

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144503.9 + phase: 0 /pseudo
(346 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
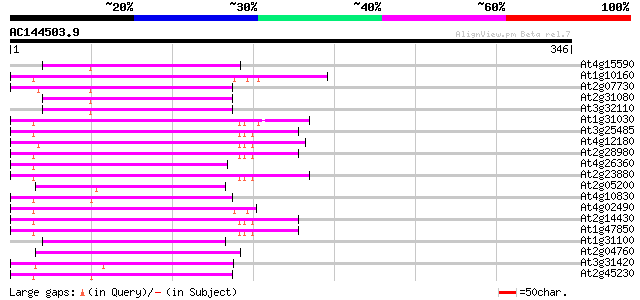
Score E
Sequences producing significant alignments: (bits) Value
At4g15590 reverse transcriptase like protein 68 6e-12
At1g10160 putative reverse transcriptase 68 8e-12
At2g07730 putative non-LTR retroelement reverse transcriptase 66 2e-11
At2g31080 putative non-LTR retroelement reverse transcriptase 65 5e-11
At3g32110 non-LTR reverse transcriptase, putative 65 7e-11
At1g31030 F17F8.5 64 2e-10
At3g25485 unknown protein 59 5e-09
At4g12180 putative reverse transcriptase 57 1e-08
At2g28980 putative non-LTR retroelement reverse transcriptase 57 1e-08
At4g26360 putative protein 56 3e-08
At2g23880 putative non-LTR retroelement reverse transcriptase 56 3e-08
At2g05200 putative non-LTR retroelement reverse transcriptase 55 4e-08
At4g10830 putative protein 55 7e-08
At4g02490 55 7e-08
At2g14430 putative non-LTR retroelement reverse transcriptase 55 7e-08
At1g47850 hypothetical protein 55 7e-08
At1g31100 hypothetical protein 54 1e-07
At2g04760 putative non-LTR retroelement reverse transcriptase 54 1e-07
At3g31420 hypothetical protein 54 2e-07
At2g45230 putative non-LTR retroelement reverse transcriptase 52 4e-07
>At4g15590 reverse transcriptase like protein
Length = 929
Score = 68.2 bits (165), Expect = 6e-12
Identities = 40/124 (32%), Positives = 63/124 (50%), Gaps = 2/124 (1%)
Query: 21 QFGFMPGRLTMEAIYLLRRVIERYRTDK--KDLHLVFIDLEKTYDRVPREILWKALEKKG 78
Q F+PGRL+ + I +++ + R K K L+ +DLEK YDR+ + L + LE G
Sbjct: 353 QASFIPGRLSFDNIVVVQEAVHSMRRKKGRKGWMLLKLDLEKAYDRIRWDFLAETLEAAG 412
Query: 79 VRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGSTLSPYLFTLVLDVLTEHIQE 138
+ +I+ I + G S+ T+ F G+ QG +SPYLF L ++ L I+
Sbjct: 413 LSEGWIKRIMECVAGPEMSLLWNGEKTDSFTPERGLRQGDPISPYLFVLCIERLCHQIET 472
Query: 139 LAPR 142
R
Sbjct: 473 AVGR 476
>At1g10160 putative reverse transcriptase
Length = 1118
Score = 67.8 bits (164), Expect = 8e-12
Identities = 60/216 (27%), Positives = 95/216 (43%), Gaps = 20/216 (9%)
Query: 1 KLWERVIERRLRK--EARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLH-LVFID 57
K+ R+I RRL+ + V NQ GF+ GRL E + L +++ + D + + +D
Sbjct: 534 KVITRIISRRLKLFIDQAVQANQVGFIKGRLLCENVLLASELVDNFEADGETTRGCLQVD 593
Query: 58 LEKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQG 117
+ K YD V E L L+ + + +I I AS S+ F G+ QG
Sbjct: 594 ISKAYDNVNWEFLINILKALDLPLVFIHWIWVCISSASYSIAFNGELIGFFQGKKGIRQG 653
Query: 118 STLSPYLFTLVLDVLTEHIQ--------ELAPRCML-------FADVVLV--GESREEVN 160
+S +LF LV+DVL++ + L P C+ FAD VLV + +
Sbjct: 654 DPMSSHLFVLVMDVLSKSLDLGALNGLFNLHPNCLAPIITHLSFADDVLVFSDGAASSIA 713
Query: 161 GRLETWRQALEAYGFRLSRSTTEYMECNFSGRRSRS 196
