

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144515.13 + phase: 0
(381 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
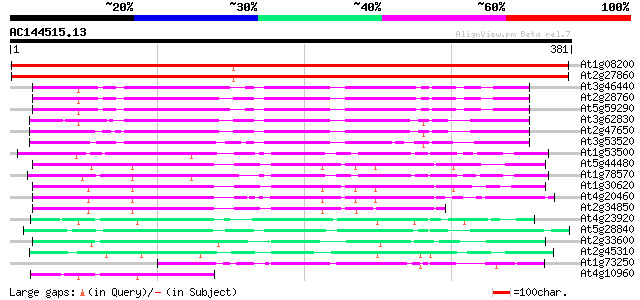
Score E
Sequences producing significant alignments: (bits) Value
At1g08200 unknown protein 603 e-173
At2g27860 putative dTDP-glucose 4-6-dehydratase 593 e-170
At3g46440 dTDP-glucose 4-6-dehydratases-like protein 110 2e-24
At2g28760 putative nucleotide-sugar dehydratase 107 1e-23
At5g59290 dTDP-glucose 4-6-dehydratase - like protein 106 2e-23
At3g62830 dTDP-glucose 4-6-dehydratase homolog D18 99 5e-21
At2g47650 putative nucleotide-sugar dehydratase 97 1e-20
At3g53520 UDP-glucuronic acid decarboxylase (UXS1) 96 2e-20
At1g53500 dTDP-D-glucose 4,6-dehydratase, putative 68 1e-11
At5g44480 putative protein 60 2e-09
At1g78570 Similar to dTDP-D-glucose 4,6-dehydratase 60 3e-09
At1g30620 UDP-galactose 4-epimerase-like protein 57 2e-08
At4g20460 UDP-glucose 4-epimerase - like protein 54 1e-07
At2g34850 putative UDP-galactose-4-epimerase 52 4e-07
At4g23920 UDPglucose 4-epimerase - like protein 50 3e-06
At5g28840 GDP-mannose 3",5"-epimerase 47 1e-05
At2g33600 putative cinnamoyl-CoA reductase 47 2e-05
At2g45310 putative nucleotide sugar epimerase 45 7e-05
At1g73250 GDP-4-keto-6-deoxy-D-mannose-3,5-epimerase-4-reductase... 45 7e-05
At4g10960 UDP-galactose 4-epimerase - like protein 45 9e-05
>At1g08200 unknown protein
Length = 389
Score = 603 bits (1554), Expect = e-173
Identities = 283/381 (74%), Positives = 339/381 (88%), Gaps = 3/381 (0%)
Query: 2 DRVNLDGKPIVPISICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSH-P 60
DR++LDGKPI P++IC+IG GGFIGSHL EKLM+ET HK + +DV ++K+ HLL+
Sbjct: 6 DRLDLDGKPIKPMTICMIGAGGFIGSHLCEKLMTETPHKVLALDVYNDKIKHLLEPDTVQ 65
Query: 61 WANRIEFHQMNIKNDSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIK 120
WA RI+FH++NIK+DSRLE L+K +DLTINLAAICTPADYNTRPLDTIYSNFIDA+PV+K
Sbjct: 66 WAGRIQFHRINIKHDSRLEGLIKMADLTINLAAICTPADYNTRPLDTIYSNFIDALPVVK 125
Query: 121 FCTENNKRLIHFSTCEVFGKTIGSFLPEEY--RKEPQYYKLKEDVSPCIFGPVHKQRWSY 178
+C+ENNKRLIHFSTCEV+GKTIGSFLP+++ R++P++Y LKED+SPCIFG + KQRWSY
Sbjct: 126 YCSENNKRLIHFSTCEVYGKTIGSFLPKDHPLRQDPEFYVLKEDISPCIFGSIEKQRWSY 185
Query: 179 ACAKQMTDRLIYAEHAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLL 238
ACAKQ+ +RL+YAE AENGL+FTIVRP+NWIGPRMDFIPG+DGPS+GVPRVLACFSNNLL
Sbjct: 186 ACAKQLIERLVYAEGAENGLEFTIVRPFNWIGPRMDFIPGIDGPSEGVPRVLACFSNNLL 245
Query: 239 RGEPLKLVDGGHSQRTFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAELM 298
R EPLKLVDGG SQRTF+YIKDAIEAV+LMI+NP+RANGHIFNVGNP+NEV+V+QLAE+M
Sbjct: 246 RREPLKLVDGGESQRTFIYIKDAIEAVLLMIENPERANGHIFNVGNPNNEVTVRQLAEMM 305
Query: 299 IKVYAKVAGVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLLD 358
+VYAKV+G T+DVSS+ FYG+GYDDSD+RIPDMTII +QLGW PKTSL DLL+
Sbjct: 306 TEVYAKVSGETAIESPTIDVSSKEFYGEGYDDSDKRIPDMTIINRQLGWNPKTSLWDLLE 365
Query: 359 STLQYQHQTYSHAIKKELSKP 379
STL YQH TY+ AIKK SKP
Sbjct: 366 STLTYQHTTYAEAIKKATSKP 386
>At2g27860 putative dTDP-glucose 4-6-dehydratase
Length = 389
Score = 593 bits (1530), Expect = e-170
Identities = 279/381 (73%), Positives = 336/381 (87%), Gaps = 3/381 (0%)
Query: 2 DRVNLDGKPIVPISICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSH-P 60
+RV+LDGKPI P++IC+IG GGFIGSHL EKL++ET HK + +DV ++K+ HLL+
Sbjct: 6 NRVDLDGKPIQPLTICMIGAGGFIGSHLCEKLLTETPHKVLALDVYNDKIKHLLEPDTVE 65
Query: 61 WANRIEFHQMNIKNDSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIK 120
W+ RI+FH++NIK+DSRLE LVK +DL INLAAICTPADYNTRPLDTIYSNFIDA+PV+K
Sbjct: 66 WSGRIQFHRINIKHDSRLEGLVKMADLIINLAAICTPADYNTRPLDTIYSNFIDALPVVK 125
Query: 121 FCTENNKRLIHFSTCEVFGKTIGSFLPEEY--RKEPQYYKLKEDVSPCIFGPVHKQRWSY 178
