

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144538.7 + phase: 1 /pseudo
(632 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
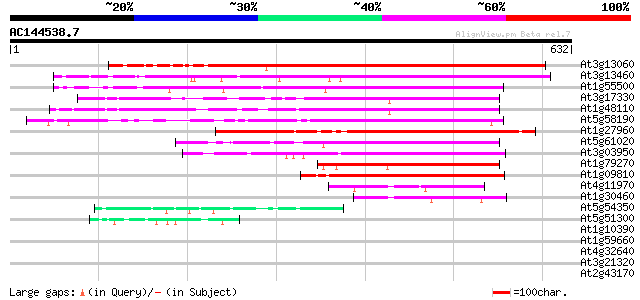
Score E
Sequences producing significant alignments: (bits) Value
At3g13060 unknown protein 556 e-158
At3g13460 unknown protein 374 e-104
At1g55500 unknown protein 348 4e-96
At3g17330 unknown protein 314 1e-85
At1g48110 unknown protein 311 5e-85
At5g58190 unknown protein 305 4e-83
At1g27960 hypothetical protein 305 7e-83
At5g61020 unknown protein 284 9e-77
At3g03950 unknown protein 273 2e-73
At1g79270 unknown protein 271 1e-72
At1g09810 unknown protein 263 2e-70
At4g11970 unknown protein 79 8e-15
At1g30460 hypothetical protein 64 2e-10
At5g54350 unknown protein 44 4e-04
At5g51300 unknown protein 42 8e-04
At1g10390 unknown protein 40 0.003
At1g59660 nucleoporin 98-like protein 39 0.007
At4g32640 putative protein 39 0.009
At3g21320 hypothetical protein 38 0.015
At2g43170 unknown protein 38 0.015
>At3g13060 unknown protein
Length = 539
Score = 556 bits (1434), Expect = e-158
Identities = 295/505 (58%), Positives = 373/505 (73%), Gaps = 32/505 (6%)
Query: 112 MPYGPYSPVTSPLPSVGGDAQLYSPQQFPYT--PPYYNQLVPPPNLPYLNSPTPVSQPEL 169
MPYGPYSP SPLPS G QLYSPQQFP++ PYY Q+VPP ++ Y+ SPT QPEL
Sbjct: 1 MPYGPYSPAASPLPSEG---QLYSPQQFPFSGASPYYQQVVPP-SMQYITSPT---QPEL 53
Query: 170 TNLLGIDQQVESNFFGPRASYPS--VGSFARGSFPVAPGSFSFHESQQGFDGSRSGGLWS 227
T+L+G+DQQ ++ GPR SY +G F G+ P + F E QQGFDG G+WS
Sbjct: 54 TSLVGVDQQGDN--IGPRQSYHPHPIGPF-NGNQP----NLGFPEWQQGFDG----GIWS 102
Query: 228 DSSKPSERQRSFMPLSPSVPQQPIGSVRSFGPSAGMASHQQRSLYGFGSSSNPYGRGYLS 287
D SKPS+ R +SP++ QP+GS S+G + M S +QRS YGFGS SN Y RGY+
Sbjct: 103 DWSKPSDMHRHSSSISPALSPQPLGSYGSYGQNIPMGSQRQRSFYGFGSGSNSYNRGYMH 162
Query: 288 N----PGSSFGGSAISGLSANDRSFLSLENSRRHGR--ETASFCRCNGTLDILSEQNRGP 341
+ GS++G IS + ++ ++ ++NSR GR + + NGT DIL+EQNRGP
Sbjct: 163 SGGRGQGSNYGSRLISNVGMGNQGWIGVDNSRGRGRVSDPSLGGAYNGTFDILNEQNRGP 222
Query: 342 RASKLKNHISSE-NNAIDGSKNNASTAKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVH 400
RASK K + E ++A D KNN +AK +ES N DF TD+ +AK F+IKSYSEDNVH
Sbjct: 223 RASKPKTQVLEELDSAADSKKNNKGSAKEHEES-NNADFVTDYTNAKLFIIKSYSEDNVH 281
Query: 401 KSIKYGVWASTPNGNRKLDAAYCQAKEKQDACRIFLFFSVNASAQFCGVAEMVGPVNFDK 460
KSIKY VWASTPNGN+KLDAAY +AK++++ C +FL FSVNAS+QFCGVAEMVGPV+F+K
Sbjct: 282 KSIKYNVWASTPNGNKKLDAAYREAKDEKEPCPLFLLFSVNASSQFCGVAEMVGPVDFEK 341
Query: 461 SVDFWQQDKWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEML 520
SVD+WQQDKWSGQFPVKWHIIKDVPNSQFRHI+LENNDNKPVTNSRDTQEVKL+QGIEML
Sbjct: 342 SVDYWQQDKWSGQFPVKWHIIKDVPNSQFRHIILENNDNKPVTNSRDTQEVKLEQGIEML 401
Query: 521 TIFKNYETDTSILDDFDFYEDRQKAMQERKARQQSSIMPTGLVGG-SEHRSSSDS-TGDF 578
IFKNY+ DTSILDDF FYE+R+K +Q+RKAR+Q S+ G+V G +EH+ +S + DF
Sbjct: 402 KIFKNYDADTSILDDFGFYEEREKIIQDRKARRQPSLPSAGVVAGENEHKPASAALPTDF 461
Query: 579 IKQVSKNFSLVVRLDDNNNEVIAAN 603
+K +SK+F+ VVRLD+ + E + A+
Sbjct: 462 MKNMSKSFAQVVRLDEGSKEAVKAS 486
>At3g13460 unknown protein
Length = 608
Score = 374 bits (960), Expect = e-104
Identities = 231/611 (37%), Positives = 330/611 (53%), Gaps = 68/611 (11%)
Query: 50 YPPNVYAPQAQAFYYGGFDNGNGEWDEYPSYVNNGGIEIGSPGVYNENQSLVFHSGYGFN 109
Y PN Y P YY + +G EW +YP+Y N G+++ S G+Y EN ++V+ GYG+
Sbjct: 13 YVPNPYNPYQ---YYNVYGSGQ-EWTDYPAYTNPEGVDMNS-GIYGENGTVVYPQGYGYA 67
Query: 110 PQMPYGPYSPVTSPLPSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPEL 169
PYSP TSP P +GG+ QLY QQ+ Y P Y+ P + PY +S +QP+L
Sbjct: 68 AY----PYSPATSPAPQLGGEGQLYGAQQYQY-PNYF-----PNSGPYASSVATPTQPDL 117
Query: 170 TNLLGIDQQVESNFFGPRASYPSVGSFARGSFPV------------------APG---SF 208
+ + AS + + GS PV APG +
Sbjct: 118 SANKPAGVKTLPADSNNVASAAGITKGSNGSAPVKPTNQATLNTSSNLYGMGAPGGGLAA 177
Query: 209 SFHESQQGFDGSRSGGLWSDSSKPSERQR--------SFMPLSPSVPQQPIGSVRSFGPS 260
+ + + ++G + W D SK S+ QR S S +VP + RS S
Sbjct: 178 GYQDPRYAYEGYYAPVPWHDGSKYSDVQRPVSGSGVASSYSKSSTVPSSRNQNYRS--NS 235
