

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144538.13 + phase: 0 /pseudo
(1574 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
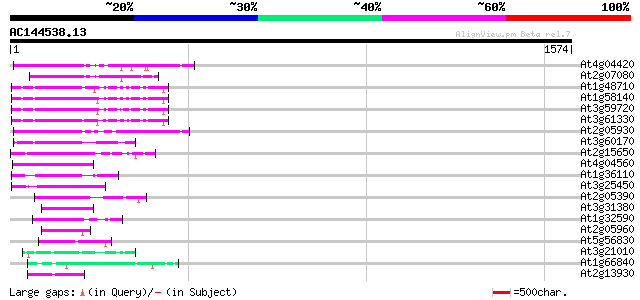
Score E
Sequences producing significant alignments: (bits) Value
At4g04420 putative transposon protein 166 1e-40
At2g07080 putative gag-protease polyprotein 141 3e-33
At1g48710 hypothetical protein 128 3e-29
At1g58140 hypothetical protein 127 4e-29
At3g59720 copia-type reverse transcriptase-like protein 127 5e-29
At3g61330 copia-type polyprotein 127 7e-29
At2g05930 copia-like retroelement pol polyprotein 126 9e-29
At3g60170 putative protein 109 1e-23
At2g15650 putative retroelement pol polyprotein 108 2e-23
At4g04560 putative transposon protein 105 3e-22
At1g36110 hypothetical protein 94 8e-19
At3g25450 hypothetical protein 93 1e-18
At2g05390 putative retroelement pol polyprotein 93 1e-18
At3g31380 hypothetical protein 86 1e-16
At1g32590 hypothetical protein, 5' partial 86 1e-16
At2g05960 putative retroelement pol polyprotein 77 6e-14
At5g56830 unknown protein 57 8e-08
At3g21010 unknown protein 54 9e-07
At1g66840 hypothetical protein 54 9e-07
At2g13930 putative retroelement pol polyprotein 53 1e-06
>At4g04420 putative transposon protein
Length = 1008
Score = 166 bits (419), Expect = 1e-40
Identities = 141/576 (24%), Positives = 255/576 (43%), Gaps = 88/576 (15%)
Query: 12 LFEGTDYYYWKGKMELFLQS*DNHMWTVVENGNYIPY--DEDLNE--KKKEDWSEEEGKR 67
+ E +Y +WK KM ++ W G P ED + K K+ W++ E +
Sbjct: 15 MLEKGNYGHWKVKMRALIRGLGKEAWIATSIGWKAPVIKGEDGEDVLKTKDQWNDAEEAK 74
Query: 68 MLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGVRKFEL 127
N +A + +++ ++ R+Q C++AKE WD L +EGTS VK +RID+ +FE
Sbjct: 75 AKANSRALSLIFNFVNQNQFKRIQNCESAKEAWDKLAKAYEGTSSVKRSRIDMLASQFEN 134
Query: 128 FEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRSWRHIVTAITEAKDL 187
M+ETE I+E G+ + I +E +L K + + V+K+LRCLP + TA+ + D
Sbjct: 135 LSMEETENIEEFSGKISAIASEAHNLGKKYKDKKLVKKLLRCLPSRFESKRTAMGTSLDT 194
Query: 188 KKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDNQQQELDEEDH 247
+ E+++G L+A+E+ + K SK +AL + + N+ QEL
Sbjct: 195 DSIDFEEVVGMLQAYELEITSGK-GGYSKGLALAASAKK----------NEIQEL----- 238
Query: 248 QDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKS----QVTCYGCYKLGHYKNEC 303
+D + ++ + R ++R + K+ F R + D+S ++ C+ C GH K EC
Sbjct: 239 KDTMSMMAKDFSRAMRRVE--KKGF-GRNQGTDRYRDRSSKRDEIQCHECQGYGHIKAEC 295
Query: 304 PLNKRKSSN--------------LQTNQKSFMTTWDEPEEQTNEDEQADLC--------- 340
P KRK + + K + E E +N+ + D
Sbjct: 296 PSLKRKDLKCSECNGLGHTKFDCVGSKSKPDKSCSSESESDSNDGDSEDYIKGFVSFVGI 355
Query: 341 ---------LMAKSDNEEVNLDPCSSCEKTEHL---FDNLFYSSQMLEQE---------- 378
A ++E+ + D S EK ++ F L+ S ML +E
Sbjct: 356 IEEKDESSDSEADGEDEDNSADEDSDIEKDVNINEEFRKLYDSWLMLSKEKVAWLEEKLK 415
Query: 379 ----CERLRNE-------NLELRKEKEILEKENKDLIKNIQHSESKEVSENKNNSQKENI 427
E+L+ E N EL ++ + E++N++L + + + N ++I
Sbjct: 416 VQELTEKLKGELTAANQKNSELIQKCSVAEEKNRELSQELSDTRKNIHMLNSGTKDLDSI 475
Query: 428 ILKENVLKLKNDISNFVKSTETFQKIMGSQVSVFDKACIGFKTN----QKQKLYENFFIP 483
+ V K + + T + S+ + K+ GF++N +++Y+N
Sbjct: 476 LAAGRVGKSNFGLGYNGAGSGTKTNFVRSEAAAPTKSQTGFRSNYDAVPARRVYQN-HDH 534
Query: 484 KKNKNTCERYKCTYCEKYGHLEPFCFRKRKRLASQK 519
+++ T Y+C YC ++GH++ +C+R RL K
Sbjct: 535 YQSRRTVTGYECYYCGRHGHIQRYCYRYAARLNKLK 570
>At2g07080 putative gag-protease polyprotein
Length = 627
Score = 141 bits (355), Expect = 3e-33
Identities = 106/382 (27%), Positives = 176/382 (45%), Gaps = 45/382 (11%)
Query: 55 KKKEDWSEEEGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVK 114
K K +W+ EE + N +A + + +E+ +Q CK+AK+ WDTL+ HEGTS VK
Sbjct: 43 KPKANWTAEEKLQSKFNARAMKAIFNGVDEDEFKLIQGCKSAKQAWDTLQKSHEGTSSVK 102
Query: 115 ETRIDIGVRKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRSW 174
TR+D +FE +M+ E I + + + + NE + K + + V+K+LRCLP +
