

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144484.11 - phase: 0
(796 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
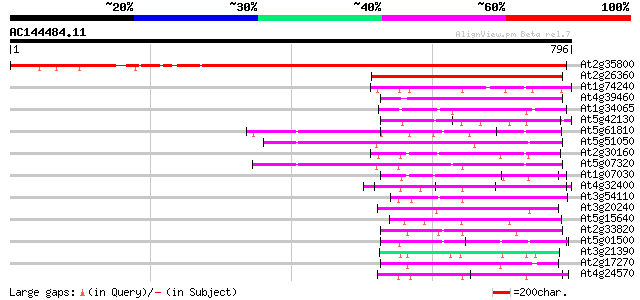
Score E
Sequences producing significant alignments: (bits) Value
At2g35800 putative mitochondrial carrier protein 994 0.0
At2g26360 mitochondrial carrier like protein 379 e-105
At1g74240 mitochondrial carrier like protein 131 1e-30
At4g39460 mitochondrial carrier like protein 124 2e-28
At1g34065 allinase like protein 124 3e-28
At5g42130 mitochondrial carrier protein-like 114 2e-25
At5g61810 peroxisomal Ca-dependent solute carrier - like protein 109 7e-24
At5g51050 calcium-binding transporter-like protein 104 2e-22
At2g30160 putative mitochondrial carrier protein 103 5e-22
At5g07320 peroxisomal Ca-dependent solute carrier-like protein 100 3e-21
At1g07030 unknown protein 99 7e-21
At4g32400 adenylate translocator (brittle-1) - like protein 93 5e-19
At3g54110 uncoupling protein (ucp/PUMP) 83 7e-16
At3g20240 mitochondrial carrier protein, putative 82 1e-15
At5g15640 putative mitochondrial carrier protein 77 5e-14
At2g33820 mitochondrial basic amino acid carrier (BAC1) 75 1e-13
At5g01500 unknown protein 70 6e-12
At3g21390 unknown protein 70 6e-12
At2g17270 putative mitochondrial phosphate translocator protein 70 6e-12
At4g24570 putative mitochondrial uncoupling protein 69 1e-11
>At2g35800 putative mitochondrial carrier protein
Length = 823
Score = 994 bits (2571), Expect = 0.0
Identities = 538/833 (64%), Positives = 639/833 (76%), Gaps = 65/833 (7%)
Query: 1 MVLDNDPVESFFNSIQVMKES-LSPLEVGFRKAAKDFEHCF--------------AKNKT 45
MV ND +E+ FNSIQ++K+ L P+E+G +KAA+D E+C+ +N+
Sbjct: 1 MVSKNDHIETLFNSIQLVKDVVLLPIELGVKKAARDIENCWISKERDLGLVFRSSGRNRK 60
Query: 46 QGVCLIAQVKDGGDFQI-CDV---KKKKGLSMK-VPLKAFLGKFSQN--SEKLNKTQVV- 97
+ + + D + C V ++KKGLS+K +P+K+ G FS N SEKL++ V
Sbjct: 61 KRIVATPEFDDNATNNVQCVVVTDERKKGLSIKKIPVKSLFGMFSPNLVSEKLSRGNDVV 120
Query: 98 ---------KENESSCSNCLKFSVTWSLLVSGFIQSLPIPFKSVKKRGQKVCDEDSHKEK 148
K+++ SC++C KF++TWSLLVSGF+ + PIPFK KKR K+ D+++ K
Sbjct: 121 VAKKDKSLEKKDDDSCTDCFKFAMTWSLLVSGFVHAFPIPFKIGKKRIHKMGDDENSLRK 180
Query: 149 CSCMKPSLSPCEMKHNESKGRTIKEKVVK-------RKDGKEHVSLECVIGFIFDQLSHT 201
+SK + K V+ K+G S+EC +GF+ + L+
Sbjct: 181 HGL-------------KSKAAFVSRKEVRCQSVESVEKEGNPF-SIECAVGFVVEMLAQN 226