Query: 261 AGMASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGLS---------ANDRSFLSL 311
+ HQ S+ G+G++ Y R Y + +G + S L N R + +
Sbjct: 236 HYTSVHQPSSVTGYGTAQGYYNRMYQNKLYGQYGSTGRSALGYGSSGYDSRTNGRGWAAT 295
Query: 312 ENSRRH-GRETASFCRCNGTLDILSEQNRGPRASKLKNHISSENNAID-----GSKNNAS 365
+N R GR + + +D L+E NRGPRA KN + +++++ G N
Sbjct: 296 DNKYRSWGRGNSYYYGNENNVDGLNELNRGPRAKGTKNQKGNLDDSLEVKEQTGESNVTE 355
Query: 366 TAKFQD-------ESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKL 418
+ + E N+ DF D+ +A FF+IKSYSED+VHKSIKY VWASTPNGN+KL
Sbjct: 356 VGEADNTCVVPDREQYNKEDFPVDYANAMFFIIKSYSEDDVHKSIKYNVWASTPNGNKKL 415
Query: 419 DAAYCQAKEKQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKW 478
AAY +A++K C IFLFFSVNAS QF G+AEM GPV+F+ +V++WQQDKW+G FP+KW
Sbjct: 416 AAAYQEAQQKAGGCPIFLFFSVNASGQFVGLAEMTGPVDFNTNVEYWQQDKWTGSFPLKW 475
Query: 479 HIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDF 538
HI+KDVPNS +HI LENN+NKPVTNSRDTQEVKL+QG++++ IFK + + T ILDDF F
Sbjct: 476 HIVKDVPNSLLKHITLENNENKPVTNSRDTQEVKLEQGLKIVKIFKEHSSKTCILDDFSF 535
Query: 539 YEDRQKAMQERKARQQSSIMPTGLVGGSEHRSSSDSTGDFIKQVSKNFSLVVRLDDNNNE 598
YE RQK + E+KA+Q + V + S++ + + S V++ +N +
Sbjct: 536 YEVRQKTILEKKAKQTQKQVSEEKVTDEKKESATAESASKESPAAVQTSSDVKVAENGSV 595
Query: 599 VIAANRDKLAS 609
D +A+
Sbjct: 596 AKPVTGDVVAN 606
>At1g55500 unknown protein
Length = 592
Score = 348 bits (894), Expect = 4e-96
Identities = 216/534 (40%), Positives = 292/534 (54%), Gaps = 61/534 (11%)
Query: 50 YPPNVYAPQAQAFYYGGFDNGNGEWDEYPSYVNNGGIEIGSPGVYNENQSLVFHSGYGFN 109
Y PNVY Q +YYG +Y Y N+ +++ S GYG+
Sbjct: 72 YVPNVYQ---QPYYYG-------YGSDYTGYTNSESVDMTS--------------GYGYA 107
Query: 110 PQMPYGPYSPVTSPLPSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPEL 169
PYSP TSP P +GGD QLY QQ+ Y P + + P+ +S +Q +L
Sbjct: 108 AF----PYSPATSPAPQLGGDGQLYGAQQYQYPFP-----LTASSGPFASSVPASTQSKL 158
Query: 170 TNLLGIDQQ---VESNFFGPRASYPSVGSFARGSFPVAPG-SFSFHESQQGFDGSRSGGL 225
+ + + G P S G+ + G + + + + +DG +
Sbjct: 159 STNKAANSASAGIPKGMNGSAPVKPLNQSALYGNSALGGGLAAGYQDPRYSYDGFYTPVS 218
Query: 226 WSDSSKPSERQRSF---------------MPLSPSVPQQPIGSVRSFGPSAGMASHQQRS 270
W D S S+ QRS +P + + S A M + +
Sbjct: 219 WHDGSNFSDVQRSVSGSGVASSYSKANNNVPATRNQNSSSNSHYTSMYQPASMTGYAAQG 278
Query: 271 LYGFGSSSNPYGRGYLSNPGSSFG-GSAISGLSANDRSFLSLENSRRHGRETASFCRCNG 329
Y S + YG+ Y S S G GS+ G N+R +L+ +N R S+ N
Sbjct: 279 YYDRVSPNKSYGQ-YGSTVRSGMGYGSSGYGSRTNERGWLNTDNKYRSRGRGNSYFYGNE 337
Query: 330 TLDILSEQNRGPRA--SKLKNHISSEN---NAIDGSKNNAS-TAKFQD-ESLNRPDFATD 382
+D L+E NRGPRA +K +SSE D S + T D E NR DF +
Sbjct: 338 NIDGLNELNRGPRAKGTKATEEVSSEEVKKQTFDESNTEETVTCVLPDREECNRDDFPVE 397
Query: 383 FKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKLDAAYCQAKEKQDACRIFLFFSVNA 442
+KDAKFF+IKSYSED+VHKSIKY VWASTPNGN+KLDAAY +A++K C +FLFFSVNA
Sbjct: 398 YKDAKFFIIKSYSEDDVHKSIKYNVWASTPNGNKKLDAAYQEAQQKSSGCPVFLFFSVNA 457
Query: 443 SAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPV 502
S QF G+AEM GPV+F+K++++WQQDKW+G FP+KWHI+KDVPNS +HI LE N+NKPV
Sbjct: 458 SGQFIGLAEMKGPVDFNKNIEYWQQDKWTGSFPLKWHILKDVPNSLLKHITLEYNENKPV 517
Query: 503 TNSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDFYEDRQKAMQERKARQQSS 556
TNSRDTQEVKL+QG++++ IFK + + T ILDDF FYE RQK + E+KA+QQ S
Sbjct: 518 TNSRDTQEVKLEQGLKVVKIFKEHNSKTCILDDFSFYEARQKTILEKKAKQQQS 571
>At3g17330 unknown protein
Length = 595
Score = 314 bits (804), Expect = 1e-85
Identities = 193/483 (39%), Positives = 261/483 (53%), Gaps = 66/483 (13%)
Query: 77 YPSYVNNG-GIEIGSPGVYNENQSLVFHS-GYGFNPQMPYGPYSPVTSPLPSVGGDAQLY 134
YP+Y ++G +G G NEN + ++ YG+ Q PY PY+P P S+G D+
Sbjct: 17 YPAYYSSGYDSSVGLQGGQNENAPYICYTPSYGY-AQSPYNPYNPYI-PGASIGVDSSFV 74
Query: 135 SPQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPELTNLLGIDQQVESNFFGPRASYPSVG 194
QQ+ PPY + P +PY+ P VS +L+ Q + G
Sbjct: 75 GFQQYYSDPPYESAASSPTYVPYVIQPDMVSNSSTDSLVANGGQSDGR-----------G 123