+C+ENNKRLIHFSTCEV+GKTIGSFLP+++ R +P +Y LKED+SPCIFG + KQRWSY
Sbjct: 126 YCSENNKRLIHFSTCEVYGKTIGSFLPKDHPLRDDPAFYVLKEDISPCIFGSIEKQRWSY 185
Query: 179 ACAKQMTDRLIYAEHAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLL 238
ACAKQ+ +RL+YAE AENGL+FTIVRP+NWIGPRMDFIPG+DGPS+GVPRVLACFSNNLL
Sbjct: 186 ACAKQLIERLVYAEGAENGLEFTIVRPFNWIGPRMDFIPGIDGPSEGVPRVLACFSNNLL 245
Query: 239 RGEPLKLVDGGHSQRTFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAELM 298
R EPLKLVDGG SQRTF+YI DAIEAV+LMI+NP+RANGHIFNVGNP+NEV+V+QLAE+M
Sbjct: 246 RREPLKLVDGGESQRTFVYINDAIEAVLLMIENPERANGHIFNVGNPNNEVTVRQLAEMM 305
Query: 299 IKVYAKVAGVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLLD 358
+VYAKV+G T+DVSS+ FYG+GYDDSD+RIPDMTII +QLGW PKTSL DLL+
Sbjct: 306 TEVYAKVSGEGAIESPTVDVSSKEFYGEGYDDSDKRIPDMTIINRQLGWNPKTSLWDLLE 365
Query: 359 STLQYQHQTYSHAIKKELSKP 379
STL YQH+TY+ A+KK SKP
Sbjct: 366 STLTYQHRTYAEAVKKATSKP 386
>At3g46440 dTDP-glucose 4-6-dehydratases-like protein
Length = 341
Score = 110 bits (274), Expect = 2e-24
Identities = 95/340 (27%), Positives = 154/340 (44%), Gaps = 51/340 (15%)
Query: 16 ICLIGGGGFIGSHLTEKLMSETSHKAIVID--VSSEKVNHLLDKSHPWANRIEFHQMNIK 73
I + GG GFIGSHL +KLM ++ IV D + K N HP R E + ++
Sbjct: 31 ILISGGAGFIGSHLVDKLMENEKNEVIVADNYFTGSKDNLKKWIGHP---RFELIRHDV- 86
Query: 74 NDSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENNKRLIHFS 133
E L+ D +LA +P Y P+ TI +N I + ++ R++ S
Sbjct: 87 ----TEPLLIEVDQIYHLACPASPIFYKYNPVKTIKTNVIGTLNMLGLAKRVGARILLTS 142
Query: 134 TCEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDRLIYAEH 193
T EV+G + PE Y +V+P R Y K++ + L++ H
Sbjct: 143 TSEVYGDPLIHPQPESY---------WGNVNPI------GVRSCYDEGKRVAETLMFDYH 187
Query: 194 AENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRGEPLKLVDGGHSQR 253
++G++ I R +N GPRM+ G RV++ F LRGE L + G R
Sbjct: 188 RQHGIEIRIARIFNTYGPRMNIDDG---------RVVSNFIAQALRGEALTVQKPGTQTR 238
Query: 254 TFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAELMIKVYAKVAGVPESSL 313
+F Y+ D ++ +M +++ D N+GNP E ++ +LAE + ++ P +
Sbjct: 239 SFCYVSDMVDGLMRLMEGDDTGP---INIGNP-GEFTMVELAETVKELIN-----PSIEI 289
Query: 314 STLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSL 353
++ + DD +R PD+T + LGW+PK L
Sbjct: 290 KMVENTP--------DDPRQRKPDITKAKEVLGWEPKVKL 321
>At2g28760 putative nucleotide-sugar dehydratase
Length = 343
Score = 107 bits (266), Expect = 1e-23
Identities = 93/340 (27%), Positives = 152/340 (44%), Gaps = 51/340 (15%)
Query: 16 ICLIGGGGFIGSHLTEKLMSETSHKAIVID--VSSEKVNHLLDKSHPWANRIEFHQMNIK 73
I + GG GFIGSHL +KLM ++ IV D + K N HP R E + ++
Sbjct: 33 ILVTGGAGFIGSHLVDKLMQNEKNEVIVADNYFTGSKDNLKKWIGHP---RFELIRHDV- 88
Query: 74 NDSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENNKRLIHFS 133
E L D +LA +P Y P+ TI +N I + ++ R++ S
Sbjct: 89 ----TEPLFVEVDQIYHLACPASPIFYKYNPVKTIKTNVIGTLNMLGLAKRVGARILLTS 144
Query: 134 TCEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDRLIYAEH 193
T EV+G + PQ +V+P R Y K++ + L++ H
Sbjct: 145 TSEVYGDPL---------VHPQTESYWGNVNPI------GVRSCYDEGKRVAETLMFDYH 189
Query: 194 AENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRGEPLKLVDGGHSQR 253
++G++ I R +N GPRM+ G RV++ F LRGE L + G R
Sbjct: 190 RQHGIEIRIARIFNTYGPRMNIDDG---------RVVSNFIAQALRGEALTVQKPGTQTR 240
Query: 254 TFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAELMIKVYAKVAGVPESSL 313
+F Y+ D +E +M +++ N+GNP E ++ +LAE + ++ P+ +
Sbjct: 241 SFCYVSDMVEGLMRLMEGDQTGP---INIGNP-GEFTMVELAETVKELIK-----PDVEI 291
Query: 314 STLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSL 353
++ + DD +R PD++ + LGW+PK L
Sbjct: 292 KMVENTP--------DDPRQRKPDISKAKEVLGWEPKVKL 323
>At5g59290 dTDP-glucose 4-6-dehydratase - like protein
