

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144539.12 + phase: 0
(417 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
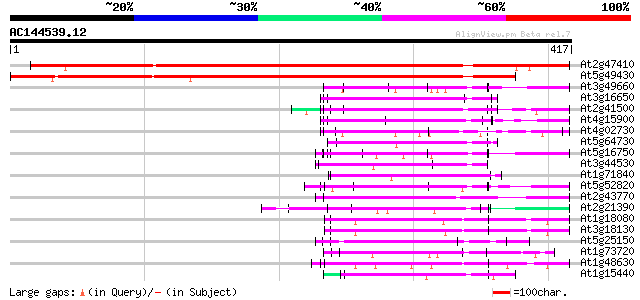
Score E
Sequences producing significant alignments: (bits) Value
At2g47410 putative WD-40 repeat protein 523 e-149
At5g49430 WD-40 repeat protein-like 486 e-138
At3g49660 putative WD-40 repeat - protein 79 4e-15
At3g16650 PP1/PP2A phosphatases pleiotropic regulator PRL2 79 4e-15
At2g41500 putative U4/U6 small nuclear ribonucleoprotein 79 6e-15
At4g15900 PRL1 protein 78 1e-14
At4g02730 putative WD-repeat protein 77 2e-14
At5g64730 unknown protein 73 3e-13
At5g16750 WD40-repeat protein 72 7e-13
At3g44530 WD repeat domain protein 72 7e-13
At1g71840 unknown protein (At1g71840) 71 1e-12
At5g52820 Notchless protein homolog 71 1e-12
At2g43770 putative splicing factor 71 1e-12
At2g21390 coatomer alpha subunit 69 4e-12
At1g18080 putative guanine nucleotide-binding protein beta sub... 68 1e-11
At3g18130 protein kinase C-receptor/G-protein, putative 66 4e-11
At5g25150 transcription initiation factor IID-associated factor-... 65 5e-11
At1g73720 unknown protein 65 5e-11
At1g48630 unknown protein 65 9e-11
At1g15440 unknown protein 64 1e-10
>At2g47410 putative WD-40 repeat protein
Length = 1389
Score = 523 bits (1348), Expect = e-149
Identities = 273/420 (65%), Positives = 314/420 (74%), Gaps = 27/420 (6%)
Query: 16 SLKHLSSSKAPEKTQPDVANQNHDM-----DVDVDIGEVYFLIMRFLSAGPCHKTCSHLW 70
S + S + Q +V Q HD D+D+D+ EVYFLI+ FLS GPC +T HL
Sbjct: 7 STPFMEPSNLAKLVQGNVPLQPHDSHSSLTDLDMDLREVYFLILHFLSIGPCERTFGHLR 66
Query: 71 NELLENQLLPRRYHAWYSRSGASSGVPHDDGQSFPLEDYNKLAERYPHIEKDHLVKLLKQ 130
+E+LE LLPRRYH+W+SRSG SG DDG S PL Y+ L ERYPHIEKDHLVKLLKQ
Sbjct: 67 DEILEKGLLPRRYHSWWSRSGIYSGRADDDGISLPLS-YDNLIERYPHIEKDHLVKLLKQ 125
Query: 131 LLLNKASLSPGMSTGNPPNAADVPTLLGRGSFSLLSYDGDKVNEEVKPPPPYMRWPHTKA 190
L+LN + S GN PNAADVPTLLG G+FSL+ + +++ + Y+RWPH A
Sbjct: 126 LILNPSFPSHMRVEGNAPNAADVPTLLGSGTFSLVDRSNNIESQKARHVASYLRWPHMHA 185
Query: 191 NQVHGLHLREIGGGLPRHHRAPSIRAACYAIAKPSTMVQKMQNIKRIRGHCNAVYCAIFD 250
+QV GL LREIGGG +HHRAPSI +AC+AIAKPSTMVQKMQNIK++RGH NAVYCAIFD
Sbjct: 186 DQVRGLSLREIGGGFRKHHRAPSILSACHAIAKPSTMVQKMQNIKKLRGHRNAVYCAIFD 245
Query: 251 RSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVW 310
RSGRYVITGSDDRLVKIWSMETA LASCRGH GDITDLAVSSNNALVAS+SND++IRVW
Sbjct: 246 RSGRYVITGSDDRLVKIWSMETALCLASCRGHEGDITDLAVSSNNALVASASNDFVIRVW 305
Query: 311 RLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDARYTHSTPRLYVPK 370
RLPDG+PISVLRGHTGAVTAIAFSPR +SSDDGTCRIWDARY+ PR+YVP
Sbjct: 306 RLPDGMPISVLRGHTGAVTAIAFSPRQ-------ASSDDGTCRIWDARYSQWLPRIYVPS 358
Query: 371 PSDST---------GRSSGPSSNT-----MPQSHQIFCCAFNANGTVFVTGSSDNLARVY 416
PSD+ G S+ SSNT QSHQI CCA+NANGT+FVTGSSD+ ARVY
Sbjct: 359 PSDANTGSLEVSLLGESTFTSSNTGSTSNASQSHQILCCAYNANGTIFVTGSSDSNARVY 418
>At5g49430 WD-40 repeat protein-like
Length = 1576
Score = 486 bits (1252), Expect = e-138
Identities = 250/382 (65%), Positives = 295/382 (76%), Gaps = 15/382 (3%)
Query: 1 MALQKYVPSGDAPTVSLKHLS-SSKAPEKTQ---PDVANQNHDMDVDVDIGEVYFLIMRF 56
MAL+K P GD+ ++ +K L+ S K P Q PD+ Q+ ++D+D+ EVYFL++
Sbjct: 1 MALRKNTPKGDSVSLPMKPLNFSRKLPGNVQIPDPDIV-QSVAPNIDLDLREVYFLMLHL 59
Query: 57 LSAGPCHKTCSHLWNELLENQLLPRRYHAWYSRSGASSGVPHDDGQSFPLEDYNKLAERY 116
LS+GPC KT + L +ELLE++LLPRRYHAWYSRSG SG +DDG SFPL +Y +LA+RY
Sbjct: 60 LSSGPCQKTYALLRHELLEHELLPRRYHAWYSRSGLPSGDENDDGNSFPL-NYTELAKRY 118
Query: 117 PHIEKDHLVKLLKQLLL--NKASLSPGMSTGNPPNAADVPTLLGRGSFSLLSYDGDKVNE 174
H++KDHLV+LLKQL+ N+ + S G+ GN A VPTLLG GSFSLLS D + V
Sbjct: 119 SHVKKDHLVELLKQLVFVSNRPNPSRGIGDGNKMIGAGVPTLLGTGSFSLLSSDKEIVGS 178
Query: 175 EVKPPPPYMRWPHTKANQVHGLHLREIGGGLPRHHRAPSIRAACYAIAKPSTMVQKMQNI 234
++KPPP MRWPH A+QV G+ LREIGGG RHHRAPSIRAACY IAKPS+MVQKMQNI
Sbjct: 179 DLKPPPIGMRWPHMHADQVRGISLREIGGGFARHHRAPSIRAACYVIAKPSSMVQKMQNI 238
Query: 235 KRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSN 294
KR+RGH NAVYCAI DRSGRYVITGSDDRLVK+WSM+TAY LASCRGH GDITDLAVSSN
Sbjct: 239 KRLRGHRNAVYCAILDRSGRYVITGSDDRLVKVWSMDTAYCLASCRGHEGDITDLAVSSN 298
Query: 295 NALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRI 354
N +AS+SND +IRVWRLPDGLP+SVLRGHTGAVTAIAFSPRP SSDDGTCRI
Sbjct: 299 NIFIASASNDCVIRVWRLPDGLPVSVLRGHTGAVTAIAFSPRP-------GSSDDGTCRI 351
Query: 355 WDARYTHSTPRLYVPKPSDSTG 376
WDAR PR+YVP+P G
Sbjct: 352 WDARGAQFAPRIYVPRPPSPDG 373
>At3g49660 putative WD-40 repeat - protein
Length = 317
Score = 79.3 bits (194), Expect = 4e-15
Identities = 47/183 (25%), Positives = 85/183 (45%), Gaps = 21/183 (11%)
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSS 293
++ GH N + F R++++ SDD+ +K+W +ET + + GH + +
Sbjct: 64 VQEFTGHENGISDVAFSSDARFIVSASDDKTLKLWDVETGSLIKTLIGHTNYAFCVNFNP 123
Query: 294 NNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCR 353
+ ++ S S D +R+W + G + VL H+ VTA+ F+ R ++ ++SSS DG CR
Sbjct: 124 QSNMIVSGSFDETVRIWDVTTGKCLKVLPAHSDPVTAVDFN-RDGSL--IVSSSYDGLCR 180
Query: 354 IWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAFNANGTVFVTGSSDNLA 413
IWD+ H L ++ + F+ NG + G+ DN
Sbjct: 181 IWDSGTGHCVKTL------------------IDDENPPVSFVRFSPNGKFILVGTLDNTL 222
Query: 414 RVY 416
R++
Sbjct: 223 RLW 225
Score = 55.8 bits (133), Expect = 4e-08
Identities = 32/110 (29%), Positives = 53/110 (48%), Gaps = 6/110 (5%)
Query: 249 FDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGD---ITDLAVSSNNALVASSSNDY 305
F +G++++ G+ D +++W++ +A L + GHV I+ +N + S S D
Sbjct: 206 FSPNGKFILVGTLDNTLRLWNISSAKFLKTYTGHVNAQYCISSAFSVTNGKRIVSGSEDN 265
Query: 306 IIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIW 355
+ +W L + L GHT V +A P N + S S D T RIW
Sbjct: 266 CVHMWELNSKKLLQKLEGHTETVMNVACHPTENLI---ASGSLDKTVRIW 312
Score = 53.1 bits (126), Expect = 3e-07
Identities = 25/80 (31%), Positives = 41/80 (51%), Gaps = 3/80 (3%)
Query: 234 IKRIRGHCNAVYC---AIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLA 290
+K GH NA YC A +G+ +++GS+D V +W + + L GH + ++A
Sbjct: 233 LKTYTGHVNAQYCISSAFSVTNGKRIVSGSEDNCVHMWELNSKKLLQKLEGHTETVMNVA 292
Query: 291 VSSNNALVASSSNDYIIRVW 310
L+AS S D +R+W
Sbjct: 293 CHPTENLIASGSLDKTVRIW 312
Score = 48.1 bits (113), Expect = 9e-06
