

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144516.5 - phase: 0
(332 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
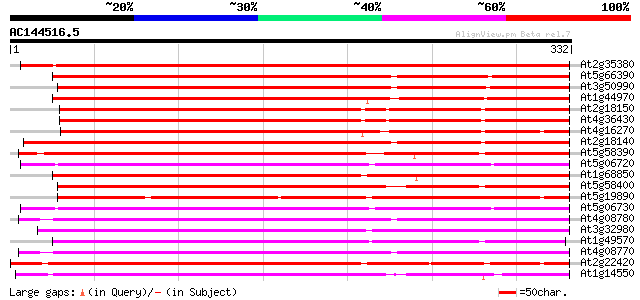
Score E
Sequences producing significant alignments: (bits) Value
At2g35380 putative peroxidase 430 e-121
At5g66390 peroxidase (emb|CAA66964.1) 319 1e-87
At3g50990 peroxidase-like protein 315 3e-86
At1g44970 peroxidase like protein 315 3e-86
At2g18150 putative peroxidase 308 2e-84
At4g36430 peroxidase like protein 302 1e-82
At4g16270 peroxidase like protein 294 5e-80
At2g18140 putative peroxidase 289 1e-78
At5g58390 peroxidase 278 3e-75
At5g06720 peroxidase (emb|CAA68212.1) 276 9e-75
At1g68850 peroxidase ATP23a 270 1e-72
At5g58400 peroxidase 263 1e-70
At5g19890 peroxidase ATP N 263 1e-70
At5g06730 peroxidase 263 1e-70
At4g08780 peroxidase C2 precursor like protein 260 8e-70
At3g32980 peroxidase, putative 259 2e-69
At1g49570 peroxidase like protein 258 4e-69
At4g08770 peroxidase C2 precursor like protein 256 1e-68
At2g22420 putative peroxidase 256 2e-68
At1g14550 peroxidase, putative 256 2e-68
>At2g35380 putative peroxidase
Length = 336
Score = 430 bits (1105), Expect = e-121
Identities = 212/325 (65%), Positives = 264/325 (81%), Gaps = 1/325 (0%)
Query: 7 LLILITILSNIHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFH 66
L++L I +++ G LL +YKE CPLAE+IV+HN+ VAVLKDPR+AASLLRL FH
Sbjct: 12 LIVLYAITTSVLGDFGEPLL-KGFYKESCPLAEEIVKHNIEVAVLKDPRMAASLLRLQFH 70
Query: 67 DCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILA 126
DCFV+GCDASVLLD+ M SEKQA PN+NSLRGFEVID IKYLLE+ CPLTVSC+DILA
Sbjct: 71 DCFVLGCDASVLLDTHGDMLSEKQATPNLNSLRGFEVIDYIKYLLEEACPLTVSCSDILA 130
Query: 127 MVARDAVELRGGPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDL 186
+ ARD+V LRGGP WEV LGR+DSL++SF+GAN FIPAPNSSL++LI NFKQQGL+I+DL
Sbjct: 131 LAARDSVFLRGGPWWEVLLGRRDSLKASFAGANQFIPAPNSSLDSLIINFKQQGLNIQDL 190
Query: 187 VVLSGSHTIGRARCLSFRQRIYETKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKFAP 246
+ LSG+HTIG+ARC+SF+QRI + E D ++R++TFRR+L S C + RD++ +P
Sbjct: 191 IALSGAHTIGKARCVSFKQRIVQPNMEQTFYVDEFRRHSTFRRVLGSQCKDSSRDNELSP 250
Query: 247 LDFQTPKRFDNQYFINIIEGKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKS 306
LD +TP FDN YFIN++EG+GLL SDNVL+S+D +G I ++VW YA N+ LFF F +S
Sbjct: 251 LDIKTPAYFDNHYFINLLEGRGLLISDNVLVSEDHEGEIFQKVWEYAVNQDLFFIDFVES 310
Query: 307 MIKMGNINVLTGSEGEIRRNCRFVN 331
M+KMGNINVLTG EGEIR NCRFVN
Sbjct: 311 MLKMGNINVLTGIEGEIRENCRFVN 335
>At5g66390 peroxidase (emb|CAA66964.1)
Length = 336
Score = 319 bits (818), Expect = 1e-87
Identities = 165/306 (53%), Positives = 216/306 (69%), Gaps = 5/306 (1%)
Query: 26 LVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDSVEGM 85
L ++Y + CP A++IV+ VA A DPR+ ASLLRLHFHDCFV GCDAS+LLDS +
Sbjct: 33 LFPQFYDQSCPKAQEIVQSIVAKAFEHDPRMPASLLRLHFHDCFVKGCDASILLDSSGTI 92
Query: 86 TSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVARDAVELRGGPRWEVWL 145
SEK++ PN NS RGFE+I++IK+ LE+ECP TVSCADILA+ ARD+ + GGP WEV L
Sbjct: 93 ISEKRSNPNRNSARGFELIEEIKHALEQECPETVSCADILALAARDSTVITGGPSWEVPL 152
Query: 146 GRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGRARCLSFRQ 205
GR+D+ +S SG+N IPAPN++ +T++ FK+QGLD+ DLV LSGSHTIG +RC SFRQ
Sbjct: 153 GRRDARGASLSGSNNDIPAPNNTFQTILTKFKRQGLDLVDLVSLSGSHTIGNSRCTSFRQ 212
Query: 206 RIYETKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKFAPLDFQTPKRFDNQYFINIIE 265
R+Y + Y T +L+ CP +G D LDF TP +FDN YF N+I
Sbjct: 213 RLYNQSGNGKPDMTLSQYYAT---LLRQRCPRSGGDQTLFFLDFATPFKFDNHYFKNLIM 269
