

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144539.1 + phase: 0
(163 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
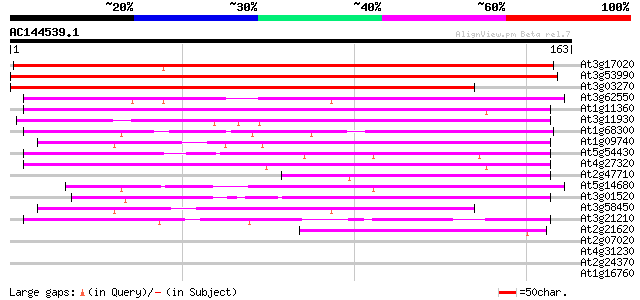
Score E
Sequences producing significant alignments: (bits) Value
At3g17020 unknown protein 196 5e-51
At3g53990 unknown protein 151 2e-37
At3g03270 unknown protein 140 3e-34
At3g62550 unknown protein 84 3e-17
At1g11360 unknown protein 79 8e-16
At3g11930 unknown protein 79 1e-15
At1g68300 unknown protein 77 5e-15
At1g09740 putative ER6 protein 76 9e-15
At5g54430 unknown protein 67 4e-12
At4g27320 unknown protein 65 2e-11
At2g47710 unknown protein 57 6e-09
At5g14680 unknown protein 54 3e-08
At3g01520 unknown protein 52 1e-07
At3g58450 putative protein 49 9e-07
At3g21210 unknown protein 48 2e-06
At2g21620 RD2 protein (RD2) 44 4e-05
At2g07020 putative protein kinase 35 0.018
At4g31230 putative protein 32 0.20
At2g24370 putative protein kinase 30 0.44
At1g16760 hypothetical protein 30 0.57
>At3g17020 unknown protein
Length = 163
Score = 196 bits (498), Expect = 5e-51
Identities = 91/158 (57%), Positives = 117/158 (73%), Gaps = 1/158 (0%)
Query: 2 ASARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPE-EYEHGEMQLWEVTGSPL 60
+ RR+G+A+DFS CS KA W +DN+V++GD+LILI I + YE GEMQLWE GSP
Sbjct: 4 SGGRRIGVAVDFSDCSKKALSWAIDNVVRDGDHLILITIAHDMNYEEGEMQLWETVGSPF 63
Query: 61 TPLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPL 120
P+ EF ++ + KKY +K D E L I TA +K + V++K+YWGD REK+C A EQ+PL
Sbjct: 64 IPMSEFSDAAVMKKYALKPDAETLDIVNTAARKKTITVVMKIYWGDPREKICAAAEQIPL 123
Query: 121 DGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKS 158
L MGNRGLG L+R IMGSVSN+VVNN +CPVTVVK+
Sbjct: 124 SSLVMGNRGLGGLKRMIMGSVSNHVVNNVACPVTVVKA 161
>At3g53990 unknown protein
Length = 160
Score = 151 bits (381), Expect = 2e-37
Identities = 73/159 (45%), Positives = 103/159 (63%)
Query: 1 MASARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPL 60
M R +GIAMDFS S A +W ++N+ +GD + +I P + LW +GSPL
Sbjct: 1 MPKDRNIGIAMDFSESSKNALKWAIENLADKGDTIYIIHTLPLSGDESRNSLWFKSGSPL 60
Query: 61 TPLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPL 120
PL EF ++ +KY +KTD L + T QK+V V+ K+YWGDAREKL +A++ + L
Sbjct: 61 IPLAEFREPEIMEKYGVKTDIACLDMLDTGSRQKEVHVVTKLYWGDAREKLVDAVKDLKL 120
Query: 121 DGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKSS 159
D + MG+RGL L+R IMGSVS++V+ +A CPVTVVK +
Sbjct: 121 DSIVMGSRGLSALQRIIMGSVSSFVIQHAPCPVTVVKDN 159
>At3g03270 unknown protein
Length = 201
Score = 140 bits (353), Expect = 3e-34
Identities = 65/135 (48%), Positives = 93/135 (68%)
Query: 1 MASARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPL 60
M AR +G+ MD+SP S A +W +N++++GD +ILI ++P+ +H L+E TGSPL
Sbjct: 1 MGKARTVGVGMDYSPTSKLALRWAAENLLEDGDTVILIHVQPQNADHTRKILFEETGSPL 60
