

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144503.4 - phase: 0
(366 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
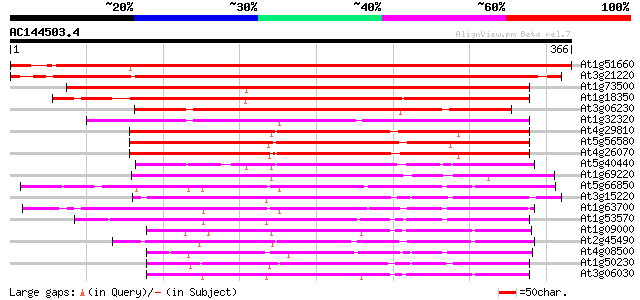
Score E
Sequences producing significant alignments: (bits) Value
At1g51660 unknown protein 509 e-144
At3g21220 MAP kinase kinase 5 474 e-134
At1g73500 unknown protein 284 6e-77
At1g18350 MAP kinase kinase 5, putative 276 1e-74
At3g06230 putative MAP kinase 209 1e-54
At1g32320 MAP kinase, putative 204 8e-53
At4g29810 MAP kinase kinase 2 202 3e-52
At5g56580 protein kinase MEK1 homolog 199 2e-51
At4g26070 MEK1 199 2e-51
At5g40440 MAP kinase kinase 3 (ATMKK3) 169 2e-42
At1g69220 serine/threonine kinase (SIK1) 142 3e-34
At5g66850 MAP protein kinase like protein 137 1e-32
At3g15220 putative MAP kinase 136 2e-32
At1g63700 putative protein kinase 135 3e-32
At1g53570 MEK kinase MAP3Ka, putative 134 8e-32
At1g09000 NPK1-related protein kinase 1S (ANP1) 131 5e-31
At2g45490 putative protein kinase 129 3e-30
At4g08500 MEKK1/MAP kinase kinase kinase 128 6e-30
At1g50230 hypothetical protein 126 2e-29
At3g06030 NPK1-related protein kinase 3 126 2e-29
>At1g51660 unknown protein
Length = 366
Score = 509 bits (1310), Expect = e-144
Identities = 260/374 (69%), Positives = 293/374 (77%), Gaps = 20/374 (5%)
Query: 1 MRPIQLPPPTSTGSATASGSPNNNNSRPQRRRRDHLTLPLPQRDTNLAVPLPLPPSGGSG 60
MRPIQ PP S SRP RRR LTLPLPQRD +LAVPLPLPP+ G
Sbjct: 1 MRPIQSPPGVSVPV----------KSRP--RRRPDLTLPLPQRDVSLAVPLPLPPTSGGS 48
Query: 61 GGGNGSGSGSGGASQQL-------VIPFSELERLNRIGSGSGGTVYKVVHRINGRAYALK 113
GG +GS SGG++ +S+L R NRIGSG+GGTVYKV+HR + R YALK
Sbjct: 49 GGSSGSAPSSGGSASSTNTNSSIEAKNYSDLVRGNRIGSGAGGTVYKVIHRPSSRLYALK 108
Query: 114 VIYGHHEESVRRQIHREIQILRDVDDVNVVKCHEMYDHNAEIQVLLEYMDGGSLEGKHIP 173
VIYG+HEE+VRRQI REI+ILRDV+ NVVKCHEM+D N EIQVLLE+MD GSLEG H+
Sbjct: 109 VIYGNHEETVRRQICREIEILRDVNHPNVVKCHEMFDQNGEIQVLLEFMDKGSLEGAHVW 168
Query: 174 QENQLADVARQILRGLAYLHRRHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPC 233
+E QLAD++RQIL GLAYLH RHIVHRDIKPSNLLINS K VKIADFGV RIL QTMDPC
Sbjct: 169 KEQQLADLSRQILSGLAYLHSRHIVHRDIKPSNLLINSAKNVKIADFGVSRILAQTMDPC 228
Query: 234 NSSVGTIAYMSPERINTDINDGQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLM 293
NSSVGTIAYMSPERINTD+N G+YD YAGDIWSLGVSILEFY+GRFPF V RQGDWASLM
Sbjct: 229 NSSVGTIAYMSPERINTDLNQGKYDGYAGDIWSLGVSILEFYLGRFPFPVSRQGDWASLM 288
Query: 294 CAICMSQPPEAPTTASPEFRDFVSRCLQRDPSRRWTASRLLSHPFLVRNGSNHNQSPPNM 353
CAICMSQPPEAP TASPEFR F+S CLQR+P +R +A +LL HPF++R + N+SP N+
Sbjct: 289 CAICMSQPPEAPATASPEFRHFISCCLQREPGKRRSAMQLLQHPFILRASPSQNRSPQNL 348
Query: 354 HQLLPPP-PRSQSS 366
HQLLPPP P S SS
Sbjct: 349 HQLLPPPRPLSSSS 362
>At3g21220 MAP kinase kinase 5
Length = 348
Score = 474 bits (1221), Expect = e-134
Identities = 243/360 (67%), Positives = 276/360 (76%), Gaps = 19/360 (5%)
Query: 1 MRPIQLPPPTSTGSATASGSPNNNNSRPQRRRRDHLTLPLPQRDTNLAVPLPLPPSGGSG 60
M+PIQ P + SP N + R+R L+LPLP RD LAVPLPLPP S
Sbjct: 1 MKPIQSP--------SGVASPMKN----RLRKRPDLSLPLPHRDVALAVPLPLPPPSSSS 48
Query: 61 GGGNGSGSGSGGASQQLVIPFSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHE 120
S + S S + SELER+NRIGSG+GGTVYKV+H R +ALKVIYG+HE
Sbjct: 49 SAPASSSAISTNISAAKSL--SELERVNRIGSGAGGTVYKVIHTPTSRPFALKVIYGNHE 106