Query: 202 LQSLDQGINGLQEKNDELECGK-ASLDSAPFGHVNAFTSFLEGHKVDVNGFLGNLNFAKV 260
LQ LDQ I E +E C K AS + +P + E K+DVNGFLGNL FA+V
Sbjct: 227 LQKLDQFIQDSSE--NESCCSKEASSNDSPL-----IFNIWEARKLDVNGFLGNLMFARV 279
Query: 261 GGVPSSVAGEEIASQNEMGDSANDET--KEESVGISAQKVASNIFSIPLTNVERLKTTLS 318
G V S + G + +E GD +N T KEES S Q +A+ + SIPL+NVERLK+TLS
Sbjct: 280 GDVASGIGGLT-SHISEDGDESNVSTAGKEESAVDSPQNLATGLLSIPLSNVERLKSTLS 338
Query: 319 TVSLTELIEMLPQLGKTTKDHPDKKKLFSVQDFFRYTESEGRRFFEELDRDGDGQVTLED 378
T+SLTELIE+LPQLG+ ++DHPDKKKL SVQDFFRYTESEGRRFFEELDRDGDG+VTLED
Sbjct: 339 TISLTELIELLPQLGRPSRDHPDKKKLISVQDFFRYTESEGRRFFEELDRDGDGKVTLED 398
Query: 379 LEIAMRRRKLPRRYAKEFMSRTRSHLFSRSFGWKQFLSFMEQKEPTILRAYTSLCLTKSG 438
LEIAMRRRKLPRRYAKEFM R RSHLFS+SFGWKQFLS MEQKEPTILRAYTSLCLTKSG
Sbjct: 399 LEIAMRRRKLPRRYAKEFMRRARSHLFSKSFGWKQFLSLMEQKEPTILRAYTSLCLTKSG 458
Query: 439 TLKKSEILESLKNSGLPANEDNAAAMMRFLNADTEESISYGHFRNFMLLLPSDRLQEDPR 498
TLKKSEIL SL N+GLPANE+NA AMMRFL ADTEESISYGHFRNFM+LLP +RLQ+DPR
Sbjct: 459 TLKKSEILASLNNAGLPANEENAIAMMRFLKADTEESISYGHFRNFMVLLPYERLQDDPR 518
Query: 499 SIWFEAATVVAVPPSVEIPAGSVLRSALAGGLSCALSCALLHPVDSIKTRVQASSMSFPE 558
+IWFEAATVVAV P V +PAG VL+SALAGGL+ ALS +L+HP+D+IKTRVQAS++SFPE
Sbjct: 519 NIWFEAATVVAVAPPVALPAGDVLKSALAGGLASALSTSLMHPIDTIKTRVQASTLSFPE 578
Query: 559 IIAKLPEIGTRGLYRGSIPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASF 618
+IAKLPEIG RG+YRGSIPAILGQFSSHGLRTGIFEASKLVL+N APNLPE+QVQSIASF
Sbjct: 579 VIAKLPEIGVRGVYRGSIPAILGQFSSHGLRTGIFEASKLVLINFAPNLPEIQVQSIASF 638
Query: 619 CSTFLGTAVRIPCEVLKQRLQAGLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYV 678
CST LGTAVRIPCEVLKQRLQAG+FNNVGEA+VGTW+QDG GFFRGTGATLCREVP YV
Sbjct: 639 CSTLLGTAVRIPCEVLKQRLQAGMFNNVGEAIVGTWKQDGPSGFFRGTGATLCREVPLYV 698
Query: 679 AGMGLYAESKKGVQKLLGRELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTA-QGRSV 737
GMGLYAESKK V + LGRELEAWETIAVGA+SGG+AAVVTTPFDVMKTRMMTA GR +
Sbjct: 699 VGMGLYAESKKMVAQALGRELEAWETIAVGAVSGGIAAVVTTPFDVMKTRMMTATPGRPI 758
Query: 738 SMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAMNKNDEA 790
SMS+V SILR+EGPLGLFKGAVPRFFW+APLGAMNFAGYELA+KAM KN++A
Sbjct: 759 SMSMVVVSILRNEGPLGLFKGAVPRFFWVAPLGAMNFAGYELAKKAMQKNEDA 811
>At2g26360 mitochondrial carrier like protein
Length = 369
Score = 379 bits (973), Expect = e-105
Identities = 191/273 (69%), Positives = 226/273 (81%), Gaps = 2/273 (0%)