Query: 61 TPLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPL 120
PL EF +L K+Y + DPEVL + T KKV V+ KVYWGD REKLC+A+E + L
Sbjct: 61 IPLEEFREVNLSKQYGLAYDPEVLDVLDTLSRAKKVKVVAKVYWGDPREKLCDAVENLKL 120
Query: 121 DGLTMGNRGLGTLRR 135
D + +G+RGLG+L+R
Sbjct: 121 DSIVLGSRGLGSLKR 135
>At3g62550 unknown protein
Length = 162
Score = 84.3 bits (207), Expect = 3e-17
Identities = 52/163 (31%), Positives = 86/163 (51%), Gaps = 15/163 (9%)
Query: 5 RRLGIAMDFSPCSIKAFQWTVDNIVKEGDN--LILIIIRPE--EYEHGEMQLWEVTGSPL 60
R++ +A+D S S++A W++DN+ G N LIL+ ++P Y + + VTG P+
Sbjct: 7 RKIVVAVDESEESMEALSWSLDNLFPYGSNNTLILLYVKPPLPVYSSLDAAGFIVTGDPV 66
Query: 61 TPLGEFINSDLPKKYEIKTDPEVLKIATTAIE--QKKVVVLVKVYWGDAREKLCEAIEQV 118
L KKYE + V+ + T + + + + +V GDA+E +C A++++
Sbjct: 67 AAL---------KKYEYELVESVMARSRTVYQDYESDINIERRVGRGDAKEVICNAVQKL 117
Query: 119 PLDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKSSGQ 161
+D L MG G +RA++GSVS Y CPV +VK Q
Sbjct: 118 RVDMLVMGTHDYGFFKRALLGSVSEYCAKRVKCPVVIVKKQAQ 160
>At1g11360 unknown protein
Length = 242
Score = 79.3 bits (194), Expect = 8e-16
Identities = 49/156 (31%), Positives = 81/156 (51%), Gaps = 3/156 (1%)
Query: 5 RRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPLTPLG 64
R++GIA+D S S A QW V N ++ GD ++L+ ++P +G P
Sbjct: 38 RKIGIAVDLSDESAYAVQWAVQNYLRSGDAVVLLHVQPTSVLYGADWGAMDLSPQWDPNN 97
Query: 65 EFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPLDGLT 124
E L ++I T+ + +A +E + V D +E+LC +E++ L L
Sbjct: 98 EESQRKLEDDFDIVTNKKASDVAQPLVEADIPFKIHIVKDHDMKERLCLEVERLGLSTLI 157
Query: 125 MGNRGLGTLRRAI---MGSVSNYVVNNASCPVTVVK 157
MG+RG G +R+ +GSVS+Y V++ +CPV VV+
Sbjct: 158 MGSRGFGATKRSSKGRLGSVSDYSVHHCACPVVVVR 193
>At3g11930 unknown protein
Length = 200
Score = 79.0 bits (193), Expect = 1e-15
Identities = 51/168 (30%), Positives = 88/168 (52%), Gaps = 18/168 (10%)
Query: 3 SARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGS---- 58
+ +R+ +A+D S S A QW +D+ +L+ E E G + + V
Sbjct: 31 TTKRMVVAIDESDSSFYALQWVIDHFSN-----LLLTTAAAEAESGMLTVIHVQSPFNHF 85
Query: 59 ---PLTPLGE---FINSDL---PKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDARE 109
P P G + +S + KK + +T +L A K++ V G+A+E
Sbjct: 86 AAFPAGPGGATAVYASSSMIESVKKAQQETSAALLSRALQMCRAKQIRTETLVLEGEAKE 145
Query: 110 KLCEAIEQVPLDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVK 157
+CEA+E++ +D L +G+RGLG ++RA +GSVS+Y ++A+CP+ +VK
Sbjct: 146 MICEAVEKMHVDLLVVGSRGLGKIKRAFLGSVSDYCAHHANCPILIVK 193
>At1g68300 unknown protein
Length = 160
Score = 76.6 bits (187), Expect = 5e-15
Identities = 54/159 (33%), Positives = 87/159 (53%), Gaps = 15/159 (9%)
Query: 5 RRLGIAMDFSPCSIKAFQWTVDNIVKE--GDNLILIIIRPEEYEHGEMQLWEVTGSPLTP 62
+++ +A+D S CS +A QWT+ + ++IL +P H ++ + P
Sbjct: 10 KQVMVAIDESECSKRALQWTLVYLKDSLADSDIILFTAQP----HLDLSCVYASSYGAAP 65