Query: 121 ESVRRQIHREIQILRDVDDVNVVKCHEMYDHNAEIQVLLEYMDGGSLEGKHIPQENQLAD 180
++VRRQI REI+ILR VD NVVKCH+M+DHN EIQVLLE+MD GSLEG HI QE +LAD
Sbjct: 107 DTVRRQICREIEILRSVDHPNVVKCHDMFDHNGEIQVLLEFMDQGSLEGAHIWQEQELAD 166
Query: 181 VARQILRGLAYLHRRHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTI 240
++RQIL GLAYLHRRHIVHRDIKPSNLLINS K VKIADFGV RIL QTMDPCNSSVGTI
Sbjct: 167 LSRQILSGLAYLHRRHIVHRDIKPSNLLINSAKNVKIADFGVSRILAQTMDPCNSSVGTI 226
Query: 241 AYMSPERINTDINDGQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICMSQ 300
AYMSPERINTD+N G+YD YAGD+WSLGVSILEFY+GRFPFAV RQGDWASLMCAICMSQ
Sbjct: 227 AYMSPERINTDLNHGRYDGYAGDVWSLGVSILEFYLGRFPFAVSRQGDWASLMCAICMSQ 286
Query: 301 PPEAPTTASPEFRDFVSRCLQRDPSRRWTASRLLSHPFLVRNGSNHNQSPPNMHQLLPPP 360
PPEAP TAS EFR FVS CLQ DP +RW+A +LL HPF+++ PN+ Q+LPPP
Sbjct: 287 PPEAPATASQEFRHFVSCCLQSDPPKRWSAQQLLQHPFILKATGG-----PNLRQMLPPP 341
>At1g73500 unknown protein
Length = 310
Score = 284 bits (726), Expect = 6e-77
Identities = 147/306 (48%), Positives = 201/306 (65%), Gaps = 4/306 (1%)
Query: 38 LPLPQRDTNLAVPLPLPPSGGSGGGGNGSGSGSGGASQQLVIPFSELERLNRIGSGSGGT 97
+ L + L + LPLPP + S + + + I +LE+LN +G G+GG
Sbjct: 1 MALVRERRQLNLRLPLPPISDRRFSTSSSSATTTTVAGCNGISACDLEKLNVLGCGNGGI 60
Query: 98 VYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVDDVNVVKCHEMYDHNA--EI 155
VYKV H+ YALK + G + RQ+ RE++ILR D VVKCH +++ E+
Sbjct: 61 VYKVRHKTTSEIYALKTVNGDMDPIFTRQLMREMEILRRTDSPYVVKCHGIFEKPVVGEV 120
Query: 156 QVLLEYMDGGSLEG-KHIPQENQLADVARQILRGLAYLHRRHIVHRDIKPSNLLINSRKQ 214
+L+EYMDGG+LE + E +LA A+QIL+GL+YLH IVHRDIKP+NLL+NS+ +
Sbjct: 121 SILMEYMDGGTLESLRGGVTEQKLAGFAKQILKGLSYLHALKIVHRDIKPANLLLNSKNE 180
Query: 215 VKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDINDGQYDAYAGDIWSLGVSILEF 274
VKIADFGV +IL +++D CNS VGT AYMSPER +++ + G D YAGDIWS G+ +LE
Sbjct: 181 VKIADFGVSKILVRSLDSCNSYVGTCAYMSPERFDSESSGGSSDIYAGDIWSFGLMMLEL 240
Query: 275 YMGRFP-FAVGRQGDWASLMCAICMSQPPEAPTTASPEFRDFVSRCLQRDPSRRWTASRL 333
+G FP G++ DWA+LMCA+C +PP AP S EFR FV CL++D S+RWTA +L
Sbjct: 241 LVGHFPLLPPGQRPDWATLMCAVCFGEPPRAPEGCSEEFRSFVECCLRKDSSKRWTAPQL 300
Query: 334 LSHPFL 339
L+HPFL
Sbjct: 301 LAHPFL 306
>At1g18350 MAP kinase kinase 5, putative
Length = 307
Score = 276 bits (706), Expect = 1e-74
Identities = 150/315 (47%), Positives = 202/315 (63%), Gaps = 20/315 (6%)
Query: 29 QRRRRDHLTLPLPQRDTNLAVPLPLPPSGGSGGGGNGSGSGSGGASQQLVIPFSELERLN 88
++RR+ +L LP+P L+V LP S +G I S++E+L+
Sbjct: 5 RKRRQINLRLPVPP----LSVHLPWFSFASSTAPVINNG-----------ISASDVEKLH 49
Query: 89 RIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVDDVNVVKCHEM 148
+G GS G VYKV H+ G YALK + G + RQ+ RE++ILR D VV+C +
Sbjct: 50 VLGRGSSGIVYKVHHKTTGEIYALKSVNGDMSPAFTRQLAREMEILRRTDSPYVVRCQGI 109
Query: 149 YDHN--AEIQVLLEYMDGGSLEG-KHIPQENQLADVARQILRGLAYLHRRHIVHRDIKPS 205
++ E+ +L+EYMDGG+LE + E QLA +RQIL+GL+YLH IVHRDIKP+
Sbjct: 110 FEKPIVGEVSILMEYMDGGNLESLRGAVTEKQLAGFSRQILKGLSYLHSLKIVHRDIKPA 169
Query: 206 NLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDINDGQYDAYAGDIW 265
NLL+NSR +VKIADFGV +I+ +++D CNS VGT AYMSPER ++ + D YAGDIW
Sbjct: 170 NLLLNSRNEVKIADFGVSKIITRSLDYCNSYVGTCAYMSPERFDSAAGENS-DVYAGDIW 228
Query: 266 SLGVSILEFYMGRFP-FAVGRQGDWASLMCAICMSQPPEAPTTASPEFRDFVSRCLQRDP 324