Query: 266 GKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKSMIKMGNINVLTGSEGEIRR 325
KGLL SD +L +++ ++ V YA N++ FF+ FAKSM+KMGNI+ LTG++GEIRR
Sbjct: 270 YKGLLSSDEILFTKNKQS--KELVELYAENQEAFFEQFAKSMVKMGNISPLTGAKGEIRR 327
Query: 326 NCRFVN 331
CR VN
Sbjct: 328 ICRRVN 333
>At3g50990 peroxidase-like protein
Length = 336
Score = 315 bits (806), Expect = 3e-86
Identities = 160/303 (52%), Positives = 211/303 (68%), Gaps = 5/303 (1%)
Query: 29 EYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDSVEGMTSE 88
++Y+ CP A+ IV+ VA A DPR+AAS+LRLHFHDCFV GCDASVLLDS M SE
Sbjct: 36 QFYENSCPNAQAIVQSYVANAYFNDPRMAASILRLHFHDCFVNGCDASVLLDSSGTMESE 95
Query: 89 KQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVARDAVELRGGPRWEVWLGRK 148
K++ N +S RGFEVID+IK LE ECP TVSCAD+LA+VARD++ + GGP WEV+LGR+
Sbjct: 96 KRSNANRDSARGFEVIDEIKSALENECPETVSCADLLALVARDSIVICGGPSWEVYLGRR 155
Query: 149 DSLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGRARCLSFRQRIY 208
D+ E+S G+ IP+P S+L+T++ F QGLD+ DLV L GSHTIG +RC+ FRQR+Y
Sbjct: 156 DAREASLIGSMENIPSPESTLQTILTMFNFQGLDLTDLVALLGSHTIGNSRCIGFRQRLY 215
Query: 209 ETKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKFAPLDFQTPKRFDNQYFINIIEGKG 268
+ Y + +LQ CP++G D LD+ TP +FDN Y+ N++ +G
Sbjct: 216 NHTGNNDPDQTLNQDYAS---MLQQGCPISGNDQNLFNLDYVTPTKFDNYYYKNLVNFRG 272
Query: 269 LLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKSMIKMGNINVLTGSEGEIRRNCR 328
LL SD +L +Q ++ + V YA NE FF+ FAKSM+KMGNI+ LTG++GEIRR CR
Sbjct: 273 LLSSDEILFTQSIE--TMEMVKYYAENEGAFFEQFAKSMVKMGNISPLTGTDGEIRRICR 330
Query: 329 FVN 331
VN
Sbjct: 331 RVN 333
>At1g44970 peroxidase like protein
Length = 346
Score = 315 bits (806), Expect = 3e-86
Identities = 163/308 (52%), Positives = 214/308 (68%), Gaps = 8/308 (2%)
Query: 26 LVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDSVEGM 85
L ++Y+ CP A++IV + A+ K+PR+AASLLRLHFHDCFV GCDAS+LLD +
Sbjct: 45 LYPQFYQFSCPQADEIVMTVLEKAIAKEPRMAASLLRLHFHDCFVQGCDASILLDDSATI 104
Query: 86 TSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVARDAVELRGGPRWEVWL 145
SEK AGPN NS+RGF+VID+IK LE+ CP TVSCADILA+ AR + L GGP WE+ L
Sbjct: 105 RSEKNAGPNKNSVRGFQVIDEIKAKLEQACPQTVSCADILALAARGSTILSGGPSWELPL 164
Query: 146 GRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGRARCLSFRQ 205
GR+DS +S +GAN IPAPNS+++ L+ F+++GL+ EDLV LSG HTIG ARC +F+Q
Sbjct: 165 GRRDSRTASLNGANTNIPAPNSTIQNLLTMFQRKGLNEEDLVSLSGGHTIGVARCTTFKQ 224
Query: 206 RIYET--KQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKFAPLDFQTPKRFDNQYFINI 263
R+Y + +R Y L+SICP TG D+ +PLD +P RFDN YF +
Sbjct: 225 RLYNQNGNNQPDETLERSYYYG-----LRSICPPTGGDNNISPLDLASPARFDNTYFKLL 279
Query: 264 IEGKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKSMIKMGNINVLTGSEGEI 323
+ GKGLL SD VL++ ++ G+ V YA +E+LFF FAKSM+ MGNI LTG GEI
Sbjct: 280 LWGKGLLTSDEVLLTGNV-GKTGALVKAYAEDERLFFQQFAKSMVNMGNIQPLTGFNGEI 338
Query: 324 RRNCRFVN 331
R++C +N
Sbjct: 339 RKSCHVIN 346
>At2g18150 putative peroxidase
Length = 338
Score = 308 bits (790), Expect = 2e-84
Identities = 160/302 (52%), Positives = 207/302 (67%), Gaps = 5/302 (1%)
Query: 30 YYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDSVEGMTSEK 89
+Y+ CP AE+IVR VA AV ++ R+AASL+RLHFHDCFV GCD S+LLD+ + +EK
Sbjct: 40 FYRSSCPRAEEIVRSVVAKAVARETRMAASLMRLHFHDCFVQGCDGSLLLDTSGSIVTEK 99
Query: 90 QAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVARDAVELRGGPRWEVWLGRKD 149
+ PN S RGFEV+D+IK LE ECP TVSCAD L + ARD+ L GGP W V LGR+D
Sbjct: 100 NSNPNSRSARGFEVVDEIKAALENECPNTVSCADALTLAARDSSVLTGGPSWMVPLGRRD 159