Query: 195 SFARGSFPVAPGSFSFHESQQGFDGSRSGGLWSDSSKPSERQRSFMPLSPSVPQQPIGSV 254
S R +GS GL D+ K + + P P
Sbjct: 124 SMQR-------------------NGSAIAGLPKDAPKSTTTGQYKQPGIPK--------- 155
Query: 255 RSFGPSAGMASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGLSANDRSFLSLENS 314
S ++H + + ++ PYG+ ++N G S+I+ S S + ++
Sbjct: 156 ---NVSTTASAHSLQGKTAYANTLLPYGKSDIAN-----GVSSIASYSYKPCS--KIYDA 205
Query: 315 RRHGRETASFCRCNGTLDILSEQNRGPRASKLKNH-ISSENNAIDGSKNNASTAKFQDES 373
R T S SEQNRG R + +N I G+ + +
Sbjct: 206 RGDNNTTGS--------TYTSEQNRGSRTRRSRNQLIVKAYTTKAGNADAEGNIVINPDR 257
Query: 374 LNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKLDAAYCQAK----EKQ 429
N+ DF+ ++ DA+FFVIKSYSED+VHKSIKYGVW+ST NGN+KL + Y A+ EK
Sbjct: 258 YNKEDFSIEYSDARFFVIKSYSEDDVHKSIKYGVWSSTLNGNKKLQSVYEDAQRIATEKS 317
Query: 430 DACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKWHIIKDVPNSQF 489
C IFLFFSVN+S FCGVAEM GPV+FD+ +DFWQQDKWSG FPVKWHIIKDVPNS F
Sbjct: 318 RECPIFLFFSVNSSGLFCGVAEMTGPVSFDRDMDFWQQDKWSGSFPVKWHIIKDVPNSYF 377
Query: 490 RHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDFYEDRQKAMQER 549
RHI+L NN+NKPVTNSRDTQE+ L+QG+E+L +FK++ TS+LDDF +YEDRQ+ MQE
Sbjct: 378 RHIILHNNENKPVTNSRDTQEIILKQGLEVLKLFKHHAEKTSLLDDFMYYEDRQRLMQEE 437
Query: 550 KAR 552
+AR
Sbjct: 438 RAR 440
>At1g48110 unknown protein
Length = 639
Score = 311 bits (798), Expect = 5e-85
Identities = 199/515 (38%), Positives = 269/515 (51%), Gaps = 44/515 (8%)
Query: 46 DQPLYPPNVYAPQAQAFYYGGFDNGNGEWDEYPSYVNNGGIEIGSPGVYNENQSLVFHS- 104
DQ +Y P + +Y G+++ G+W+ + + G E+ G NEN + ++
Sbjct: 13 DQGMYYP---VDASYGYYCTGYESP-GDWENHQMFFGVDGSEVQYTGGQNENSPYICYTP 68
Query: 105 GYGFNPQMPYGPYSPVTSPLPSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPV 164
YG+ Q PY P++P P S+G D+ +PQQF PPY + P +PY P V
Sbjct: 69 SYGY-AQSPYNPFNPYI-PGASIGVDSAFVAPQQFYSIPPYQSVATSPTFVPYAIQPEIV 126
Query: 165 SQPELTNLL--GIDQQVESNFFGPRASYPSVGSFARGSFPVAPGSFSFHESQQGFDGSRS 222
S +L+ G + S+ G R + + + + P P S
Sbjct: 127 SNSSTNSLVETGSANRGRSDGRGSRQRSGTATAGLQRNDPKLPAGNSL------------ 174
Query: 223 GGLWSDSSKPSERQRSFMPLSPSVPQQPIGSVRSFGPSAGMASHQQRSLYGFGSSSNPYG 282
G S+ +P+ Q + S G R G + SSS
Sbjct: 175 -GKISEKPRPNSGQSRQSEMDKSDSTSSSGQARQ-----GRVTSVSAQPVDVVSSSRV-- 226
Query: 283 RGYLSNPGSSFGGSAISGLSANDRSFLSLENSRRHGRETASFCRCNGTLDILSEQNRGPR 342
SSF I+ ND S ++ N+ + N D + EQNRG R
Sbjct: 227 --------SSFRQLDIAPPQLNDFSKIATNNNNIRPKLYGG--HANIIPDTVREQNRGRR 276
Query: 343 ASKLKNH-ISSENNAIDGSKNNASTAKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVHK 401
+ L N I G+ + N+ D D+ +AKFFVIKSYSED+VHK
Sbjct: 277 SRALGNQLIVKAYTTKAGNADAEGNIVINPSQYNKEDLRIDYSNAKFFVIKSYSEDDVHK 336
Query: 402 SIKYGVWASTPNGNRKLDAAYCQAK----EKQDACRIFLFFSVNASAQFCGVAEMVGPVN 457
SIKY VW+ST +GN+KL +AY A+ EK C IFLFFSVNAS FCG+AEM GPV+
Sbjct: 337 SIKYNVWSSTLHGNKKLQSAYEDAQRIATEKSCECPIFLFFSVNASGLFCGMAEMTGPVS 396
Query: 458 FDKSVDFWQQDKWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGI 517
FDK +DFWQQDKWSG FPVKWHIIKDVPNS FRHI+L+NN+NKPVTNSRDTQE+ L+QG+
Sbjct: 397 FDKDMDFWQQDKWSGSFPVKWHIIKDVPNSYFRHIILQNNENKPVTNSRDTQEIMLKQGL 456
Query: 518 EMLTIFKNYETDTSILDDFDFYEDRQKAMQERKAR 552
E+L IFK++ TS+LDDF +YE RQ+ MQ+ + R
Sbjct: 457 EVLKIFKDHMERTSLLDDFVYYESRQRVMQDERTR 491
>At5g58190 unknown protein
Length = 528
Score = 305 bits (782), Expect = 4e-83
Identities = 212/561 (37%), Positives = 286/561 (50%), Gaps = 105/561 (18%)
Query: 20 NSSQGTTTIGHSRDITSQSGSFG---SGADQPLYPPNVYAPQAQAFYY------GGFDNG 70
N Q + R+I S S S S P Y NV ++Y G+ G
Sbjct: 27 NGVQQQVSHDDGRNIVSSSYSSSVVPSPDPAPYY--NVLQTPTNLYHYDWDIANSGYAQG 84
Query: 71 NGEWDEYPSYVNNGGIEIGSPGV-YNENQSLVF-HSGYGFNPQMPYGPYSPVTSPLPSVG 128
WD YP Y + P V YN+N SL++ + GYGFNP PSV
Sbjct: 85 INTWDGYPRYATTTPEAMHVPPVVYNDNSSLMYQYPGYGFNPY-------------PSVM 131
Query: 129 GDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPELTNLLGIDQQVESNFFGPRA 188
+ Q+ P +P YY P P P + L D
Sbjct: 132 LEGQI------PVSPAYY--------------PQPFGAPSAMHYLPSD------------ 159