S GV ILE ++G FP G++ DWA+LMC +C +PP AP S EFR FV CL+++
Sbjct: 229 SFGVMILELFVGHFPLLPQGQRPDWATLMCVVCFGEPPRAPEGCSDEFRSFVDCCLRKES 288
Query: 325 SRRWTASRLLSHPFL 339
S RWTAS+LL HPFL
Sbjct: 289 SERWTASQLLGHPFL 303
>At3g06230 putative MAP kinase
Length = 293
Score = 209 bits (533), Expect = 1e-54
Identities = 108/251 (43%), Positives = 168/251 (66%), Gaps = 13/251 (5%)
Query: 82 SELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVDDVN 141
+ L+R++ +GSG+GGTV+KV + YALK + +E+ REI+ILR V+
Sbjct: 51 ANLDRISVLGSGNGGTVFKVKDKTTSEIYALKKV----KENWDSTSLREIEILRMVNSPY 106
Query: 142 VVKCHEMYDH-NAEIQVLLEYMDGGSLEGKHIPQENQLADVARQILRGLAYLHRRHIVHR 200
V KCH+++ + + E+ +L++YMD GSLE E QLA ++RQ+L G YLH IVHR
Sbjct: 107 VAKCHDIFQNPSGEVSILMDYMDLGSLESLRGVTEKQLALMSRQVLEGKNYLHEHKIVHR 166
Query: 201 DIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDIN----DGQ 256
DIKP+NLL +S+++VKIADFGV +I+ ++++ CNS VGT AYMSPER++++ + + +
Sbjct: 167 DIKPANLLRSSKEEVKIADFGVSKIVVRSLNKCNSFVGTFAYMSPERLDSEADGVTEEDK 226
Query: 257 YDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICMSQPPEAPTTASPEFRDFV 316
+ YAGDIWS G+++LE +G +P D A+++CA+C +PP+AP S + + F+
Sbjct: 227 SNVYAGDIWSFGLTMLEILVGYYPML----PDQAAIVCAVCFGEPPKAPEECSDDLKSFM 282
Query: 317 SRCLQRDPSRR 327
CL++ S R
Sbjct: 283 DCCLRKKASER 293
>At1g32320 MAP kinase, putative
Length = 305
Score = 204 bits (518), Expect = 8e-53
Identities = 119/295 (40%), Positives = 173/295 (58%), Gaps = 13/295 (4%)
Query: 51 LPLPPSGGSGGGGNGSGSGSGGASQQLVIPFSELERLNRIGSGSGGTVYKVVHRINGRAY 110
L +PP G + + S S + ++LE+L+ +G GSGGTVYK HR Y
Sbjct: 15 LSIPPLIYHGTAFSVASSSSSSPETSPIQTLNDLEKLSVLGQGSGGTVYKTRHRRTKTLY 74
Query: 111 ALKVIYGHHEESVRRQIHREIQILRDVDDVNVVKCHEMYDHNAEIQVLLEYMDGGSLEGK 170
ALKV+ ++ + E IL+ ++ ++KC+ ++ ++ ++E M+ GSL
Sbjct: 75 ALKVL----RPNLNTTVTVEADILKRIESSFIIKCYAVFVSLYDLCFVMELMEKGSLHDA 130
Query: 171 HIPQ----ENQLADVARQILRGLAYLHRRHIVHRDIKPSNLLINSRKQVKIADFGVGRIL 226
+ Q E ++ +A +IL+GL YL + IVH DIKPSNLLIN + +VKIADFG RI+
Sbjct: 131 LLAQQVFSEPMVSSLANRILQGLRYLQKMGIVHGDIKPSNLLINKKGEVKIADFGASRIV 190
Query: 227 NQTMDPCNSSVGTIAYMSPERINTD-INDGQYDAYAGDIWSLGVSILEFYMGRFPFA-VG 284
S GT AYMSPER++ + G +AGD+WSLGV +LE Y+GR+P VG
Sbjct: 191 ---AGGDYGSNGTCAYMSPERVDLEKWGFGGEVGFAGDVWSLGVVVLECYIGRYPLTKVG 247
Query: 285 RQGDWASLMCAICMSQPPEAPTTASPEFRDFVSRCLQRDPSRRWTASRLLSHPFL 339
+ DWA+L CAIC ++ + P + S EFRDFV RCL++D +R T LL H F+
Sbjct: 248 DKPDWATLFCAICCNEKVDIPVSCSLEFRDFVGRCLEKDWRKRDTVEELLRHSFV 302
>At4g29810 MAP kinase kinase 2
Length = 363
Score = 202 bits (513), Expect = 3e-52
Identities = 120/272 (44%), Positives = 165/272 (60%), Gaps = 17/272 (6%)
Query: 79 IPFSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVD 138
+ S+L+ + IG GS G V V H+ G+ +ALKVI + +E++R+ I +E++I +
Sbjct: 65 LSLSDLDMVKVIGKGSSGVVQLVQHKWTGQFFALKVIQLNIDEAIRKAIAQELKINQSSQ 124
Query: 139 DVNVVKCHEMYDHNAEIQVLLEYMDGGSLEG-----KHIPQENQLADVARQILRGLAYLH 193
N+V ++ + N I ++LEYMDGGSL K IP ++ L+ + RQ+L+GL YLH
Sbjct: 125 CPNLVTSYQSFYDNGAISLILEYMDGGSLADFLKSVKAIP-DSYLSAIFRQVLQGLIYLH 183
Query: 194 R-RHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDI 252
RHI+HRD+KPSNLLIN R +VKI DFGV ++ T N+ VGT YMSPERI
Sbjct: 184 HDRHIIHRDLKPSNLLINHRGEVKITDFGVSTVMTNTAGLANTFVGTYNYMSPERI---- 239