Sbjct: 103 RTRLDHIATQFEYLKMEPDEKIVKFSSKISALANEAEVMGKTYKDQKLVKKLLRCLPPKF 162
Query: 175 RHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMN 234
+ A + K+ DL+G LK E+ +DK SK IA +Q S
Sbjct: 163 AAHKAVMRVAGNTDKISFVDLVGMLKLEEMKADQDK-VKPSKNIAFNADQGS-------- 213
Query: 235 FDNQQQELDEEDHQDQIILLTRKLQRMIQRRD---QNKRNFPARKENAKTELDKSQVTCY 291
Q QE+ +D + LL R + ++R + + R +R EN K ++ CY
Sbjct: 214 --EQFQEI-----KDGMALLARNFGKALKRVEIDGERSRGRFSRSENDDLR-KKKEIQCY 265
Query: 292 GCYKLGHYKNECPLNKRKS-----------------SNLQTNQKSFMTTWDEPEEQTNED 334
C GH K ECP+ KRK + + +KS ++ D + E+
Sbjct: 266 ECGGFGHIKPECPITKRKEMKCLKCKGVGHTKFECPNKSKLKEKSLISFSDSESDDEGEE 325
Query: 335 EQADLCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRKEKE 394
+ MA SD+ + D S C++ + D ++L +L + L+L KEK
Sbjct: 326 LLNFVAFMASSDSSKFMSDTDSDCDEELNPKDKY----RVLYDSWVQLSKDKLKLVKEKL 381
Query: 395 ILEKENKDLIKNIQHSESKEVS 416
LE + + N+ + +++S
Sbjct: 382 TLEAK----LANVSTEDKQKLS 399
Score = 33.1 bits (74), Expect = 1.3
Identities = 52/223 (23%), Positives = 90/223 (40%), Gaps = 40/223 (17%)
Query: 340 CLMAKS--DNEEVNLDPCSSCEKT--EHLFDNLFYSSQMLEQECERLRNENLELRKEKEI 395
C AK D + + + SS ++T +H+ Y +++ + ++ L E E+
Sbjct: 81 CKSAKQAWDTLQKSHEGTSSVKRTRLDHIATQFEYLKMEPDEKIVKFSSKISALANEAEV 140
Query: 396 LEKENKD--LIKNI------QHSESKEVSENKNNSQKENIILKENVLKL----------- 436
+ K KD L+K + + + K V N+ K + + +LKL
Sbjct: 141 MGKTYKDQKLVKKLLRCLPPKFAAHKAVMRVAGNTDKISFVDLVGMLKLEEMKADQDKVK 200
Query: 437 --KNDISNFVKSTETFQKIMGSQVSV---FDKACI-----GFKTNQKQKLYENFFIPKKN 486
KN N + +E FQ+I + F KA G ++ + EN + KK
Sbjct: 201 PSKNIAFNADQGSEQFQEIKDGMALLARNFGKALKRVEIDGERSRGRFSRSENDDLRKKK 260
Query: 487 KNTCERYKCTYCEKYGHLEPFC-FRKRKRLASQKFKN-NYTNF 527
+ +C C +GH++P C KRK + K K +T F
Sbjct: 261 E-----IQCYECGGFGHIKPECPITKRKEMKCLKCKGVGHTKF 298
>At1g48710 hypothetical protein
Length = 1352
Score = 128 bits (321), Expect = 3e-29
Identities = 120/474 (25%), Positives = 209/474 (43%), Gaps = 77/474 (16%)
Query: 6 SSNRP---PLFEGTDYYYWKGKMELFLQS*DNHMWTVVENGNYIPYDEDLNEKKKEDWSE 62
S+N P P+ ++Y W +M+ L + D +W +VE G P +E + ++D
Sbjct: 3 SNNVPFQVPVLTKSNYDNWSLRMKAILGAHD--VWEIVEKGFIEPENEGSLSQTQKDGLR 60
Query: 63 EEGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGV 122
+ KR + KA + L + +++V E +AKE W+ L+ ++G VK+ R+
Sbjct: 61 DSRKR---DKKALCLIYQGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLR 117
Query: 123 RKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRSWRHIVTAIT 182
+FE +MKE E + + + R + N L+ + + K+LR L + HIVT I
Sbjct: 118 GEFEALQMKEGELVSDYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIE 177
Query: 183 EAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFD------ 236
E KDL+ + +E L+GSL+A+E +K K +I N + +E ++
Sbjct: 178 ETKDLEAMTIEQLLGSLQAYE-----EKKKKKEDIIEQVLNMQITKEENGQSYQRRGGGQ 232
Query: 237 ----------NQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKS 286
N + ED+ +Q R + + R K + K+ DKS
Sbjct: 233 VRGRGRGGYGNGRGWRPHEDNTNQ-------------RGENSSRG--RGKGHPKSRYDKS 277
Query: 287 QVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMA--K 344
V CY C K GHY +EC + SN + +K+ + EE+ E+ D+ LMA K
Sbjct: 278 SVKCYNCGKFGHYASEC----KAPSNKKFEEKA-----NYVEEKIQEE---DMLLMASYK 325
Query: 345 SDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDLI 404
D +E N + H+ M + E +R N+ L E ++ K +++
Sbjct: 326 KDEQEENHKWYLDSGASNHMCGR----KSMFAELDESVRG-NVALGDESKMEVKGKGNIL 380
Query: 405 KNIQHSESKEVSE-NKNNSQKENII-------------LKENVLKLKNDISNFV 444
+++ + + +S S K NI+ LK+N L +++ SN +
Sbjct: 381 IRLKNGDHQFISNVYYIPSMKTNILSLGQLLEKGYDIRLKDNNLSIRDQESNLI 434
>At1g58140 hypothetical protein
Length = 1320
Score = 127 bits (320), Expect = 4e-29
Identities = 118/467 (25%), Positives = 209/467 (44%), Gaps = 63/467 (13%)
Query: 6 SSNRP---PLFEGTDYYYWKGKMELFLQS*DNHMWTVVENGNYIPYDEDLNEKKKEDWSE 62
S+N P P+ ++Y W +M+ L + D +W +VE G P +E + ++D
Sbjct: 3 SNNVPFQVPVLTKSNYDNWSLRMKAILGAHD--VWEIVEKGFIEPENEGSLSQTQKDGLR 60