Length = 342
Score = 106 bits (265), Expect = 2e-23
Identities = 92/340 (27%), Positives = 154/340 (45%), Gaps = 51/340 (15%)
Query: 16 ICLIGGGGFIGSHLTEKLMSETSHKAIVID--VSSEKVNHLLDKSHPWANRIEFHQMNIK 73
I + GG GFIGSHL +KLM ++ +V D + K N HP R E + ++
Sbjct: 32 ILISGGAGFIGSHLVDKLMENEKNEVVVADNYFTGSKENLKKWIGHP---RFELIRHDV- 87
Query: 74 NDSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENNKRLIHFS 133
E L+ D +LA +P Y P+ TI +N I + ++ R++ S
Sbjct: 88 ----TEPLLIEVDRIYHLACPASPIFYKYNPVKTIKTNVIGTLNMLGLAKRVGARILLTS 143
Query: 134 TCEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDRLIYAEH 193
T EV+G + PE Y +V+P R Y K++ + L++ H
Sbjct: 144 TSEVYGDPLIHPQPESY---------WGNVNPI------GVRSCYDEGKRVAETLMFDYH 188
Query: 194 AENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRGEPLKLVDGGHSQR 253
++G++ I R +N GPRM+ G RV++ F LRGE L + G R
Sbjct: 189 RQHGIEIRIARIFNTYGPRMNIDDG---------RVVSNFIAQALRGEALTVQKPGTQTR 239
Query: 254 TFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAELMIKVYAKVAGVPESSL 313
+F Y+ D ++ ++ +++ D N+GNP E ++ +LAE + ++ P +
Sbjct: 240 SFCYVSDMVDGLIRLMEGNDTGP---INIGNP-GEFTMVELAETVKELIN-----PSIEI 290
Query: 314 STLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSL 353
++ + DD +R PD++ + LGW+PK L
Sbjct: 291 KMVENTP--------DDPRQRKPDISKAKEVLGWEPKVKL 322
>At3g62830 dTDP-glucose 4-6-dehydratase homolog D18
Length = 445
Score = 98.6 bits (244), Expect = 5e-21
Identities = 91/345 (26%), Positives = 152/345 (43%), Gaps = 58/345 (16%)
Query: 14 ISICLIGGGGFIGSHLTEKLMSETSHKAIVID--VSSEKVNHLLDKSHPWANRIEFHQMN 71
+ + + GG GF+GSHL ++LM+ IV+D + K N + S+P I H +
Sbjct: 119 LRVVVTGGAGFVGSHLVDRLMAR-GDTVIVVDNFFTGRKENVMHHFSNPNFEMIR-HDV- 175
Query: 72 IKNDSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENNKRLIH 131
+E ++ D +LA +P Y P+ TI +N + + ++ R +
Sbjct: 176 ------VEPILLEVDQIYHLACPASPVHYKFNPVKTIKTNVVGTLNMLGLAKRVGARFLL 229
Query: 132 FSTCEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDRLIYA 191
ST EV+G + + PQ +V+P R Y K+ + L
Sbjct: 230 TSTSEVYGDPL---------QHPQVETYWGNVNPI------GVRSCYDEGKRTAETLTMD 274
Query: 192 EHAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRGEPLKLVDGGHS 251
H ++ I R +N GPRM G RV++ F LR EPL + G
Sbjct: 275 YHRGANVEVRIARIFNTYGPRMCIDDG---------RVVSNFVAQALRKEPLTVYGDGKQ 325
Query: 252 QRTFLYIKDAIEAVMLMIDNPDRANGHI--FNVGNPDNEVSVKQLAELMIKVYAKVAGVP 309
R+F ++ D +E +M +++ H+ FN+GNP E ++ +LA+++
Sbjct: 326 TRSFQFVSDLVEGLMRLMEGE-----HVGPFNLGNP-GEFTMLELAKVV----------- 368
Query: 310 ESSLSTLDVSSEV-FYGKGYDDSDRRIPDMTIITKQLGWKPKTSL 353
T+D ++ + F DD +R PD+T + LGW+PK SL
Sbjct: 369 ---QETIDPNANIEFRPNTEDDPHKRKPDITKAKELLGWEPKVSL 410
>At2g47650 putative nucleotide-sugar dehydratase
Length = 443
Score = 97.1 bits (240), Expect = 1e-20
Identities = 86/343 (25%), Positives = 154/343 (44%), Gaps = 54/343 (15%)
Query: 14 ISICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSHPWANRIEFHQMNIK 73
+ + + GG GF+GSHL ++LM+ + +V + + + +++ N F I+
Sbjct: 121 LRVVVTGGAGFVGSHLVDRLMARGDNVIVVDNFFTGRKENVMHHF----NNPNFEM--IR 174
Query: 74 NDSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENNKRLIHFS 133
+D +E ++ D +LA +P Y P+ TI +N + + ++ R + S
Sbjct: 175 HDV-VEPILLEVDQIYHLACPASPVHYKFNPVKTIKTNVVGTLNMLGLAKRVGARFLLTS 233
Query: 134 TCEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDRLIYAEH 193
T EV+G + + PQ +V+P R Y K+ + L H
Sbjct: 234 TSEVYGDPL---------QHPQVETYWGNVNPI------GVRSCYDEGKRTAETLTMDYH 278
Query: 194 AENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRGEPLKLVDGGHSQR 253
++ I R +N GPRM G RV++ F LR EPL + G R
Sbjct: 279 RGANVEVRIARIFNTYGPRMCIDDG---------RVVSNFVAQALRKEPLTVYGDGKQTR 329
Query: 254 TFLYIKDAIEAVMLMIDNPDRANGHI--FNVGNPDNEVSVKQLAELMIKVYAKVAGVPES 311