Query: 253 NDGQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQGD-WAS---LMCAICMSQPPEAPT-T 307
G DIWSLG+ +LE G+FP+A Q + W S LM AI PP P+
Sbjct: 240 -VGNKYGNKSDIWSLGLVVLECATGKFPYAPPNQEETWTSVFELMEAIVDQPPPALPSGN 298
Query: 308 ASPEFRDFVSRCLQRDPSRRWTASRLLSHPFL 339
SPE F+S CLQ+DP+ R +A L+ HPFL
Sbjct: 299 FSPELSSFISTCLQKDPNSRSSAKELMEHPFL 330
>At5g56580 protein kinase MEK1 homolog
Length = 356
Score = 199 bits (507), Expect = 2e-51
Identities = 121/273 (44%), Positives = 167/273 (60%), Gaps = 18/273 (6%)
Query: 79 IPFSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVD 138
I +LE + IG GSGG V V H+ G+ +A+KVI + +E +R+QI +E++I +
Sbjct: 65 ITAEDLETVKVIGKGSGGVVQLVRHKWVGKFFAMKVIQMNIQEEIRKQIVQELKINQASS 124
Query: 139 DV-NVVKCHEMYDHNAEIQVLLEYMDGGSL-----EGKHIPQENQLADVARQILRGLAYL 192
+VV C+ + HN ++LEYMD GSL + K I E LA V +Q+L GL YL
Sbjct: 125 QCPHVVVCYHSFYHNGAFSLVLEYMDRGSLADVIRQVKTI-LEPYLAVVCKQVLLGLVYL 183
Query: 193 HR-RHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTD 251
H RH++HRDIKPSNLL+N + +VKI+DFGV L +M ++ VGT YMSPERI+
Sbjct: 184 HNERHVIHRDIKPSNLLVNHKGEVKISDFGVSASLASSMGQRDTFVGTYNYMSPERISGS 243
Query: 252 INDGQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQ----GDWASLMCAICMSQPPEAPTT 307
D Y+ DIWSLG+S+LE +GRFP+ + L+ AI + PP AP+
Sbjct: 244 TYD-----YSSDIWSLGMSVLECAIGRFPYLESEDQQNPPSFYELLAAIVENPPPTAPSD 298
Query: 308 A-SPEFRDFVSRCLQRDPSRRWTASRLLSHPFL 339
SPEF FVS C+Q+DP R ++ LLSHPF+
Sbjct: 299 QFSPEFCSFVSACIQKDPPARASSLDLLSHPFI 331
>At4g26070 MEK1
Length = 354
Score = 199 bits (506), Expect = 2e-51
Identities = 119/273 (43%), Positives = 167/273 (60%), Gaps = 19/273 (6%)
Query: 79 IPFSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVD 138
+ ++LE + IG GS G V V H++ + +ALKVI + EES R I +E++I
Sbjct: 63 LSLADLEVIKVIGKGSSGNVQLVKHKLTQQFFALKVIQLNTEESTCRAISQELRINLSSQ 122
Query: 139 DVNVVKCHEMYDHNAEIQVLLEYMDGGSLE------GKHIPQENQLADVARQILRGLAYL 192
+V C++ + HN + ++LE+MDGGSL GK +P EN L+ + +++LRGL Y+
Sbjct: 123 CPYLVSCYQSFYHNGLVSIILEFMDGGSLADLLKKVGK-VP-ENMLSAICKRVLRGLCYI 180
Query: 193 HR-RHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTD 251
H R I+HRD+KPSNLLIN R +VKI DFGV +IL T NS VGT YMSPERI+
Sbjct: 181 HHERRIIHRDLKPSNLLINHRGEVKITDFGVSKILTSTSSLANSFVGTYPYMSPERIS-- 238
Query: 252 INDGQYDAYAGDIWSLGVSILEFYMGRFPFA-VGRQGDWAS---LMCAICMSQPPEAPTT 307
G + DIWSLG+ +LE G+FP+ + W+S L+ AI + PP AP+
Sbjct: 239 ---GSLYSNKSDIWSLGLVLLECATGKFPYTPPEHKKGWSSVYELVDAIVENPPPCAPSN 295
Query: 308 A-SPEFRDFVSRCLQRDPSRRWTASRLLSHPFL 339
SPEF F+S+C+Q+DP R +A LL H F+
Sbjct: 296 LFSPEFCSFISQCVQKDPRDRKSAKELLEHKFV 328
>At5g40440 MAP kinase kinase 3 (ATMKK3)
Length = 520
Score = 169 bits (428), Expect = 2e-42
Identities = 114/276 (41%), Positives = 154/276 (55%), Gaps = 31/276 (11%)
Query: 83 ELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVDDVNV 142
E+ IGSG+ V + +H N R ALK I E R+Q+ EI+ L +
Sbjct: 82 EMRVFGAIGSGASSVVQRAIHIPNHRILALKKI-NIFEREKRQQLLTEIRTLCEAP---- 136
Query: 143 VKCHE-MYD-HNA-------EIQVLLEYMDGGSLEG-----KHIPQENQLADVARQILRG 188
CHE + D H A +I + LEYM+GGSL K IP E L+ + ++L+G
Sbjct: 137 --CHEGLVDFHGAFYSPDSGQISIALEYMNGGSLADILKVTKKIP-EPVLSSLFHKLLQG 193
Query: 189 LAYLHR-RHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPER 247
L+YLH RH+VHRDIKP+NLLIN + + KI DFG+ L +M C + VGT+ YMSPER