Identities = 36/166 (21%), Positives = 62/166 (36%), Gaps = 43/166 (25%)
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSS 293
IK + GH N +C F+ +++GS D V+IW + T L H +T + +
Sbjct: 106 IKTLIGHTNYAFCVNFNPQSNMIVSGSFDETVRIWDVTTGKCLKVLPAHSDPVTAVDFNR 165
Query: 294 NNALVASSSNDYIIRVWRLPDGL-----------PISVLR-------------------- 322
+ +L+ SSS D + R+W G P+S +R
Sbjct: 166 DGSLIVSSSYDGLCRIWDSGTGHCVKTLIDDENPPVSFVRFSPNGKFILVGTLDNTLRLW 225
Query: 323 ------------GHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWD 356
GH A I+ + +++S S+D +W+
Sbjct: 226 NISSAKFLKTYTGHVNAQYCISSAFSVTNGKRIVSGSEDNCVHMWE 271
Score = 47.4 bits (111), Expect = 1e-05
Identities = 33/140 (23%), Positives = 58/140 (40%), Gaps = 27/140 (19%)
Query: 282 HVGDITDLAVSSNNALVASSSNDYIIRVWRL-----PDGLPISVLRGHTGAVTAIAFSPR 336
H ++ + SS+ L+AS+S D IR + + P P+ GH ++ +AFS
Sbjct: 23 HNRAVSSVKFSSDGRLLASASADKTIRTYTINTINDPIAEPVQEFTGHENGISDVAFSSD 82
Query: 337 PNAVYQLLSSSDDGTCRIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFCCA 396
+ +S+SDD T ++WD + + ++ FC
Sbjct: 83 ARFI---VSASDDKTLKLWDV-------------------ETGSLIKTLIGHTNYAFCVN 120
Query: 397 FNANGTVFVTGSSDNLARVY 416
FN + V+GS D R++
Sbjct: 121 FNPQSNMIVSGSFDETVRIW 140
>At3g16650 PP1/PP2A phosphatases pleiotropic regulator PRL2
Length = 479
Score = 79.3 bits (194), Expect = 4e-15
Identities = 41/127 (32%), Positives = 62/127 (48%), Gaps = 3/127 (2%)
Query: 232 QNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAV 291
+N + ++GH V FD S + TGS DR +KIW + T + GH+G + LAV
Sbjct: 161 KNYRVLQGHLGWVRSVAFDPSNEWFCTGSADRTIKIWDVATGVLKLTLTGHIGQVRGLAV 220
Query: 292 SSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGT 351
S+ + + S+ +D ++ W L I GH V +A P + V L+ D
Sbjct: 221 SNRHTYMFSAGDDKQVKCWDLEQNKVIRSYHGHLHGVYCLALHPTLDVV---LTGGRDSV 277
Query: 352 CRIWDAR 358
CR+WD R
Sbjct: 278 CRVWDIR 284
Score = 57.0 bits (136), Expect = 2e-08
Identities = 34/126 (26%), Positives = 55/126 (42%), Gaps = 4/126 (3%)
Query: 237 IRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNA 296
+ GH V Y+ + DD+ VK W +E + S GH+ + LA+
Sbjct: 208 LTGHIGQVRGLAVSNRHTYMFSAGDDKQVKCWDLEQNKVIRSYHGHLHGVYCLALHPTLD 267
Query: 297 LVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWD 356
+V + D + RVW + + I VL + + +A P Q+++ S D T + WD
Sbjct: 268 VVLTGGRDSVCRVWDIRTKMQIFVLPHDSDVFSVLARPTDP----QVITGSHDSTIKFWD 323
Query: 357 ARYTHS 362
RY S
Sbjct: 324 LRYGKS 329
Score = 50.4 bits (119), Expect = 2e-06
Identities = 27/105 (25%), Positives = 47/105 (44%), Gaps = 1/105 (0%)
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSS 293
I+ GH + VYC + V+TG D + ++W + T + H D+ +
Sbjct: 247 IRSYHGHLHGVYCLALHPTLDVVLTGGRDSVCRVWDIRTKMQI-FVLPHDSDVFSVLARP 305
Query: 294 NNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPN 338
+ V + S+D I+ W L G ++ + H V A+A P+ N
Sbjct: 306 TDPQVITGSHDSTIKFWDLRYGKSMATITNHKKTVRAMALHPKEN 350
Score = 38.1 bits (87), Expect = 0.009
Identities = 39/180 (21%), Positives = 72/180 (39%), Gaps = 21/180 (11%)
Query: 240 HCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAV--SSNNAL 297
H + V+ + + VITGS D +K W + S+A+ H + +A+ N+ +
Sbjct: 294 HDSDVFSVLARPTDPQVITGSHDSTIKFWDLRYGKSMATITNHKKTVRAMALHPKENDFV 353
Query: 298 VASSSNDYIIRVWRLPDG-LPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWD 356
AS+ N I+ + LP G ++L + A+A N +++ D G WD
Sbjct: 354 SASADN---IKKFSLPKGEFCHNMLSLQRDIINAVAV----NEDGVMVTGGDKGGLWFWD 406
Query: 357 ARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAFNANGTVFVTGSSDNLARVY 416
+ H+ R S +G I+ ++ G+ VT D +++
Sbjct: 407 WKSGHNFQRAETIVQPGSLESEAG-----------IYAACYDQTGSRLVTCEGDKTIKMW 455
>At2g41500 putative U4/U6 small nuclear ribonucleoprotein
Length = 554
Score = 78.6 bits (192), Expect = 6e-15
Identities = 44/129 (34%), Positives = 62/129 (47%), Gaps = 3/129 (2%)
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSS 293
++ GH + + F SG+Y+ T S D+ ++W + T L GH + +A
Sbjct: 332 LQTFEGHLDRLARVAFHPSGKYLGTTSYDKTWRLWDINTGAELLLQEGHSRSVYGIAFQQ 391
Query: 294 NNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCR 353
+ AL AS D + RVW L G I V +GH V ++ FSP Y L S +D CR
Sbjct: 392 DGALAASCGLDSLARVWDLRTGRSILVFQGHIKPVFSVNFSPNG---YHLASGGEDNQCR 448
Query: 354 IWDARYTHS 362
IWD R S
Sbjct: 449 IWDLRMRKS 457
Score = 57.4 bits (137), Expect = 1e-08
Identities = 31/120 (25%), Positives = 53/120 (43%), Gaps = 2/120 (1%)
Query: 239 GHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALV 298
GH +VY F + G + D L ++W + T S+ +GH+ + + S N +
Sbjct: 379 GHSRSVYGIAFQQDGALAASCGLDSLARVWDLRTGRSILVFQGHIKPVFSVNFSPNGYHL 438
Query: 299 ASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDAR 358
AS D R+W L + ++ H V+ + + P+ Y L ++S D IW R
Sbjct: 439 ASGGEDNQCRIWDLRMRKSLYIIPAHANLVSQVKYEPQEG--YFLATASYDMKVNIWSGR 496
Score = 52.0 bits (123), Expect = 6e-07
Identities = 52/211 (24%), Positives = 82/211 (38%), Gaps = 29/211 (13%)
Query: 210 RAPSIRAACY--AIAKPSTMVQKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKI 267
R I A C + K M Q I ++ H +F + T S DR K+
Sbjct: 265 RDGKILATCSLSGVTKLWEMPQVTNTIAVLKDHKERATDVVFSPVDDCLATASADRTAKL 324
Query: 268 WSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGA 327
W + L + GH+ + +A + + ++S D R+W + G + + GH+ +
Sbjct: 325 WKTDGTL-LQTFEGHLDRLARVAFHPSGKYLGTTSYDKTWRLWDINTGAELLLQEGHSRS 383
Query: 328 VTAIAFSPRPNAVYQLLSSSDDGTCRIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMP 387
V IAF + A+ S D R+WD R TGRS +
Sbjct: 384 VYGIAFQ-QDGAL--AASCGLDSLARVWDLR----------------TGRSI-----LVF 419
Query: 388 QSH--QIFCCAFNANGTVFVTGSSDNLARVY 416
Q H +F F+ NG +G DN R++
Sbjct: 420 QGHIKPVFSVNFSPNGYHLASGGEDNQCRIW 450
Score = 50.1 bits (118), Expect = 2e-06
Identities = 42/169 (24%), Positives = 72/169 (41%), Gaps = 24/169 (14%)
Query: 249 FDRSGRYVITGSDDRLVKIWSM-ETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYII 307
F R G+ + T S + K+W M + ++A + H TD+ S + +A++S D
Sbjct: 263 FSRDGKILATCSLSGVTKLWEMPQVTNTIAVLKDHKERATDVVFSPVDDCLATASADRTA 322
Query: 308 RVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDARYTHSTPRLY 367
++W+ DG + GH + +AF P L ++S D T R+WD
Sbjct: 323 KLWK-TDGTLLQTFEGHLDRLARVAFHPSGK---YLGTTSYDKTWRLWDI---------- 368
Query: 368 VPKPSDSTGRSSGPSSNTMPQSHQIFCCAFNANGTVFVTGSSDNLARVY 416
+TG S ++ AF +G + + D+LARV+