G L + G ++R TE + + R+RS
Sbjct: 714 GILTILDDFRQGSGLGINREKTELLLDGGNFARNRS 749
>At2g07730 putative non-LTR retroelement reverse transcriptase
Length = 970
Score = 66.2 bits (160), Expect = 2e-11
Identities = 47/141 (33%), Positives = 69/141 (48%), Gaps = 4/141 (2%)
Query: 1 KLWERVIERRLRKEAR--VTDNQFGFMPGRLTMEAIYLLRRVIERYRTDK--KDLHLVFI 56
K+ ++I RL+K + Q F+PGRL+++ I L++ + R K K L+ +
Sbjct: 229 KIITKMIVNRLKKVILKLIGPAQANFIPGRLSIDNIVLVQEAVHSMRRKKGRKGWMLLKL 288
Query: 57 DLEKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQ 116
DLEK YDR+ L + L G+ + I S SV G T+ F G+ Q
Sbjct: 289 DLEKAYDRIRWNFLQETLVATGLSEVWTHRIMAGVTDPSMSVLWNGGKTDSFVPARGLRQ 348
Query: 117 GSTLSPYLFTLVLDVLTEHIQ 137
G LSPYLF L L+ L I+
Sbjct: 349 GDPLSPYLFVLCLERLCHLIE 369
>At2g31080 putative non-LTR retroelement reverse transcriptase
Length = 1231
Score = 65.1 bits (157), Expect = 5e-11
Identities = 43/119 (36%), Positives = 60/119 (50%), Gaps = 2/119 (1%)
Query: 21 QFGFMPGRLTMEAIYLLRRVIERYRTDK--KDLHLVFIDLEKTYDRVPREILWKALEKKG 78
Q F+PGRL+++ I L++ + R K K L+ +DLEK YDRV + L + LE G
Sbjct: 401 QASFIPGRLSIDNIVLVQEAVHSMRRKKGRKGWMLLKLDLEKAYDRVRWDFLQETLEAAG 460
Query: 79 VRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGSTLSPYLFTLVLDVLTEHIQ 137
+ + I S SV T+ F G+ QG LSPYLF L L+ L I+
Sbjct: 461 LSEGWTSRIMAGVTDPSMSVLWNGERTDSFVPARGLRQGDPLSPYLFVLCLERLCHLIE 519
>At3g32110 non-LTR reverse transcriptase, putative
Length = 1911
Score = 64.7 bits (156), Expect = 7e-11
Identities = 40/119 (33%), Positives = 63/119 (52%), Gaps = 2/119 (1%)
Query: 21 QFGFMPGRLTMEAIYLLRRVIERYRTDK--KDLHLVFIDLEKTYDRVPREILWKALEKKG 78
Q F+PGRL+ + I +++ V+ R K K L+ +DLEK YDR+ ++L L+ G
Sbjct: 1077 QTSFIPGRLSTDNIVVVQEVVHSMRRKKGVKGWMLLKLDLEKAYDRIRWDLLEDTLKAAG 1136
Query: 79 VRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGSTLSPYLFTLVLDVLTEHIQ 137
+ +++ I EG S + T+ F G+ QG LSPYLF L ++ L I+
Sbjct: 1137 LPGTWVQWIMKCVEGPSMRLLWNGEKTDAFKPLRGLRQGDPLSPYLFVLCIERLCHLIE 1195
>At1g31030 F17F8.5
Length = 872
Score = 63.5 bits (153), Expect = 2e-10
Identities = 57/207 (27%), Positives = 93/207 (44%), Gaps = 23/207 (11%)
Query: 1 KLWERVIERRLRK--EARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVF-ID 57
K+ ++I RL+ + +NQ F+ RL +E + L +++ Y D ID
Sbjct: 187 KVISKIIANRLKLLLPRFIAENQSAFVKDRLLIENLLLATELVKDYHKDSISARCAIKID 246
Query: 58 LEKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQG 117
+ K +D V L L +I I AS SV+ F G+ QG
Sbjct: 247 ISKAFDSVQWSFLTNTLVAMNFSPTFIHWINLCITTASFSVQVNGDLVGYFQSKRGLRQG 306
Query: 118 STLSPYLFTLVLDVLTEHIQELA--------PRC-------MLFADVVLV---GESREEV 159
+LSPYLF + +DVL++ + + A P+C + FAD ++V G++R +
Sbjct: 307 CSLSPYLFVICMDVLSKMLDKAAGVRKFGFHPKCQRLGLTHLSFADDLMVLSDGKTR-SI 365
Query: 160 NGRLETWRQALEAYGFRLS-RSTTEYM 185
G LE + + + G R+S +T YM