Query: 150 SLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGRARCLSFRQRIYE 209
S +S SG+N IPAPN++ T++ F QGLD+ D+V LSGSHTIG +RC SFRQR+Y
Sbjct: 160 STSASLSGSNNNIPAPNNTFNTIVTRFNNQGLDLTDVVALSGSHTIGFSRCTSFRQRLY- 218
Query: 210 TKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKFAPLDFQTPKRFDNQYFINIIEGKGL 269
Q + + DR ++ L+ CP +G D + LD + RFDN YF N+IE GL
Sbjct: 219 -NQSGNGSPDRTLE-QSYAANLRQRCPRSGGDQNLSELDINSAGRFDNSYFKNLIENMGL 276
Query: 270 LGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKSMIKMGNINVLTGSEGEIRRNCRF 329
L SD VL S + + R+ V YA +++ FF+ FA+SMIKMGNI+ LTGS GEIR+NCR
Sbjct: 277 LNSDEVLFSS--NEQSRELVKKYAEDQEEFFEQFAESMIKMGNISPLTGSSGEIRKNCRK 334
Query: 330 VN 331
+N
Sbjct: 335 IN 336
>At4g36430 peroxidase like protein
Length = 331
Score = 302 bits (774), Expect = 1e-82
Identities = 161/302 (53%), Positives = 206/302 (67%), Gaps = 5/302 (1%)
Query: 30 YYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDSVEGMTSEK 89
YY CP +IVR VA AV ++ R+AASLLRLHFHDCFV GCD S+LLDS + +EK
Sbjct: 34 YYAHSCPQVNEIVRSVVAKAVARETRMAASLLRLHFHDCFVQGCDGSLLLDSSGRVATEK 93
Query: 90 QAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVARDAVELRGGPRWEVWLGRKD 149
+ PN S RGF+V+D+IK LEK+CP TVSCAD+L + ARD+ L GGP W V LGR+D
Sbjct: 94 NSNPNSKSARGFDVVDQIKAELEKQCPGTVSCADVLTLAARDSSVLTGGPSWVVPLGRRD 153
Query: 150 SLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGRARCLSFRQRIYE 209
S +S S +N IPAPN++ +T+++ F +QGLDI DLV LSGSHTIG +RC SFRQR+Y
Sbjct: 154 SRSASLSQSNNNIPAPNNTFQTILSKFNRQGLDITDLVALSGSHTIGFSRCTSFRQRLY- 212
Query: 210 TKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKFAPLDFQTPKRFDNQYFINIIEGKGL 269
Q + + D +F L+ CP +G D + LD + FDN YF N+IE KGL
Sbjct: 213 -NQSGNGSPDMTLE-QSFAANLRQRCPKSGGDQILSVLDIISAASFDNSYFKNLIENKGL 270
Query: 270 LGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKSMIKMGNINVLTGSEGEIRRNCRF 329
L SD VL S + + R+ V YA ++ FF+ FA+SMIKMGNI+ LTGS GEIR+NCR
Sbjct: 271 LNSDQVLFSS--NEKSRELVKKYAEDQGEFFEQFAESMIKMGNISPLTGSSGEIRKNCRK 328
Query: 330 VN 331
+N
Sbjct: 329 IN 330
>At4g16270 peroxidase like protein
Length = 362
Score = 294 bits (752), Expect = 5e-80
Identities = 158/303 (52%), Positives = 196/303 (64%), Gaps = 11/303 (3%)
Query: 31 YKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDSVEGMTSEKQ 90
Y+ CP AE IV V VL+DPR+AASLLRLHFHDCFV GCDASVLLD EG+ EK
Sbjct: 69 YRNSCPEAESIVYSWVETTVLEDPRMAASLLRLHFHDCFVNGCDASVLLDDTEGLVGEKT 128
Query: 91 AGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVARDAVELRGGPRWEVWLGRKDS 150
A PN+NSLRGFEVID IK +E CP TVSCADILAM ARD+V + GGPRWEV +GRKDS
Sbjct: 129 APPNLNSLRGFEVIDSIKSDIESVCPETVSCADILAMAARDSVVVSGGPRWEVEVGRKDS 188
Query: 151 LESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGRARCLSFRQRI--Y 208
+S A +P+PNS++ TLI+ F+ GL D+V LSG HT+G+ARC SF R+
Sbjct: 189 RTASKQAATNGLPSPNSTVSTLISTFQNLGLSQTDMVALSGGHTLGKARCTSFTARLQPL 248
Query: 209 ETKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKFAPLDFQTPKRFDNQYFINIIEGKG 268
+T Q +H + F LQ +C G LD TP FDNQY++N++ G+G
Sbjct: 249 QTGQPANHGDN-----LEFLESLQQLCSTVGPSVGITQLDLVTPSTFDNQYYVNLLSGEG 303
Query: 269 LLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKSMIKMGNINVLTGSEGEIRRNCR 328
LL SD L Q D R V YA+++ +FF+ F +M+KMG I GS EIR+NCR
Sbjct: 304 LLPSDQALAVQ--DPGTRAIVETYATDQSVFFEDFKNAMVKMGGIP--GGSNSEIRKNCR 359
Query: 329 FVN 331
+N
Sbjct: 360 MIN 362
>At2g18140 putative peroxidase
Length = 337
Score = 289 bits (740), Expect = 1e-78
Identities = 155/324 (47%), Positives = 205/324 (62%), Gaps = 6/324 (1%)