Sbjct: 369 ------NTGAELLLQEG---HSRSVYGIAFQQDGALAASCGLDSLARVW 408
Score = 43.5 bits (101), Expect = 2e-04
Identities = 26/125 (20%), Positives = 54/125 (42%), Gaps = 4/125 (3%)
Query: 232 QNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAV 291
++I +GH V+ F +G ++ +G +D +IW + SL H ++ +
Sbjct: 414 RSILVFQGHIKPVFSVNFSPNGYHLASGGEDNQCRIWDLRMRKSLYIIPAHANLVSQVKY 473
Query: 292 SSNNA-LVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDG 350
+A++S D + +W D + L GH V ++ + + + + S D
Sbjct: 474 EPQEGYFLATASYDMKVNIWSGRDFSLVKSLAGHESKVASLDITADSSCI---ATVSHDR 530
Query: 351 TCRIW 355
T ++W
Sbjct: 531 TIKLW 535
>At4g15900 PRL1 protein
Length = 486
Score = 77.8 bits (190), Expect = 1e-14
Identities = 41/127 (32%), Positives = 61/127 (47%), Gaps = 3/127 (2%)
Query: 232 QNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAV 291
+N + I+GH V FD S + TGS DR +KIW + T + GH+ + LAV
Sbjct: 167 KNYRVIQGHLGWVRSVAFDPSNEWFCTGSADRTIKIWDVATGVLKLTLTGHIEQVRGLAV 226
Query: 292 SSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGT 351
S+ + + S+ +D ++ W L I GH V +A P + LL+ D
Sbjct: 227 SNRHTYMFSAGDDKQVKCWDLEQNKVIRSYHGHLSGVYCLALHPTLDV---LLTGGRDSV 283
Query: 352 CRIWDAR 358
CR+WD R
Sbjct: 284 CRVWDIR 290
Score = 60.1 bits (144), Expect = 2e-09
Identities = 33/123 (26%), Positives = 54/123 (43%), Gaps = 3/123 (2%)
Query: 237 IRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNA 296
+ GH V Y+ + DD+ VK W +E + S GH+ + LA+
Sbjct: 214 LTGHIEQVRGLAVSNRHTYMFSAGDDKQVKCWDLEQNKVIRSYHGHLSGVYCLALHPTLD 273
Query: 297 LVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWD 356
++ + D + RVW + + I L GH V ++ P Q+++ S D T + WD
Sbjct: 274 VLLTGGRDSVCRVWDIRTKMQIFALSGHDNTVCSVFTRPTDP---QVVTGSHDTTIKFWD 330
Query: 357 ARY 359
RY
Sbjct: 331 LRY 333
Score = 58.2 bits (139), Expect = 8e-09
Identities = 32/118 (27%), Positives = 54/118 (45%), Gaps = 3/118 (2%)
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSS 293
I+ GH + VYC + ++TG D + ++W + T + + GH + +
Sbjct: 253 IRSYHGHLSGVYCLALHPTLDVLLTGGRDSVCRVWDIRTKMQIFALSGHDNTVCSVFTRP 312
Query: 294 NNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGT 351
+ V + S+D I+ W L G +S L H +V A+ P+ NA S+S D T
Sbjct: 313 TDPQVVTGSHDTTIKFWDLRYGKTMSTLTHHKKSVRAMTLHPKENA---FASASADNT 367
Score = 45.4 bits (106), Expect = 6e-05
Identities = 35/137 (25%), Positives = 58/137 (41%), Gaps = 22/137 (16%)
Query: 280 RGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNA 339
+GH+G + +A +N + S D I++W + G+ L GH V +A S R
Sbjct: 173 QGHLGWVRSVAFDPSNEWFCTGSADRTIKIWDVATGVLKLTLTGHIEQVRGLAVSNRHTY 232
Query: 340 VYQLLSSSDDGTCRIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAFNA 399
++ S+ DD + WD + R Y G SG ++C A +
Sbjct: 233 MF---SAGDDKQVKCWDLEQ-NKVIRSY-------HGHLSG-----------VYCLALHP 270
Query: 400 NGTVFVTGSSDNLARVY 416
V +TG D++ RV+
Sbjct: 271 TLDVLLTGGRDSVCRVW 287
Score = 41.2 bits (95), Expect = 0.001
Identities = 36/135 (26%), Positives = 63/135 (46%), Gaps = 9/135 (6%)
Query: 230 KMQNIKRIRGHCNAVYCAIFDR-SGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITD 288
KMQ I + GH N V C++F R + V+TGS D +K W + ++++ H +
Sbjct: 292 KMQ-IFALSGHDNTV-CSVFTRPTDPQVVTGSHDTTIKFWDLRYGKTMSTLTHHKKSVRA 349
Query: 289 LAVSSNNALVASSSNDYIIRVWRLPDG-LPISVLRGHTGAVTAIAFSPRPNAVYQLLSSS 347
+ + AS+S D + + LP G ++L + A+A N +++
Sbjct: 350 MTLHPKENAFASASADN-TKKFSLPKGEFCHNMLSQQKTIINAMAV----NEDGVMVTGG 404
Query: 348 DDGTCRIWDARYTHS 362
D+G+ WD + HS
Sbjct: 405 DNGSIWFWDWKSGHS 419
>At4g02730 putative WD-repeat protein
Length = 333
Score = 76.6 bits (187), Expect = 2e-14
Identities = 54/187 (28%), Positives = 90/187 (47%), Gaps = 25/187 (13%)
Query: 232 QNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAV 291
+++K + GH A+ C F G + + S D+ + +WS + GH I+DLA
Sbjct: 34 RHLKTLEGHTAAISCVKFSNDGNLLASASVDKTMILWSATNYSLIHRYEGHSSGISDLAW 93
Query: 292 SSNNALVASSSNDYIIRVW--RLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDD 349
SS++ S+S+D +R+W R P + VLRGHT V + F+P N + +S S D
Sbjct: 94 SSDSHYTCSASDDCTLRIWDARSPYEC-LKVLRGHTNFVFCVNFNPPSNLI---VSGSFD 149
Query: 350 GTCRIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAFNANGTVFVTGSS 409
T RIW+ + T R+ +++MP I FN +G++ V+ S
Sbjct: 150 ETIRIWEVK-TGKCVRMI--------------KAHSMP----ISSVHFNRDGSLIVSASH 190
Query: 410 DNLARVY 416
D +++
Sbjct: 191 DGSCKIW 197
Score = 60.1 bits (144), Expect = 2e-09
Identities = 46/182 (25%), Positives = 91/182 (49%), Gaps = 26/182 (14%)
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSS 293
+K +RGH N V+C F+ +++GS D ++IW ++T + + H I+ + +
Sbjct: 121 LKVLRGHTNFVFCVNFNPPSNLIVSGSFDETIRIWEVKTGKCVRMIKAHSMPISSVHFNR 180
Query: 294 NNALVASSSNDYIIRVWRLPDGLPI-SVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTC 352
+ +L+ S+S+D ++W +G + +++ + AV+ FS PN + L+++ D
Sbjct: 181 DGSLIVSASHDGSCKIWDAKEGTCLKTLIDDKSPAVSFAKFS--PNGKFILVATLD---- 234
Query: 353 RIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFC--CAFN-ANGTVFVTGSS 409
ST +L + +TG+ + ++++FC AF+ NG V+GS
Sbjct: 235 ---------STLKL----SNYATGKFLKVYTG---HTNKVFCITSAFSVTNGKYIVSGSE 278
Query: 410 DN 411
DN
Sbjct: 279 DN 280
Score = 47.8 bits (112), Expect = 1e-05
Identities = 24/83 (28%), Positives = 42/83 (49%), Gaps = 5/83 (6%)
Query: 234 IKRIRGHCNAVYC---AIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLA 290
+K GH N V+C A +G+Y+++GS+D V +W ++ L GH + ++
Sbjct: 248 LKVYTGHTNKVFCITSAFSVTNGKYIVSGSEDNCVYLWDLQARNILQRLEGHTDAVISVS 307
Query: 291 VSSNNALVASSSN--DYIIRVWR 311
++SS N D IR+W+
Sbjct: 308 CHPVQNEISSSGNHLDKTIRIWK 330
Score = 45.1 bits (105), Expect = 7e-05
Identities = 35/119 (29%), Positives = 53/119 (44%), Gaps = 10/119 (8%)
Query: 243 AVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGD---ITDLAVSSNNALVA 299
AV A F +G++++ + D +K+ + T L GH IT +N +
Sbjct: 215 AVSFAKFSPNGKFILVATLDSTLKLSNYATGKFLKVYTGHTNKVFCITSAFSVTNGKYIV 274
Query: 300 SSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSD---DGTCRIW 355
S S D + +W L + L GHT AV +++ P N + SSS D T RIW
Sbjct: 275 SGSEDNCVYLWDLQARNILQRLEGHTDAVISVSCHPVQNEI----SSSGNHLDKTIRIW 329
>At5g64730 unknown protein
Length = 299
Score = 73.2 bits (178), Expect = 3e-13
Identities = 39/126 (30%), Positives = 65/126 (50%), Gaps = 4/126 (3%)
Query: 237 IRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNA 296