+F ++ D +E +M +++ H+ FN+GNP E ++ +LA+++
Sbjct: 330 SFQFVSDLVEGLMRLMEGE-----HVGPFNLGNP-GEFTMLELAKVV------------- 370
Query: 312 SLSTLDVSSEV-FYGKGYDDSDRRIPDMTIITKQLGWKPKTSL 353
T+D ++++ F DD +R PD+T + LGW+PK +L
Sbjct: 371 -QETIDPNAKIEFRPNTEDDPHKRKPDITKAKELLGWEPKVAL 412
>At3g53520 UDP-glucuronic acid decarboxylase (UXS1)
Length = 433
Score = 96.3 bits (238), Expect = 2e-20
Identities = 91/344 (26%), Positives = 157/344 (45%), Gaps = 58/344 (16%)
Query: 14 ISICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSHPWAN-RIEFHQMNI 72
+ I + GG GF+GSHL +KL+ ++ + + + +L+ H ++N R E + ++
Sbjct: 120 LRIVVTGGAGFVGSHLVDKLIGRGDEVIVIDNFFTGRKENLV---HLFSNPRFELIRHDV 176
Query: 73 KNDSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENNKRLIHF 132
+E ++ D +LA +P Y P+ TI +N + + ++ R +
Sbjct: 177 -----VEPILLEVDQIYHLACPASPVHYKYNPVKTIKTNVMGTLNMLGLAKRVGARFLLT 231
Query: 133 STCEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDRLIYAE 192
ST EV+G P E+ ++ Y+ +V+P +R Y K+ + L
Sbjct: 232 STSEVYGD------PLEHPQKETYW---GNVNPI------GERSCYDEGKRTAETLAMDY 276
Query: 193 HAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRGEPLKLVDGGHSQ 252
H G++ I R +N GPRM G RV++ F +R P+ + G
Sbjct: 277 HRGAGVEVRIARIFNTYGPRMCLDDG---------RVVSNFVAQTIRKHPMTVYGDGKQT 327
Query: 253 RTFLYIKDAIEAVMLMIDNPDRANGHI--FNVGNPDNEVSVKQLAELMIKVYAKVAGVPE 310
R+F Y+ D + V LM N H+ FN+GNP E ++ +LAE++ +V
Sbjct: 328 RSFQYVSD-LGLVALM------ENDHVGPFNLGNP-GEFTMLELAEVVKEV--------- 370
Query: 311 SSLSTLDVSSEV-FYGKGYDDSDRRIPDMTIITKQLGWKPKTSL 353
+D S+ + F DD +R PD++ +QL W+PK SL
Sbjct: 371 -----IDPSATIEFKPNTADDPHKRKPDISKAKEQLNWEPKISL 409
>At1g53500 dTDP-D-glucose 4,6-dehydratase, putative
Length = 667
Score = 67.8 bits (164), Expect = 1e-11
Identities = 91/369 (24%), Positives = 156/369 (41%), Gaps = 54/369 (14%)
Query: 6 LDGKPIVPISICLIGGGGFIGSHLTEKLMSETSHKAIVI----DVSSEKVNHLLDKSHPW 61
+D P +I + G GFI SH+ +L+ IV+ D S+ N LD S
Sbjct: 1 MDDTTYKPKNILITGAAGFIASHVANRLIRNYPDYKIVVLDKLDYCSDLKN--LDPSFSS 58
Query: 62 ANRIEFHQMNIKNDSRLETLVKASDL-TINLAAICTPADYNT-RPLDTIYSNFIDAIPVI 119
N +F + +I +D + L+ ++ TI A T D + + +N ++
Sbjct: 59 PN-FKFVKGDIASDDLVNYLLITENIDTIMHFAAQTHVDNSFGNSFEFTKNNIYGTHVLL 117
Query: 120 KFC--TENNKRLIHFSTCEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWS 177
+ C T +R IH ST EV+G+T + + S + P +
Sbjct: 118 EACKVTGQIRRFIHVSTDEVYGET-----------DEDAAVGNHEASQLL--PTNP---- 160
Query: 178 YACAKQMTDRLIYAEHAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNL 237
Y+ K + L+ A GL R N GP P +P+ +
Sbjct: 161 YSATKAGAEMLVMAYGRSYGLPVITTRGNNVYGPNQF-------PEKMIPKFILL----A 209
Query: 238 LRGEPLKLVDGGHSQRTFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAEL 297
+ G+PL + G + R++LY +D EA +++ + GH++NVG E V +A
Sbjct: 210 MSGKPLPIHGDGSNVRSYLYCEDVAEAFEVVLHKGE--IGHVYNVGT-KRERRVIDVARD 266
Query: 298 MIKVYAKVAGVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLL 357
+ K++ K PESS+ V + F + Y D+++ K+LGW+ +T+ +D L
Sbjct: 267 ICKLFGK---DPESSIQF--VENRPFNDQRYFLDDQKL-------KKLGWQERTNWEDGL 314
Query: 358 DSTLQYQHQ 366
T+ + Q
Sbjct: 315 KKTMDWYTQ 323
>At5g44480 putative protein
Length = 436
Score = 60.1 bits (144), Expect = 2e-09
Identities = 85/369 (23%), Positives = 149/369 (40%), Gaps = 59/369 (15%)
Query: 16 ICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHL--LDKSHPWANRIEFHQMNIK 73
+ + GG G+IGSH +L+ ++ IV ++S + + L + P R++F ++
Sbjct: 97 VLVTGGAGYIGSHAALRLLRDSYRVTIVDNLSRGNLGAVKTLQQLFPQTGRLQFIYADLG 156
Query: 74 NDSRLETLV--KASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENN-KRLI 130
+ +E + A D ++ AA+ + PL ++ + + V++ + K+LI
Sbjct: 157 DPLAVEKIFSENAFDAVMHFAAVAYVGESTLYPLKYYHNITSNTLGVLEAMARHKVKKLI 216