Query: 9 ILITILSNIHTLRGSEL-LVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHD 67
+ + I N G++ L ++Y+ CP AE+IVR VA A ++ R+AASL+RLHFHD
Sbjct: 17 LTLCICDNASNFGGNKRNLFPDFYRSSCPRAEEIVRSVVAKAFERETRMAASLMRLHFHD 76
Query: 68 CFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAM 127
CFV GCD S+LLD+ + +EK + PN S RGFEV+D+IK LE ECP TVSCAD L +
Sbjct: 77 CFVQGCDGSLLLDTSGSIVTEKNSNPNSRSARGFEVVDEIKAALENECPNTVSCADALTL 136
Query: 128 VARDAVELRGGPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLV 187
ARD+ L GGP W V LGR+DS +S + N +P P++ +T+ F +GL++ DLV
Sbjct: 137 AARDSSVLTGGPSWTVPLGRRDSATASRAKPNKDLPEPDNLFDTIFLRFSNEGLNLTDLV 196
Query: 188 VLSGSHTIGRARCLSFRQRIYETKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKFAPL 247
LSGSHTIG +RC SFRQR+Y K Y IL+ CP +G D + L
Sbjct: 197 ALSGSHTIGFSRCTSFRQRLYNQSGSGSPDTTLEKSYAA---ILRQRCPRSGGDQNLSEL 253
Query: 248 DFQTPKRFDNQYFINIIEGKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKSM 307
D + RFDN YF N+IE GLL SD VL S + + R+ V YA +++ FF+ FA+SM
Sbjct: 254 DINSAGRFDNSYFKNLIENMGLLNSDQVLFSS--NEQSRELVKKYAEDQEEFFEQFAESM 311
Query: 308 IKMGNINVLTGSEGEIRRNCRFVN 331
IKMG I+ LTGS GEIR+ CR +N
Sbjct: 312 IKMGKISPLTGSSGEIRKKCRKIN 335
>At5g58390 peroxidase
Length = 316
Score = 278 bits (711), Expect = 3e-75
Identities = 157/329 (47%), Positives = 209/329 (62%), Gaps = 20/329 (6%)
Query: 6 LLLILITILSNIHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHF 65
+LL++I +L++ + L ++YKE CP +VR V AV ++PR+ ASLLRL F
Sbjct: 5 VLLMMIMMLAS----QSEAQLNRDFYKESCPSLFLVVRRVVKRAVAREPRMGASLLRLFF 60
Query: 66 HDCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADIL 125
HDCFV GCD S+LLD EK +GP+ NS+RGFEVIDKIK+ +EK CP VSCADIL
Sbjct: 61 HDCFVNGCDGSLLLDDTPSFLGEKTSGPSNNSVRGFEVIDKIKFKVEKMCPGIVSCADIL 120
Query: 126 AMVARDAVELRGGPRWEVWLGRKDSLESSFSGANL-FIPAPNSSLETLINNFKQQGLDIE 184
A+ ARD+V L GGP W V LGR+DS ++F+ AN IP P ++L LIN FK QGL
Sbjct: 121 AITARDSVLLLGGPGWSVKLGRRDSTTANFAAANSGVIPPPITTLSNLINRFKAQGLSTR 180
Query: 185 DLVVLSGSHTIGRARCLSFRQRIYETKQEYHHAYDRYKRYTTFRRILQSICPVT--GRDD 242
D+V LSG+HTIGRA+C++FR RIY T+F + CP T D+
Sbjct: 181 DMVALSGAHTIGRAQCVTFRNRIYNAS----------NIDTSFAISKRRNCPATSGSGDN 230
Query: 243 KFAPLDFQTPKRFDNQYFINIIEGKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDS 302
K A LD ++P RFD+ ++ ++ KGLL SD VL + +G V Y+ N F+
Sbjct: 231 KKANLDVRSPDRFDHGFYKQLLSKKGLLTSDQVLFN---NGPTDSLVIAYSHNLNAFYRD 287
Query: 303 FAKSMIKMGNINVLTGSEGEIRRNCRFVN 331
FA++MIKMG+I+ LTGS G+IR+NCR N
Sbjct: 288 FARAMIKMGDISPLTGSNGQIRQNCRRPN 316
>At5g06720 peroxidase (emb|CAA68212.1)
Length = 335
Score = 276 bits (707), Expect = 9e-75
Identities = 153/325 (47%), Positives = 192/325 (59%), Gaps = 5/325 (1%)
Query: 7 LLILITILSNIHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFH 66
++ LI I+S+I ++L +Y CP A IVR + A+ D R+ ASL+RLHFH
Sbjct: 14 IISLIVIVSSIFGTSSAQLNA-TFYSGTCPNASAIVRSTIQQALQSDTRIGASLIRLHFH 72
Query: 67 DCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILA 126
DCFV GCDAS+LLD + SEK AGPNVNS RGF V+D IK LE CP VSC+D+LA
Sbjct: 73 DCFVNGCDASILLDDTGSIQSEKNAGPNVNSARGFNVVDNIKTALENACPGVVSCSDVLA 132
Query: 127 MVARDAVELRGGPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDL 186
+ + +V L GGP W V LGR+DSL ++ +GAN IP+P SL + F GL+ DL
Sbjct: 133 LASEASVSLAGGPSWTVLLGRRDSLTANLAGANSSIPSPIESLSNITFKFSAVGLNTNDL 192
Query: 187 VVLSGSHTIGRARCLSFRQRIYETKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKFAP 246
V LSG+HT GRARC F R++ +T LQ +CP G
Sbjct: 193 VALSGAHTFGRARCGVFNNRLFNFSGT---GNPDPTLNSTLLSTLQQLCPQNGSASTITN 249