++GH AV A F+ G Y +T DR +++W+ + + + H ++ D+ V+S+NA
Sbjct: 14 LKGHEGAVLAARFNGDGNYALTCGKDRTIRLWNPHRGILIKTYKSHGREVRDVHVTSDNA 73
Query: 297 LVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWD 356
S D + W + G I RGH G V A+ F+ + V +S+ D + R+WD
Sbjct: 74 KFCSCGGDRQVYYWDVSTGRVIRKFRGHDGEVNAVKFNDSSSVV---VSAGFDRSLRVWD 130
Query: 357 ARYTHS 362
R +HS
Sbjct: 131 CR-SHS 135
Score = 57.4 bits (137), Expect = 1e-08
Identities = 33/114 (28%), Positives = 56/114 (48%), Gaps = 5/114 (4%)
Query: 244 VYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDI--TDLAVSSNNALVASS 301
V C G V+ G D +++ T L +GH+ TD +++++A V
Sbjct: 187 VNCISISNDGNCVLAGCLDSTLRLLDRTTGELLQVYKGHISKSFKTDCCLTNSDAHVIGG 246
Query: 302 SNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIW 355
S D ++ W L D +S R H VT++++ P+ + +L+SS DGT R+W
Sbjct: 247 SEDGLVFFWDLVDAKVLSKFRAHDLVVTSVSYHPKEDC---MLTSSVDGTIRVW 297
Score = 37.4 bits (85), Expect = 0.015
Identities = 21/60 (35%), Positives = 31/60 (51%)
Query: 252 SGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVWR 311
S +VI GS+D LV W + A L+ R H +T ++ + +SS D IRVW+
Sbjct: 239 SDAHVIGGSEDGLVFFWDLVDAKVLSKFRAHDLVVTSVSYHPKEDCMLTSSVDGTIRVWK 298
Score = 31.6 bits (70), Expect = 0.85
Identities = 43/188 (22%), Positives = 69/188 (35%), Gaps = 40/188 (21%)
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGD-------- 285
I++ RGH V F+ S V++ DR +++W CR H +
Sbjct: 95 IRKFRGHDGEVNAVKFNDSSSVVVSAGFDRSLRVW---------DCRSHSVEPVQIIDTF 145
Query: 286 -ITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLL 344
T ++V + S D +R + + G +S G +
Sbjct: 146 LDTVMSVVLTKTEIIGGSVDGTVRTFDMRIGREMSDNLGQP---------------VNCI 190
Query: 345 SSSDDGTCRIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAFNANGTVF 404
S S+DG C + A ST RL +TG + +S + CC N++ V
Sbjct: 191 SISNDGNCVL--AGCLDSTLRLL----DRTTGELLQVYKGHISKSFKTDCCLTNSDAHV- 243
Query: 405 VTGSSDNL 412
+ GS D L
Sbjct: 244 IGGSEDGL 251
>At5g16750 WD40-repeat protein
Length = 876
Score = 71.6 bits (174), Expect = 7e-13
Identities = 35/120 (29%), Positives = 59/120 (49%), Gaps = 1/120 (0%)
Query: 237 IRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNA 296
I G + + + + + R +++W +ET + S +GH G + +A ++
Sbjct: 56 IEGESDTLTALALSPDDKLLFSAGHSRQIRVWDLETLKCIRSWKGHEGPVMGMACHASGG 115
Query: 297 LVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWD 356
L+A++ D + VW + G RGH G V++I F P N L+S SDD T R+WD
Sbjct: 116 LLATAGADRKVLVWDVDGGFCTHYFRGHKGVVSSILFHPDSNKNI-LISGSDDATVRVWD 174
Score = 63.9 bits (154), Expect = 2e-10
Identities = 47/190 (24%), Positives = 83/190 (42%), Gaps = 35/190 (18%)
Query: 239 GHCNAVYCAIF-DRSGRYVITGSDDRLVKIWSME----------TAYSLASCRGHVGDIT 287
GH + F +S + ++GS DR +K+WS++ + + H DI
Sbjct: 444 GHNGDILAVAFAKKSFSFFVSGSGDRTLKVWSLDGISEDSEEPINLKTRSVVAAHDKDIN 503
Query: 288 DLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSS 347
+AV+ N++LV + S D +WRLPD + + L+GH + ++ FS V +++S
Sbjct: 504 SVAVARNDSLVCTGSEDRTASIWRLPDLVHVVTLKGHKRRIFSVEFSTVDQCV---MTAS 560
Query: 348 DDGTCRIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMP-QSHQIFCCAFNANGTVFVT 406
D T +IW S G T + + +F +GT FV+
Sbjct: 561 GDKTVKIW--------------------AISDGSCLKTFEGHTSSVLRASFITDGTQFVS 600
Query: 407 GSSDNLARVY 416
+D L +++
Sbjct: 601 CGADGLLKLW 610
Score = 57.8 bits (138), Expect = 1e-08
Identities = 30/120 (25%), Positives = 54/120 (45%), Gaps = 3/120 (2%)
Query: 237 IRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNA 296
+ H + R+ V TGS+DR IW + + + +GH I + S+ +
Sbjct: 495 VAAHDKDINSVAVARNDSLVCTGSEDRTASIWRLPDLVHVVTLKGHKRRIFSVEFSTVDQ 554
Query: 297 LVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWD 356
V ++S D +++W + DG + GHT +V +F Q +S DG ++W+
Sbjct: 555 CVMTASGDKTVKIWAISDGSCLKTFEGHTSSVLRASFITDGT---QFVSCGADGLLKLWN 611
Score = 52.0 bits (123), Expect = 6e-07
Identities = 36/135 (26%), Positives = 62/135 (45%), Gaps = 9/135 (6%)
Query: 228 VQKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDIT 287
++ ++ I+ +GH V SG + T DR V +W ++ + RGH G ++
Sbjct: 89 LETLKCIRSWKGHEGPVMGMACHASGGLLATAGADRKVLVWDVDGGFCTHYFRGHKGVVS 148
Query: 288 DLAV--SSNNALVASSSNDYIIRVWRL----PDGLPISVLRGHTGAVTAIAFSPRPNAVY 341
+ SN ++ S S+D +RVW L + ++++ H AVT+IA S
Sbjct: 149 SILFHPDSNKNILISGSDDATVRVWDLNAKNTEKKCLAIMEKHFSAVTSIALS---EDGL 205
Query: 342 QLLSSSDDGTCRIWD 356
L S+ D +WD
Sbjct: 206 TLFSAGRDKVVNLWD 220
Score = 48.5 bits (114), Expect = 7e-06
Identities = 27/93 (29%), Positives = 46/93 (49%), Gaps = 2/93 (2%)
Query: 263 RLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLR 322
R+ + +M +Y LA + V + SS N L+ + S D +R+W I V
Sbjct: 384 RVYDVATMSCSYVLAGHKEVVLSLDTCVSSSGNVLIVTGSKDKTVRLWNATSKSCIGVGT 443
Query: 323 GHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIW 355
GH G + A+AF+ + + + +S S D T ++W
Sbjct: 444 GHNGDILAVAFAKKSFSFF--VSGSGDRTLKVW 474
Score = 45.8 bits (107), Expect = 4e-05
Identities = 24/123 (19%), Positives = 51/123 (40%), Gaps = 3/123 (2%)
Query: 233 NIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVS 292
++ ++GH ++ F + V+T S D+ VKIW++ L + GH + +
Sbjct: 533 HVVTLKGHKRRIFSVEFSTVDQCVMTASGDKTVKIWAISDGSCLKTFEGHTSSVLRASFI 592
Query: 293 SNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTC 352
++ S D ++++W + I+ H V A+A + + + D
Sbjct: 593 TDGTQFVSCGADGLLKLWNVNTSECIATYDQHEDKVWALAVGKKTE---MIATGGGDAVI 649
Query: 353 RIW 355
+W
Sbjct: 650 NLW 652
Score = 45.8 bits (107), Expect = 4e-05
Identities = 21/77 (27%), Positives = 39/77 (50%)
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSS 293
+K GH ++V A F G ++ D L+K+W++ T+ +A+ H + LAV
Sbjct: 576 LKTFEGHTSSVLRASFITDGTQFVSCGADGLLKLWNVNTSECIATYDQHEDKVWALAVGK 635
Query: 294 NNALVASSSNDYIIRVW 310
++A+ D +I +W
Sbjct: 636 KTEMIATGGGDAVINLW 652
Score = 31.2 bits (69), Expect = 1.1
Identities = 26/109 (23%), Positives = 47/109 (42%), Gaps = 4/109 (3%)
Query: 248 IFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYII 307
I G ++ D + + S +++ ++ G +T LA+S ++ L+ S+ + I
Sbjct: 26 IVSSDGSFIACACGDVINIVDSTDSSVK-STIEGESDTLTALALSPDDKLLFSAGHSRQI 84
Query: 308 RVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWD 356
RVW L I +GH G V +A L ++ D +WD
Sbjct: 85 RVWDLETLKCIRSWKGHEGPVMGMACHASGGL---LATAGADRKVLVWD 130