Query: 131 HFSTCEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDRLIY 190
+ STC +G EP+ + ED P Y AK+M + +I
Sbjct: 217 YSSTCATYG-------------EPEKMPITEDTPQVPINP-------YGKAKKMAEDMIL 256
Query: 191 AEHAENGLKFTIVRPYNWIGP----RMDFIPGVDGPSDGVPRVL-ACF--SNNLLRGEPL 243
+ + I+R +N IG R+ P + G R+ ACF + + G +
Sbjct: 257 DFSKNSDMAVMILRYFNVIGSDPGGRLGEAPRPELREQG--RISGACFDAARGFIPGLQV 314
Query: 244 KLVD----GGHSQRTFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAELMI 299
K D G R ++ + D ++A + ++ I+NVG SVK+ E
Sbjct: 315 KGTDYKTSDGTCIRDYIDVTDLVDAHVKALEKAQPRKVGIYNVGTGKGR-SVKEFVEACK 373
Query: 300 K---VYAKVAGVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPK-TSLDD 355
K V KV +P +V S D T I K L W + T+L D
Sbjct: 374 KATGVEIKVDFLPRRPGDYAEVYS----------------DPTKILKDLNWTARFTNLQD 417
Query: 356 LLDSTLQYQ 364
L ++Q
Sbjct: 418 SLQVAWRWQ 426
>At1g78570 Similar to dTDP-D-glucose 4,6-dehydratase
Length = 669
Score = 59.7 bits (143), Expect = 3e-09
Identities = 79/361 (21%), Positives = 151/361 (40%), Gaps = 52/361 (14%)
Query: 13 PISICLIGGGGFIGSHLTEKLM-SETSHKAIVIDVSS--EKVNHLLDKSHPWANRIEFHQ 69
P +I + G GFI SH+ +L+ S +K +V+D + +L H + +F +
Sbjct: 6 PKNILITGAAGFIASHVANRLIRSYPDYKIVVLDKLDYCSNLKNLNPSKH--SPNFKFVK 63
Query: 70 MNIKNDSRLETLV--KASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFC--TEN 125
+I + + L+ + D ++ AA + + +N +++ C T
Sbjct: 64 GDIASADLVNHLLITEGIDTIMHFAAQTHVDNSFGNSFEFTKNNIYGTHVLLEACKVTGQ 123
Query: 126 NKRLIHFSTCEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMT 185
+R IH ST EV+G+T L + P + Y+ K
Sbjct: 124 IRRFIHVSTDEVYGETDEDALVGNHEASQLL-------------PTNP----YSATKAGA 166
Query: 186 DRLIYAEHAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRGEPLKL 245
+ L+ A GL R N GP P +P+ + +RG+ L +
Sbjct: 167 EMLVMAYGRSYGLPVITTRGNNVYGPNQF-------PEKLIPKFILL----AMRGQVLPI 215
Query: 246 VDGGHSQRTFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAELMIKVYAKV 305
G + R++LY +D EA +++ + GH++N+G E V +A+ + K++
Sbjct: 216 HGDGSNVRSYLYCEDVAEAFEVVLHKGE--VGHVYNIGT-KKERRVNDVAKDICKLFNM- 271
Query: 306 AGVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLLDSTLQYQH 365
PE+++ +D + F + Y D+++ K+LGW +T+ ++ L T+ +
Sbjct: 272 --DPEANIKFVD--NRPFNDQRYFLDDQKL-------KKLGWSERTTWEEGLKKTMDWYT 320
Query: 366 Q 366
Q
Sbjct: 321 Q 321
>At1g30620 UDP-galactose 4-epimerase-like protein
Length = 419
Score = 57.0 bits (136), Expect = 2e-08
Identities = 82/369 (22%), Positives = 149/369 (40%), Gaps = 59/369 (15%)
Query: 16 ICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVN--HLLDKSHPWANRIEFHQMNIK 73
+ + GG G+IGSH +L+ E+ IV ++S + +L + P R++F ++
Sbjct: 73 VLVTGGAGYIGSHAALRLLKESYRVTIVDNLSRGNLAAVRILQELFPEPGRLQFIYADLG 132
Query: 74 NDSRLETLV--KASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIK-FCTENNKRLI 130
+ + + A D ++ AA+ + PL ++ + + V++ K LI
Sbjct: 133 DAKAVNKIFTENAFDAVMHFAAVAYVGESTQFPLKYYHNITSNTLVVLETMAAHGVKTLI 192
Query: 131 HFSTCEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDRLIY 190
+ STC +G EP + E+ P Y AK+M + +I
Sbjct: 193 YSSTCATYG-------------EPDIMPITEETPQVPINP-------YGKAKKMAEDIIL 232
Query: 191 AEHAENGLKFTIVRPYNWIGP----RMDFIPGVDGPSDGVPRVL-ACF--SNNLLRGEPL 243
+ + I+R +N IG R+ P + G R+ ACF + ++ G +
Sbjct: 233 DFSKNSDMAVMILRYFNVIGSDPEGRLGEAPRPELREHG--RISGACFDAARGIMPGLQI 290
Query: 244 KLVD----GGHSQRTFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAELMI 299
K D G R ++ + D ++A + + I+NVG SVK+ E
Sbjct: 291 KGTDYKTADGTCVRDYIDVTDLVDAHVKALQKAKPRKVGIYNVGTGKGS-SVKEFVEACK 349
Query: 300 K---VYAKVAGVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPK-TSLDD 355
K V K+ +P + Y + Y D + I K+L W K T+L +
Sbjct: 350 KATGVEIKIDYLPRRAGD---------YAEVYSDPSK-------IRKELNWTAKHTNLKE 393