Query: 63 EEGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGV 122
+ KR + KA + L + +++V E +AKE W+ L+ ++G VK+ R+
Sbjct: 61 DSRKR---DKKALCLIYQGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLR 117
Query: 123 RKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRSWRHIVTAIT 182
+FE +MKE E + + + R + N L+ + + K+LR L + HIVT I
Sbjct: 118 GEFEALQMKEGELVSDYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIE 177
Query: 183 EAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDNQQQEL 242
E KDL+ + +E L+GSL+A+E ++ K +++ ++ +E Q + Q +
Sbjct: 178 ETKDLEAMTIEQLLGSLQAYEEK-KKKKEDIVEQVLNMQITKEENGQSYQRRGGGQVRGR 236
Query: 243 DE---------EDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKSQVTCYGC 293
H+D QR + + R K + K+ DKS V CY C
Sbjct: 237 GRGGYGNGRGWRPHEDNTN----------QRGENSSRG--RGKGHPKSRYDKSSVKCYNC 284
Query: 294 YKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMA--KSDNEEVN 351
K GHY +EC + SN + +K+ + EE+ E+ D+ LMA K D +E N
Sbjct: 285 GKFGHYASEC----KAPSNKKFEEKA-----NYVEEKIQEE---DMLLMASYKKDEQEEN 332
Query: 352 LDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDLIKNIQHSE 411
+ H+ M + E +R N+ L E ++ K +++ +++ +
Sbjct: 333 HKWYLDSGASNHMCGR----KSMFAELDESVRG-NVALGDESKMEVKGKGNILIRLKNGD 387
Query: 412 SKEVSE-NKNNSQKENII-------------LKENVLKLKNDISNFV 444
+ +S S K NI+ LK+N L +++ SN +
Sbjct: 388 HQFISNVYYIPSMKTNILSLGQLLEKGYDIRLKDNNLSIRDQESNLI 434
>At3g59720 copia-type reverse transcriptase-like protein
Length = 1272
Score = 127 bits (319), Expect = 5e-29
Identities = 119/467 (25%), Positives = 207/467 (43%), Gaps = 63/467 (13%)
Query: 6 SSNRP---PLFEGTDYYYWKGKMELFLQS*DNHMWTVVENGNYIPYDEDLNEKKKEDWSE 62
S+N P P+ ++Y W +M+ L + D +W +VE G P +E + ++D
Sbjct: 3 SNNVPFQVPVLTKSNYDNWSLRMKAILGAHD--VWEIVEKGFIEPENEGSLSQTQKDGLR 60
Query: 63 EEGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGV 122
+ KR + KA + L + +++V E +AKE W+ L+ ++G VK+ R+
Sbjct: 61 DSRKR---DKKALCLIYQGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLR 117
Query: 123 RKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRSWRHIVTAIT 182
+FE +MKE E + + + R + N L+ + + K+LR L + HIVT I
Sbjct: 118 GEFEALQMKEGELVSDYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIE 177
Query: 183 EAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDNQQQEL 242
E KDL+ + +E L+GSL+A+E ++ K +++ ++ +E Q + Q +
Sbjct: 178 ETKDLEAMTIEQLLGSLQAYEEK-KKKKEDIVEQVLNMQITKEENGQSYQRRGGGQVRGR 236
Query: 243 DE---------EDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKSQVTCYGC 293
H+D QR + + R K + K+ DKS V CY C
Sbjct: 237 GRGGYGNGRGWRPHEDNTN----------QRGENSSRG--RGKGHPKSRYDKSSVKCYNC 284
Query: 294 YKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMA--KSDNEEVN 351
K GHY +EC + SN K F + EE+ E+ D+ LMA K D +E N
Sbjct: 285 GKFGHYASEC----KAPSN-----KKFKEKANYVEEKIQEE---DMLLMASYKKDEQEEN 332
Query: 352 LDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDLIKNIQHSE 411
+ H+ M + E +R N+ L E ++ K +++ +++ +
Sbjct: 333 HKWYLDSGASNHMCGR----KSMFAELDESVRG-NVALGDESKMEVKGKGNILIRLKNGD 387
Query: 412 SKEVSE-NKNNSQKENII-------------LKENVLKLKNDISNFV 444
+ +S S K NI+ LK+N L +++ SN +
Sbjct: 388 HQFISNVYYIPSMKTNILSLGQLLEKGYDIRLKDNNLSIRDKESNLI 434
>At3g61330 copia-type polyprotein
Length = 1352
Score = 127 bits (318), Expect = 7e-29
Identities = 117/467 (25%), Positives = 208/467 (44%), Gaps = 63/467 (13%)
Query: 6 SSNRP---PLFEGTDYYYWKGKMELFLQS*DNHMWTVVENGNYIPYDEDLNEKKKEDWSE 62
S+N P P+ ++Y W +M+ L + D +W +VE G P +E + ++D
Sbjct: 3 SNNVPFQVPVLTKSNYDNWSLRMKAILGAHD--VWEIVEKGFIEPENEGSLSQTQKDGLR 60
Query: 63 EEGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGV 122
+ KR + KA + L + +++V E +AKE W+ L+ ++G VK+ R+
Sbjct: 61 DSRKR---DKKALCLIYQGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLR 117
Query: 123 RKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRSWRHIVTAIT 182