Query: 189 SYPSVGSFARGSFPVAPGSFSFHESQQGFDGSRSGGLWSDSSKPSERQRSFMPLSPSVPQ 248
+ P S ++ + G G D S L+ +P
Sbjct: 160 --------------IDPTSAAYMIPYGQYGGGNYSGNQGDIS-----------LTSHIPY 194
Query: 249 QPIGSVRSFGPSAGMASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGLSANDRSF 308
+ GP AS Q +L+G G +S+ GY S S ++R
Sbjct: 195 PQTMGI--LGPYDHNAS--QVALHGSGVASSSSLGGYYHVGSYQNPSSTPSYYGVDNRVR 250
Query: 309 LSLENSRRHGRETASFCRCNGTLDILSEQNRGPRAS-KLKNHISSEN-NAIDGSKNNAST 366
L+ + +R ++ S + T D+ NRGPRAS ++K+ SS+ + I S +++ST
Sbjct: 251 LTPDIGKRREKDQGSI---SSTSDLYG--NRGPRASSRVKSKNSSKPCSTIGDSASDSST 305
Query: 367 AKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKLDAAYCQAK 426
A N P+F TD+K+AKFF++KS+SEDNVH+SIKY VWASTP+GN+KLD AY A+
Sbjct: 306 AGPNPSLYNHPEFVTDYKNAKFFIVKSFSEDNVHRSIKYNVWASTPHGNKKLDTAYRDAE 365
Query: 427 EKQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKWHIIKDVPN 486
+ C IFLFFSVNAS QFCGV+EMVGPV+F+K +WQQD+WSGQFPVKWHI+KD+PN
Sbjct: 366 KMGGKCPIFLFFSVNASGQFCGVSEMVGPVDFEKDAGYWQQDRWSGQFPVKWHIVKDIPN 425
Query: 487 SQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDFYED----- 541
++F HI+L+NNDNKPVT+SRD+QEVKL+QGIEML IFK YE TSILDDF +Y++
Sbjct: 426 NRFCHILLQNNDNKPVTHSRDSQEVKLRQGIEMLRIFKEYEAHTSILDDFGYYDELEGQK 485
Query: 542 ------RQKAMQERKARQQSS 556
R+KA +E + +Q S
Sbjct: 486 VGEDGTRKKAGEEETSVEQLS 506
>At1g27960 hypothetical protein
Length = 539
Score = 305 bits (780), Expect = 7e-83
Identities = 176/364 (48%), Positives = 233/364 (63%), Gaps = 20/364 (5%)
Query: 232 PSERQRSFMPLSPSVP-QQPIGSVRSFGPSAGMASHQQRSLYGFGSSSNPYGR--GYLSN 288
P + F PL+ + P + +GS M F S++N R Y
Sbjct: 188 PFSGENMFPPLASTRPCMRHLGSAELLANDFNMGPAHGGLHDAFESAANLSYREQAYAQC 247
Query: 289 PGSSFGGSAISGLSANDRSFLSLENSRRHGRETASFCRCNGTLDILSEQNRGPRASKLKN 348
S F S+ S ND + L+ +R F C LD+L+E NRGPRAS+L
Sbjct: 248 KRSPFSASSSSPTWENDYNLPPLDEARSESYN--DFSHCPAMLDMLTESNRGPRASRL-- 303
Query: 349 HISSENNAIDGSKNNASTAKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVW 408
+S++ I + + +F + L + F+DAKFFVIKSYSEDNVHKSIK+ VW
Sbjct: 304 --NSKSKMISYDRVD----RFCQQEL-----LSQFRDAKFFVIKSYSEDNVHKSIKHCVW 352
Query: 409 ASTPNGNRKLDAAYCQAKEKQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQD 468
AST NGN+KLDAAY +AK+K AC +FL FSVNAS+QFCGVAEMVGPV+F+ SV++WQQD
Sbjct: 353 ASTKNGNKKLDAAYREAKKKDVACPVFLLFSVNASSQFCGVAEMVGPVDFNTSVEYWQQD 412
Query: 469 KWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYET 528
+WSG FPV+W I+KDVPNS FRHI++E+NDNKPVTNSRDTQEV L++GIEML IF + E
Sbjct: 413 RWSGHFPVQWLIVKDVPNSLFRHIIIESNDNKPVTNSRDTQEVGLEKGIEMLDIFISCEM 472
Query: 529 DTSILDDFDFYEDRQKAMQERKARQQSSIMPTGLVGGSEHRSSSDSTGDFIKQVSKNFSL 588
+SILDDF+FYE+RQ A+Q+RKARQ++ + L S S DF++++SK+F+
Sbjct: 473 RSSILDDFNFYEERQIAIQDRKARQRAVLESLALSATSAPTYSLHD--DFVREMSKHFAE 530
Query: 589 VVRL 592
+ L
Sbjct: 531 ALAL 534
>At5g61020 unknown protein
Length = 495
Score = 284 bits (727), Expect = 9e-77
Identities = 158/374 (42%), Positives = 221/374 (58%), Gaps = 37/374 (9%)
Query: 188 ASYPSVGSFARGSFPVAPGSFSFHESQQGFDGSRSGGLWSDSSKPSERQRSFMPLSPSVP 247
A+Y + GS+ +G++ ++ + G+ GS + G + Q ++
Sbjct: 82 AAYNAKGSYGKGAYAYGYYPPAYQYPRHGYTGSYASG-------KTNLQYQYLT------ 128
Query: 248 QQPIGSVRSFGPSAGMASHQQRSLYGF-GSSSNPYGRGYLSNPGSSFGGSAISGLSANDR 306
QQ G G S G S YG G +N YG G + + A +
Sbjct: 129 QQ--GRSAGNGQSYGGYMDNIYSNYGMCGPYTNGYGYGSYGYDSWKY----MPNWYAVNN 182
Query: 307 SFLSLENSRRHGRETASFCRCNGTLDILSEQNRGPRASKLKNHISS--------ENNAID 358
++ +G+E ++ L+E NRGPRA + S E +
Sbjct: 183 TYKPRNGYHGYGKEN---------IEGLNEMNRGPRAKGFNSQDGSKVMAVSLKEQRVTE 233
Query: 359 GSKNNASTAKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKL 418
K + + + N+ DF + +AKF+VIKSYSED++HKSIKY VW+STPNGN+KL
Sbjct: 234 TEKLSEDVSLLDPKDYNKIDFPETYTEAKFYVIKSYSEDDIHKSIKYSVWSSTPNGNKKL 293
Query: 419 DAAYCQAKEKQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKW 478
DA+Y +AK+K D C +FL FSVN S QF G+AEMVGPV+F+K+V++WQQDKW G FPVKW
Sbjct: 294 DASYNEAKQKSDGCPVFLLFSVNTSGQFVGLAEMVGPVDFNKTVEYWQQDKWIGCFPVKW 353