Query: 356 LLDSTLQYQ 364
L++ ++Q
Sbjct: 394 SLETAWRWQ 402
>At4g20460 UDP-glucose 4-epimerase - like protein
Length = 379
Score = 54.3 bits (129), Expect = 1e-07
Identities = 84/372 (22%), Positives = 158/372 (41%), Gaps = 54/372 (14%)
Query: 16 ICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVN--HLLDKSHPWANRIEFHQMNIK 73
+ + GG G+IGSH +L+ ++ IV ++S + +L P R++F ++
Sbjct: 40 VLVTGGAGYIGSHAALRLLKDSYRVTIVDNLSRGNLGAVKVLQGLFPEPGRLQFIYADLG 99
Query: 74 NDSRLETLV--KASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENN-KRLI 130
+ ++ + A D ++ AA+ + PL ++ + + V++ + K+LI
Sbjct: 100 DAKAVDKIFSENAFDAVMHFAAVAYVGESTLDPLKYYHNITSNTLVVLEAVARHKVKKLI 159
Query: 131 HFSTCEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDRLIY 190
+ STC +G EP + E V+P + P++ Y AK+M + +I
Sbjct: 160 YSSTCATYG-------------EPDKMPIVE-VTPQV--PIN----PYGKAKKMAEDMIL 199
Query: 191 AEHAENGLKFTIVRPYNWIGP----RMDFIPGVDGPSDGVPRVL-ACF--SNNLLRGEPL 243
+ + I+R +N IG R+ P + G R+ ACF + ++ G +
Sbjct: 200 DFSKNSDMAVMILRYFNVIGSDPEGRLGEAPKPELREHG--RISGACFDAARGVIPGLQV 257
Query: 244 KLVD----GGHSQRTFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAELMI 299
K D G R ++ + D ++A + ++ N I+NVG SVK+ E
Sbjct: 258 KGTDYKTGDGTCVRDYIDVTDLVDAHVKALEKAKPRNVGIYNVGTGKGR-SVKEFVE--- 313
Query: 300 KVYAKVAGVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPK-TSLDDLLD 358
K GV D+ + F + D D I + L W + T+L + L+
Sbjct: 314 -ACKKATGV--------DIKVD-FLPRRPGDYAEVYSDPAKILRDLNWSARYTNLQESLE 363
Query: 359 STLQYQHQTYSH 370
++Q +T+ H
Sbjct: 364 VAWKWQ-KTHPH 374
>At2g34850 putative UDP-galactose-4-epimerase
Length = 385
Score = 52.4 bits (124), Expect = 4e-07
Identities = 66/298 (22%), Positives = 123/298 (41%), Gaps = 41/298 (13%)
Query: 16 ICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVN--HLLDKSHPWANRIEFHQMNIK 73
+ + GG G+IGSH +L+ ++ IV ++S + +L + P +++F ++
Sbjct: 40 VLVTGGAGYIGSHAALRLLKDSYRVTIVDNLSRGNLGAVKILQQLFPEPGKLQFIYADLG 99
Query: 74 NDSRLETLV--KASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIK-FCTENNKRLI 130
+ + + + A D ++ AA+ + PL ++ + + V++ K LI
Sbjct: 100 DANAVNKIFSENAFDAVMHFAAVAYVGESTQFPLKYYHNITSNTLVVLETMAAHGVKTLI 159
Query: 131 HFSTCEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDRLIY 190
+ STC +G EP+ + E+ P Y AK+M + +I
Sbjct: 160 YSSTCATYG-------------EPEKMPITEETPQVPINP-------YGKAKKMAEDIIL 199
Query: 191 AEHAENGLKFTIVRPYNWIGP----RMDFIPGVDGPSDGVPRVL-ACFS-------NNLL 238
+ + I+R +N IG R+ P + G R+ ACF +
Sbjct: 200 DFSKNSIMAVMILRYFNVIGSDPEGRLGEAPRPELSEHG--RISGACFDAARGIIPGLQI 257
Query: 239 RGEPLKLVDGGHSQRTFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAE 296
+G K VD G R ++ + D ++A + ++ IFNVG SVK+ E
Sbjct: 258 KGTDYKTVD-GTCVRDYIDVTDLVDAHVKALEKAKPRKVGIFNVGTGKGS-SVKEFVE 313
>At4g23920 UDPglucose 4-epimerase - like protein
Length = 350
Score = 49.7 bits (117), Expect = 3e-06
Identities = 76/365 (20%), Positives = 148/365 (39%), Gaps = 65/365 (17%)
Query: 15 SICLIGGGGFIGSHLTEKLMSETSHKAIVID-------VSSEKVNHLLDKSHPWANRIEF 67
S+ + GG G+IGSH +L+ E + A+V+D S ++V L ++ NR+ F
Sbjct: 4 SVLVTGGAGYIGSHTVLQLL-EGGYSAVVVDNYDNSSAASLQRVKKLAGEN---GNRLSF 59
Query: 68 HQMNIKNDSRLETLVKAS--DLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTEN 125
HQ+++++ LE + + D I+ A + + +PL +N + + +++ +
Sbjct: 60 HQVDLRDRPALEKIFSETKFDAVIHFAGLKAVGESVEKPLLYYNNNIVGTVTLLEVMAQY 119
Query: 126 N-KRLIHFSTCEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQM 184
K L+ S+ V+G P+E PC Y K
Sbjct: 120 GCKNLVFSSSATVYG------WPKEV--------------PCTEESPISATNPYGRTKLF 159
Query: 185 TDRLIYAEH-AENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRG-EP 242
+ + H +++ K ++R +N +G G D GVP L + + G P