+FE +MKE E + + + R + N L+ + + K+LR L + HIVT I
Sbjct: 118 GEFEALQMKEGELVSDYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIE 177
Query: 183 EAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDNQQQEL 242
E KDL+ + +E L+GSL+A+E ++ K +++ ++ +E Q + Q +
Sbjct: 178 ETKDLEAMTIEQLLGSLQAYEE-KKKKKEDIAEQVLNMQITKEENGQSYQRRGGGQVRGR 236
Query: 243 DE---------EDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKSQVTCYGC 293
H+D QR + + R K + K+ DKS V CY C
Sbjct: 237 GRGGYGNGRGWRPHED----------NTNQRGENSSRG--RGKGHPKSRYDKSSVKCYNC 284
Query: 294 YKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMA--KSDNEEVN 351
K GHY +EC + SN + +K+ EE+ E+ D+ LMA K D ++ N
Sbjct: 285 GKFGHYASEC----KAPSNKKFEEKAHYV-----EEKIQEE---DMLLMASYKKDEQKEN 332
Query: 352 LDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDLIKNIQHSE 411
+ H+ M + E +R N+ L E ++ K +++ +++ +
Sbjct: 333 HKWYLDSGASNHMCGR----KSMFAELDESVRG-NVALGDESKMEVKGKGNILIRLKNGD 387
Query: 412 SKEVSE-NKNNSQKENII-------------LKENVLKLKNDISNFV 444
+ +S S K NI+ LK+N L +++ SN +
Sbjct: 388 HQFISNVYYIPSMKTNILSLGQLLEKGYDIRLKDNNLSIRDQESNLI 434
>At2g05930 copia-like retroelement pol polyprotein
Length = 916
Score = 126 bits (317), Expect = 9e-29
Identities = 116/505 (22%), Positives = 222/505 (42%), Gaps = 46/505 (9%)
Query: 12 LFEGTDYYYWKGKMELFLQS*DNHMWTVVENGNYIPY--DEDLNE--KKKEDWSEEEGKR 67
+ E +Y +WK KM ++ W G P ED + K ++ W++ E +
Sbjct: 27 ILEKGNYGHWKVKMRALIRGLGKEAWIATSIGWKAPVIKGEDGEDVLKTEDQWNDAEEAK 86
Query: 68 MLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGVRKFEL 127
N +A + ++++ ++ ++Q C++AKE WD L +EGTS VK +RID+ +FE
Sbjct: 87 ATANSRALSLIFNSVNQNQFKQIQNCESAKEAWDKLAKAYEGTSSVKRSRIDMLASQFEN 146
Query: 128 FEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRSWRHIVTAITEAKDL 187
M+ETE I+E G+ + I +E +L K + + V+K+LRCLP + TA+ + D
Sbjct: 147 LTMEETENIEEFSGKISAIASEAHNLGKKYKDKKLVKKLLRCLPSRFESKRTAMGTSLDT 206
Query: 188 KKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDNQQQELDEEDH 247
+ E+++G +A+E+ + +S K +K SL ++ D + E H
Sbjct: 207 NSIDFEEVVGMFQAYEL----EITSGKGGYGHIKAECPSLKRK-----DLKCSECKGLGH 257
Query: 248 QDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKSQVTCYGCYKLGHYKNECPLNK 307
I+ ++ P R ++++E D + G ++
Sbjct: 258 --------------IKFDCVGSKSKPDRSCSSESESDSND---------GDSEDYIKGFV 294
Query: 308 RKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMAKSDNEEVNLDP---CSSCEKTEHL 364
++ +S + D +E + DE +D+ K + E L S EK L
Sbjct: 295 SFVGIIEEKDESSDSEADGEDEDNSADEDSDIEKDVKINEEFRKLYDSWLMLSKEKVAWL 354
Query: 365 FDNLFYS--SQMLEQECERLRNENLELRKEKEILEKENKDLIKNIQHSESKEVSENKNNS 422
+ L ++ L+ E +N EL ++ + E++N++L + + + K N
Sbjct: 355 EEKLKVQELTEKLKGELTAANQKNSELTQKCSVAEEKNRELSQELSDTRKKIHMLNSGTK 414
Query: 423 QKENIILKENVLKLKNDISNFVKSTETFQKIMGSQVSVFDKACIGFKTN----QKQKLYE 478
++I+ V K + + T + S+ K+ GF++N +++Y+
Sbjct: 415 DLDSILAAGRVGKSNFGLGYNGVGSGTKTNFVRSEAGAPTKSQTGFRSNYDAVPARRMYQ 474
Query: 479 NFFIPKKNKNTCERYKCTYCEKYGH 503
N ++ T Y+C YC ++GH
Sbjct: 475 N-HDHYHSRRTVTGYECYYCGRHGH 498
>At3g60170 putative protein
Length = 1339
Score = 109 bits (273), Expect = 1e-23
Identities = 87/345 (25%), Positives = 153/345 (44%), Gaps = 62/345 (17%)
Query: 11 PLFEGTDYYYWKGKMELFLQS*DNHMWTVVENGNYIPYDEDLNEKKKEDWSEEEGKRMLL 70
P F+G Y +W ME FL+S +W +VE G + + + EE K L
Sbjct: 13 PRFDGY-YDFWSMTMENFLRS--RELWRLVEEGIPAIVVGTTPVSEAQRSAVEEAK--LK 67
Query: 71 NFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGVRKFELFEM 130
+ K K FL A+ RE + + + +K IW+++K ++G++ VK ++ ++FEL M
Sbjct: 68 DLKVKNFLFQAIDREILETILDKSTSKAIWESMKKKYQGSTKVKRAQLQALRKEFELLAM 127
Query: 131 KETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRSWRHIVTAITEAKDLKKL 190
KE E ID GR ++N++++ + V KILR L + ++V +I E+ DL L
Sbjct: 128 KEGEKIDTFLGRTLTVVNKMKTNGEVMEQSTIVSKILRSLTPKFNYVVCSIEESNDLSTL 187
Query: 191 RLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDNQQQELDEEDHQDQ 250
+++L GSL HE Q L+ ++Q
Sbjct: 188 SIDELHGSLLVHE------------------------------------QRLNGHVQEEQ 211
Query: 251 IILLTRKLQRMIQRRDQN--KRNFPARKENAKTELDKSQVTCYGCYKLGHYKNECPLNKR 308
+ +T + +R Q R + + + + ++ +++ V CY C+ LGH++ ECP
Sbjct: 212 ALKVTHE-ERPSQGRGRGVFRGSRGRGRGRGRSGTNRAIVECYKCHNLGHFQYECP---- 266
Query: 309 KSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMAKSDNEEVNLD 353
W++ +E+ +L LMA + + N D
Sbjct: 267 --------------EWEKNANYAELEEEEELLLMAYVEQNQANRD 297
>At2g15650 putative retroelement pol polyprotein
Length = 1347
Score = 108 bits (271), Expect = 2e-23
Identities = 97/423 (22%), Positives = 185/423 (42%), Gaps = 58/423 (13%)
Query: 3 EGGSSNRPPLFEGTDYYYWKGKMELFLQS*DNHMWTVVENGNYIPYD----EDLNEKKKE 58
+G S P+F+G Y +W KM ++ +W+VVE G +P + E+ E +
Sbjct: 2 QGSSHQVIPIFDGEKYDFWSIKMATIFRT--RKLWSVVEEG--VPVEPVQAEETPETARA 57
Query: 59 DWSEEEGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKETRI 118
EE + + A L A++ + + R+ ++KE WD LK ++G+ V+ ++
Sbjct: 58 KTLREEA--VTNDTMALQILQTAVTDQIFSRIAAASSSKEAWDVLKDEYQGSPQVRLVKL 115
Query: 119 DIGVRKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRSWRHIV 178
R++E +M + + I + ++ +L + T + ++KIL LP + IV
Sbjct: 116 QSLRREYENLKMYDNDNIKTFTDKLIVLEIQLTYHGEKKTNTQLIQKILISLPAKFDSIV 175
Query: 179 TAITEAKDLKKLRLEDLIGSLKAHE--VLLQEDKSSNKSKMIALKTNQESLNQELEMNFD 236
+ + + +DL L + +L+G LKA E V +E+ + + + K + Q+ N
Sbjct: 176 SVLEQTRDLDALTMSELLGILKAQEARVTAREESTKEGAFYVRSKGRESGFKQDNTNNRV 235
Query: 237 NQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKSQVTCYGCYKL 296
NQ ++ + + ++ R P ++ K + +S + CY C K+
Sbjct: 236 NQDKK-------------WCGFHKSSKHTEEECREKPKNDDHGKNK--RSNIKCYKCGKI 280
Query: 297 GHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMAKSDNEEVNL---- 352
GHY NEC ++ +++ EE NED L + S+ E L
Sbjct: 281 GHYANECRSKNKERAHVTLE-----------EEDVNEDHM----LFSASEEESTTLREDV 325
Query: 353 ----DPCSS-CEKTEHLFDNLFYSSQMLEQECERLRNENLELRKEK---EILEKENKDLI 404
C++ K E F N+ S ++ R+RN ++ + K ++ + K +I
Sbjct: 326 WLVDSGCTNHMTKEERYFSNINKSIKV----PIRVRNGDIVMTAGKGDITVMTRHGKRII 381
Query: 405 KNI 407
KN+
Sbjct: 382 KNV 384
>At4g04560 putative transposon protein
Length = 590
Score = 105 bits (261), Expect = 3e-22
Identities = 62/229 (27%), Positives = 121/229 (52%), Gaps = 4/229 (1%)
Query: 9 RPPLFEGTDYYYWKGKMELFLQS*DNHMWTVVENGNYIPYDEDLNE----KKKEDWSEEE 64
+P + Y YWK ++ +QS D W VE+G P +D K + +W +E
Sbjct: 344 KPLKLDAEHYGYWKVLIKRSIQSIDMDAWFAVEDGWMPPTTKDAKRDIVSKSRTEWIADE 403
Query: 65 GKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGVRK 124
N +A + +L R ++ +VQ C +AKE+W+ L+V E T++VK TR+D+ +
Sbjct: 404 KTAANHNSQALSVIFGSLLRNKFTQVQGCLSAKEVWEILQVSFECTNNVKRTRLDMLASE 463
Query: 125 FELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRSWRHIVTAITEA 184
FE M+ E++D+ G+ + I E L K + + V+K LR LP ++ +AI +
Sbjct: 464 FENLTMEAEESVDDFNGKLSSITQEAVVLGKTYKDKKMVKKFLRSLPDKFQSHKSAIDVS 523
Query: 185 KDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEM 233
+ +L+ + ++G ++A++ +E +S + A++ + ++ ++ +M
Sbjct: 524 LNSDQLKFDQVVGMMQAYDTDKEEILNSYATYFGAIEDDDHTVEEDAQM 572
Score = 43.1 bits (100), Expect = 0.001
Identities = 40/181 (22%), Positives = 76/181 (41%), Gaps = 26/181 (14%)
Query: 11 PLFEGTDYYYWKGKMELFLQS*DNHMWTVVENGNYIPYDEDLNEKKKEDWSEEEGKRMLL 70
P+F +Y +W+ KM+ Q+ +W +V+ G P E + + L
Sbjct: 8 PIFNKENYGFWRIKMKTIFQT--KKLWEIVDEGVPKPPAEGDHSPEAVQQKTRCEAASLK 65
Query: 71 NFKAKLFLTMALSREEYDRVQECKNAK-EIWDTLKVHHEGTSHVKETRIDIGVRKFELFE 129
+ A L A+S + R+ +A + W+ +E +
Sbjct: 66 DLTALQILQTAVSDSIFPRIAPASSALGKPWE-----------------------YENLK 102
Query: 130 MKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRSWRHIVTAITEAKDLKK 189
MKE++ I+ + + N+LR ++ + ++ V+KIL LP+ + IV + + KDL
Sbjct: 103 MKESDNINTFMTKLIEMGNQLRVHGEEKSDYQIVQKILISLPKRFDIIVAMMKQTKDLTS 162
Query: 190 L 190
L
Sbjct: 163 L 163
>At1g36110 hypothetical protein
Length = 745