Query: 247 LDFQTPKRFDNQYFINIIEGKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKS 306
LD TP FDN YF N+ GLL SD L S I V +ASN+ LFF +FA+S
Sbjct: 250 LDLSTPDAFDNNYFANLQSNDGLLQSDQELFSTTGSSTI-AIVTSFASNQTLFFQAFAQS 308
Query: 307 MIKMGNINVLTGSEGEIRRNCRFVN 331
MI MGNI+ LTGS GEIR +C+ VN
Sbjct: 309 MINMGNISPLTGSNGEIRLDCKKVN 333
>At1g68850 peroxidase ATP23a
Length = 336
Score = 270 bits (689), Expect = 1e-72
Identities = 138/309 (44%), Positives = 194/309 (62%), Gaps = 6/309 (1%)
Query: 26 LVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDSVEGM 85
L +YYK CP D+++ + V +DPR AA ++RLHFHDCFV GCD SVLLD E +
Sbjct: 30 LTLDYYKSTCPTVFDVIKKEMECIVKEDPRNAAIIIRLHFHDCFVQGCDGSVLLDETETL 89
Query: 86 TSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVARDAVELRGGPRWEVWL 145
EK+A PN+NSL+G++++D+IK ++E ECP VSCAD+L + ARDA L GGP W+V +
Sbjct: 90 QGEKKASPNINSLKGYKIVDRIKNIIESECPGVVSCADLLTIGARDATILVGGPYWDVPV 149
Query: 146 GRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGRARCLSFRQ 205
GRKDS +S+ A +P P L ++I F QGL +ED+V L G+HTIG+A+C +FR
Sbjct: 150 GRKDSKTASYELATTNLPTPEEGLISIIAKFYSQGLSVEDMVALIGAHTIGKAQCRNFRS 209
Query: 206 RIYETKQEYHHAYDRYKRYTTFRRILQSICPVTG--RDDKFAPLDFQTPKRFDNQYFINI 263
RIY ++ T+ L+ ICP + D +D TP FDN + +
Sbjct: 210 RIY---GDFQVTSALNPVSETYLASLREICPASSGEGDSNVTAIDNVTPNLFDNSIYHTL 266
Query: 264 IEGKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKSMIKMGNI-NVLTGSEGE 322
+ G+GLL SD + + + R+ V YA + FF+ F+KSM+KMGNI N + ++GE
Sbjct: 267 LRGEGLLNSDQEMYTSLFGIQTRRIVSKYAEDPVAFFEQFSKSMVKMGNILNSESLADGE 326
Query: 323 IRRNCRFVN 331
+RRNCRFVN
Sbjct: 327 VRRNCRFVN 335
>At5g58400 peroxidase
Length = 325
Score = 263 bits (671), Expect = 1e-70
Identities = 149/307 (48%), Positives = 193/307 (62%), Gaps = 18/307 (5%)
Query: 29 EYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDSVEGMTSE 88
++Y + CP VR V V K+ R+AASLLRL FHDCFV GCDAS+LLD E
Sbjct: 33 DFYSDSCPSLLPTVRRVVQREVAKERRIAASLLRLFFHDCFVNGCDASILLDDTRSFLGE 92
Query: 89 KQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVARDAVELRGGPRWEVWLGRK 148
K AGPN NS+RG+EVID IK +E+ CP VSCADILA+ ARD+V L GG W V LGR+
Sbjct: 93 KTAGPNNNSVRGYEVIDAIKSRVERLCPGVVSCADILAITARDSVLLMGGRGWSVKLGRR 152
Query: 149 DSLESSFSGANL-FIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGRARCLSFRQRI 207
DS+ +SFS AN +P P S+L+ LIN F+ GL D+V LSG+HTIG+ARC++FR RI
Sbjct: 153 DSITASFSTANSGVLPPPTSTLDNLINLFRANGLSPRDMVALSGAHTIGQARCVTFRSRI 212
Query: 208 Y-ETKQEYHHAYDRYKRYTTFRRILQSICP-VTGR-DDKFAPLDFQTPKRFDNQYFINII 264
Y T + A R + CP TG D+ A LD +TP++FD YF+ ++
Sbjct: 213 YNSTNIDLSFALSRRRS-----------CPAATGSGDNNAAILDLRTPEKFDGSYFMQLV 261
Query: 265 EGKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKSMIKMGNINVLTGSEGEIR 324
+GLL SD VL + G V Y+ + + F+ F +MIKMG+I+ LTGS G+IR
Sbjct: 262 NHRGLLTSDQVLFN---GGSTDSIVVSYSRSVQAFYRDFVAAMIKMGDISPLTGSNGQIR 318
Query: 325 RNCRFVN 331
R+CR N
Sbjct: 319 RSCRRPN 325
>At5g19890 peroxidase ATP N
Length = 328
Score = 263 bits (671), Expect = 1e-70
Identities = 146/304 (48%), Positives = 190/304 (62%), Gaps = 10/304 (3%)
Query: 29 EYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDSVEGMTSE 88
+ Y + CP IVR VA+A+ + R+AASL+RLHFHDCFV GCDAS+LLD G SE
Sbjct: 33 DIYAKSCPNLVQIVRKQVAIALKAEIRMAASLIRLHFHDCFVNGCDASLLLD---GADSE 89
Query: 89 KQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVARDAVELRGGPRWEVWLGRK 148
K A PN+NS RGFEVID IK +E CP VSCADIL + ARD+V L GGP W V LGRK
Sbjct: 90 KLAIPNINSARGFEVIDTIKAAVENACPGVVSCADILTLAARDSVVLSGGPGWRVALGRK 149
Query: 149 DSLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGRARCLSFRQRIY 208