Query: 479 HIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDF 538
H +KD+PNS RHI LENN+NKPVTNSRDTQEVKL+QGI+++ IFK++ + T ILDDF+F
Sbjct: 354 HFVKDIPNSSLRHITLENNENKPVTNSRDTQEVKLEQGIKVIKIFKDHASKTCILDDFEF 413
Query: 539 YEDRQKAMQERKAR 552
YE+RQK +QERK++
Sbjct: 414 YENRQKIIQERKSK 427
>At3g03950 unknown protein
Length = 425
Score = 273 bits (699), Expect = 2e-73
Identities = 158/397 (39%), Positives = 218/397 (54%), Gaps = 50/397 (12%)
Query: 195 SFARGSFPVAPGSFSFHESQQGFDGSRSGGLWSDSSKPSERQRSFMPLSPSVPQQPIGSV 254
+F +GSF G + S G + +P+ + + S+ P G +
Sbjct: 35 NFTKGSFQHPYGHAPYGASSHGSE-----------RRPNMNAGNLLNGGDSIGSYPWGYI 83
Query: 255 RSFGPSAGMASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGLSANDRSFLS---- 310
+ PS G + +G+ +SN +L NP SS + L ND + +
Sbjct: 84 PANYPSGGYPDPR----FGYDRNSNHSSFSHLMNPHSSQEVPSFDQLGYNDHLYSNHGLY 139
Query: 311 ------LENSRRHG-------------------RETASFCRCNG----TLDILSEQNRGP 341
+++ +G R+T SF G D L+E RGP
Sbjct: 140 GLYGNVIDSGHAYGTFGYDSWKLGRGWYPVDGYRKTRSFNHGRGYSDEKADRLNELCRGP 199
Query: 342 RASKLKNHISSENNAIDGSKNNASTAKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVHK 401
R+S KN ++ +D K + S Q N +F F AKFFVIKSYSED+VH
Sbjct: 200 RSSDFKNPQVLNSSMLDAMKQDVSAVDLQ--RYNGENFPESFVKAKFFVIKSYSEDDVHN 257
Query: 402 SIKYGVWASTPNGNRKLDAAYCQAKEKQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKS 461
IKYG W+STP GN+KL+AAY +AKE C ++L FSVNAS QF G+AEMVGPV+F+K+
Sbjct: 258 CIKYGAWSSTPTGNKKLNAAYYEAKENSQECPVYLLFSVNASGQFVGLAEMVGPVDFNKT 317
Query: 462 VDFWQQDKWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLT 521
+++WQQDKW G FPVKWHIIKD+PNS RHI L NN+NKPVTNSRDTQEV L+ G +++
Sbjct: 318 MEYWQQDKWIGCFPVKWHIIKDIPNSLLRHITLANNENKPVTNSRDTQEVNLEHGTKIIK 377
Query: 522 IFKNYETDTSILDDFDFYEDRQKAMQERKARQQSSIM 558
IFK Y + T ILDD+ FYE RQK ++++K +Q+ +
Sbjct: 378 IFKEYMSKTCILDDYKFYETRQKIIRDKKIKQKKQAL 414
>At1g79270 unknown protein
Length = 528
Score = 271 bits (692), Expect = 1e-72
Identities = 131/216 (60%), Positives = 167/216 (76%), Gaps = 10/216 (4%)
Query: 347 KNHISS---ENNAIDGSKNNAST---AKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVH 400
++HI++ E+ ++D N S + + + N P F T +++A FFVIKSYSED++H
Sbjct: 276 QDHITNGECESCSLDAEGNERSNGVGSVIRRDQYNLPSFQTKYEEAIFFVIKSYSEDDIH 335
Query: 401 KSIKYGVWASTPNGNRKLDAAYCQ----AKEKQDACRIFLFFSVNASAQFCGVAEMVGPV 456
KSIKY VW+ST NGN+KLD+AY + A +K C +FLFFSVNAS QFCGVAEM+G V
Sbjct: 336 KSIKYNVWSSTLNGNKKLDSAYQESQKKAADKSGKCPVFLFFSVNASGQFCGVAEMIGRV 395
Query: 457 NFDKSVDFWQQDKWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQG 516
+++KS++FWQQDKW+G FPVKWHIIKDVPN Q RHI+LENN+NKPVTNSRDTQEV+L QG
Sbjct: 396 DYEKSMEFWQQDKWTGYFPVKWHIIKDVPNPQLRHIILENNENKPVTNSRDTQEVRLPQG 455
Query: 517 IEMLTIFKNYETDTSILDDFDFYEDRQKAMQERKAR 552
E+L IFKNY TSILDDFDFYE+R+K M ++K R
Sbjct: 456 NEVLNIFKNYAAKTSILDDFDFYENREKVMVQKKLR 491
>At1g09810 unknown protein
Length = 381
Score = 263 bits (672), Expect = 2e-70
Identities = 129/234 (55%), Positives = 177/234 (75%), Gaps = 7/234 (2%)
Query: 328 NGTLDILSEQNRGPRASKLKNHISSENNAIDGSKNNASTAKFQDESLNRPDFATDFKDAK 387
NG D L E GPRA+ K SE++ + +NN+ + E N PDF TD++DAK
Sbjct: 134 NGESDYLVELKCGPRANA-KTRPPSESSPL--KQNNSFALALRREMYNLPDFQTDYEDAK 190
Query: 388 FFVIKSYSEDNVHKSIKYGVWASTPNGNRKLDAAYCQAKEK--QDACR--IFLFFSVNAS 443
FFVIKSYSED+VHKSIKY VW+ST NGN+KLDAA+ A+ K +D + IFLFFSVNAS
Sbjct: 191 FFVIKSYSEDDVHKSIKYSVWSSTINGNKKLDAAFRDAETKTLEDGKKRPIFLFFSVNAS 250
Query: 444 AQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPVT 503
QF G+AEMVG V+F+K +DFWQ DKWSG FPV+WH++KD+PN + RHI+L+NN++KPVT
Sbjct: 251 RQFVGLAEMVGYVDFNKDLDFWQVDKWSGFFPVEWHVVKDIPNWELRHIILDNNEDKPVT 310
Query: 504 NSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDFYEDRQKAMQERKARQQSSI 557
++RDT E+KL++G++ML+IFK Y T +LDD DFYE+R+K+++ +K + +++
Sbjct: 311 HTRDTHEIKLKEGLQMLSIFKKYSAVTFLLDDMDFYEEREKSLRAKKEHKPATL 364
>At4g11970 unknown protein
Length = 444
Score = 79.0 bits (193), Expect = 8e-15