>At3g44530 WD repeat domain protein
Length = 1051
Score = 71.6 bits (174), Expect = 7e-13
Identities = 41/143 (28%), Positives = 72/143 (49%), Gaps = 20/143 (13%)
Query: 230 KMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWS-----------------MET 272
K + + +R H +V C + ++ RYV +GSDD++++I +E
Sbjct: 48 KERLLATLRDHFGSVNCVRWAKNSRYVASGSDDQVIQIHERKPGSGTTEFGSGEAPDVEN 107
Query: 273 AYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIA 332
++ + RGH D+ DL S +++++AS S D + +W + G+ +VLRGH V +
Sbjct: 108 WKAVMTLRGHTADVVDLNWSPDDSMLASGSLDNTVHIWNMRTGMCTTVLRGHLSLVKGVT 167
Query: 333 FSPRPNAVYQLLSSSDDGTCRIW 355
+ P + + S SDD T IW
Sbjct: 168 WDPIGSFI---ASQSDDKTVIIW 187
Score = 43.9 bits (102), Expect = 2e-04
Identities = 23/87 (26%), Positives = 39/87 (44%)
Query: 228 VQKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDIT 287
V+ + + +RGH V + + +GS D V IW+M T RGH+ +
Sbjct: 105 VENWKAVMTLRGHTADVVDLNWSPDDSMLASGSLDNTVHIWNMRTGMCTTVLRGHLSLVK 164
Query: 288 DLAVSSNNALVASSSNDYIIRVWRLPD 314
+ + +AS S+D + +WR D
Sbjct: 165 GVTWDPIGSFIASQSDDKTVIIWRTSD 191
>At1g71840 unknown protein (At1g71840)
Length = 407
Score = 71.2 bits (173), Expect = 1e-12
Identities = 42/130 (32%), Positives = 64/130 (48%), Gaps = 5/130 (3%)
Query: 239 GHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGH---VGDITDLAVSSNN 295
GH V C F G+ + TGSDD + +W+ +T S+ +GH +T L ++SN+
Sbjct: 195 GHNLNVTCGDFTPDGKLICTGSDDASLIVWNPKTCESIHIVKGHPYHTEGLTCLDINSNS 254
Query: 296 ALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIW 355
+L S S D + + + G +S L HT +V + FSP + + D IW
Sbjct: 255 SLAISGSKDGSVHIVNIVTGKVVSSLNSHTDSVECVKFSPSSATIPLAATGGMDKKLIIW 314
Query: 356 DARYTHSTPR 365
D + HSTPR
Sbjct: 315 DLQ--HSTPR 322
Score = 42.7 bits (99), Expect = 4e-04
Identities = 31/120 (25%), Positives = 55/120 (45%), Gaps = 5/120 (4%)
Query: 238 RGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNAL 297
+G A+ C+ D + V TG D +W + A GH ++ LA S + L
Sbjct: 70 KGELYALACSPTDAT--LVATGGGDDKAFLWKIGNGDWAAELPGHKDSVSCLAFSYDGQL 127
Query: 298 VASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDA 357
+AS D +++++ G VL G + + + PR + V L+ S+D + +W+A
Sbjct: 128 LASGGLDGVVQIFDASSGTLKCVLDGPGAGIEWVRWHPRGHIV---LAGSEDCSLWMWNA 184
Score = 39.7 bits (91), Expect = 0.003
Identities = 29/127 (22%), Positives = 55/127 (42%), Gaps = 7/127 (5%)
Query: 234 IKRIRGHCNAVYCAIFDRSGRYV---ITGSDDRLVKIWSMETAYSLASCRGHVGDITDLA 290
+ + H ++V C F S + TG D+ + IW ++ + C H +T L
Sbjct: 277 VSSLNSHTDSVECVKFSPSSATIPLAATGGMDKKLIIWDLQHSTPRFICE-HEEGVTSLT 335
Query: 291 VSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDG 350
+ +A+ + + +W G + GH AV AI+ S + + +S S D
Sbjct: 336 WIGTSKYLATGCANGTVSIWDSLLGNCVHTYHGHQDAVQAISVSTNTDFI---VSVSVDN 392
Query: 351 TCRIWDA 357
T R++++
Sbjct: 393 TARVFES 399
>At5g52820 Notchless protein homolog
Length = 473
Score = 70.9 bits (172), Expect = 1e-12
Identities = 47/150 (31%), Positives = 71/150 (47%), Gaps = 13/150 (8%)
Query: 220 AIAKPSTMVQKMQNIKRIR----------GHCNAVYCAIFDRSGRYVITGSDDRLVKIWS 269
++ K T+V + Q + RIR GH AV C F G+ + +GS D V++W
Sbjct: 78 SVEKVLTIVYQQQAVFRIRPVNRCSQTIAGHAEAVLCVSFSPDGKQLASGSGDTTVRLWD 137
Query: 270 METAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDG-LPISVLRGHTGAV 328
+ T L +C+GH + +A S + + S S I W G L S L GH +
Sbjct: 138 LYTETPLFTCKGHKNWVLTVAWSPDGKHLVSGSKSGEICCWNPKKGELEGSPLTGHKKWI 197
Query: 329 TAIAFSP--RPNAVYQLLSSSDDGTCRIWD 356
T I++ P + + ++SS DG RIWD
Sbjct: 198 TGISWEPVHLSSPCRRFVTSSKDGDARIWD 227
Score = 62.0 bits (149), Expect = 6e-10
Identities = 34/121 (28%), Positives = 60/121 (49%), Gaps = 3/121 (2%)
Query: 235 KRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSN 294
KR+ GH V F G+++ + S D+ V++W+ T + RGHVG + ++ S++
Sbjct: 354 KRLTGHQQLVNHVYFSPDGKWIASASFDKSVRLWNGITGQFVTVFRGHVGPVYQVSWSAD 413
Query: 295 NALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRI 354
+ L+ S S D +++W + L GH V A+ +SP V +S D ++
Sbjct: 414 SRLLLSGSKDSTLKIWEIRTKKLKQDLPGHADEVFAVDWSPDGEKV---VSGGKDRVLKL 470
Query: 355 W 355
W
Sbjct: 471 W 471
Score = 51.2 bits (121), Expect = 1e-06
Identities = 39/162 (24%), Positives = 71/162 (43%), Gaps = 23/162 (14%)
Query: 256 VITGSDDRLVKIWSMETAYSLAS-CRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPD 314
+++GSDD + +W + GH + + S + +AS+S D +R+W
Sbjct: 332 LVSGSDDFTMFLWEPSVSKQPKKRLTGHQQLVNHVYFSPDGKWIASASFDKSVRLWNGIT 391
Query: 315 GLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDARYTHSTPRLYVPKPSDS 374
G ++V RGH G V +++S LLS S D T +IW+ R T +L P
Sbjct: 392 GQFVTVFRGHVGPVYQVSWSADSRL---LLSGSKDSTLKIWEIR----TKKLKQDLPG-- 442
Query: 375 TGRSSGPSSNTMPQSHQIFCCAFNANGTVFVTGSSDNLARVY 416
+ ++F ++ +G V+G D + +++
Sbjct: 443 -------------HADEVFAVDWSPDGEKVVSGGKDRVLKLW 471
Score = 45.1 bits (105), Expect = 7e-05
Identities = 21/80 (26%), Positives = 37/80 (46%)
Query: 232 QNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAV 291
Q + RGH VY + R +++GS D +KIW + T GH ++ +
Sbjct: 393 QFVTVFRGHVGPVYQVSWSADSRLLLSGSKDSTLKIWEIRTKKLKQDLPGHADEVFAVDW 452
Query: 292 SSNNALVASSSNDYIIRVWR 311
S + V S D ++++W+
Sbjct: 453 SPDGEKVVSGGKDRVLKLWK 472
Score = 37.0 bits (84), Expect = 0.020
Identities = 26/100 (26%), Positives = 44/100 (44%), Gaps = 5/100 (5%)
Query: 254 RYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLP 313
R +T S D +IW + S+ GH +T + + ++ + S D I++W
Sbjct: 212 RRFVTSSKDGDARIWDITLKKSIICLSGHTLAVTCVKWGGDG-IIYTGSQDCTIKMWETT 270
Query: 314 DGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCR 353
G I L+GH + ++A S Y L + + D T R
Sbjct: 271 QGKLIRELKGHGHWINSLALSTE----YVLRTGAFDHTGR 306
>At2g43770 putative splicing factor
Length = 343
Score = 70.9 bits (172), Expect = 1e-12
Identities = 48/184 (26%), Positives = 79/184 (42%), Gaps = 23/184 (12%)
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSME-TAYSLASCRGHVGDITDLAVS 292
I + GH +AVY F+ +G + +GS DR + +W + + +GH I DL +
Sbjct: 46 IMLLSGHPSAVYTMKFNPAGTLIASGSHDREIFLWRVHGDCKNFMVLKGHKNAILDLHWT 105
Query: 293 SNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTC 352
S+ + + S+S D +R W + G I + H+ V + P ++S SDDGT
Sbjct: 106 SDGSQIVSASPDKTVRAWDVETGKQIKKMAEHSSFVNSCC--PTRRGPPLIISGSDDGTA 163
Query: 353 RIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAFNANGTVFVTGSSDNL 412
++WD R + T P +QI +F+ TG DN
Sbjct: 164 KLWDMRQRGAI--------------------QTFPDKYQITAVSFSDAADKIFTGGVDND 203
Query: 413 ARVY 416
+V+
Sbjct: 204 VKVW 207
Score = 54.3 bits (129), Expect = 1e-07
Identities = 47/190 (24%), Positives = 81/190 (41%), Gaps = 16/190 (8%)
Query: 228 VQKMQNIKRIRGHCNAVY-CAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDI 286
V+ + IK++ H + V C R +I+GSDD K+W M ++ + I
Sbjct: 125 VETGKQIKKMAEHSSFVNSCCPTRRGPPLIISGSDDGTAKLWDMRQRGAIQTFPDKY-QI 183
Query: 287 TDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSS 346
T ++ S + + D ++VW L G L GH +T ++ S P+ Y L +
Sbjct: 184 TAVSFSDAADKIFTGGVDNDVKVWDLRKGEATMTLEGHQDTITGMSLS--PDGSYLLTNG 241
Query: 347 SDDGTCRIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAFNANGTVFVT 406
D+ C +WD R Y P+ + G N + C+++ +GT
Sbjct: 242 MDNKLC-VWDM-------RPYAPQ-NRCVKIFEGHQHNF---EKNLLKCSWSPDGTKVTA 289
Query: 407 GSSDNLARVY 416
GSSD + ++
Sbjct: 290 GSSDRMVHIW 299
Score = 38.9 bits (89), Expect = 0.005
Identities = 26/116 (22%), Positives = 46/116 (39%), Gaps = 8/116 (6%)
Query: 228 VQKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASC----RGHV 283
++K + + GH + + G Y++T D + +W M C GH
Sbjct: 209 LRKGEATMTLEGHQDTITGMSLSPDGSYLLTNGMDNKLCVWDMRPYAPQNRCVKIFEGHQ 268
Query: 284 GD----ITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSP 335
+ + + S + V + S+D ++ +W I L GHTG+V F P
Sbjct: 269 HNFEKNLLKCSWSPDGTKVTAGSSDRMVHIWDTTSRRTIYKLPGHTGSVNECVFHP 324
Score = 37.7 bits (86), Expect = 0.012
Identities = 25/114 (21%), Positives = 48/114 (41%), Gaps = 7/114 (6%)
Query: 249 FDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIR 308
F + + TG D VK+W + + + GH IT +++S + + + ++ D +
Sbjct: 188 FSDAADKIFTGGVDNDVKVWDLRKGEATMTLEGHQDTITGMSLSPDGSYLLTNGMDNKLC 247
Query: 309 VWRL----PDGLPISVLRG--HTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWD 356
VW + P + + G H + S P+ ++ + S D IWD
Sbjct: 248 VWDMRPYAPQNRCVKIFEGHQHNFEKNLLKCSWSPDGT-KVTAGSSDRMVHIWD 300
Score = 35.8 bits (81), Expect = 0.045
Identities = 17/63 (26%), Positives = 29/63 (45%), Gaps = 1/63 (1%)
Query: 242 NAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASS 301
N + C+ + G V GS DR+V IW + ++ GH G + + ++ S
Sbjct: 274 NLLKCS-WSPDGTKVTAGSSDRMVHIWDTTSRRTIYKLPGHTGSVNECVFHPTEPIIGSC 332
Query: 302 SND 304
S+D
Sbjct: 333 SSD 335
>At2g21390 coatomer alpha subunit
Length = 1218
Score = 69.3 bits (168), Expect = 4e-12
Identities = 50/169 (29%), Positives = 76/169 (44%), Gaps = 12/169 (7%)
Query: 188 TKANQVHGLHLREIGGGLPRHHRAPSIRAACYAIAKPSTMVQKMQNIKRIRGHCNAVYCA 247
TK+N+V GL H + P I A+ ++ + I R H V
Sbjct: 7 TKSNRVKGLSF---------HPKRPWILASLHSGVIQLWDYRMGTLIDRFDEHEGPVRGV 57
Query: 248 IFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYII 307
F S ++G DD +K+W+ +T L + GH+ I + N + S+S+D I
Sbjct: 58 HFHNSQPLFVSGGDDYKIKVWNYKTHRCLFTLLGHLDYIRTVQFHHENPWIVSASDDQTI 117
Query: 308 RVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWD 356
R+W ISVL GH V +F P+ + V +S+S D T R+WD
Sbjct: 118 RIWNWQSRTCISVLTGHNHYVMCASFHPKEDLV---VSASLDQTVRVWD 163
Score = 52.4 bits (124), Expect = 5e-07
Identities = 32/116 (27%), Positives = 54/116 (45%), Gaps = 5/116 (4%)
Query: 237 IRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMET--AYSLASCRGHVGDITDLAVSSN 294
+ GH V A F + +++G+DDR VK+W M A+ + + RGH+ +++ + +
Sbjct: 200 LEGHDRGVNWASFHPTLPLIVSGADDRQVKLWRMNETKAWEVDTLRGHMNNVSSVMFHAK 259
Query: 295 NALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDG 350
++ S+S D IRVW I R +A P N L + D+G
Sbjct: 260 QDIIVSNSEDKSIRVWDATKRTGIQTFRREHDRFWILAVHPEINL---LAAGHDNG 312
Score = 46.2 bits (108), Expect = 3e-05
Identities = 38/179 (21%), Positives = 71/179 (39%), Gaps = 32/179 (17%)
Query: 208 HHRAPSIRAACYAIAKPSTMVQKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKI 267
HH P I +A Q I + GH + V CA F V++ S D+ V++
Sbjct: 102 HHENPWIVSASDDQTIRIWNWQSRTCISVLTGHNHYVMCASFHPKEDLVVSASLDQTVRV 161
Query: 268 WSMETAYSLASC---------------------------RGHVGDITDLAVSSNNALVAS 300
W + ++ GH + + L+ S
Sbjct: 162 WDIGALKKKSASPADDLMRFSQMNSDLFGGVDAIVKYVLEGHDRGVNWASFHPTLPLIVS 221
Query: 301 SSNDYIIRVWRLPD--GLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDA 357
++D +++WR+ + + LRGH V+++ F + + + +S+S+D + R+WDA
Sbjct: 222 GADDRQVKLWRMNETKAWEVDTLRGHMNNVSSVMFHAKQDII---VSNSEDKSIRVWDA 277
Score = 43.5 bits (101), Expect = 2e-04
Identities = 32/162 (19%), Positives = 61/162 (36%), Gaps = 22/162 (13%)
Query: 255 YVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPD 314
+++ ++++W + H G + + ++ L S +DY I+VW
Sbjct: 23 WILASLHSGVIQLWDYRMGTLIDRFDEHEGPVRGVHFHNSQPLFVSGGDDYKIKVWNYKT 82
Query: 315 GLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDARYTHSTPRLYVPKPSDS 374
+ L GH + + F + +S+SDD T RIW+
Sbjct: 83 HRCLFTLLGHLDYIRTVQFHHENPWI---VSASDDQTIRIWN------------------ 121
Query: 375 TGRSSGPSSNTMPQSHQIFCCAFNANGTVFVTGSSDNLARVY 416
+S S +H + C +F+ + V+ S D RV+
Sbjct: 122 -WQSRTCISVLTGHNHYVMCASFHPKEDLVVSASLDQTVRVW 162
Score = 33.5 bits (75), Expect = 0.22
Identities = 22/94 (23%), Positives = 42/94 (44%), Gaps = 1/94 (1%)
Query: 230 KMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDL 289
K + +RGH N V +F +++ S+D+ +++W + + R L
Sbjct: 237 KAWEVDTLRGHMNNVSSVMFHAKQDIIVSNSEDKSIRVWDATKRTGIQTFRREHDRFWIL 296
Query: 290 AVSSNNALVASSSNDYIIRVWRLPDGLPISVLRG 323
AV L+A+ ++ +I V++L P L G
Sbjct: 297 AVHPEINLLAAGHDNGMI-VFKLERERPAFALSG 329
>At1g18080 putative guanine nucleotide-binding protein beta
subunit
Length = 327
Score = 67.8 bits (164), Expect = 1e-11
Identities = 49/183 (26%), Positives = 91/183 (48%), Gaps = 13/183 (7%)
Query: 239 GHCNAVYCAIFDRSGRY--VITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNA 296
GH + V C F + +++ S D+ VK+W++ ++ GH G ++ +AVS + +
Sbjct: 148 GHRDWVSCVRFSPNTLQPTIVSASWDKTVKVWNLSNCKLRSTLAGHTGYVSTVAVSPDGS 207
Query: 297 LVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWD 356
L AS D ++ +W L +G + L ++ + A+ FSP Y L ++++ G +IWD
Sbjct: 208 LCASGGKDGVVLLWDLAEGKKLYSLEANS-VIHALCFSPNR---YWLCAATEHG-IKIWD 262
Query: 357 ARYTHSTPRLYVP-KPSDSTGRSSGPSSNTMPQSHQIFCCAFN--ANGTVFVTGSSDNLA 413
L V K +SGP++ + I+C + N A+G+ +G +D +
Sbjct: 263 LESKSIVEDLKVDLKAEAEKADNSGPAAT---KRKVIYCTSLNWSADGSTLFSGYTDGVI 319
Query: 414 RVY 416