Query: 514 VEIPAGSVLRSALAGGLSCALSCALLHPVDSIKTRVQASS-MSFPEIIAKLPEIGTRGLY 572
V + G +L+SALAGG+SCA S L+HPVD++KT+VQAS+ +SF EI++K+PEIG RGLY
Sbjct: 86 VGLDVGHLLKSALAGGISCAFSAFLMHPVDTVKTQVQASTTLSFLEILSKIPEIGARGLY 145
Query: 573 RGSIPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCE 632
+GSIPA++GQF+SHGLRT I+EASKL L VAP L ++QVQSIASF T LGT +RIPCE
Sbjct: 146 KGSIPAVVGQFASHGLRTSIYEASKLALPLVAPTLLDIQVQSIASFIGTVLGTTLRIPCE 205
Query: 633 VLKQRLQAGLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQ 692
VLKQRLQA F+N+ EA V TW Q+GLKG FRGTG TL REVPFYVAGMGLY +SKK V+
Sbjct: 206 VLKQRLQANQFDNIVEATVSTWHQEGLKGLFRGTGVTLLREVPFYVAGMGLYNQSKKVVE 265
Query: 693 KLLGRELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTA-QGRSVSMSIVAFSILRHEG 751
+ LGRELE WE IAVGALSGG AV+TTPFDV+KTRMMTA QG +SM + A+SIL HEG
Sbjct: 266 RQLGRELEPWEAIAVGALSGGFTAVLTTPFDVIKTRMMTAPQGVELSMLMAAYSILTHEG 325
Query: 752 PLGLFKGAVPRFFWIAPLGAMNFAGYELARKAM 784
PL +KGAVPRFFW APLGA+N AGYEL +KAM
Sbjct: 326 PLAFYKGAVPRFFWTAPLGALNLAGYELLQKAM 358
>At1g74240 mitochondrial carrier like protein
Length = 364
Score = 131 bits (330), Expect = 1e-30
Identities = 100/339 (29%), Positives = 155/339 (45%), Gaps = 60/339 (17%)
Query: 513 SVEIPAGS----VLRSALAGGLSCALSCALLHPVDSIKTRVQA-----SSMSFPEIIAKL 563
SVEI A V R L GG++ A ++HPVD++KTR+Q+ ++ I+ L
Sbjct: 20 SVEIKATHDQFFVWREFLWGGIAGAFGEGMMHPVDTLKTRLQSQIIMNATQRQKSILQML 79
Query: 564 PEI----GTRGLYRGSIPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFC 619
+ G +G YRG P + G ++ G E++K + P+L IA
Sbjct: 80 RTVWVGDGLKGFYRGIAPGVTGSLATGATYFGFIESTKKWIEESHPSLAGHWAHFIAGAV 139
Query: 620 STFLGTAVRIPCEVLKQRLQA-------------------------GLFNNVGEALVGTW 654
LG+ + +PCEV+KQR+Q G + + +A W
Sbjct: 140 GDTLGSFIYVPCEVIKQRMQIQGTSSSWSSYISRNSVPVQPRGDMYGYYTGMFQAGCSIW 199
Query: 655 QQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELEAW---------ETI 705
++ G KG + G +TL R+VPF GL +G++ L + + + E +
Sbjct: 200 KEQGPKGLYAGYWSTLARDVPF----AGLMVVFYEGLKDLTDQGKKKFPQYGVNSSIEGL 255
Query: 706 AVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMS---IVAFSILRHEGPLGLFKGAVPR 762
+G L+GGL+A +TTP DV+KTR+ QG ++ I R EGP G F+G+VPR
Sbjct: 256 VLGGLAGGLSAYLTTPLDVVKTRLQ-VQGSTIKYKGWLDAVGQIWRKEGPQGFFRGSVPR 314
Query: 763 FFWIAPLGAMNFAGYELAR-----KAMNKNDEAKTGNLE 796
W P A+ F E R K+ N N+ ++E
Sbjct: 315 VMWYLPASALTFMAVEFLRDNFREKSNNNNNVVSNLSIE 353
>At4g39460 mitochondrial carrier like protein