Identities = 56/189 (29%), Positives = 86/189 (44%), Gaps = 21/189 (11%)
Query: 360 SKNNASTAKFQDESLNRPDFATDFKDAK---------FFVIKSYSEDNVHKSIKYGVWAS 410
SK + +K + N PD K K +F+IKS + DN+ S++ G+WA+
Sbjct: 37 SKEDHKLSKVDVDRRNFPDQLESAKANKNSKPGYRTRYFIIKSLNYDNIQVSVEKGIWAT 96
Query: 411 TPNGNRKLDAAYCQAKEKQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQ--- 467
L+ A+ ++ R+ L FSVN S F G AEM+ PV + + W Q
Sbjct: 97 QVMNEPILEGAFHKSG------RVILIFSVNMSGFFQGYAEMLSPVGWRRD-QIWSQGGG 149
Query: 468 --DKWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKN 525
+ W F VKW + ++P + H+ ND KPV SRD QE+ G + +
Sbjct: 150 KNNPWGRSFKVKWLRLSELPFQKTLHLKNPLNDYKPVKISRDCQELPEDIGEALCELLDA 209
Query: 526 YETDTSILD 534
D +L+
Sbjct: 210 NSCDDGLLN 218
>At1g30460 hypothetical protein
Length = 678
Score = 64.3 bits (155), Expect = 2e-10
Identities = 49/181 (27%), Positives = 83/181 (45%), Gaps = 16/181 (8%)
Query: 388 FFVIKSYSEDNVHKSIKYGVWASTPNGNRKLDAAYCQAKEKQDACRIFLFFSVNASAQFC 447
+FV+KS + +N S++ GVWA+ + KL+ A+ + + L FSVN + F
Sbjct: 286 YFVVKSNNRENFELSVQQGVWATQRSNEAKLNEAFDSVE------NVILIFSVNRTRHFQ 339
Query: 448 GVAEMVGPVNFDKSVDFWQQDKWSGQ----FPVKWHIIKDVPNSQFRHIVLENNDNKPVT 503
G A+M + W+ + + Q F VKW + ++ + R++ N+N PV
Sbjct: 340 GCAKMTSRIGGYIGGGNWKHEHGTAQYGRNFSVKWLKLCELSFHKTRNLRNPYNENLPVK 399
Query: 504 NSRDTQEVKLQQGIEMLTIFKNYETDT-----SILDDFDFYEDRQKAMQERKARQQSSIM 558
SRD QE++ G E L E D+ SI + E++ K + + I+
Sbjct: 400 ISRDCQELEPSVG-EQLASLLYLEPDSELMAISIAAEAKREEEKAKGVNPESRAENPDIV 458
Query: 559 P 559
P
Sbjct: 459 P 459
>At5g54350 unknown protein
Length = 279
Score = 43.5 bits (101), Expect = 4e-04
Identities = 71/296 (23%), Positives = 116/296 (38%), Gaps = 46/296 (15%)
Query: 96 ENQSLVFHSGYGFNPQMPYGPYSPVTSPLPSVGGDAQLYSPQQFPYTPPYYNQLVPPPNL 155
ENQS+ S + P P + S + +V + +P+Q P + +P P L
Sbjct: 2 ENQSVSSSS----SSTTPKKPDQKLKSAVVTVEEKETMNNPEQRPPQEVVFENTIPDPEL 57
Query: 156 PYLNSPTPVSQPELTNLLGI---DQQVESNFFGPRASYPSVGSFARGSF----PVAPGSF 208
P V + E N L D+ ++S G S P SF SF P GS
Sbjct: 58 PVFT--LEVVRTERDNELKYPPKDKNLDSVLSG-GTSLPPFQSFPVPSFGLFGPAYGGSS 114
Query: 209 SFHESQQGFDGSRSGGLWSD-------SSKPSERQRSFMPLSPSVPQQPIGSVRSFGPSA 261
S ++ D + WS+ ++ PS ++ F P+ P VP+ + + P
Sbjct: 115 STNDI--NLDLTLGPSSWSNTNPSSWSNTDPSFYRKLFTPVLPCVPR---NNFVAGNPGN 169
Query: 262 GMASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGLSANDRSFLSLENSRRHGRET 321
AS +SL+ G + + GS S++ +S S+E ++
Sbjct: 170 SSASTHSKSLFSMGKNQ------------AMENGSG----SSHSKSLFSMEKNKAVEPPV 213
Query: 322 ASFCRCNGTLDIL-SEQNRGPRASKLKNHISSENNAIDGSKNNASTAKFQDESLNR 376
A NGT D+L G +S LK+ IS N + +N+ + D ++ R
Sbjct: 214 AP---PNGTTDLLFPSPENGSGSSHLKSLISKGENKVVVPENDDDDDDYTDTTIGR 266
>At5g51300 unknown protein
Length = 804
Score = 42.4 bits (98), Expect = 8e-04
Identities = 53/202 (26%), Positives = 73/202 (35%), Gaps = 49/202 (24%)
Query: 91 PGVYNENQSLVFHSGYGFNPQMPYGP----YSPVTSPLPSVGGDAQLYSPQQFPYTPPYY 146
PG Y Q GY P +P+GP YSP P P G Y P + PPY
Sbjct: 585 PGAYPSQQYAT--GGYSTAP-VPWGPPVPSYSPYALPPPPPGS----YHPVHGQHMPPYG 637
Query: 147 NQLVPPPNLPYLNSPTP--------VSQPELTNLLGID-------QQVESNFFG------ 185
Q PPP P++ P S+P+ + G+ + N +G
Sbjct: 638 MQYPPPP--PHVTQAPPPGTTQNPSSSEPQQSFPPGVQADSGAATSSIPPNVYGSSVTAM 695
Query: 186 ----PRASYPSVGSFARGSFPVAPGSFSFHESQQGFDGSRSGGLWSDSSKPSERQRS--- 238
P SYPS + P AP S + H G W+++ S S
Sbjct: 696 PGQPPYMSYPSYYNAVPPPTPPAPASSTDHSQNMG------NMPWANNPSVSTPDHSQGL 749
Query: 239 -FMPLSPSVPQQP-IGSVRSFG 258
P +P+ P P +G +S G
Sbjct: 750 VNAPWAPNPPMPPTVGYSQSMG 771
>At1g10390 unknown protein
Length = 1041
Score = 40.4 bits (93), Expect = 0.003
Identities = 78/314 (24%), Positives = 116/314 (36%), Gaps = 46/314 (14%)
Query: 19 SNSSQGTTTIGHSRDITSQSGS--FGSGADQPLYPPNVYAPQAQAFYYGGFDNGNGEWDE 76
+N T G S +QSGS FGS + P +P A +G +
Sbjct: 32 NNPFAPATPFGTSAPFAAQSGSSIFGSTSTGVFGAPQTSSPFASTPTFGASSS------- 84
Query: 77 YPSYVNNGGIEIGSPGVYNENQSLVFHSGYGFNPQMPYGPYSPVTSPLPSVGGDAQ-LYS 135
P++ N+ SP S F GF Q P G +P ++P + +Q +