Sbjct: 194 LSYLHGVRHLVHRDIKPANLLINLKGEPKITDFGISAGLENSMAMCATFVGTVTYMSPER 253
Query: 248 INTDINDGQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICMSQPPEAPTT 307
I D +Y DIWSLG+++ E G FP+ + +G +LM I P P
Sbjct: 254 IRNDSY-----SYPADIWSLGLALFECGTGEFPY-IANEGP-VNLMLQILDDPSPTPPKQ 306
Query: 308 A-SPEFRDFVSRCLQRDPSRRWTASRLLSHPFLVRN 342
SPEF F+ CLQ+DP R TA +LLSHPF+ ++
Sbjct: 307 EFSPEFCSFIDACLQKDPDARPTADQLLSHPFITKH 342
>At1g69220 serine/threonine kinase (SIK1)
Length = 836
Score = 142 bits (358), Expect = 3e-34
Identities = 95/286 (33%), Positives = 146/286 (50%), Gaps = 21/286 (7%)
Query: 80 PFSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVDD 139
P ++ E LN +G GS G+VYK A+KVI E +I EI++L+ +
Sbjct: 245 PTTKYEFLNELGKGSYGSVYKARDLKTSEIVAVKVISLTEGEEGYEEIRGEIEMLQQCNH 304
Query: 140 VNVVKCHEMYDHNAEIQVLLEYMDGGSLEG-----KHIPQENQLADVARQILRGLAYLHR 194
NVV+ Y + +++EY GGS+ + +E Q+A + R+ L+GLAYLH
Sbjct: 305 PNVVRYLGSYQGEDYLWIVMEYCGGGSVADLMNVTEEALEEYQIAYICREALKGLAYLHS 364
Query: 195 RHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDIND 254
+ VHRDIK N+L+ + +VK+ DFGV L +TM N+ +GT +M+PE I + D
Sbjct: 365 IYKVHRDIKGGNILLTEQGEVKLGDFGVAAQLTRTMSKRNTFIGTPHWMAPEVIQENRYD 424
Query: 255 GQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICMSQPPEAPTTASPE--- 311
G+ D+W+LGVS +E G P + M + M AP E
Sbjct: 425 GKV-----DVWALGVSAIEMAEGLPPRSS------VHPMRVLFMISIEPAPMLEDKEKWS 473
Query: 312 --FRDFVSRCLQRDPSRRWTASRLLSHPFLVRNGSNHNQSPPNMHQ 355
F DFV++CL ++P R TA+ +L H F+ R + + P + +
Sbjct: 474 LVFHDFVAKCLTKEPRLRPTAAEMLKHKFVERCKTGASAMSPKIEK 519
>At5g66850 MAP protein kinase like protein
Length = 533
Score = 137 bits (344), Expect = 1e-32
Identities = 116/368 (31%), Positives = 184/368 (49%), Gaps = 32/368 (8%)
Query: 8 PPTSTGSATASGSPNNNNSRPQRRRRDHLTLPLPQRDTNLAVPLPLPPSGGSGGGGNGSG 67
P + S +S P ++ S P R + +RD PLPLPP G S
Sbjct: 87 PSSPLHSVDSSAPPRDSVSSPLHPRLS-TDVTNGRRDCCNVHPLPLPP-----GATCSSS 140
Query: 68 SGSGGASQQLVIPF------SELERLNRIGSGSGGTVYKVVHRINGRAYALKVI--YGHH 119
S + S Q + S+ ++ IG G+ G+VY + G A+K + +
Sbjct: 141 SAASVPSPQAPLKLDSFPMNSQWKKGKLIGRGTFGSVYVASNSETGALCAMKEVELFPDD 200
Query: 120 EESVR--RQIHREIQILRDVDDVNVVKCHEMYDHNAEIQVLLEYMDGGSLEGKHIPQ--- 174
+S +Q+ +EI++L ++ N+V+ + LEY+ GS+ K+I
Sbjct: 201 PKSAECIKQLEQEIKLLSNLQHPNIVQYFGSETVEDRFFIYLEYVHPGSIN-KYIRDHCG 259
Query: 175 ---ENQLADVARQILRGLAYLHRRHIVHRDIKPSNLLINSRKQVKIADFGVGR-ILNQTM 230
E+ + + R IL GLAYLH + VHRDIK +NLL+++ VK+ADFG+ + + Q
Sbjct: 260 TMTESVVRNFTRHILSGLAYLHNKKTVHRDIKGANLLVDASGVVKLADFGMAKHLTGQRA 319
Query: 231 DPCNSSVGTIAYMSPERINTDI-NDGQYD-AYAGDIWSLGVSILEFYMGRFPFAVGRQGD 288
D S G+ +M+PE + + D D A+A DIWSLG +I+E + G+ P++ + +
Sbjct: 320 D--LSLKGSPYWMAPELMQAVMQKDSNPDLAFAVDIWSLGCTIIEMFTGKPPWS---EFE 374
Query: 289 WASLMCAICMSQPPEAPTTASPEFRDFVSRCLQRDPSRRWTASRLLSHPFLVRNGSNHNQ 348
A+ M + PP P + SPE +DF+ C QR+P+ R TAS LL H FL + +
Sbjct: 375 GAAAMFKVMRDSPP-IPESMSPEGKDFLRLCFQRNPAERPTASMLLEHRFLKNSLQPTSP 433
Query: 349 SPPNMHQL 356
S ++ QL
Sbjct: 434 SNSDVSQL 441
>At3g15220 putative MAP kinase
Length = 690
Score = 136 bits (343), Expect = 2e-32
Identities = 91/284 (32%), Positives = 144/284 (50%), Gaps = 18/284 (6%)
Query: 81 FSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVDDV 140
FS++E IG GS G VYK + + A+KVI E I +EI +L