Sbjct: 366 EGILEVFDEFCKRSGLRISLEKSTLYM 392
>At3g25485 unknown protein
Length = 979
Score = 58.5 bits (140), Expect = 5e-09
Identities = 51/198 (25%), Positives = 87/198 (43%), Gaps = 20/198 (10%)
Query: 1 KLWERVIERRLRK--EARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDK-KDLHLVFID 57
K+ ++I +RL++ + NQ F+ RL ++ + L +++ Y + D + ID
Sbjct: 456 KVISKLIAKRLKEILPQFIAGNQSAFVKDRLLIQNLLLATEIVKDYHKESVSDRCAIKID 515
Query: 58 LEKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQG 117
+ K +D V L L +I I AS SV+ F + G+ QG
Sbjct: 516 ISKAFDSVQWSFLENVLHSLNFSQEFIHWIMLCITTASFSVQVNGELVGFFNSSRGLRQG 575
Query: 118 STLSPYLFTLVLDVLTEHIQELA--------PRC-------MLFAD--VVLVGESREEVN 160
+LSPYLF + +DVL++ + A PRC + FAD +VL V
Sbjct: 576 CSLSPYLFVIAMDVLSKMLDRAAGFKKFGYHPRCQTIGLTHLTFADDLMVLSDGKVRSVE 635
Query: 161 GRLETWRQALEAYGFRLS 178
G + + + + G ++S
Sbjct: 636 GIVSVFDEFAKKSGLKIS 653
>At4g12180 putative reverse transcriptase
Length = 662
Score = 57.4 bits (137), Expect = 1e-08
Identities = 51/202 (25%), Positives = 88/202 (43%), Gaps = 20/202 (9%)
Query: 1 KLWERVIERRLRKEAR--VTDNQFGFMPGRLTMEAIYLLRRVIERYRTDK-KDLHLVFID 57
K+ ++I RL++ + NQ F+ RL +E + L +++ Y D + + ID
Sbjct: 58 KVISKIIANRLKRVLPQFIAGNQSAFIKDRLLIENLLLATELVKDYHKDSVSERCAIKID 117
Query: 58 LEKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQG 117
+ K +D V L L ++ I AS SV+ F G+ QG
Sbjct: 118 ISKAFDSVQWSFLRNVLLTLDFPQEFVHWIMLCVTTASFSVQVNRELAGYFNSLRGLRQG 177
Query: 118 STLSPYLFTLVLDVLTEHIQELA--------PRC-------MLFAD--VVLVGESREEVN 160
+L+PYLF +V+DVL++ + A P+C + FAD +VL +
Sbjct: 178 CSLTPYLFVIVMDVLSKKLDRAAGLRKFGYHPKCKNLGLTHLSFADDIMVLTDGKLRSLE 237
Query: 161 GRLETWRQALEAYGFRLSRSTT 182
G +E + + G ++S + T
Sbjct: 238 GIVEVFDSFAKQSGLKISMAKT 259
>At2g28980 putative non-LTR retroelement reverse transcriptase
Length = 1529
Score = 57.4 bits (137), Expect = 1e-08
Identities = 50/198 (25%), Positives = 86/198 (43%), Gaps = 20/198 (10%)
Query: 1 KLWERVIERRLRK--EARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVF-ID 57
K+ +++ RL+ + + NQ F+ RL ME + L +++ Y + ID
Sbjct: 837 KVISKILANRLKLLLPSFILQNQSAFVKERLLMENVLLATELVKDYHKESVTPRCAMKID 896
Query: 58 LEKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQG 117
+ K +D V + L LE + IK A+ SV+ F + G+ QG
Sbjct: 897 ISKAFDSVQWQFLLNTLEALNFPETFRHWIKLCISTATFSVQVNGELAGFFGSSRGLRQG 956
Query: 118 STLSPYLFTLVLDVLTEHIQELA--------PRC-------MLFAD--VVLVGESREEVN 160
LSPYLF + ++VL+ I E A P+C + FAD +V V + +
Sbjct: 957 CALSPYLFVICMNVLSHMIDEAAVHRNIGYHPKCEKIGLTHLCFADDLMVFVDGHQWSIE 1016
Query: 161 GRLETWRQALEAYGFRLS 178
G + +++ G ++S
Sbjct: 1017 GVINVFKEFAGRSGLQIS 1034
>At4g26360 putative protein
Length = 1141
Score = 56.2 bits (134), Expect = 3e-08
Identities = 40/137 (29%), Positives = 64/137 (46%), Gaps = 3/137 (2%)