Query: 63 LGEFINS--DLPKKYEIKTDPEVLKI-ATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVP 119
+ E INS + K + E KI A T + +KV+ +G+ +E +CEA E++
Sbjct: 66 I-ELINSLQESHKNAGLNRLDEGTKICAETGVTPRKVLE-----FGNPKEAICEAAEKLG 119
Query: 120 LDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKS 158
+D L +G+ G G L+R +GSVSNY VNNA CPV VV++
Sbjct: 120 VDMLVVGSHGKGALQRTFLGSVSNYCVNNAKCPVLVVRT 158
>At1g09740 putative ER6 protein
Length = 171
Score = 75.9 bits (185), Expect = 9e-15
Identities = 47/160 (29%), Positives = 84/160 (52%), Gaps = 18/160 (11%)
Query: 9 IAMDFSPCSIKAFQWTVDNIV----KEGDNLILIIIRPEEYEHGEMQLWEVTGSPLT-PL 63
+A+D S S++A +W +DN+ + +++ ++P + SP T P
Sbjct: 12 VAVDGSEVSMEALRWALDNLKLSSSSSDSSFVVLHVQPSPSVAAGV-------SPGTIPF 64
Query: 64 GEFINSDLP------KKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQ 117
G ++P ++++ + +L+ A+ +K V V +V GD + K+CEA+E
Sbjct: 65 GGPSGLEVPAFTAAIEQHQKRITDTILEHASQICAEKSVNVKTQVVIGDPKYKICEAVEN 124
Query: 118 VPLDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVK 157
+ D L MG+R G ++R +GSVSNY N+A CPV ++K
Sbjct: 125 LHADLLVMGSRAYGRIKRMFLGSVSNYCTNHAHCPVVIIK 164
>At5g54430 unknown protein
Length = 250
Score = 67.0 bits (162), Expect = 4e-12
Identities = 50/163 (30%), Positives = 83/163 (50%), Gaps = 17/163 (10%)
Query: 5 RRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPLTPLG 64
R++G+A+D S S A +W VD+ ++ GD ++L+ + P L+ PL PL
Sbjct: 48 RKIGVAVDLSEESSFAVRWAVDHYIRPGDAVVLLHVSPTSV------LFGADWGPL-PLK 100
Query: 65 EFINSDLPKKYEIKTDPEVL---KIATTAIEQKKVVVLVKVYW---GDAREKLCEAIEQV 118
I + + D + K+A A K++ K++ D RE+LC IE++
Sbjct: 101 TQIEDPNAQPQPSQEDFDAFTSTKVADLAKPLKELGFPYKIHIVKDHDMRERLCLEIERL 160
Query: 119 PLDGLTMGNRGLGTLRR----AIMGSVSNYVVNNASCPVTVVK 157
L + MG+RG G ++ +GSVS+Y V++ CPV VV+
Sbjct: 161 GLSAVIMGSRGFGAEKKRGSDGKLGSVSDYCVHHCVCPVVVVR 203
>At4g27320 unknown protein
Length = 260
Score = 64.7 bits (156), Expect = 2e-11
Identities = 47/160 (29%), Positives = 77/160 (47%), Gaps = 7/160 (4%)
Query: 5 RRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPLTPLG 64
R++G+A+D S S A +W VD+ ++ GD ++++ + P G +P P
Sbjct: 45 RKIGVAVDLSEESAFAVRWAVDHYIRPGDAVVILHVSPTSVLFGADWGPLPLQTPPPPSA 104
Query: 65 EFINSDLPK----KYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPL 120
PK ++ T +V +A E + V D RE+LC E++ L
Sbjct: 105 ATDPGAQPKPSQEDFDAFTSSKVADLAKPLKEAGFPHKIHIVKDHDMRERLCLETERLNL 164
Query: 121 DGLTMGNRGLGTLRRAI---MGSVSNYVVNNASCPVTVVK 157
+ MG+RG G +R +GSVS+Y V++ CPV VV+
Sbjct: 165 SAVIMGSRGFGAEKRGSDGKLGSVSDYCVHHCVCPVVVVR 204
>At2g47710 unknown protein
Length = 162
Score = 56.6 bits (135), Expect = 6e-09
Identities = 27/86 (31%), Positives = 49/86 (56%), Gaps = 8/86 (9%)
Query: 80 DPEVLKIATTAIEQKKVV--------VLVKVYWGDAREKLCEAIEQVPLDGLTMGNRGLG 131
D ++ A +E+ K + +++V+ GDAR LCE +++ L +G+ G G