Sbjct: 160 IEEICRDVHRSDSEWKIILLRYFNPVGAHPSGYIGED--PLGVPNNLMPYVQQVAVGRRP 217
Query: 243 LKLVDG-------GHSQRTFLYIKDAIEAVMLMIDNPD--RANGHIFNVGNPDNEVSVKQ 293
V G G R ++++ D + + + D + + ++N+G N SV +
Sbjct: 218 HLTVFGTDYKTKDGTGVRDYIHVMDLADGHIAALRKLDDLKISCEVYNLGT-GNGTSVLE 276
Query: 294 LAELMIKVYAKVAG--VPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKT 351
M+ + K +G +P +EV Y + ++R +L WK K
Sbjct: 277 ----MVAAFEKASGKKIPLVMAGRRPGDAEVVYA-STEKAER----------ELNWKAKN 321
Query: 352 SLDDL 356
++++
Sbjct: 322 GIEEM 326
>At5g28840 GDP-mannose 3",5"-epimerase
Length = 377
Score = 47.4 bits (111), Expect = 1e-05
Identities = 83/375 (22%), Positives = 149/375 (39%), Gaps = 54/375 (14%)
Query: 10 PIVPISICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSHPWANRIEFHQ 69
P + I + G GGFI SH+ +L E + VI +K H+ + EFH
Sbjct: 24 PSENLKISITGAGGFIASHIARRLKHEGHY---VIASDWKKNEHMTEDMF----CDEFHL 76
Query: 70 MNIKNDSRLETLVKASDLTINLAAICTPADY-NTRPLDTIYSNFIDAIPVIKFCTENN-K 127
++++ + + D NLAA + + +Y+N + + +I+ N K
Sbjct: 77 VDLRVMENCLKVTEGVDHVFNLAADMGGMGFIQSNHSVIMYNNTMISFNMIEAARINGIK 136
Query: 128 RLIHFSTCEVFGKTIGSFLPEEYRKEPQYYKLKE-DVSPCIFGPVHKQRWSYACAKQMTD 186
R + S+ ++ PE + E LKE D P + + +Y K T+
Sbjct: 137 RFFYASSACIY--------PEFKQLETTNVSLKESDAWPA------EPQDAYGLEKLATE 182
Query: 187 RLIYAEHAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRG-EPLKL 245
L + + G++ I R +N GP + G + A F + ++
Sbjct: 183 ELCKHYNKDFGIECRIGRFHNIYGPFGTW-------KGGREKAPAAFCRKAQTSTDRFEM 235
Query: 246 VDGGHSQRTFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAELMIKVYAKV 305
G R+F +I + +E V+ + + R N+G+ D VS+ ++AE++
Sbjct: 236 WGDGLQTRSFTFIDECVEGVLRLTKSDFR---EPVNIGS-DEMVSMNEMAEMV------- 284
Query: 306 AGVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLLDSTLQYQH 365
LS + + + G + R D +I ++LGW P L + L T +
Sbjct: 285 -------LSFEEKKLPIHHIPGPEGVRGRNSDNNLIKEKLGWAPNMRLKEGLRITYFW-- 335
Query: 366 QTYSHAIKKELSKPS 380
I+KE +K S
Sbjct: 336 --IKEQIEKEKAKGS 348
>At2g33600 putative cinnamoyl-CoA reductase
Length = 321
Score = 47.0 bits (110), Expect = 2e-05
Identities = 74/362 (20%), Positives = 139/362 (37%), Gaps = 68/362 (18%)
Query: 16 ICLIGGGGFIGSHLTEKLMSETSH-KAIVIDVSSEKVNHL--LDKSHPWANRIEFHQMNI 72
+C+ G GGF+GS + L+S V D +EK HL LDK+ ++++ + ++
Sbjct: 9 VCVTGAGGFLGSWVVNHLLSRDYFVHGTVRDPGNEKYAHLKKLDKA---GDKLKLFKADL 65
Query: 73 KNDSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENN-KRLIH 131
N L++ + ++A A +D I + V+K C E KR+++
Sbjct: 66 LNYGSLQSAIAGCSGVFHVACPVPSASVPNPEVDLIAPAVDGTLNVLKACVEAKVKRVVY 125
Query: 132 FSTCEVFGK----TIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDR 187
S+ + L E + Y K E+ W Y+ +K +
Sbjct: 126 VSSVSAVAMNPMWSKSQVLDETAWSDQDYCKKTEN-------------W-YSLSKTRAES 171
Query: 188 LIYAEHAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRGEPLKLVD 247
+ GL V P +GP VL + N LKL+
Sbjct: 172 EAFEFAKRTGLDLVSVCPTLVLGP-----------------VLQQHTVNASSLVLLKLLK 214
Query: 248 GGH-----SQRTFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAELMIKVY 302
G+ +R + ++D +A++L+ + + A G +G+ E +++AE + +Y
Sbjct: 215 EGYESRNNQERHLVDVRDVAQALLLVYEKAE-AEGRYICIGHTVRE---QEVAEKLKSLY 270
Query: 303 AKVAGVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLLDSTLQ 362
Y K Y ++D ++ + ++LGW + + L+DS
Sbjct: 271 LNYN-----------------YPKRYIEADGKVKVSSEKLQKLGWTYRPLEETLVDSVES 313
Query: 363 YQ 364
Y+
Sbjct: 314 YR 315
>At2g45310 putative nucleotide sugar epimerase
Length = 437
Score = 45.1 bits (105), Expect = 7e-05
Identities = 76/383 (19%), Positives = 145/383 (37%), Gaps = 77/383 (20%)
Query: 14 ISICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSHPWANR-------IE 66
I++ + G GF+G+H++ L + + N D S A R I
Sbjct: 97 ITVLVTGAAGFVGTHVSAALKRRGDGV-----IGLDNFNDYYDPSLKRARRALLERSGIF 151