D L ++ + AN +P+P L+ +I F L+I D+V LSG+HT G+A+C F R++
Sbjct: 150 DGLVANQNSAN-NLPSPFEPLDAIIAKFVAVNLNITDVVALSGAHTFGQAKCAVFSNRLF 208
Query: 209 ETKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKFAPLDFQTPKRFDNQYFINIIEGKG 268
T+ LQ++CP+ G + APLD T FDN YF N++EGKG
Sbjct: 209 NFT---GLGNPDATLETSLLSNLQTVCPLGGNSNITAPLDRSTTDTFDNNYFKNLLEGKG 265
Query: 269 LLGSDNVLISQDLD-GRIRKQVWGYASNEKLFFDSFAKSMIKMGNINVLTGSEGEIRRNC 327
LL SD +L S DL +K V Y+ ++ LFF F +MI+MGNI+ G+ GE+R NC
Sbjct: 266 LLSSDQILFSSDLAVNTTKKLVEAYSRSQSLFFRDFTCAMIRMGNIS--NGASGEVRTNC 323
Query: 328 RFVN 331
R +N
Sbjct: 324 RVIN 327
>At5g06730 peroxidase
Length = 358
Score = 263 bits (671), Expect = 1e-70
Identities = 142/325 (43%), Positives = 189/325 (57%), Gaps = 5/325 (1%)
Query: 7 LLILITILSNIHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFH 66
++ LI I+S++ ++L +Y CP A IVR + A+ D R+ SL+RLHFH
Sbjct: 15 IISLIVIVSSLFGTSSAQLNA-TFYSGTCPNASAIVRSTIQQALQSDARIGGSLIRLHFH 73
Query: 67 DCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILA 126
DCFV GCD S+LLD + SEK A N NS RGF V+D IK LE CP VSC+DILA
Sbjct: 74 DCFVNGCDGSLLLDDTSSIQSEKNAPANANSTRGFNVVDSIKTALENACPGIVSCSDILA 133
Query: 127 MVARDAVELRGGPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDL 186
+ + +V L GGP W V LGR+D L ++ SGAN +P+P L + + F GL D+
Sbjct: 134 LASEASVSLAGGPSWTVLLGRRDGLTANLSGANSSLPSPFEGLNNITSKFVAVGLKTTDV 193
Query: 187 VVLSGSHTIGRARCLSFRQRIYETKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKFAP 246
V LSG+HT GR +C++F R++ +T LQ +CP G +
Sbjct: 194 VSLSGAHTFGRGQCVTFNNRLFNFNGT---GNPDPTLNSTLLSSLQQLCPQNGSNTGITN 250
Query: 247 LDFQTPKRFDNQYFINIIEGKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKS 306
LD TP FDN YF N+ GLL SD L S + V +ASN+ LFF++F +S
Sbjct: 251 LDLSTPDAFDNNYFTNLQSNNGLLQSDQELFSNTGSATV-PIVNSFASNQTLFFEAFVQS 309
Query: 307 MIKMGNINVLTGSEGEIRRNCRFVN 331
MIKMGNI+ LTGS GEIR++C+ VN
Sbjct: 310 MIKMGNISPLTGSSGEIRQDCKVVN 334
>At4g08780 peroxidase C2 precursor like protein
Length = 346
Score = 260 bits (664), Expect = 8e-70
Identities = 144/327 (44%), Positives = 192/327 (58%), Gaps = 11/327 (3%)
Query: 6 LLLILITILSNIHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHF 65
LLL+L LS+ L +Y + CP DIV + + A+ DPR+AAS+LRLHF
Sbjct: 11 LLLLLQVSLSHAQ-------LSPSFYDKTCPQVFDIVTNTIVNALRSDPRIAASILRLHF 63
Query: 66 HDCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADIL 125
HDCFV GCDAS+LLD+ +EK A N NS RGF+VIDK+K +EK CP TVSCAD+L
Sbjct: 64 HDCFVNGCDASILLDNTTSFRTEKDAFGNANSARGFDVIDKMKAAIEKACPRTVSCADML 123
Query: 126 AMVARDAVELRGGPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLD-IE 184
A+ A++++ L GGP W V GR+DSL AN +P P+S+L+ L + FK GLD
Sbjct: 124 AIAAKESIVLAGGPSWMVPNGRRDSLRGFMDLANDNLPGPSSTLKQLKDRFKNVGLDRSS 183
Query: 185 DLVVLSGSHTIGRARCLSFRQRIYETKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKF 244
DLV LSG HT G+++C R+Y + K Y L+ CP G
Sbjct: 184 DLVALSGGHTFGKSQCQFIMDRLYNFGETGLPDPTLDKSYLA---TLRKQCPRNGNQSVL 240
Query: 245 APLDFQTPKRFDNQYFINIIEGKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFA 304
D +TP FDN+Y++N+ E KGL+ SD L S V YA + FFD+F
Sbjct: 241 VDFDLRTPTLFDNKYYVNLKENKGLIQSDQELFSSPDAADTLPLVRAYADGQGTFFDAFV 300
Query: 305 KSMIKMGNINVLTGSEGEIRRNCRFVN 331
K++I+M +++ LTG +GEIR NCR VN
Sbjct: 301 KAIIRMSSLSPLTGKQGEIRLNCRVVN 327
>At3g32980 peroxidase, putative
Length = 352
Score = 259 bits (661), Expect = 2e-69
Identities = 140/316 (44%), Positives = 187/316 (58%), Gaps = 4/316 (1%)
Query: 17 IHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDAS 76