Sbjct: 15 FSQIEL---IGRGSFGDVYKAFDKDLNKEVAIKVIDLEESEDEIEDIQKEISVLSQCRCP 71
Query: 141 NVVKCHEMYDHNAEIQVLLEYMDGGS----LEGKHIPQENQLADVARQILRGLAYLHRRH 196
+ + + Y H ++ +++EYM GGS L+ + E +A + R +L + YLH
Sbjct: 72 YITEYYGSYLHQTKLWIIMEYMAGGSVADLLQSNNPLDETSIACITRDLLHAVEYLHNEG 131
Query: 197 IVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDINDGQ 256
+HRDIK +N+L++ VK+ADFGV L +T+ + VGT +M+PE I N
Sbjct: 132 KIHRDIKAANILLSENGDVKVADFGVSAQLTRTISRRKTFVGTPFWMAPEVIQ---NSEG 188
Query: 257 YDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICMSQPPEAPTTASPEFRDFV 316
Y+ A DIWSLG++++E G P A ++ I PP+ S + ++FV
Sbjct: 189 YNEKA-DIWSLGITVIEMAKGEPPLADLHP---MRVLFIIPRETPPQLDEHFSRQVKEFV 244
Query: 317 SRCLQRDPSRRWTASRLLSHPFLVRNGSNHNQSPPNMHQLLPPP 360
S CL++ P+ R +A L+ H F+ N +SP + ++ P
Sbjct: 245 SLCLKKAPAERPSAKELIKHRFI----KNARKSPKLLERIRERP 284
>At1g63700 putative protein kinase
Length = 883
Score = 135 bits (341), Expect = 3e-32
Identities = 111/344 (32%), Positives = 175/344 (50%), Gaps = 29/344 (8%)
Query: 9 PTSTGSATASGSPNNNNSRPQRRRRDHLTLPLPQRDTNLAVPLPLPPSGGSGGGGNGSGS 68
P + GS T S + +++R Q R LPLP L + P S + S
Sbjct: 334 PRAGGSTTGSPTRRLDDNRQQSHR-----LPLPP----LLISNTCPFSPTYSAATSPSVP 384
Query: 69 GSGGASQQLVIPFSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRR--- 125
S ++ V P S ++ +G GS G VY + +G A+K + ++ R
Sbjct: 385 RSPARAEATVSPGSRWKKGRLLGMGSFGHVYLGFNSESGEMCAMKEVTLCSDDPKSRESA 444
Query: 126 -QIHREIQILRDVDDVNVVKCHEMYDHNAEIQVLLEYMDGGSLEGKHIPQ-----ENQLA 179
Q+ +EI +L + N+V+ + + ++ + LEY+ GGS+ K + + EN +
Sbjct: 445 QQLGQEISVLSRLRHQNIVQYYGSETVDDKLYIYLEYVSGGSIY-KLLQEYGQFGENAIR 503
Query: 180 DVARQILRGLAYLHRRHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGT 239
+ +QIL GLAYLH ++ VHRDIK +N+L++ +VK+ADFG+ + + P S G+
Sbjct: 504 NYTQQILSGLAYLHAKNTVHRDIKGANILVDPHGRVKVADFGMAKHITAQSGPL-SFKGS 562
Query: 240 IAYMSPERINTDINDGQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICMS 299
+M+PE I A DIWSLG ++LE + P++ Q + M I S
Sbjct: 563 PYWMAPEVIKNSNGSN----LAVDIWSLGCTVLEMATTKPPWS---QYEGVPAMFKIGNS 615
Query: 300 QP-PEAPTTASPEFRDFVSRCLQRDPSRRWTASRLLSHPFLVRN 342
+ P+ P S E +DFV +CLQR+P+ R TA++LL H F VRN
Sbjct: 616 KELPDIPDHLSEEGKDFVRKCLQRNPANRPTAAQLLDHAF-VRN 658
>At1g53570 MEK kinase MAP3Ka, putative
Length = 609
Score = 134 bits (337), Expect = 8e-32
Identities = 100/306 (32%), Positives = 162/306 (52%), Gaps = 18/306 (5%)
Query: 43 RDTNLAVPLPLPPSGGSGGGGNGSGSGSGGASQQLVIPFSELERLNRIGSGSGGTVYKVV 102
R ++ PLP PP+ + GS GG + FS ++ +GSG+ G VY
Sbjct: 174 RSSSECHPLPRPPTSPTSPSAV-HGSRIGGGYETSPSGFSTWKKGKFLGSGTFGQVYLGF 232
Query: 103 HRINGRAYALKVIYGHHEESVRR----QIHREIQILRDVDDVNVVKCHEMYDHNAEIQVL 158
+ G+ A+K + ++ + Q+++EI +L + N+V+ + + V
Sbjct: 233 NSEKGKMCAIKEVKVISDDQTSKECLKQLNQEINLLNQLCHPNIVQYYGSELSEETLSVY 292
Query: 159 LEYMDGGS----LEGKHIPQENQLADVARQILRGLAYLHRRHIVHRDIKPSNLLINSRKQ 214
LEY+ GGS L+ E + + RQIL GLAYLH R+ VHRDIK +N+L++ +
Sbjct: 293 LEYVSGGSIHKLLKDYGSFTEPVIQNYTRQILAGLAYLHGRNTVHRDIKGANILVDPNGE 352
Query: 215 VKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDINDGQYDAYAGDIWSLGVSILEF 274
+K+ADFG+ + + S G+ +M+PE + ++ Y +A DIWSLG +ILE
Sbjct: 353 IKLADFGMAKHVT-AFSTMLSFKGSPYWMAPEVV---MSQNGY-THAVDIWSLGCTILEM 407
Query: 275 YMGRFPFAVGRQGDWASLMCAICMSQ-PPEAPTTASPEFRDFVSRCLQRDPSRRWTASRL 333