Sbjct: 85 -PAFGNSTPAFGASPA------SSPFGGSSGFG-QKPLGFSTPQSNPFGNSTQQSQPAFG 136
Query: 136 PQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPEL----TNLLGIDQQVESNFFGPRASYP 191
F + P+ P P S S P T G + FG S P
Sbjct: 137 NTSFGSSTPFGATNTPAFGAPSTPSFGATSTPSFGASSTPAFGA---TNTPAFGASNS-P 192
Query: 192 SVGSFARGSFPVAPG-SFSFHESQQGFDGSRSGGLWSDSSKPSERQRSFMPLSPSVPQQP 250
S G+ +F +P +F + G G SGG + S+ P+ + P
Sbjct: 193 SFGATNTPAFGASPTPAFGSTGTTFGNTGFGSGGAFGASNTPAFG-------ASGTPAFG 245
Query: 251 IGSVRSFGPSAGMASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGLSANDRSFLS 310
+FG S+ A FG+SS P G S P +FGGS+ A++ S S
Sbjct: 246 ASGTPAFGASSTPA---------FGASSTP-AFGASSTP--AFGGSSTPSFGASNTSSFS 293
Query: 311 LENSRRHGRETASF 324
+S G+ T++F
Sbjct: 294 FGSSPAFGQSTSAF 307
Score = 30.0 bits (66), Expect = 4.1
Identities = 67/300 (22%), Positives = 94/300 (31%), Gaps = 89/300 (29%)
Query: 16 SKFSNSSQGTTTIGHSRDITSQSGSFGSGADQPLYPPNVYAPQAQAFYYGG--------- 66
S F + T T G S S G G+ Y P V A G
Sbjct: 317 SPFGGAQASTPTFGGSGFGQSTFGGQQGGSRAVPYAPTVEADTGTGTQPAGKLESISAMP 376
Query: 67 -FDNGNGE---WDEYPSYVNNGGIEIG-SPGVYNENQSLVFHSGYGFNPQMPYGPYSPVT 121
+ N E W++Y G + G SPG ++G+G +P P P+SP
Sbjct: 377 AYKEKNYEELRWEDYQRGDKGGPLPAGQSPG----------NAGFGISPSQP-NPFSP-- 423
Query: 122 SPLPSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPELTNLLGIDQQVES 181
P + Q P P+ +S + +
Sbjct: 424 ---------------------SPAFGQTSANPTNPFSSSTS------------------T 444
Query: 182 NFFGPRASYPSVGSFARGSFPVAPGSFSFHESQQGFDGSRSGGLWSDSSKPSERQRSFMP 241
N F P+ P++ S SF A +F GS G+ S S+
Sbjct: 445 NPFAPQT--PTIAS---SSFGTATSNF----------GSSPFGVTSSSN--------LFG 481
Query: 242 LSPSVPQQPIGSVRSFGPSAGMASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGL 301
S GS +FG + S GFGSSS+ +G +FG S S L
Sbjct: 482 SGSSTTTSVFGSSSAFGTTTPSPLFGSSSTPGFGSSSSIFGSAPGQGATPAFGNSQPSTL 541
>At1g59660 nucleoporin 98-like protein
Length = 997
Score = 39.3 bits (90), Expect = 0.007
Identities = 55/235 (23%), Positives = 87/235 (36%), Gaps = 37/235 (15%)
Query: 74 WDEYPSYVNNGGIEIG-SP-----GVYNENQSLVFHSGYGFNPQMPYGPYSPVTSPLPSV 127
W++Y G G SP GV N S +F + F+ Q P P +P + P+
Sbjct: 378 WEDYQRGDKGGQRSTGQSPEGAGFGVTNSQPS-IFSTSPAFS-QTPVNPTNPFSQTTPTS 435
Query: 128 GGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPELTNLLGIDQQVESNFFGPR 187
+ +SP T P + Q P +++ T V + Q + S+ FG
Sbjct: 436 NTN---FSPSFSQPTTPSFGQPTTPSFRSTVSNTTSVFGSSSSLTTNTSQPLGSSIFGST 492
Query: 188 ASYPSVGSFARGSFPVAPGSFSFHESQQGFDGSRSGGLWSDSSKPSERQRSFMPLSPSVP 247
++ S F+ G GF+ S+S L+ S PS Q + S + P
Sbjct: 493 PAHGSTPGFSIG----------------GFNNSQSSPLF--GSNPSFAQNTTPAFSQTSP 534
Query: 248 QQPIGSVRSFGPSAGMASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGLS 302
FG + A Q S++G ++ S P + FG + S S
Sbjct: 535 --------LFGQNTTPALGQSSSVFGQNTNPALVQSNTFSTPSTGFGNTFSSSSS 581
Score = 34.3 bits (77), Expect = 0.22
Identities = 73/308 (23%), Positives = 113/308 (35%), Gaps = 40/308 (12%)
Query: 41 FGSGADQPLYPPNVYAP-QAQAFYYGGFDNGNGEWDEY---------PSYVNNGGIEIG- 89
FGS + P ++ +P Q G N N + + P G G
Sbjct: 2 FGSSNNNPFGQSSISSPFGTQTHSLFGQTNNNASNNPFATKPFGTSTPFGAQTGSSMFGG 61
Query: 90 -SPGVYNENQSLVFHSGYGFNPQMPYGPYSPV--TSPLPSVGGDAQLYSPQQFPYTPPYY 146
S GV+ Q+ S +G +PQ +G + S PS G S F T +
Sbjct: 62 TSTGVFGAPQT---SSPFGASPQA-FGSSTQAFGASSTPSFGS-----SNSPFGGTSTFG 112
Query: 147 NQ---LVPPPNLPYLNSPTPVSQPELTNLLGIDQQVESNFFGPRASYPSVGSFARGSFPV 203
+ L P + P+ S T SQP N S FG + P+ G+ + +F V
Sbjct: 113 QKSFGLSTPQSSPF-GSTTQQSQPAFGN----STFGSSTPFGASTT-PAFGASSTPAFGV 166
Query: 204 APGSFSFHESQQGFDGSRSGGLWSDSSKPSERQRSFMPLSPSVPQQPIGSVRSFGPSAGM 263
+ S + GF + + G S+ + S + P S FG S+
Sbjct: 167 SNTSGFGATNTPGFGATNTTGFGGSSTPGFGASSTPAFGSTNTPAFGASSTPLFGSSSSP 226
Query: 264 ASHQQRSLYGFGSSSNPYGRGYLSNPGS-------SFGGSAISGLSANDRSFLSLENSRR 316
A + FGSS N +G S+ G+ +FG S S A+ + +S
Sbjct: 227 AFGASPAP-AFGSSGNAFGNNTFSSGGAFGSSSTPTFGASNTSAFGASSSPSFNFGSSPA 285
Query: 317 HGRETASF 324
G+ T++F