Query: 1 KLWERVIERRLRK--EARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLH-LVFID 57
K+ R++ RL+K ++ +Q F+PGRL E + L ++ Y L ++ +D
Sbjct: 512 KVIARLLTDRLQKLLSCVISPSQSAFLPGRLLAENVLLATEMVHGYNWRNISLRGMLKVD 571
Query: 58 LEKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQG 117
L K +D V E + AL GV +I I + +V F G+ QG
Sbjct: 572 LRKAFDSVRWEFIIAALLALGVPTKFINWIHQCISTPTFTVSVNGCCGGFFKSAKGLRQG 631
Query: 118 STLSPYLFTLVLDVLTE 134
LSPYLF L ++V ++
Sbjct: 632 DPLSPYLFVLAMEVFSK 648
>At2g23880 putative non-LTR retroelement reverse transcriptase
Length = 1216
Score = 55.8 bits (133), Expect = 3e-08
Identities = 51/206 (24%), Positives = 85/206 (40%), Gaps = 21/206 (10%)
Query: 1 KLWERVIERRLRK--EARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVF-ID 57
K +++ RL++ + NQ F+ RL +E + L +++ Y D ID
Sbjct: 261 KAISKILANRLKRILPKFIVGNQSAFVKDRLLIENVLLATELVKDYHKDSISTRCAMKID 320
Query: 58 LEKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQG 117
+ K +D + L L +I I AS S++ F G+ QG
Sbjct: 321 ISKAFDSLQWSFLTHVLAAMNFPGEFIHWISLCMSTASFSIQVNGELAGYFRSARGLRQG 380
Query: 118 STLSPYLFTLVLDVLTEHIQELA--------PRC-------MLFAD--VVLVGESREEVN 160
+LSPYLF + +DVL+ + + A PRC + FAD ++L V+
Sbjct: 381 CSLSPYLFVISMDVLSRMLDKAAGAREFGYHPRCKTLGLTHLCFADDLMILTDGKIRSVD 440
Query: 161 GRLETWRQALEAYGFRL-SRSTTEYM 185
G ++ Q G ++ TT Y+
Sbjct: 441 GIVKVLNQFAAKLGLKICMEKTTLYL 466
>At2g05200 putative non-LTR retroelement reverse transcriptase
Length = 1229
Score = 55.5 bits (132), Expect = 4e-08
Identities = 36/120 (30%), Positives = 55/120 (45%), Gaps = 3/120 (2%)
Query: 17 VTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLH---LVFIDLEKTYDRVPREILWKA 73
+++NQ F+PGR+ + + + V+ RT H V D+ K YDRV + L K
Sbjct: 428 ISENQSAFVPGRVISDNVLITHEVLHFLRTSSAKKHCSMAVKTDMSKAYDRVEWDFLKKV 487
Query: 74 LEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGSTLSPYLFTLVLDVLT 133
L++ G +I + + S S T G+ QG LSP LF L +VL+
Sbjct: 488 LQRFGFHSIWIDWVLECVTSVSYSFLINGTPQGKVVPTRGLRQGDPLSPCLFILCTEVLS 547
>At4g10830 putative protein
Length = 1294
Score = 54.7 bits (130), Expect = 7e-08
Identities = 40/142 (28%), Positives = 64/142 (44%), Gaps = 5/142 (3%)
Query: 1 KLWERVIERRLRK--EARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKK---DLHLVF 55
K+ + + RL+ +A V+D+Q F+PGRL + + + ++ +T K+ V
Sbjct: 877 KIISKCLVERLKGHLDAIVSDSQAAFIPGRLVNDNVMIAHEMMHSLKTRKRVSQSYMAVK 936
Query: 56 IDLEKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVH 115
D+ K YDRV L + G +I+ I + + SV G+
Sbjct: 937 TDVSKAYDRVEWNFLETTMRLFGFSETWIKWIMGAVKSVNYSVLVNGIPHGTIQPQRGIR 996
Query: 116 QGSTLSPYLFTLVLDVLTEHIQ 137
QG LSPYLF L D+L I+
Sbjct: 997 QGDPLSPYLFILCADILNHLIK 1018
>At4g02490
Length = 657
Score = 54.7 bits (130), Expect = 7e-08
Identities = 47/170 (27%), Positives = 74/170 (42%), Gaps = 18/170 (10%)