+ P++ Q + + + I S+ PE P S + ++F+ CLQR+P+ R TAS+L
Sbjct: 408 ATSKPPWS---QFEGVAAIFKIGNSKDTPEIPDHLSNDAKNFIRLCLQRNPTVRPTASQL 464
Query: 334 LSHPFL 339
L HPFL
Sbjct: 465 LEHPFL 470
>At1g09000 NPK1-related protein kinase 1S (ANP1)
Length = 666
Score = 131 bits (330), Expect = 5e-31
Identities = 87/264 (32%), Positives = 145/264 (53%), Gaps = 19/264 (7%)
Query: 90 IGSGSGGTVYKVVHRINGRAYALK--VIYGHHEESVRRQIH-----REIQILRDVDDVNV 142
IG G+ GTVY ++ +G A+K +I + + Q H E+++L+++ N+
Sbjct: 75 IGRGAFGTVYMGMNLDSGELLAVKQVLIAANFASKEKTQAHIQELEEEVKLLKNLSHPNI 134
Query: 143 VKCHEMYDHNAEIQVLLEYMDGGSLEG---KHIP-QENQLADVARQILRGLAYLHRRHIV 198
V+ + + +LLE++ GGS+ K P E+ + RQ+L GL YLH I+
Sbjct: 135 VRYLGTVREDDTLNILLEFVPGGSISSLLEKFGPFPESVVRTYTRQLLLGLEYLHNHAIM 194
Query: 199 HRDIKPSNLLINSRKQVKIADFGVGRILNQ--TMDPCNSSVGTIAYMSPERINTDINDGQ 256
HRDIK +N+L++++ +K+ADFG + + + TM S GT +M+PE I + G
Sbjct: 195 HRDIKGANILVDNKGCIKLADFGASKQVAELATMTGAKSMKGTPYWMAPEVI---LQTGH 251
Query: 257 YDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICMSQPPEAPTTASPEFRDFV 316
+++ DIWS+G +++E G+ P++ + A S PP P T S + +DF+
Sbjct: 252 --SFSADIWSVGCTVIEMVTGKAPWSQQYKEVAAIFFIGTTKSHPP-IPDTLSSDAKDFL 308
Query: 317 SRCLQRDPSRRWTASRLLSHPFLV 340
+CLQ P+ R TAS LL HPF++
Sbjct: 309 LKCLQEVPNLRPTASELLKHPFVM 332
>At2g45490 putative protein kinase
Length = 288
Score = 129 bits (323), Expect = 3e-30
Identities = 88/282 (31%), Positives = 141/282 (49%), Gaps = 20/282 (7%)
Query: 68 SGSGGASQQLVIPFSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEES--VRR 125
S +G +Q + E+ R +G G G VY + ALKVI+ E +
Sbjct: 8 SDAGNTEKQWSLADFEIGR--PLGKGKFGRVYLAREAKSKYIVALKVIFKEQIEKYKIHH 65
Query: 126 QIHREIQILRDVDDVNVVKCHEMYDHNAEIQVLLEYMDGGSLEGK-----HIPQENQLAD 180
Q+ RE++I + N+++ + N I ++LEY GG L G H+ E Q A
Sbjct: 66 QLRREMEIQTSLRHPNILRLFGWFHDNERIFLILEYAHGGELYGVLKQNGHLT-EQQAAT 124
Query: 181 VARQILRGLAYLHRRHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTI 240
+ + LAY H + ++HRDIKP NLL++ ++KIADFG Q+ + + GT+
Sbjct: 125 YIASLSQALAYCHGKCVIHRDIKPENLLLDHEGRLKIADFGWS---VQSSNKRKTMCGTL 181
Query: 241 AYMSPERINTDINDGQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICMSQ 300
Y++PE + +D YA D W+LG+ EF G PF Q D + I +S
Sbjct: 182 DYLAPEMVENRDHD-----YAVDNWTLGILCYEFLYGNPPFEAESQKDTFKRILKIDLSF 236
Query: 301 PPEAPTTASPEFRDFVSRCLQRDPSRRWTASRLLSHPFLVRN 342
P S E ++ +S+ L +DPS+R + +++ HP++V+N
Sbjct: 237 P--LTPNVSEEAKNLISQLLVKDPSKRLSIEKIMQHPWIVKN 276
>At4g08500 MEKK1/MAP kinase kinase kinase
Length = 608
Score = 128 bits (321), Expect = 6e-30
Identities = 89/261 (34%), Positives = 141/261 (53%), Gaps = 19/261 (7%)
Query: 90 IGSGSGGTVYKVVHRINGRAYALKVI----YGHHEESVRRQIHREIQILRDVDDVNVVKC 145
+G GS G+VY+ + +G +A+K + G + +Q+ EI++L + N+V+
Sbjct: 339 LGRGSFGSVYEGISG-DGDFFAVKEVSLLDQGSQAQECIQQLEGEIKLLSQLQHQNIVRY 397
Query: 146 HEMYDHNAEIQVLLEYMDGGSLEGKHIPQENQLAD-----VARQILRGLAYLHRRHIVHR 200
+ + + LE + GSL + Q QL D RQIL GL YLH + +HR
Sbjct: 398 RGTAKDGSNLYIFLELVTQGSL--LKLYQRYQLRDSVVSLYTRQILDGLKYLHDKGFIHR 455
Query: 201 DIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDINDGQYDAY 260
DIK +N+L+++ VK+ADFG+ ++ + S GT +M+PE IN +DG Y +
Sbjct: 456 DIKCANILVDANGAVKLADFGLAKV--SKFNDIKSCKGTPFWMAPEVINRKDSDG-YGSP 512