Query: 67 FHQMNIKNDSRLETLVKASDLT--INLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTE 124
+ +I + L L K T ++LAA P ++SN + +++ C
Sbjct: 152 IVEGDINDVELLRKLFKIVSFTHVMHLAAQAGVRYAMENPSSYVHSNIAGFVNLLEICKS 211
Query: 125 NNKR--LIHFSTCEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAK 182
N + ++ S+ V+G K P K K D + YA K
Sbjct: 212 VNPQPAIVWASSSSVYGLN---------TKVPFSEKDKTDQPASL----------YAATK 252
Query: 183 QMTDRLIYAEHAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRGEP 242
+ + + + + GL T +R + V GP F+ ++L+G+
Sbjct: 253 KAGEEIAHTYNHIYGLSLTGLRFFT-----------VYGPWGRPDMAYFFFTKDILKGKS 301
Query: 243 LKLVDG---GHSQRTFLYIKDAIEAVMLMIDNPDRANG-----------HIFNVGN--PD 286
+ + + G R F YI D ++ + +D +++ G +FN+GN P
Sbjct: 302 ISIFESANHGTVARDFTYIDDIVKGCLAALDTAEKSTGSGGKKRGPAQLRVFNLGNTSPV 361
Query: 287 NEVSVKQLAELMIKVYAKVAGVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLG 346
+ ++ E +KV AK +L + + +V + ++++ ++LG
Sbjct: 362 PVSDLVRILERQLKVKAK------KNLIKMPRNGDVPFTHA---------NISLAQRELG 406
Query: 347 WKPKTSLDDLLDSTLQYQHQTYS 369
+KP T L L +++ YS
Sbjct: 407 YKPTTDLQTGLKKFVRWYLSYYS 429
>At1g73250 GDP-4-keto-6-deoxy-D-mannose-3,5-epimerase-4-reductase
(GER1)
Length = 312
Score = 45.1 bits (105), Expect = 7e-05
Identities = 72/273 (26%), Positives = 113/273 (41%), Gaps = 53/273 (19%)
Query: 101 NTRPLDTIYSNFIDAIPVIKFCTENN-KRLIHFSTCEVFGKTIGSFLPEEYRKEPQYYKL 159
NT P D I N VI E+ K+L+ + ++ K F P+ P+ L
Sbjct: 75 NTYPADFIGVNLQIQTNVIHSAYEHGVKKLLFLGSSCIYPK----FAPQPI---PESALL 127
Query: 160 KEDVSPCIFGPVHKQRWSYACAKQMTDRLIYAEHAENGLKFTIVRPYNWIGPRMDFIPGV 219
+ P W YA AK + A ++G P N GP +F P
Sbjct: 128 TASLEPT-------NEW-YAIAKIAGIKTCQAYRIQHGWDAISGMPTNLYGPNDNFHPE- 178
Query: 220 DGPSDGVPRVLACFSNNLLRGEPLKLVDG-GHSQRTFLYIKDAIEAVMLMIDNPDRANG- 277
S +P ++ F + G +V G G R FL++ D +A + ++D R +G
Sbjct: 179 --NSHVLPALMRRFHEAKVNGAEEVVVWGTGSPLREFLHVDDLADACVFLLD---RYSGL 233
Query: 278 -HIFNVGNPDNEVSVKQLAELMIKVYAKVAGVPESSLSTLDVSSEVFYGK-GYD-----D 330
H+ N+G+ EV++++LAEL+ +V F GK G+D
Sbjct: 234 EHV-NIGS-GQEVTIRELAELVKEVVG-------------------FEGKLGWDCTKPDG 272
Query: 331 SDRRIPDMTIITKQLGWKPKTSLDDLLDSTLQY 363
+ R++ D + + LGW PK SL D L T +
Sbjct: 273 TPRKLMDSSKLAS-LGWTPKVSLRDGLSQTYDW 304
>At4g10960 UDP-galactose 4-epimerase - like protein
Length = 351
Score = 44.7 bits (104), Expect = 9e-05
Identities = 33/135 (24%), Positives = 67/135 (49%), Gaps = 14/135 (10%)
Query: 15 SICLIGGGGFIGSHLTEKLMSETSHKAIVID-------VSSEKVNHLLDKSHPWANRIEF 67
++ + GG G+IGSH +L+ + +V+D VS ++V L + R+ F
Sbjct: 5 NVLVSGGAGYIGSHTVLQLLL-GGYSVVVVDNLDNSSAVSLQRVKKLAAEH---GERLSF 60
Query: 68 HQMNIKNDSRLETLVKAS--DLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTEN 125
HQ+++++ S LE + + D I+ A + + +PL +N + I +++ ++
Sbjct: 61 HQVDLRDRSALEKIFSETKFDAVIHFAGLKAVGESVEKPLLYYNNNLVGTITLLEVMAQH 120
Query: 126 N-KRLIHFSTCEVFG 139
K L+ S+ V+G
Sbjct: 121 GCKNLVFSSSATVYG 135
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.137 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,840,871
Number of Sequences: 26719
Number of extensions: 382484
Number of successful extensions: 953
Number of sequences better than 10.0: 48
Number of HSP's better than 10.0 without gapping: 14
Number of HSP's successfully gapped in prelim test: 34
Number of HSP's that attempted gapping in prelim test: 887
Number of HSP's gapped (non-prelim): 58
length of query: 381
length of database: 11,318,596
effective HSP length: 101
effective length of query: 280
effective length of database: 8,619,977
effective search space: 2413593560
effective search space used: 2413593560
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC144515.13