+H+ S L +Y CP IVR + + DPR+AAS+LRLHFHDCFV GCDAS
Sbjct: 22 LHSSISSAQLTPTFYDNTCPSVFTIVRDTIVNELRSDPRIAASILRLHFHDCFVNGCDAS 81
Query: 77 VLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVARDAVELR 136
+LLD+ +EK A PN NS RGF VID++K +E CP TVSCADIL + A+ AV L
Sbjct: 82 ILLDNTTSFRTEKDAAPNANSARGFPVIDRMKAAVETACPRTVSCADILTIAAQQAVNLA 141
Query: 137 GGPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLD-IEDLVVLSGSHTI 195
GGP W V LGR+DSL++ F+ AN +PAP +L L +F+ GLD DLV LSG HT
Sbjct: 142 GGPSWRVPLGRRDSLQAFFALANTNLPAPFFTLPQLKASFQNVGLDRPSDLVALSGGHTF 201
Query: 196 GRARCLSFRQRIYETKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKFAPLDFQTPKRF 255
G+ +C R+Y + TT+ + L+ CP G D +TP F
Sbjct: 202 GKNQCQFIMDRLYNFS---NTGLPDPTLNTTYLQTLRGQCPRNGNQTVLVDFDLRTPTVF 258
Query: 256 DNQYFINIIEGKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKSMIKMGNINV 315
DN+Y++N+ E KGL+ +D L S V YA + FF++F ++M +MGNI
Sbjct: 259 DNKYYVNLKELKGLIQTDQELFSSPNATDTIPLVREYADGTQKFFNAFVEAMNRMGNITP 318
Query: 316 LTGSEGEIRRNCRFVN 331
LTG++G+IR+NCR VN
Sbjct: 319 LTGTQGQIRQNCRVVN 334
>At1g49570 peroxidase like protein
Length = 350
Score = 258 bits (658), Expect = 4e-69
Identities = 143/305 (46%), Positives = 184/305 (59%), Gaps = 5/305 (1%)
Query: 26 LVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDSVEGM 85
L + +Y CP + IV+ V A D R+AASLLRLHFHDCFV GCD S+LL+ E
Sbjct: 48 LNYRFYDRSCPRLQTIVKSGVWRAFKDDSRIAASLLRLHFHDCFVNGCDGSILLNDSEDF 107
Query: 86 TSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVARDAVELRGGPRWEVWL 145
EK A PN NS+RGFEVI+ IK +E CPLTVSCADI+A+ AR+AV L GGP W V L
Sbjct: 108 KGEKNAQPNRNSVRGFEVIEDIKSDIESSCPLTVSCADIVALAAREAVVLTGGPFWPVPL 167
Query: 146 GRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGRARCLSFRQ 205
GR+DSL +S AN +P+P +LE + F GLD++D+VVLSG+HTIG A+C +
Sbjct: 168 GRRDSLTASEQAANTNLPSPFEALENITAKFVTLGLDLKDVVVLSGAHTIGFAQCFVIKH 227
Query: 206 RIYETKQEYHHAYDRYKRYTTFRRILQSICP-VTGRDDKFAPLDFQTPKRFDNQYFINII 264
R++ K + L+ CP V D K A LD + +FDN Y++N++
Sbjct: 228 RLFNFKGS-GQPDPNLAASSALLSKLKDTCPNVDSSDSKLAALDAASSVKFDNAYYVNLM 286
Query: 265 EGKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKSMIKMGNINVLTGSEGEIR 324
GLL SD L++ D V Y+ N LF FA SM+KMGNI V+TGS+G IR
Sbjct: 287 NNIGLLDSDQTLMT---DPTAAALVKSYSENPYLFSRDFAVSMVKMGNIGVMTGSDGVIR 343
Query: 325 RNCRF 329
C F
Sbjct: 344 GKCGF 348
>At4g08770 peroxidase C2 precursor like protein
Length = 346
Score = 256 bits (654), Expect = 1e-68
Identities = 145/327 (44%), Positives = 188/327 (57%), Gaps = 11/327 (3%)
Query: 6 LLLILITILSNIHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHF 65
LLL++ LS+ L +Y + CP DI + A+ DPR+AAS+LRLHF
Sbjct: 11 LLLLIQVSLSHAQ-------LSPSFYDKTCPQVFDIATTTIVNALRSDPRIAASILRLHF 63
Query: 66 HDCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADIL 125
HDCFV GCDAS+LLD+ +EK A N NS RGF+VIDK+K +EK CP TVSCAD+L
Sbjct: 64 HDCFVNGCDASILLDNTTSFRTEKDAFGNANSARGFDVIDKMKAAVEKACPKTVSCADLL 123
Query: 126 AMVARDAVELRGGPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLD-IE 184
A+ A+++V L GGP W V GR+DSL AN +PAP +L L + FK GLD
Sbjct: 124 AIAAQESVVLAGGPSWRVPNGRRDSLRGFMDLANDNLPAPFFTLNQLKDRFKNVGLDRAS 183
Query: 185 DLVVLSGSHTIGRARCLSFRQRIYETKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKF 244
DLV LSG HT G+ +C R+Y K Y + L+ CP G
Sbjct: 184 DLVALSGGHTFGKNQCQFIMDRLYNFSNTGLPDPTLDKSYLS---TLRKQCPRNGNQSVL 240
Query: 245 APLDFQTPKRFDNQYFINIIEGKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFA 304
D +TP FDN+Y++N+ E KGL+ SD L S V YA + FFD+FA
Sbjct: 241 VDFDLRTPTLFDNKYYVNLKENKGLIQSDQELFSSPDASDTLPLVREYADGQGKFFDAFA 300