Score = 93.6 bits (231), Expect = 8e-19
Identities = 76/303 (25%), Positives = 124/303 (40%), Gaps = 59/303 (19%)
Query: 6 SSNRPPLFEGTDYYYWKGKMELFLQS*DNHMWTVVENGNYIPYDEDLNEKKKEDWSEEEG 65
SS + P+ T+Y W +M++ N MW +E G ++G
Sbjct: 15 SSTQCPMLTSTNYTVWAIRMQIARSV--NKMWETIEPGI------------------DDG 54
Query: 66 KRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGVRKF 125
+ N A+ + ++ +V AK +WD++K + G VKE R+ + +F
Sbjct: 55 YK---NTMARGLIFQSIPESLTLQVGTLATAKLVWDSIKTRYVGADRVKEARLQTLMAEF 111
Query: 126 ELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLP-RSWRHIVTAITEA 184
E +MKE+E ID GR + +L + + V+K L LP R + HI+ ++ +
Sbjct: 112 EKMKMKESEKIDVFAGRLAELATRSDALGSNIETSKLVKKFLNALPLRKYIHIIASLEQV 171
Query: 185 KDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDNQQQELDE 244
DL ED++G +K +E E +D ++Q
Sbjct: 172 LDLNNTSFEDIVGRIKVYE----------------------------ERVWDGEEQ---- 199
Query: 245 EDHQDQIILLTRKLQR--MIQRRDQNKRNFPAR-KENAKTELDKSQVTCYGCYKLGHYKN 301
ED Q +++ Q R F R + + D S+VTCY C KLGHY +
Sbjct: 200 EDDQGKLMYANTDTQDSWYASRGRGQGGGFNGRGRGRGRGSRDTSKVTCYRCDKLGHYAS 259
Query: 302 ECP 304
CP
Sbjct: 260 NCP 262
>At3g25450 hypothetical protein
Length = 1343
Score = 92.8 bits (229), Expect = 1e-18
Identities = 65/267 (24%), Positives = 127/267 (47%), Gaps = 25/267 (9%)
Query: 4 GGSSNRPPLFEGTDYYYWKGKMELFLQS*DNHMWTVVENGNYIPYDEDLNEKKKEDWSEE 63
G +S + P+ +Y W +ME L+ + +W +E G+ DE+ N+
Sbjct: 16 GSASIQCPMLNSVNYTVWTMRMEAVLRV--HKLWGTIEPGSA---DEEKNDM-------- 62
Query: 64 EGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGVR 123
A+ L ++ +V + K + +W+ +K + G VKE R+ +
Sbjct: 63 ----------ARALLFQSIPESLILQVGKQKTSSAVWEAIKSRNLGAERVKEARLQTLMA 112
Query: 124 KFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPR-SWRHIVTAIT 182
+F+ +MK++ETID+ GR + I + +L +D + V+K L+ LPR + HIV A+
Sbjct: 113 EFDKLKMKDSETIDDYVGRISEITTKAAALGEDIEESKIVKKFLKSLPRKKYIHIVAALE 172
Query: 183 EAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDNQ-QQE 241
+ DLK ED+ G +K +E + +D S++ + + +E + +LE + +E
Sbjct: 173 QVLDLKTTTFEDIAGRIKTYEDRVWDDDDSHEDQGKLMTEVEEEVVDDLEEEEEEVINKE 232
Query: 242 LDEEDHQDQIILLTRKLQRMIQRRDQN 268
+ + H +L +LQ ++ + +
Sbjct: 233 IKAKSHVIDRLLKLIRLQEQKEKEEDD 259
>At2g05390 putative retroelement pol polyprotein
Length = 1307
Score = 92.8 bits (229), Expect = 1e-18
Identities = 74/320 (23%), Positives = 139/320 (43%), Gaps = 41/320 (12%)
Query: 71 NFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGVRKFELFEM 130
N A+ L ++ +V + K +K +W+ +K + G VKE ++ + +F+ M
Sbjct: 24 NDMARALLFQSVPESTILQVGKHKTSKAMWEAIKTRNLGAERVKEAKLQTLMAEFDRLNM 83
Query: 131 KETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRS-WRHIVTAITEAKDLKK 189
K+ ETIDE GR + I + SL ++ + V+K L+ LPR + HI+ A+ + DL
Sbjct: 84 KDNETIDEFVGRISEISTKSESLGEEIEESKIVKKFLKSLPRKKYIHIIAALEQILDLNT 143
Query: 190 LRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDNQQQELDEEDHQD 249
ED++G +K +E + ++ S + + + N ES ++D
Sbjct: 144 TGFEDIVGRMKTYEDRVCDEDDSPEEQGKLMYANSES-------SYDT------------ 184
Query: 250 QIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKSQVTCYGCYKLGHYKNECPLNKRK 309
R + R + + + R + DKS+V CY C K GHY +EC K
Sbjct: 185 ----------RGGRGRGRGRSSGRGRGGYGYQQRDKSKVICYRCDKTGHYASECLDRLLK 234
Query: 310 SSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMAKSDNEEVNLDPCSSCE------KTEH 363
Q Q++ +E +++ ++ + + + + CS + H
Sbjct: 235 LIKAQEQQQN-----NEDDDEIESLMMHEVVYLNERSVKPKEFEACSDNSWYLDNGASNH 289
Query: 364 LFDNLFYSSQMLEQECERLR 383
+ NL + S++ E ++R
Sbjct: 290 MTGNLQWFSKLNEMITGKVR 309
>At3g31380 hypothetical protein
Length = 262
Score = 86.3 bits (212), Expect = 1e-16
Identities = 44/151 (29%), Positives = 85/151 (56%), Gaps = 4/151 (2%)
Query: 89 RVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGVRKFELFEMKETETIDEMYGRFTIIMN 148
+V + AKE+WD++K H G VKE R+ + FE +MKE E ID+ GR + +
