SP L++ + +N +DG R+ S KK L +D E E++ + + N LN
Sbjct: 172 SPGFLNQLVQQNNNDGNRQIPGSNKKRRLPVD------EQENRGDNVANGLN 217
>At5g06910 DnaJ homologue (gb|AAB91418.1|)
Length = 284
Score = 25.0 bits (53), Expect = 7.2
Identities = 17/40 (42%), Positives = 23/40 (57%), Gaps = 8/40 (20%)
Query: 23 GVRRSAVSGK------KILLKL--DKTKEDKEAESKRNEL 54
GV R A S + K+ LKL DK ++DKEA+ K +L
Sbjct: 35 GVERRATSQEIRKAYHKLALKLHPDKNQDDKEAKDKFQQL 74
>At3g47060 FtsH metalloprotease - like protein
Length = 802
Score = 25.0 bits (53), Expect = 7.2
Identities = 13/42 (30%), Positives = 24/42 (56%)
Query: 10 VQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKR 51
V S+FL + + V++ V G ++L KL + +E+E+ R
Sbjct: 177 VPYSEFLSKVNSNQVQKVEVDGVQVLFKLRDDGKWQESETSR 218
>At3g02930 unknown protein
Length = 806
Score = 25.0 bits (53), Expect = 7.2
Identities = 13/32 (40%), Positives = 19/32 (58%)
Query: 24 VRRSAVSGKKILLKLDKTKEDKEAESKRNELL 55
V+R KKIL +L+ +KE++E K E L
Sbjct: 427 VQRLLEEKKKILSELESSKEEEEKSKKAMESL 458
>At5g52380 unknown protein
Length = 268
Score = 24.6 bits (52), Expect = 9.4
Identities = 14/50 (28%), Positives = 25/50 (50%), Gaps = 2/50 (4%)
Query: 11 QLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRN--ELLNFL 58
+L+KF G + +D S KKI D + E + K+ +++NF+
Sbjct: 218 KLTKFSGDDLEDDFTEEPKSSKKINTSDDSAQNSVEVKKKKQGPKIVNFV 267
>At5g48030 DnaJ protein-like
Length = 456
Score = 24.6 bits (52), Expect = 9.4
Identities = 13/39 (33%), Positives = 20/39 (50%)
Query: 16 LGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNEL 54
+ +N +G + A G L D K+D EAE+K E+
Sbjct: 101 VSKNAQEGEIKKAYYGLAKKLHPDMNKDDPEAETKFQEV 139
>At5g01270 hypothetical protein
Length = 771
Score = 24.6 bits (52), Expect = 9.4
Identities = 15/53 (28%), Positives = 26/53 (48%), Gaps = 1/53 (1%)
Query: 3 GERSSSPVQLSKFLGRNKDDGVRRSAV-SGKKILLKLDKTKEDKEAESKRNEL 54
G R S V+ + NK+ + +G+KI + + KTK+D ++ N L
Sbjct: 666 GRRCGSKVEFRTVISTNKELQFSVEVLFTGEKIGIGMAKTKKDAHQQAAENAL 718
>At2g42150 unknown protein
Length = 631
Score = 24.6 bits (52), Expect = 9.4
Identities = 13/40 (32%), Positives = 23/40 (57%)
Query: 19 NKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNELLNFL 58
N++D R S+++ K+ KT E+K+ SK+ +FL
Sbjct: 484 NRNDKNRDSSLNVDDSKDKVKKTDEEKKGGSKKKRAASFL 523
>At1g67590 unknown protein
Length = 347
Score = 24.6 bits (52), Expect = 9.4
Identities = 12/52 (23%), Positives = 29/52 (55%)
Query: 5 RSSSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNELLN 56
R+++PV+ + +GR+ R++ G+ + + ++ E + ES +E +N
Sbjct: 166 RTATPVRATTPVGRSPVTSPVRASQRGEAVGVVMETVTEVRRVESNNSEKVN 217
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.307 0.129 0.336
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,373,579
Number of Sequences: 26719
Number of extensions: 45870
Number of successful extensions: 167
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 12
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 153
Number of HSP's gapped (non-prelim): 19
length of query: 63
length of database: 11,318,596
effective HSP length: 39
effective length of query: 24
effective length of database: 10,276,555
effective search space: 246637320
effective search space used: 246637320
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 52 (24.6 bits)
Medicago: description of AC144541.14