Query: 1 KLWERVIERRLRK--EARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLH-LVFID 57
K+ R+I +RL + V NQ GF+ GRL E + L +++ ++ + + +D
Sbjct: 81 KVITRLISKRLNLFIDQAVQSNQVGFIKGRLLCENVLLASELVDNFQAEGDTSRGCLQVD 140
Query: 58 LEKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQG 117
L K YD V E L L+ + +I I S S+ F G+ QG
Sbjct: 141 LTKAYDNVNWEFLINILKALNLPPIFINWIWVCISTPSYSIAYNGELIGFFVGKKGIRQG 200
Query: 118 STLSPYLFTLVLDVLTEHIQ--------ELAPRCML-------FADVVLV 152
+S +LF LV+D+L + L P+C+ FAD +LV
Sbjct: 201 DPMSSHLFVLVMDILARSLDLGAVEGRFVLHPKCLAPMITHLSFADDILV 250
>At2g14430 putative non-LTR retroelement reverse transcriptase
Length = 1277
Score = 54.7 bits (130), Expect = 7e-08
Identities = 51/198 (25%), Positives = 81/198 (40%), Gaps = 20/198 (10%)
Query: 1 KLWERVIERRLRK--EARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVF-ID 57
K+ ++I RL+ + NQ F+ RL ME + L +++ Y D ID
Sbjct: 682 KVISKIIANRLKVMLPTFILQNQSAFVRERLLMENVLLATELVKDYHKDSISPRCAMKID 741
Query: 58 LEKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQG 117
+ K +D V + L LE + IK A+ SV+ F G+ QG
Sbjct: 742 ISKAFDSVQWQFLLNTLEALKFPEKFRHWIKLCISTATFSVQVNSEQAGFFGSKRGLRQG 801
Query: 118 STLSPYLFTLVLDVLTEHIQELA--------PRC-------MLFAD--VVLVGESREEVN 160
LSPYLF + ++VL+ I A P+C + FAD +V + + V
Sbjct: 802 CALSPYLFVICMNVLSHMIDVAAVHRNIGYHPKCKKLSLTHLCFADDLMVFIDGQQRSVE 861
Query: 161 GRLETWRQALEAYGFRLS 178
G + ++ G +S
Sbjct: 862 GVINIFKDFAGKSGLHIS 879
>At1g47850 hypothetical protein
Length = 799
Score = 54.7 bits (130), Expect = 7e-08
Identities = 50/198 (25%), Positives = 82/198 (41%), Gaps = 20/198 (10%)
Query: 1 KLWERVIERRLRK--EARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVF-ID 57
K+ ++I RL+ + NQ F+ RL +E + L +++ Y D ID
Sbjct: 113 KVISKIIANRLKVMLPTFILQNQSAFVRERLLIENVLLATELVKDYHKDSISPRCAMKID 172
Query: 58 LEKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQG 117
+ K +D V + L LE + IK A+ SV+ F G+ QG
Sbjct: 173 ISKAFDSVQWQFLLNTLEALNFPENFCHWIKLCISTATFSVQVNGELAGFFGSKRGLRQG 232
Query: 118 STLSPYLFTLVLDVLTEHIQELA--------PRC-------MLFAD--VVLVGESREEVN 160
LSPYLF + ++VL+ I A P+C + FAD +V + + V
Sbjct: 233 CALSPYLFVICMNVLSHMIDVAAVHRNIGYHPKCKKLSLTHLCFADDLMVFIDGQQRSVE 292
Query: 161 GRLETWRQALEAYGFRLS 178
G + +++ G +S
Sbjct: 293 GVINIFKEFAGKSGLHIS 310
>At1g31100 hypothetical protein
Length = 1090
Score = 54.3 bits (129), Expect = 1e-07
Identities = 32/114 (28%), Positives = 55/114 (48%), Gaps = 1/114 (0%)
Query: 21 QFGFMPGRLTMEAIYLLRRVIERYRTDKKDLH-LVFIDLEKTYDRVPREILWKALEKKGV 79
Q F+PGR E + L +++ Y D ++ +DL K +D + + + AL+ G+
Sbjct: 471 QSAFLPGRFLAENVLLATELVQGYNRQNIDPRGMLKVDLRKAFDSIRWDFIISALKAIGI 530
Query: 80 RIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGSTLSPYLFTLVLDVLT 133
++ I + SV T F T G+ QG+ LSP+LF L ++V +
Sbjct: 531 PDRFVYWITQCISTPTFSVCVNGNTGGFFKSTRGLRQGNPLSPFLFVLAMEVFS 584