Query: 305 KSMIKMGNINVLTGSEGEIRRNCRFVN 331
K+MI+M +++ LTG +GEIR NCR VN
Sbjct: 301 KAMIRMSSLSPLTGKQGEIRLNCRVVN 327
>At2g22420 putative peroxidase
Length = 329
Score = 256 bits (653), Expect = 2e-68
Identities = 143/331 (43%), Positives = 205/331 (61%), Gaps = 12/331 (3%)
Query: 1 MEPLRLLLILITILSNIHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASL 60
M L L++ +T+L+ + T E L +Y E CP AE IVR + A++K+ R AS+
Sbjct: 1 MSLLPHLILYLTLLTVVVT---GETLRPRFYSETCPEAESIVRREMKKAMIKEARSVASV 57
Query: 61 LRLHFHDCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVS 120
+R FHDCFV GCDAS+LLD M EK + N++SLR FEV+D IK LEK CP TVS
Sbjct: 58 MRFQFHDCFVNGCDASLLLDDTPNMLGEKLSLSNIDSLRSFEVVDDIKEALEKACPATVS 117
Query: 121 CADILAMVARDAVELRGGPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQQG 180
CADI+ M ARDAV L GGP WEV LGRKDSL +S ++ +P+P ++ LI+ F++
Sbjct: 118 CADIVIMAARDAVALTGGPDWEVKLGRKDSLTASQQDSDDIMPSPRANATFLIDLFERFN 177
Query: 181 LDIEDLVVLSGSHTIGRARCLSFRQRIYETKQEYHHAYDRYKRYTTFRRILQSICPVTGR 240
L ++D+V LSGSH+IG+ RC S R+Y + ++R+ L +CP+ G
Sbjct: 178 LSVKDMVALSGSHSIGQGRCFSIMFRLY---NQSGSGKPDPALEPSYRKKLDKLCPLGGD 234
Query: 241 DDKFAPLDFQTPKRFDNQYFINIIEGKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFF 300
++ LD TP+ FDNQYF +++ G+G L SD L + + R+ V ++ ++ FF
Sbjct: 235 ENVTGDLD-ATPQVFDNQYFKDLVSGRGFLNSDQTLYTNLV---TREYVKMFSEDQDEFF 290
Query: 301 DSFAKSMIKMGNINVLTGSEGEIRRNCRFVN 331
+FA+ M+K+G++ +G GEIR NCR VN
Sbjct: 291 RAFAEGMVKLGDLQ--SGRPGEIRFNCRVVN 319
>At1g14550 peroxidase, putative
Length = 321
Score = 256 bits (653), Expect = 2e-68
Identities = 144/331 (43%), Positives = 199/331 (59%), Gaps = 18/331 (5%)
Query: 4 LRLLLILITILSNIHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRL 63
LR +L++++I+ + L +Y + C A +R +V A+ ++ R+AASL+R+
Sbjct: 6 LRFVLMMVSIILTSSICQAQ--LSPTFYDQSCRNALSKIRSSVRTAIARERRMAASLIRM 63
Query: 64 HFHDCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCAD 123
HFHDCFV GCDAS+LL+ + SE+ A PN S+RGFEVIDK K +EK CP VSCAD
Sbjct: 64 HFHDCFVHGCDASILLEGTSTIESERDALPNFKSVRGFEVIDKAKSEVEKVCPGIVSCAD 123
Query: 124 ILAMVARDAVELRGGPRWEVWLGRKDSLESSFSGANL-FIPAPNSSLETLINNFKQQGLD 182
I+A+ ARDA E GGP+W V +GR+DS + + AN +P +L+ L F ++GL+
Sbjct: 124 IIAVAARDASEYVGGPKWAVKVGRRDSTAAFKALANSGELPGFKDTLDQLSGLFSKKGLN 183
Query: 183 IEDLVVLSGSHTIGRARCLSFRQRIYETKQEYHHAYDRYKRYTTFRRILQSICPVTGRDD 242
DLV LSG+HTIG+++C FR R+YE + + ++ RR CP G D
Sbjct: 184 TRDLVALSGAHTIGQSQCFLFRDRLYENSSDIDAGFASTRK----RR-----CPTVGGDG 234
Query: 243 KFAPLDFQTPKRFDNQYFINIIEGKGLLGSDNVLISQ--DLDGRIRKQVWGYASNEKLFF 300
A LD TP FDN Y+ N+++ KGLL +D VL DG + + Y+ N F
Sbjct: 235 NLAALDLVTPNSFDNNYYKNLMQKKGLLVTDQVLFGSGASTDGIVSE----YSKNRSKFA 290
Query: 301 DSFAKSMIKMGNINVLTGSEGEIRRNCRFVN 331
FA +MIKMGNI LTGS GEIR+ C FVN
Sbjct: 291 ADFATAMIKMGNIEPLTGSNGEIRKICSFVN 321
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.140 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,310,610
Number of Sequences: 26719
Number of extensions: 309133
Number of successful extensions: 1004
Number of sequences better than 10.0: 86
Number of HSP's better than 10.0 without gapping: 84
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 672
Number of HSP's gapped (non-prelim): 96
length of query: 332
length of database: 11,318,596
effective HSP length: 100
effective length of query: 232
effective length of database: 8,646,696
effective search space: 2006033472
effective search space used: 2006033472
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC144516.5