Sbjct: 17 QVGNLQTAKEVWDSIKTRHVGAERVKEARVQTLMADFEKMKMKEAEKIDDFAGRLSELST 76
Query: 149 ELRSLEKDFTIHERVRKILRCLPR-SWRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLL- 206
+ L + + + V+K L LPR + HI+ ++ + DL ED++G +K +E +
Sbjct: 77 KSAPLGVNIEVPKLVKKFLNSLPRKKYIHIIASLEQVLDLNNTTFEDIVGCMKVNEERVY 136
Query: 207 --QEDKSSNKSKMIALKTNQESLNQELEMNF 235
+E+ + +++K++ ++ +S N++ + +F
Sbjct: 137 DPEEETNEDQNKLMYTSSDSKSGNRDYQGDF 167
>At1g32590 hypothetical protein, 5' partial
Length = 1263
Score = 86.3 bits (212), Expect = 1e-16
Identities = 58/255 (22%), Positives = 117/255 (45%), Gaps = 32/255 (12%)
Query: 63 EEGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGV 122
E ++ + + K K +L ++ + + + + +K++W+++K ++G V+ ++
Sbjct: 17 ELAEKTVKDHKVKNYLFASIDKTILKTILQKETSKDLWESMKRKYQGNDRVQSAQLQRLR 76
Query: 123 RKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRSWRHIVTAIT 182
R FE+ EMK ETI + R I N++R+L +D + V KILR L + ++V AI
Sbjct: 77 RSFEVLEMKIGETITGYFSRVMEITNDMRNLGEDMPDSKVVEKILRTLVEKFTYVVCAIE 136
Query: 183 EAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDNQQQEL 242
E+ ++K+L ++ L SL HE L + L E + D +
Sbjct: 137 ESNNIKELTVDGLQSSLMVHEQNLSRH-----------DVEERVLKAETQWRPDGGRGRG 185
Query: 243 DEEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKSQVTCYGCYKLGHYKNE 302
R + + + R + +++ V C+ C+K+GHYK E
Sbjct: 186 GSPS------------------RGRGRGGYQGR---GRGYVNRDTVECFKCHKMGHYKAE 224
Query: 303 CPLNKRKSSNLQTNQ 317
CP +++++ ++ +
Sbjct: 225 CPSWEKEANYVEMEE 239
>At2g05960 putative retroelement pol polyprotein
Length = 1200
Score = 77.4 bits (189), Expect = 6e-14
Identities = 41/145 (28%), Positives = 79/145 (54%), Gaps = 8/145 (5%)
Query: 89 RVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGVRKFELFEMKETETIDEMYGRFTIIMN 148
+V + K +K +WD ++ + G VKE ++ + +F+ +MK+ ETIDE GR + I
Sbjct: 36 QVGKLKTSKAVWDKIQSRNLGAERVKEAKLKTLMAEFDKLKMKDNETIDECAGRLSEIST 95
Query: 149 ELRSLEKDFTIHERVRKILRCLP-RSWRHIVTAITEAKDLKKLRLEDLIGSLKAHE---- 203
+ SL +D + V+K L+ LP + + HIV A+ + DLK +D++G +K +E
Sbjct: 96 KSTSLGEDIEETKVVKKFLKSLPTKKYIHIVAALEQVLDLKNTTFKDIVGRIKTYEDKIW 155
Query: 204 ---VLLQEDKSSNKSKMIALKTNQE 225
L+++ + ++ ++ +E
Sbjct: 156 VLITCLKKEAEEEEKSVVGVEAEEE 180
>At5g56830 unknown protein
Length = 242
Score = 57.0 bits (136), Expect = 8e-08
Identities = 48/214 (22%), Positives = 95/214 (43%), Gaps = 15/214 (7%)
Query: 82 LSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGVRKFELFEMKETETIDEMYG 141
LS E + + ++KE W+ L V G + ++ +++ M + ET+ E G
Sbjct: 21 LSTEIFSWIAYATSSKEAWEILMVRSVGGPIFQARKLLDLKKEYNNTTMSDAETVHEFTG 80
Query: 142 RFTIIMNELRSLEKDFTIHERVRKILRCLPRSWRHIVTAITEAKDLKKLRLEDLIGSLKA 201
+ ++ ++S V+KIL LP + ++ + E KL + +I L
Sbjct: 81 KLLELVFRMKSCGWKVEEKILVKKILSSLPPRFDKVLVEVAE-----KLSVNVIIDCLLV 135
Query: 202 HEVLLQEDKSSNKSKMIALKTNQESLNQELEMNF---DNQQQELDEEDHQDQIILLTRKL 258
HE ++ D+ S + +I +ES E E+ + D Q ++ + Q +++ L +K
Sbjct: 136 HEYNMRPDQESFQESVIQSVNVEESFEAEAEILYSLNDITQSQVAMMERQSEVLSLVQKN 195
Query: 259 QRMIQRRD-------QNKRNFPARKENAKTELDK 285
Q+M+ + Q K + A E+ + E+ K
Sbjct: 196 QKMLMQMQKQMMMLMQQKDHVLAHSESYEEEMTK 229
>At3g21010 unknown protein
Length = 438
Score = 53.5 bits (127), Expect = 9e-07
Identities = 69/330 (20%), Positives = 127/330 (37%), Gaps = 63/330 (19%)
Query: 36 MWTVVENGNYIPYDED--------LNEKKKEDWSEEEGKRMLLNFKAKLFLTMALSREEY 87
+W VVENG +P D + ++ W + K M KA L +L+ +
Sbjct: 33 LWDVVENG--VPPDPSKVPELAATIQAEEFSKWRDLAEKDM----KALQILQSSLTDSAF 86
Query: 88 DRVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGVRKFELFEMKETETIDEMYGRFTIIM 147
+ +AK++WD+L+ + + ++ ++FE M E E I+ R I+
Sbjct: 87 RKTLSASSAKDLWDSLEKGNNEQAKLRRLE-----KQFEELSMNEGEPINPYIDRVIEIV 141
Query: 148 NELRSLEKDFTIHERVRKILRCLPRSWRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQ 207
+LR + + ++ + K+L L S+ + + + DLK + L+ + E+
Sbjct: 142 EQLRRFKIAKSDYQVIEKVLATLSGSYDDVAPVLGDLVDLKNMSLKSFV------ELFYV 195