>At2g04760 putative non-LTR retroelement reverse transcriptase
Length = 631
Score = 53.9 bits (128), Expect = 1e-07
Identities = 39/127 (30%), Positives = 55/127 (42%), Gaps = 1/127 (0%)
Query: 17 VTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVF-IDLEKTYDRVPREILWKALE 75
+ NQ F+ RL ME + L +++ Y D ID+ K +D V + L LE
Sbjct: 128 ILQNQSAFVKERLLMENVLLATELVKDYHKDSISPRCAMKIDISKAFDSVQWQFLLNTLE 187
Query: 76 KKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGSTLSPYLFTLVLDVLTEH 135
IK A+ SV+ F G+ QG LSPYLF + ++VL+
Sbjct: 188 ALNFPEKIRHWIKLCISTATFSVQVNGELAGFFGNKRGLRQGCALSPYLFVICMNVLSHM 247
Query: 136 IQELAPR 142
I E A R
Sbjct: 248 IDEAAVR 254
>At3g31420 hypothetical protein
Length = 1491
Score = 53.5 bits (127), Expect = 2e-07
Identities = 37/143 (25%), Positives = 61/143 (41%), Gaps = 5/143 (3%)
Query: 1 KLWERVIERRLRKE--ARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFI-- 56
K+ +++ RL+K +T+NQ F+PGRL + V + K+ +
Sbjct: 700 KIISKILVNRLKKHLGGAITENQAAFVPGRLITNNAIIAHEVYYALKARKRQANSYMALK 759
Query: 57 -DLEKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVH 115
D+ K YDR+ + L + + + G +I I SV G+
Sbjct: 760 TDITKAYDRLEWDFLEETMRQMGFNTKWIERIMICVTMVRFSVLINGSPHGTIKPERGIR 819
Query: 116 QGSTLSPYLFTLVLDVLTEHIQE 138
G LSPYLF L +VL+ I++
Sbjct: 820 HGDPLSPYLFILCAEVLSHMIKQ 842
>At2g45230 putative non-LTR retroelement reverse transcriptase
Length = 1374
Score = 52.4 bits (124), Expect = 4e-07
Identities = 36/142 (25%), Positives = 67/142 (46%), Gaps = 5/142 (3%)
Query: 1 KLWERVIERRLRK--EARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKK---DLHLVF 55
K+ +++ RL+K + +++ Q F+ GRL + I + ++ ++ K + +
Sbjct: 515 KVIGKLMANRLKKILPSLISETQAAFVKGRLISDNILIAHELLHALSSNNKCSEEFIAIK 574
Query: 56 IDLEKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVH 115
D+ K YDRV L KA+ G +IR I + + V + + G+
Sbjct: 575 TDISKAYDRVEWPFLEKAMRGLGFADHWIRLIMECVKSVRYQVLINGTPHGEIIPSRGLR 634
Query: 116 QGSTLSPYLFTLVLDVLTEHIQ 137
QG LSPYLF + ++L + +Q
Sbjct: 635 QGDPLSPYLFVICTEMLVKMLQ 656
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.326 0.140 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,275,854
Number of Sequences: 26719
Number of extensions: 290278
Number of successful extensions: 952
Number of sequences better than 10.0: 62
Number of HSP's better than 10.0 without gapping: 37
Number of HSP's successfully gapped in prelim test: 25
Number of HSP's that attempted gapping in prelim test: 879
Number of HSP's gapped (non-prelim): 66
length of query: 346
length of database: 11,318,596
effective HSP length: 100
effective length of query: 246
effective length of database: 8,646,696
effective search space: 2127087216
effective search space used: 2127087216
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC144503.9