Length = 325
Score = 124 bits (312), Expect = 2e-28
Identities = 81/261 (31%), Positives = 125/261 (47%), Gaps = 10/261 (3%)
Query: 526 LAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAKLPEIGTRGLYRGSIPAILGQFSS 585
+AGG + + L+P+D+IKTR+QA+ +I +GLY G I G +
Sbjct: 59 IAGGTAGVVVETALYPIDTIKTRLQAARGG--------GKIVLKGLYSGLAGNIAGVLPA 110
Query: 586 HGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVLKQRLQAGLFNN 645
L G++E +K L+ P+ A + +R+P EV+KQR+Q G F +
Sbjct: 111 SALFVGVYEPTKQKLLKTFPDHLSAVAHLTAGAIGGLAASLIRVPTEVVKQRMQTGQFTS 170
Query: 646 VGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELEAWETI 705
A+ ++G +G + G + L R++PF +Y + G +K REL E
Sbjct: 171 APSAVRMIASKEGFRGLYAGYRSFLLRDLPFDAIQFCIYEQLCLGYKKAARRELSDPENA 230
Query: 706 AVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIV--AFSILRHEGPLGLFKGAVPRF 763
+GA +G L VTTP DV+KTR+M IV +I+R EG L KG PR
Sbjct: 231 LIGAFAGALTGAVTTPLDVIKTRLMVQGSAKQYQGIVDCVQTIVREEGAPALLKGIGPRV 290
Query: 764 FWIAPLGAMNFAGYELARKAM 784
WI G++ F E ++ +
Sbjct: 291 LWIGIGGSIFFGVLESTKRTL 311
Score = 35.4 bits (80), Expect = 0.13
Identities = 28/89 (31%), Positives = 43/89 (47%), Gaps = 6/89 (6%)
Query: 524 SALAGGLSCALSCALLHPVDSIKTR--VQASSMSFPEII----AKLPEIGTRGLYRGSIP 577
+AL G + AL+ A+ P+D IKTR VQ S+ + I+ + E G L +G P
Sbjct: 229 NALIGAFAGALTGAVTTPLDVIKTRLMVQGSAKQYQGIVDCVQTIVREEGAPALLKGIGP 288
Query: 578 AILGQFSSHGLRTGIFEASKLVLVNVAPN 606
+L + G+ E++K L PN
Sbjct: 289 RVLWIGIGGSIFFGVLESTKRTLAQRRPN 317
Score = 29.3 bits (64), Expect = 9.1
Identities = 24/94 (25%), Positives = 39/94 (40%), Gaps = 16/94 (17%)
Query: 697 RELEAWETIAVGALSGGLAAVVTT----PFDVMKTRMMTAQGRSVSMSIVAFSILRHEGP 752
+ + + T+ G ++GG A VV P D +KTR+ A+G IV
Sbjct: 46 KPFDFFRTLFEGFIAGGTAGVVVETALYPIDTIKTRLQAARGGG---KIVL--------- 93
Query: 753 LGLFKGAVPRFFWIAPLGAMNFAGYELARKAMNK 786
GL+ G + P A+ YE ++ + K
Sbjct: 94 KGLYSGLAGNIAGVLPASALFVGVYEPTKQKLLK 127
>At1g34065 allinase like protein
Length = 327
Score = 124 bits (310), Expect = 3e-28
Identities = 87/275 (31%), Positives = 144/275 (51%), Gaps = 18/275 (6%)
Query: 524 SALAGGLSCALSCALLHPVDSIKTRVQASSMSFPEI-IAKLPEIGTRGLYRGSIPAILGQ 582
S + GGL+ + A L+P+D+IKTR+Q S P+ + + GLY G ++G
Sbjct: 57 SLITGGLAGVVVEAALYPIDTIKTRIQVVS---PQTNFSGWWKDNMEGLYSGLGGNLVGV 113
Query: 583 FSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAV----RIPCEVLKQRL 638
+ L G++E +K L+ V P+ + ++A + LG AV R+P EV+KQR+