RV+
Sbjct: 320 RVW 322
Score = 58.9 bits (141), Expect = 5e-09
Identities = 32/125 (25%), Positives = 61/125 (48%), Gaps = 4/125 (3%)
Query: 235 KRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSN 294
+R+ GH + V + G++ ++GS D +++W + S GH D+ +A S +
Sbjct: 57 RRLTGHSHFVEDVVLSSDGQFALSGSWDGELRLWDLAAGVSTRRFVGHTKDVLSVAFSLD 116
Query: 295 NALVASSSNDYIIRVWRLPDGLPISVL---RGHTGAVTAIAFSPRPNAVYQLLSSSDDGT 351
N + S+S D I++W ++ GH V+ + FSP ++S+S D T
Sbjct: 117 NRQIVSASRDRTIKLWNTLGECKYTISEGGEGHRDWVSCVRFSPN-TLQPTIVSASWDKT 175
Query: 352 CRIWD 356
++W+
Sbjct: 176 VKVWN 180
Score = 35.0 bits (79), Expect = 0.077
Identities = 29/92 (31%), Positives = 41/92 (44%), Gaps = 9/92 (9%)
Query: 280 RGHVGDITDLAVSSNNA-LVASSSNDYIIRVWRLPD-----GLPISVLRGHTGAVTAIAF 333
R H +T +A +NA ++ S+S D I +W+L G+ L GH+ V +
Sbjct: 12 RAHTDMVTAIATPIDNADIIVSASRDKSIILWKLTKDDKAYGVAQRRLTGHSHFVEDVVL 71
Query: 334 SPRPNAVYQLLSSSDDGTCRIWDARYTHSTPR 365
S LS S DG R+WD ST R
Sbjct: 72 SSDGQFA---LSGSWDGELRLWDLAAGVSTRR 100
>At3g18130 protein kinase C-receptor/G-protein, putative
Length = 326
Score = 65.9 bits (159), Expect = 4e-11
Identities = 48/182 (26%), Positives = 84/182 (45%), Gaps = 11/182 (6%)
Query: 239 GHCNAVYCAIFDRSGRY--VITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNA 296
GH V C F + +++ S D+ VK+W+++ S GH G + +AVS + +
Sbjct: 147 GHKEWVSCVRFSPNTLVPTIVSASWDKTVKVWNLQNCKLRNSLVGHSGYLNTVAVSPDGS 206
Query: 297 LVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWD 356
L AS D +I +W L +G + L + + ++ FSP L ++ + + RIWD
Sbjct: 207 LCASGGKDGVILLWDLAEGKKLYSLEAGS-IIHSLCFSPN----RYWLCAATENSIRIWD 261
Query: 357 ARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAFN--ANGTVFVTGSSDNLAR 414
L V S++ G + Q I+C + N A+G+ +G +D + R
Sbjct: 262 LESKSVVEDLKVDLKSEAEKNEGGVGTGN--QKKVIYCTSLNWSADGSTLFSGYTDGVVR 319
Query: 415 VY 416
V+
Sbjct: 320 VW 321
Score = 62.8 bits (151), Expect = 3e-10
Identities = 33/124 (26%), Positives = 64/124 (51%), Gaps = 3/124 (2%)
Query: 235 KRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSN 294
+R+ GH + V + G++ ++GS D +++W + T + GH D+ +A S++
Sbjct: 57 RRLTGHSHFVEDVVLSSDGQFALSGSWDGELRLWDLATGETTRRFVGHTKDVLSVAFSTD 116
Query: 295 NALVASSSNDYIIRVWRLPDGLPISVLR--GHTGAVTAIAFSPRPNAVYQLLSSSDDGTC 352
N + S+S D I++W ++ GH V+ + FSP V ++S+S D T
Sbjct: 117 NRQIVSASRDRTIKLWNTLGECKYTISEGDGHKEWVSCVRFSPN-TLVPTIVSASWDKTV 175
Query: 353 RIWD 356
++W+
Sbjct: 176 KVWN 179
Score = 33.5 bits (75), Expect = 0.22
Identities = 29/100 (29%), Positives = 44/100 (44%), Gaps = 10/100 (10%)
Query: 280 RGHVGDITDLAVSSNNA-LVASSSNDYIIRVWRLPD-----GLPISVLRGHTGAVTAIAF 333
R H +T +A +N+ ++ ++S D I +W+L G+ L GH+ V +
Sbjct: 12 RAHTDIVTAIATPIDNSDIIVTASRDKSIILWKLTKDDKSYGVAQRRLTGHSHFVEDVVL 71
Query: 334 SPRPNAVYQLLSSSDDGTCRIWDARYTHSTPRLYVPKPSD 373
S LS S DG R+WD T T R +V D
Sbjct: 72 SSDGQFA---LSGSWDGELRLWDLA-TGETTRRFVGHTKD 107
>At5g25150 transcription initiation factor IID-associated
factor-like protein
Length = 700
Score = 65.5 bits (158), Expect = 5e-11
Identities = 42/137 (30%), Positives = 63/137 (45%), Gaps = 10/137 (7%)
Query: 233 NIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVS 292
N+ +GH V+ A F G Y + S DR +IWSM+ L GH+ D+
Sbjct: 452 NLVCYKGHNYPVWDAQFSPFGHYFASCSHDRTARIWSMDRIQPLRIMAGHLSDVD---WH 508
Query: 293 SNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTC 352
N +A+ S+D +R+W + G + + GH V ++A SP + S +DGT
Sbjct: 509 PNCNYIATGSSDKTVRLWDVQTGECVRIFIGHRSMVLSLAMSPDGR---YMASGDEDGTI 565
Query: 353 RIWDARYTHSTPRLYVP 369
+WD ST R P
Sbjct: 566 MMWDL----STARCITP 578
Score = 63.9 bits (154), Expect = 2e-10
Identities = 41/148 (27%), Positives = 65/148 (43%), Gaps = 7/148 (4%)
Query: 239 GHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALV 298
GH VY A F G +V++ S D +++WS + +L +GH + D S
Sbjct: 416 GHSGPVYSATFSPPGDFVLSSSADTTIRLWSTKLNANLVCYKGHNYPVWDAQFSPFGHYF 475
Query: 299 ASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDAR 358
AS S+D R+W + P+ ++ GH V PN Y + + S D T R+WD +
Sbjct: 476 ASCSHDRTARIWSMDRIQPLRIMAGHLSDV-----DWHPNCNY-IATGSSDKTVRLWDVQ 529
Query: 359 YTHSTPRLYVPKPSDSTGRSSGPSSNTM 386
T R+++ S + P M
Sbjct: 530 -TGECVRIFIGHRSMVLSLAMSPDGRYM 556
Score = 51.6 bits (122), Expect = 8e-07
Identities = 28/106 (26%), Positives = 54/106 (50%), Gaps = 3/106 (2%)
Query: 228 VQKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDIT 287
+ ++Q ++ + GH + V + + Y+ TGS D+ V++W ++T + GH +
Sbjct: 489 MDRIQPLRIMAGHLSDVD---WHPNCNYIATGSSDKTVRLWDVQTGECVRIFIGHRSMVL 545
Query: 288 DLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAF 333
LA+S + +AS D I +W L I+ L GH V ++++
Sbjct: 546 SLAMSPDGRYMASGDEDGTIMMWDLSTARCITPLMGHNSCVWSLSY 591
Score = 40.8 bits (94), Expect = 0.001
Identities = 36/140 (25%), Positives = 47/140 (32%), Gaps = 31/140 (22%)
Query: 242 NAVYCAIFDRSGRYVITGSDDRLVKIWSM--------------------------ETAYS 275
N + C+ G V G D +K+W M +Y+
Sbjct: 353 NGLNCSSISHDGSLVAGGFSDSSIKVWDMAKIGQAGSGALQAENDSSDQSIGPNGRRSYT 412
Query: 276 LASCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSP 335
L GH G + S V SSS D IR+W + +GH V FSP
Sbjct: 413 L--LLGHSGPVYSATFSPPGDFVLSSSADTTIRLWSTKLNANLVCYKGHNYPVWDAQFSP 470
Query: 336 RPNAVYQLLSSSDDGTCRIW 355
+ S S D T RIW
Sbjct: 471 ---FGHYFASCSHDRTARIW 487
>At1g73720 unknown protein
Length = 511
Score = 65.5 bits (158), Expect = 5e-11
Identities = 43/168 (25%), Positives = 76/168 (44%), Gaps = 9/168 (5%)
Query: 240 HCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVA 299
H + V C F R + +GS D +KIW + T + H +T L+ S + + +
Sbjct: 262 HDDPVLCIDFSRDSEMLASGSQDGKIKIWRIRTGVCIRRFDAHSQGVTSLSFSRDGSQLL 321
Query: 300 SSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDARY 359
S+S D R+ L G + RGHT V F+ + +++++S D T ++WD++
Sbjct: 322 STSFDQTARIHGLKSGKLLKEFRGHTSYVNHAIFTSDGS---RIITASSDCTVKVWDSKT 378
Query: 360 THSTPRLYVPKPSDSTGRSSGPSSNTMPQS--HQIFCCAFNANGTVFV 405
T P P T S S + P++ H + C N ++++
Sbjct: 379 TDCLQTFKPPPPLRGTDASVN-SIHLFPKNTEHIVVC---NKTSSIYI 422
Score = 48.5 bits (114), Expect = 7e-06
Identities = 34/117 (29%), Positives = 58/117 (49%), Gaps = 11/117 (9%)
Query: 246 CAIFDRSGRYVITGSDDRLVKIW-------SMETAYSL-ASCRGHVGDITDLAVSSNNAL 297
CA F G+++ + S D +++W + Y S H + + S ++ +