Sbjct: 71 DADLKHTAAKVVEKAKAICQSRSVHGAVIEVFEGDARNILCEVVDKHHASILVVGSHGYG 130
Query: 132 TLRRAIMGSVSNYVVNNASCPVTVVK 157
++RA++GS S+Y ++A C V +VK
Sbjct: 131 AIKRAVLGSTSDYCAHHAHCSVMIVK 156
>At5g14680 unknown protein
Length = 175
Score = 54.3 bits (129), Expect = 3e-08
Identities = 39/151 (25%), Positives = 74/151 (48%), Gaps = 17/151 (11%)
Query: 17 SIKAFQWTVDNIVKE---GDNLILIIIRPEEYEHGEMQLWEVTGSPLTPLGEFINSDLPK 73
S KAF+WT+ IV+ G L+L+ ++ ++ E G + + SP D +
Sbjct: 27 SKKAFEWTLKKIVRSNTSGFKLLLLHVQVQD-EDGFDDMDSIYASP----------DDFR 75
Query: 74 KYEIKTDPEVLKIATTAIEQKKVVVLVKVYW---GDAREKLCEAIEQVPLDGLTMGNRGL 130
+ + + L + +++ + + W GD E +C + +V D L +G+RGL
Sbjct: 76 QMRERNKAKGLHLLEFFVKKCHDIGVGCEAWIRKGDPTELICHEVRRVRPDFLVVGSRGL 135
Query: 131 GTLRRAIMGSVSNYVVNNASCPVTVVKSSGQ 161
G ++ +G+VS + V +A CPV +K + +
Sbjct: 136 GPFQKVFVGTVSEFCVKHAECPVITIKRTAE 166
>At3g01520 unknown protein
Length = 175
Score = 52.4 bits (124), Expect = 1e-07
Identities = 38/141 (26%), Positives = 71/141 (49%), Gaps = 9/141 (6%)
Query: 19 KAFQWTVDNIVKEG--DNLILIIIRPEEYEHGEMQLWEVTGSPLTPLGEFINSDLPKKYE 76
+AF+WT++ IV+ D IL++ E G + + SP +F D+ + +
Sbjct: 29 RAFEWTLEKIVRSNTSDFKILLLHVQVVDEDGFDDVDSIYASP----EDF--RDMRQSNK 82
Query: 77 IKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPLDGLTMGNRGLGTLRRA 136
K +L+ + V + GD ++ +C+ +++V D L +G+RGLG ++
Sbjct: 83 AK-GLHLLEFFVNKCHEIGVGCEAWIKTGDPKDVICQEVKRVRPDFLVVGSRGLGRFQKV 141
Query: 137 IMGSVSNYVVNNASCPVTVVK 157
+G+VS + V +A CPV +K
Sbjct: 142 FVGTVSAFCVKHAECPVMTIK 162
>At3g58450 putative protein
Length = 174
Score = 49.3 bits (116), Expect = 9e-07
Identities = 36/140 (25%), Positives = 63/140 (44%), Gaps = 20/140 (14%)
Query: 9 IAMDFSPCSIKAFQWTVDNIV----------KEGDNLILIIIRPEEYEHGEMQLWEVTGS 58
+A+D S S A +W VD++ +EG L L+ + P ++ + S
Sbjct: 34 VAIDESKNSFDALEWAVDHLRVVISAEPETGQEGGLLTLLHVHPTYLQY-------IYPS 86
Query: 59 PLTPLGEFINSDLPKKYEIKTDPEVLKIATTAIE---QKKVVVLVKVYWGDAREKLCEAI 115
T + +P+ + + T A+E K V + GD +E +C+A+
Sbjct: 87 GGTASAVYATDSVPEPMRKAREESTTNLFTRALEICRGKMVKTETMILEGDPKEMICQAV 146
Query: 116 EQVPLDGLTMGNRGLGTLRR 135
EQ +D L +G+RGLG ++R
Sbjct: 147 EQTHVDLLVVGSRGLGMIKR 166
>At3g21210 unknown protein
Length = 686
Score = 48.1 bits (113), Expect = 2e-06
Identities = 44/160 (27%), Positives = 73/160 (45%), Gaps = 35/160 (21%)
Query: 5 RRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRP------EEYEHGEMQLWEVTGS 58
R++GIA++ S S +W VDN +++GD++I++ + P ++ + +Q T
Sbjct: 15 RKIGIAVELSEESAFTVRWAVDNYIRQGDDIIILHVSPTAGLFGADWGYYPLQ----TQP 70
Query: 59 PLTPLGEFIN-SDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQ 117
P T F +DL K + P + VK Y D RE+LC ++
Sbjct: 71 PYTTASIFSKVADLGKPLKEAGFPHTIH-------------TVKDY--DKRERLCLETQR 115
Query: 118 VPLDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVK 157