Query: 261 AGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICMSQPPEAPTTASPEFRDFVSRCL 320
A DIWSLG ++LE G+ P++ + + I PE P T S + R F+ +CL
Sbjct: 513 A-DIWSLGCTVLEMCTGQIPYS---DLEPVQALFRIGRGTLPEVPDTLSLDARLFILKCL 568
Query: 321 QRDPSRRWTASRLLSHPFLVR 341
+ +P R TA+ LL+HPF+ R
Sbjct: 569 KVNPEERPTAAELLNHPFVRR 589
>At1g50230 hypothetical protein
Length = 269
Score = 126 bits (317), Expect = 2e-29
Identities = 81/256 (31%), Positives = 130/256 (50%), Gaps = 17/256 (6%)
Query: 90 IGSGSGGTVYKVVHRINGRAYALKVIY--GHHEESVRRQIHREIQILRDVDDVNVVKCHE 147
+G GS G VYK + G+ A+K I G ++ + + +EI+ILR + N+++ +
Sbjct: 12 VGEGSFGRVYKGRRKYTGQTVAMKFIMKQGKTDKDIH-SLRQEIEILRKLKHENIIEMLD 70
Query: 148 MYDHNAEIQVLLEYMDGGSLE----GKHIPQENQLADVARQILRGLAYLHRRHIVHRDIK 203
+++ E V+ E+ G E K +P+E Q+ +A+Q+++ L YLH I+HRD+K
Sbjct: 71 SFENAREFCVVTEFAQGELFEILEDDKCLPEE-QVQAIAKQLVKALDYLHSNRIIHRDMK 129
Query: 204 PSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDINDGQYDAYAGD 263
P N+LI + VK+ DFG R ++ S GT YM+PE + + YD D
Sbjct: 130 PQNILIGAGSVVKLCDFGFARAMSTNTVVLRSIKGTPLYMAPEL----VKEQPYDRTV-D 184
Query: 264 IWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICMSQPPEAPTTASPEFRDFVSRCLQRD 323
+WSLGV + E Y+G+ PF + + P + P S F F+ L ++
Sbjct: 185 LWSLGVILYELYVGQPPFYTNS----VYALIRHIVKDPVKYPDEMSTYFESFLKGLLNKE 240
Query: 324 PSRRWTASRLLSHPFL 339
P R T L HPF+
Sbjct: 241 PHSRLTWPALREHPFV 256
>At3g06030 NPK1-related protein kinase 3
Length = 651
Score = 126 bits (316), Expect = 2e-29
Identities = 81/263 (30%), Positives = 144/263 (53%), Gaps = 19/263 (7%)
Query: 90 IGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVR-------RQIHREIQILRDVDDVNV 142
IG G+ G VY ++ +G A+K + + + R++ E+Q+L+++ N+
Sbjct: 74 IGCGAFGRVYMGMNLDSGELLAIKQVLIAPSSASKEKTQGHIRELEEEVQLLKNLSHPNI 133
Query: 143 VKCHEMYDHNAEIQVLLEYMDGGS----LEGKHIPQENQLADVARQILRGLAYLHRRHIV 198
V+ + + +L+E++ GGS LE E + +Q+L GL YLH I+
Sbjct: 134 VRYLGTVRESDSLNILMEFVPGGSISSLLEKFGSFPEPVIIMYTKQLLLGLEYLHNNGIM 193
Query: 199 HRDIKPSNLLINSRKQVKIADFGVGRILNQ--TMDPCNSSVGTIAYMSPERINTDINDGQ 256
HRDIK +N+L++++ +++ADFG + + + T++ S GT +M+PE I + G
Sbjct: 194 HRDIKGANILVDNKGCIRLADFGASKKVVELATVNGAKSMKGTPYWMAPEVI---LQTGH 250
Query: 257 YDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICMSQPPEAPTTASPEFRDFV 316
+++ DIWS+G +++E G+ P++ Q A L + PP P SPE +DF+
Sbjct: 251 --SFSADIWSVGCTVIEMATGKPPWSEQYQQFAAVLHIGRTKAHPP-IPEDLSPEAKDFL 307
Query: 317 SRCLQRDPSRRWTASRLLSHPFL 339
+CL ++PS R +A+ LL HPF+
Sbjct: 308 MKCLHKEPSLRLSATELLQHPFV 330
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.136 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,515,804
Number of Sequences: 26719
Number of extensions: 474289
Number of successful extensions: 9916
Number of sequences better than 10.0: 1133
Number of HSP's better than 10.0 without gapping: 981
Number of HSP's successfully gapped in prelim test: 154
Number of HSP's that attempted gapping in prelim test: 4031
Number of HSP's gapped (non-prelim): 3056
length of query: 366
length of database: 11,318,596
effective HSP length: 101
effective length of query: 265
effective length of database: 8,619,977
effective search space: 2284293905
effective search space used: 2284293905
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC144503.4