Sbjct: 218 CARFSPDGQFLASSSVDGFIEVWDYISGKLKKDLQYQADESFMMHDDPVLCIDFSRDSEM 277
Query: 298 VASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRI 354
+AS S D I++WR+ G+ I H+ VT+++FS + QLLS+S D T RI
Sbjct: 278 LASGSQDGKIKIWRIRTGVCIRRFDAHSQGVTSLSFSRDGS---QLLSTSFDQTARI 331
Score = 43.9 bits (102), Expect = 2e-04
Identities = 28/109 (25%), Positives = 51/109 (46%), Gaps = 6/109 (5%)
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSS 293
I+R H V F R G +++ S D+ +I +++ L RGH + +S
Sbjct: 298 IRRFDAHSQGVTSLSFSRDGSQLLSTSFDQTARIHGLKSGKLLKEFRGHTSYVNHAIFTS 357
Query: 294 NNALVASSSNDYIIRVW--RLPDGL----PISVLRGHTGAVTAIAFSPR 336
+ + + ++S+D ++VW + D L P LRG +V +I P+
Sbjct: 358 DGSRIITASSDCTVKVWDSKTTDCLQTFKPPPPLRGTDASVNSIHLFPK 406
>At1g48630 unknown protein
Length = 326
Score = 64.7 bits (156), Expect = 9e-11
Identities = 36/124 (29%), Positives = 64/124 (51%), Gaps = 3/124 (2%)
Query: 235 KRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSN 294
+R+ GH + V + G++ ++GS D +++W + T S GH D+ +A S++
Sbjct: 57 RRMTGHSHFVQDVVLSSDGQFALSGSWDGELRLWDLATGESTRRFVGHTKDVLSVAFSTD 116
Query: 295 NALVASSSNDYIIRVWRL--PDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTC 352
N + S+S D I++W IS GH V+ + FSP V ++S+S D T
Sbjct: 117 NRQIVSASRDRTIKLWNTLGECKYTISEADGHKEWVSCVRFSPN-TLVPTIVSASWDKTV 175
Query: 353 RIWD 356
++W+
Sbjct: 176 KVWN 179
Score = 62.4 bits (150), Expect = 4e-10
Identities = 50/198 (25%), Positives = 90/198 (45%), Gaps = 15/198 (7%)
Query: 225 STMVQKMQNIKRIRGHCNAVYCAIFDRSGRY--VITGSDDRLVKIWSMETAYSLASCRGH 282
+T+ + I GH V C F + +++ S D+ VK+W+++ + GH
Sbjct: 133 NTLGECKYTISEADGHKEWVSCVRFSPNTLVPTIVSASWDKTVKVWNLQNCKLRNTLAGH 192
Query: 283 VGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQ 342
G + +AVS + +L AS D +I +W L +G + L + + ++ FSP
Sbjct: 193 SGYLNTVAVSPDGSLCASGGKDGVILLWDLAEGKKLYSLEAGS-IIHSLCFSPN----RY 247
Query: 343 LLSSSDDGTCRIWDARYTHSTPRLYV--PKPSDSTGRSSGPSSNTMPQSHQIFCCAFN-- 398
L ++ + + RIWD L V ++ T S+G + T I+C + N
Sbjct: 248 WLCAATENSIRIWDLESKSVVEDLKVDLKAEAEKTDGSTGIGNKT----KVIYCTSLNWS 303
Query: 399 ANGTVFVTGSSDNLARVY 416
A+G +G +D + RV+
Sbjct: 304 ADGNTLFSGYTDGVIRVW 321
Score = 60.1 bits (144), Expect = 2e-09
Identities = 35/129 (27%), Positives = 62/129 (47%), Gaps = 7/129 (5%)
Query: 232 QNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSM--ETAYSLASCRGHVGDITDL 289
++ +R GH V F R +++ S DR +K+W+ E Y+++ GH ++ +
Sbjct: 96 ESTRRFVGHTKDVLSVAFSTDNRQIVSASRDRTIKLWNTLGECKYTISEADGHKEWVSCV 155
Query: 290 AVSSNNAL--VASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSS 347
S N + + S+S D ++VW L + + L GH+G + +A SP + S
Sbjct: 156 RFSPNTLVPTIVSASWDKTVKVWNLQNCKLRNTLAGHSGYLNTVAVSPDGSL---CASGG 212
Query: 348 DDGTCRIWD 356
DG +WD
Sbjct: 213 KDGVILLWD 221
Score = 32.0 bits (71), Expect = 0.65
Identities = 26/90 (28%), Positives = 40/90 (43%), Gaps = 9/90 (10%)
Query: 282 HVGDITDLAVSSNNA-LVASSSNDYIIRVWRLPD-----GLPISVLRGHTGAVTAIAFSP 335
H +T +A +N+ ++ +SS D I +W+L G+ + GH+ V + S
Sbjct: 14 HTDMVTAIATPVDNSDVIVTSSRDKSIILWKLTKEDKSYGVAQRRMTGHSHFVQDVVLSS 73
Query: 336 RPNAVYQLLSSSDDGTCRIWDARYTHSTPR 365
LS S DG R+WD ST R
Sbjct: 74 DGQFA---LSGSWDGELRLWDLATGESTRR 100
>At1g15440 unknown protein
Length = 900
Score = 64.3 bits (155), Expect = 1e-10
Identities = 41/138 (29%), Positives = 68/138 (48%), Gaps = 11/138 (7%)
Query: 247 AIFDRSGRYVITGSDDR-LVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDY 305
A+F+ G ++ G + +W T + +GH D+ + S ++ L+A+ ++D
Sbjct: 352 AVFNERGNWLTFGCAKLGQLLVWDWRTETYILKQQGHYFDVNCVTYSPDSQLLATGADDN 411
Query: 306 IIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDAR------- 358
++VW + G HT AVTA+ F ++ LLS+S DGT R WD +
Sbjct: 412 KVKVWNVMSGTCFITFTEHTNAVTALHFMADNHS---LLSASLDGTVRAWDFKRYKNYKT 468
Query: 359 YTHSTPRLYVPKPSDSTG 376
YT TPR +V +D +G
Sbjct: 469 YTTPTPRQFVSLTADPSG 486
Score = 46.6 bits (109), Expect = 3e-05
Identities = 35/108 (32%), Positives = 46/108 (42%), Gaps = 4/108 (3%)
Query: 250 DRSGRYVITGSDDRL-VKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIR 308
D SG V G+ D + +WS +T GH + L S L+ASSS DY +R
Sbjct: 483 DPSGDVVCAGTLDSFEIFVWSKKTGQIKDILSGHEAPVHGLMFSPLTQLLASSSWDYTVR 542
Query: 309 VWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWD 356
+W + H V +AF P QL SS+ DG WD
Sbjct: 543 LWDVFASKGTVETFRHNHDVLTVAFRPDGK---QLASSTLDGQINFWD 587
Score = 45.4 bits (106), Expect = 6e-05
Identities = 40/168 (23%), Positives = 62/168 (36%), Gaps = 49/168 (29%)
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSS 293
I + +GH V C + + + TG+DD VK+W++ + + H +T L +
Sbjct: 382 ILKQQGHYFDVNCVTYSPDSQLLATGADDNKVKVWNVMSGTCFITFTEHTNAVTALHFMA 441
Query: 294 NNALVASSSNDYIIR--------------------------------------------V 309
+N + S+S D +R V
Sbjct: 442 DNHSLLSASLDGTVRAWDFKRYKNYKTYTTPTPRQFVSLTADPSGDVVCAGTLDSFEIFV 501
Query: 310 WRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSD-DGTCRIWD 356
W G +L GH V + FSP + QLL+SS D T R+WD
Sbjct: 502 WSKKTGQIKDILSGHEAPVHGLMFSP----LTQLLASSSWDYTVRLWD 545
Score = 31.6 bits (70), Expect = 0.85
Identities = 24/116 (20%), Positives = 47/116 (39%), Gaps = 5/116 (4%)
Query: 237 IRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNA 296
+ GH V+ +F + + + S D V++W + + H D+ +A +
Sbjct: 513 LSGHEAPVHGLMFSPLTQLLASSSWDYTVRLWDVFASKGTVETFRHNHDVLTVAFRPDGK 572
Query: 297 LVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTC 352
+ASS+ D I W +G+ + + G + R +A ++S G C
Sbjct: 573 QLASSTLDGQINFWDTIEGVLMYTIEGRRDIAGGRVMTDRRSA-----ANSSSGKC 623
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.134 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,393,276
Number of Sequences: 26719
Number of extensions: 476257
Number of successful extensions: 2239
Number of sequences better than 10.0: 197
Number of HSP's better than 10.0 without gapping: 130
Number of HSP's successfully gapped in prelim test: 68
Number of HSP's that attempted gapping in prelim test: 1284
Number of HSP's gapped (non-prelim): 708
length of query: 417
length of database: 11,318,596
effective HSP length: 102
effective length of query: 315
effective length of database: 8,593,258
effective search space: 2706876270
effective search space used: 2706876270
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC144539.12