Sbjct: 114 LPASALFFGVYEPTKQKLLKVLPD----NLSAVAHLAAGALGGAVSSIVRVPTEVVKQRM 169
Query: 639 QAGLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRE 698
Q G F + +A+ ++G G + G G+ L R++PF +Y + + G + R+
Sbjct: 170 QTGQFVSAPDAVRLIIAKEGFGGMYAGYGSFLLRDLPFDALQFCVYEQLRIGYKLAARRD 229
Query: 699 LEAWETIAVGALSGGLAAVVTTPFDVMKTRMMT----AQGRSVSMSIVAFSILRHEGPLG 754
L E +GA +G + V+TTP DV+KTR+M Q + VS I +I+R EG
Sbjct: 230 LNDPENAMIGAFAGAVTGVLTTPLDVIKTRLMVQGSGTQYKGVSDCIK--TIIREEGSSA 287
Query: 755 LFKGAVPRFFWIAPLGAMNFAGYELARKAMNKNDE 789
L+KG PR WI G++ F E ++ +++ +
Sbjct: 288 LWKGMGPRVLWIGIGGSIFFGVLEKTKQILSERSQ 322
Score = 30.4 bits (67), Expect = 4.1
Identities = 20/85 (23%), Positives = 36/85 (41%), Gaps = 6/85 (7%)
Query: 702 WETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFSILRHEGPLGLFKGAVP 761
+E++ G L+G + P D +KTR+ ++ FS + GL+ G
Sbjct: 55 YESLITGGLAGVVVEAALYPIDTIKTRIQVVSPQT------NFSGWWKDNMEGLYSGLGG 108
Query: 762 RFFWIAPLGAMNFAGYELARKAMNK 786
+ P A+ F YE ++ + K
Sbjct: 109 NLVGVLPASALFFGVYEPTKQKLLK 133
>At5g42130 mitochondrial carrier protein-like
Length = 412
Score = 114 bits (286), Expect = 2e-25
Identities = 81/272 (29%), Positives = 126/272 (45%), Gaps = 18/272 (6%)
Query: 527 AGGLSCALSCALLHPVDSIKTRVQASSMS------FPEIIAKLPEIGTRGLYRGSIPAIL 580
AGGL+ A + L P+D+IKT++Q S F I+ G G Y G I+
Sbjct: 120 AGGLAGAFTYVTLLPLDAIKTKLQTKGASQVYSNTFDAIVKTFQAKGILGFYSGVSAVIV 179
Query: 581 GQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVLKQRLQA 640
G S + G E K +L P+ P + + A + +A+ +P E++ QR+QA
Sbjct: 180 GSTFSSAVYFGTCEFGKSLLSKF-PDFPTVLIPPTAGAMGNIISSAIMVPKELITQRMQA 238
Query: 641 GLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGV-QKLLGREL 699
G + L+ ++DG+ G + G ATL R +P V + K V +K L
Sbjct: 239 GASGRSYQVLLKILEKDGILGLYAGYSATLLRNLPAGVLSYSSFEYLKAAVLEKTKQSHL 298
Query: 700 EAWETIAVGALSGGLAAVVTTPFDVMKTRMMT----------AQGRSVSMSIVAFSILRH 749
E +++ GAL+G ++A +TTP DV+KTR+MT ++ IL
Sbjct: 299 EPLQSVCCGALAGAISASITTPLDVVKTRLMTQIHVEAVDKLGGAMYTGVAGTVKQILTE 358
Query: 750 EGPLGLFKGAVPRFFWIAPLGAMNFAGYELAR 781
EG +G +G PR A A+ + +E AR
Sbjct: 359 EGWVGFTRGMGPRVVHSACFSAIGYFAFETAR 390
Score = 49.7 bits (117), Expect = 6e-06
Identities = 45/182 (24%), Positives = 83/182 (44%), Gaps = 29/182 (15%)
Query: 629 IPCEVLKQRLQ----AGLFNNVGEALVGTWQQDGLKGFFRGT-----GATLCREVPFYVA 679
+P + +K +LQ + +++N +A+V T+Q G+ GF+ G G+T V F