+ L L MG GSVS++ V++ C V VV+
Sbjct: 116 LNLTALIMGFGD---------GSVSDFCVHHCVCQVVVVR 146
>At2g21620 RD2 protein (RD2)
Length = 187
Score = 43.9 bits (102), Expect = 4e-05
Identities = 25/73 (34%), Positives = 39/73 (53%), Gaps = 1/73 (1%)
Query: 85 KIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPLDGLTMGNRGLGTLRRAIMGSVSNY 144
K+A A + V + +V GDA + +C+ E+V + +G RG +R + GSVS Y
Sbjct: 94 KLAVEAYQVAMVKSVARVVEGDAGKVICKEAEKVKPAAVIVGTRGRSLVRSVLQGSVSEY 153
Query: 145 VVNNA-SCPVTVV 156
+N S PV +V
Sbjct: 154 CFHNCKSAPVIIV 166
>At2g07020 putative protein kinase
Length = 620
Score = 35.0 bits (79), Expect = 0.018
Identities = 37/147 (25%), Positives = 65/147 (44%), Gaps = 15/147 (10%)
Query: 7 LGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPLTPLGEF 66
+ IA+D S A +W VDN++ G+ L LI ++ + Q G+ G+
Sbjct: 11 VAIAIDRDKGSQAALKWAVDNLLTPGETLTLIHVKVK-------QTLANNGTQPNKSGDD 63
Query: 67 INSDLPKKYEIKTDPEVLKIATTAIEQKKV----VVLVKVYWGDAREKLCEAIEQVPLDG 122
+ +L + + + A+ I K+ VVL V DA E + E +++ +D
Sbjct: 64 V-KELFLPFRCFCTRKDVSFASNFINLLKINCEEVVLENV---DAAEGIIEYVQENAIDI 119
Query: 123 LTMGNRGLGTLRRAIMGSVSNYVVNNA 149
L +G + L+R V+N V+ A
Sbjct: 120 LVLGASKITLLKRLKAVDVTNAVIKGA 146
>At4g31230 putative protein
Length = 537
Score = 31.6 bits (70), Expect = 0.20
Identities = 11/36 (30%), Positives = 24/36 (66%)
Query: 7 LGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRP 42
+ +A+D S A +W VDN++++G ++L+ ++P
Sbjct: 18 VAVAIDRDKNSQGALKWAVDNLLQKGQTVVLVHVKP 53
>At2g24370 putative protein kinase
Length = 816
Score = 30.4 bits (67), Expect = 0.44
Identities = 11/35 (31%), Positives = 23/35 (65%)
Query: 7 LGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIR 41
+ +A+D S A +W VDN+++ G ++IL+ ++
Sbjct: 20 VAVAIDKDKSSQHALKWAVDNLLQRGQSVILVHVK 54
>At1g16760 hypothetical protein
Length = 758
Score = 30.0 bits (66), Expect = 0.57
Identities = 13/37 (35%), Positives = 21/37 (56%)
Query: 7 LGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPE 43
+ IA+D S A +WT++N+ G L LI + P+
Sbjct: 18 VAIAIDKDKSSQNAIKWTLENLATRGQTLALIHVLPK 54
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.135 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,544,768
Number of Sequences: 26719
Number of extensions: 141810
Number of successful extensions: 361
Number of sequences better than 10.0: 35
Number of HSP's better than 10.0 without gapping: 29
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 314
Number of HSP's gapped (non-prelim): 38
length of query: 163
length of database: 11,318,596
effective HSP length: 92
effective length of query: 71
effective length of database: 8,860,448
effective search space: 629091808
effective search space used: 629091808
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC144539.1