Sbjct: 286 FGQSTSAF 293
Score = 32.0 bits (71), Expect = 1.1
Identities = 35/125 (28%), Positives = 52/125 (41%), Gaps = 7/125 (5%)
Query: 188 ASYPSVGSFARGSFPVAPGSFSFHESQQGFDGSR--SGGLWSDSSKPSERQRSFMPL-SP 244
+S P GS + +F +P +F S F + SGG + SS P+ + +
Sbjct: 215 SSTPLFGSSSSPAFGASPAP-AFGSSGNAFGNNTFSSGGAFGSSSTPTFGASNTSAFGAS 273
Query: 245 SVPQQPIGSVRSFGPSAGM--ASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGG-SAISGL 301
S P GS +FG S +S + GS+ +P+G S+FGG S I G
Sbjct: 274 SSPSFNFGSSPAFGQSTSAFGSSSFGSTQSSLGSTPSPFGAQGAQASTSTFGGQSTIGGQ 333
Query: 302 SANDR 306
R
Sbjct: 334 QGGSR 338
>At4g32640 putative protein
Length = 1069
Score = 38.9 bits (89), Expect = 0.009
Identities = 63/244 (25%), Positives = 81/244 (32%), Gaps = 36/244 (14%)
Query: 51 PPNVYAPQAQAFYYGGFDNGNGEWDEYPSYVNNGGIEIGSPGVYNENQSLVFHSGYGFNP 110
PP P +Q +G + YP N + + N+ G G P
Sbjct: 6 PPGAPRPNSQ--------QNSGPPNFYPGSQGNSNALADNMQNLSLNRPPPMMPGSGPRP 57
Query: 111 QMPYGPY-SPVTSPLPSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPEL 169
P+G P PS G + SP P PP + P PVSQP
Sbjct: 58 PPPFGQSPQPFPQQSPSYGAPQRGPSPMSRPG---------PPAGMARPGGPPPVSQP-- 106
Query: 170 TNLLGIDQQVESNF-FGPRASYPSVGSFARGSFPVAPGSFSFHESQQGFDGSRSGGLWSD 228
G V N GP + PS GS R S P P + S GF G +
Sbjct: 107 ---AGFQSNVPLNRPTGPPSRQPSFGS--RPSMPGGPVA-QPAASSSGFPAFGPSGSVAA 160
Query: 229 SSKPSERQRSFMPLSPSVPQQPIGSVRSFGPSAGMASHQQRSLYGFGSSSNPYGRGYLSN 288
P R +F SP P+GS S PS + GS P G +
Sbjct: 161 GPPPGSRPMAFG--SP----PPVGSGMSMPPSGMIGGPVSNGHQMVGSGGFPRGTQF--- 211
Query: 289 PGSS 292
PG++
Sbjct: 212 PGAA 215
Score = 32.3 bits (72), Expect = 0.82
Identities = 24/63 (38%), Positives = 28/63 (44%), Gaps = 5/63 (7%)
Query: 110 PQMPYGPYSPVTSPLPSVGGDAQLYSPQQFP--YTPPYYNQLVPPPNLPYLNSPTPVSQP 167
P P+G P S LP AQ+ P FP PP Y + P PN N PT + QP
Sbjct: 265 PGAPHG--RPAVSGLPYGPPSAQVAPPLGFPGQMQPPRYG-MGPLPNQSMTNIPTAMGQP 321
Query: 168 ELT 170
T
Sbjct: 322 GAT 324
>At3g21320 hypothetical protein
Length = 540
Score = 38.1 bits (87), Expect = 0.015
Identities = 36/134 (26%), Positives = 55/134 (40%), Gaps = 27/134 (20%)
Query: 115 GPYSPVTSP-LPSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPELTNLL 173
GP P +S + V G L +P +FP + P+ + PPP N+ T V Q
Sbjct: 382 GPCPPSSSAFMVPVYGQDSLETPFRFPVSSPFSHSYFPPP-----NARTTVDQ------- 429
Query: 174 GIDQQVESNFFGPRASYPSVGSFARGSFPVAPGSFSFHESQQGFDGSRSGGLWSDSSKPS 233
+N FG + + S + P FS +SQ+ D G + SS P
Sbjct: 430 -------TNPFGQFQRWSNTSSHMTQAIP-----FSLKKSQESNDSDIHGS--TASSPPE 475
Query: 234 ERQRSFMPLSPSVP 247
+ + +PL P+ P
Sbjct: 476 KHKLEVLPLFPTEP 489
>At2g43170 unknown protein
Length = 504
Score = 38.1 bits (87), Expect = 0.015
Identities = 36/137 (26%), Positives = 51/137 (36%), Gaps = 27/137 (19%)
Query: 21 SSQGTTTIGHSRDITSQSGSFGSGADQPLYPPNVYAPQAQAFYYGGFDNGNGEWDEYPSY 80
++ T+TI + + S +G SGA PPN
Sbjct: 370 NTPATSTINMGKAMGSGTGLGRSGATAMRPPPNPMTGSGMPM------------------ 411
Query: 81 VNNGGIEIGSPGVYNENQSLVFHSGYGFNPQMPYG-PYSPVTSPLPSVGGDAQLYSPQQF 139
GG+ +GS G N+NQ + G G N P G P + ++GG Q Y Q
Sbjct: 412 --GGGMGVGSYGGMNQNQPMGMGMGAGMNQNQPMGMGMGPGMNMNMNMGGYGQGYPMQ-- 467
Query: 140 PYTPPYYNQLVPPPNLP 156
P +VP PN+P
Sbjct: 468 ----PQNPGMVPSPNMP 480
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.313 0.132 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,031,985
Number of Sequences: 26719
Number of extensions: 806826
Number of successful extensions: 2518
Number of sequences better than 10.0: 128
Number of HSP's better than 10.0 without gapping: 27
Number of HSP's successfully gapped in prelim test: 105
Number of HSP's that attempted gapping in prelim test: 2095
Number of HSP's gapped (non-prelim): 389
length of query: 632
length of database: 11,318,596
effective HSP length: 105
effective length of query: 527
effective length of database: 8,513,101
effective search space: 4486404227
effective search space used: 4486404227
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC144538.7