Query: 208 EDKSSNKSKMIALKTNQESLNQELEM----NFDNQQQELDEEDHQDQIILLTRKLQRMIQ 263
D + +S + LK N+ E E N +N +QE
Sbjct: 196 YDSITEESIHLMLKANRLRSMAEEESCVLCNKNNHKQE---------------------- 233
Query: 264 RRDQNKRNFPARKENAKTELDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTT 323
D + A K + + + C+ C + GH K K N + Q + T
Sbjct: 234 --DCSSSIPKAGKSSGARQSKPKKGKCFQCGERGH--------KAKDCNKKIQQPA--VT 281
Query: 324 WDEPEEQTNEDEQADLCLMAKSDNEEVNLD 353
D+ E ++ + D L+A + E D
Sbjct: 282 EDQIEIVEEKEIEVDFLLLAALNLNEFTYD 311
>At1g66840 hypothetical protein
Length = 607
Score = 53.5 bits (127), Expect = 9e-07
Identities = 99/470 (21%), Positives = 188/470 (39%), Gaps = 75/470 (15%)
Query: 50 EDLNEKKKEDWSEEEGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEG 109
EDL++ +E E +R+ + KAK V+ CK K + + E
Sbjct: 34 EDLHKSGRELGIYRESRRVAESAKAKA------------EVELCKAKKIVKELTLRIEES 81
Query: 110 TSHVKETRIDIGVRKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFT----------- 158
+K RIDI E E+ + G + IM EL ++++ +
Sbjct: 82 NRRLKSRRIDI--------EAVMNESRIDGNGGYVRIMRELEDMKQELSKLKLDVVYVSR 133
Query: 159 -----------IHERVRKILRCLPRSWRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQ 207
+ R+ + L+ L + A E ++ ++E L + E +
Sbjct: 134 EKVVAEKEVMELESRMEENLKLLESLKLEVDVANEEHVLVEVAKIEALKECKEVEEQREK 193
Query: 208 EDKSSNKSKMIALKTNQESLNQ-ELEMNFDNQQQE--LDEEDHQDQIILLTRKLQRMIQR 264
E K ++S K +E + + E NF+N+ E LD E + Q+ L+ ++++R +QR
Sbjct: 194 ERKEVSESLHKRKKRIREMIREIERSKNFENELAETLLDIEMLETQLKLV-KEMERKVQR 252
Query: 265 RDQNKRN----FPARKENAKTELDKSQVTCYGCYKLGHYKNE--CPLNKRKSSNLQTNQK 318
+ R+ F K+N + ++ T +L E C +N + + +
Sbjct: 253 NESMSRSKNRAFERGKDNLSVLKEVTEATEAKKAELASINAELFCLVNTMDTLRKEFDHA 312
Query: 319 SFMTTWDEPEEQTNE---DEQADLCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQML 375
T W + Q ++ + L+AK E V+ + E+ +L DNL S + L
Sbjct: 313 KKETAWLDKMIQKDDVMLERLNTKLLIAKDQLEAVS----KAEERISYLADNLTTSFEKL 368
Query: 376 EQECERLRNENLELRKEKEILEKE-----------NKDLIKNIQHSESKEVSENKNNSQK 424
+ + E + E L+LR+E I+ E K+L+ + E + +E+ +
Sbjct: 369 KSDREAAKKEELKLREEARIINNEIQKTETGFDGKEKELLSKLDELEKAKHAESLALEKL 428
Query: 425 ENIILKENVLKLKNDISNFVKSTETFQKIMGSQVSVFDKACIGFKTNQKQ 474
E ++ K + ++ + ST T + +S KAC +T +K+
Sbjct: 429 ETMVEKTMETR---EMESRRNSTITISRFEYEYLS--GKACHAEETAEKK 473
>At2g13930 putative retroelement pol polyprotein
Length = 1335
Score = 53.1 bits (126), Expect = 1e-06
Identities = 43/163 (26%), Positives = 79/163 (48%), Gaps = 9/163 (5%)
Query: 49 DEDLNEKKKEDWSEEEGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLK--VH 106
+ED ++KK D +E R+ KAK + + ++ + +++ CK A E W+TL
Sbjct: 26 EEDPEKRKKRD--ADEVARLERCDKAKNVIFLNVADKVLRKIELCKTAAEAWETLDRLFM 83
Query: 107 HEGTSHVKETRIDIGVRKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKI 166
H T++ F F+M+E + IDE F I+ +L L+ D T + +
Sbjct: 84 IRSLPHRVYTQLS-----FYTFKMQENKKIDENIDDFLKIVADLNHLQIDVTDEVQAILL 138
Query: 167 LRCLPRSWRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQED 209
L LP + +V + + +KLRL+D++ + + E L ++
Sbjct: 139 LSSLPARYDGLVETMKYSNSREKLRLDDVMVAARDKERELSQN 181
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.355 0.157 0.539
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,757,358
Number of Sequences: 26719
Number of extensions: 1175588
Number of successful extensions: 10833
Number of sequences better than 10.0: 232
Number of HSP's better than 10.0 without gapping: 47
Number of HSP's successfully gapped in prelim test: 193
Number of HSP's that attempted gapping in prelim test: 9883
Number of HSP's gapped (non-prelim): 769
length of query: 1574
length of database: 11,318,596
effective HSP length: 113
effective length of query: 1461
effective length of database: 8,299,349
effective search space: 12125348889
effective search space used: 12125348889
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 67 (30.4 bits)
Medicago: description of AC144538.13