Sbjct: 133 LPLDAIKTKLQTKGASQVYSNTFDAIVKTFQAKGILGFYSGVSAVIVGSTFSSAVYFGTC 192
Query: 680 GMGLYAESKKGVQKLLGRELEAWETIAV----GALSGGLAAVVTTPFDVMKTRMMT-AQG 734
G K L + + T+ + GA+ +++ + P +++ RM A G
Sbjct: 193 EFG----------KSLLSKFPDFPTVLIPPTAGAMGNIISSAIMVPKELITQRMQAGASG 242
Query: 735 RSVSMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAMNKNDEAKTGN 794
RS V IL +G LGL+ G P G ++++ +E + A+ ++ K +
Sbjct: 243 RSYQ---VLLKILEKDGILGLYAGYSATLLRNLPAGVLSYSSFEYLKAAV--LEKTKQSH 297
Query: 795 LE 796
LE
Sbjct: 298 LE 299
Score = 35.8 bits (81), Expect = 0.097
Identities = 30/109 (27%), Positives = 50/109 (45%), Gaps = 9/109 (8%)
Query: 685 AESKKGVQKLLGRELEAWETIAVGALSGGLAAVVT----TPFDVMKTRMMTAQGRSVSMS 740
+ S +Q L+ ++L WE +GA +GGLA T P D +KT++ T +G S S
Sbjct: 95 SRSSPRIQTLI-KQLSVWERAIIGAGAGGLAGAFTYVTLLPLDAIKTKLQT-KGASQVYS 152
Query: 741 IVAFSILR---HEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAMNK 786
+I++ +G LG + G A+ F E + ++K
Sbjct: 153 NTFDAIVKTFQAKGILGFYSGVSAVIVGSTFSSAVYFGTCEFGKSLLSK 201
>At5g61810 peroxisomal Ca-dependent solute carrier - like protein
Length = 478
Score = 109 bits (272), Expect = 7e-24
Identities = 113/475 (23%), Positives = 209/475 (43%), Gaps = 35/475 (7%)
Query: 336 TKDHPDKKK--------LFSVQDFFRYTESEGRRFFEELDRDGDGQVTLEDLEIAMRRRK 387
+K +P KK L ++++ E ++ FE D G + +E +
Sbjct: 6 SKQNPGKKPVEATMEHVLVALRETKEKREIRIQKLFEFFDNSKLGFLDDTQIEKGLSSLS 65
Query: 388 LPR--RYAKEFMSRTRSHLFSRSFGWKQFLSFMEQKEPTILRAYTSLCLTKSGTLKKSEI 445
+P RYA +F+ S+ R +++F +M+ KE + + + ++ + +G + +E+
Sbjct: 66 IPPKYRYASDFLKVCDSNRDGR-VDYQEFRRYMDAKELELYKIFQAIDIEHNGDICPAEL 124
Query: 446 LESLKNSGLPANEDNAAAMMRFLNADTEESISYGHFRNFMLLLPSDRLQEDPRSIWFEAA 505
E+L +G+ ++ A+ M ++ D I++ +R+F+LL P + E+ W E
Sbjct: 125 WEALDKAGIKIKDEELASFMEHVDKDNNGIITFEEWRDFLLLNPHEATIENIYHHW-ERV 183
Query: 506 TVVAVPPSVEIPAGSVLRSA-----LAGGLSCALSCALLHPVDSIKT--RVQASSMSFPE 558
++ + IP G + LAGG++ A+S P+D +K +VQ +++
Sbjct: 184 CLIDIGEQAVIPDGISAHAQRSKLLLAGGIAGAVSRTATAPLDRLKVALQVQRTNLGVVP 243
Query: 559 IIAKL-PEIGTRGLYRGSIPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIAS 617
I K+ E G +RG+ + ++ +E K ++ ++
Sbjct: 244 TIKKIWREDKLLGFFRGNGLNVAKVAPESAIKFAAYEMLKPIIGGADGDIGTSGRLLAGG 303
Query: 618 FCSTFLGTAVRIPCEVLKQRLQAGLFNNVGEALV-----GTWQQDGLKGFFRGTGATLCR 672
TA+ P +++K RLQ + VG + W Q+G + F+RG +L
Sbjct: 304 LAGAVAQTAI-YPMDLVKTRLQT-FVSEVGTPKLWKLTKDIWIQEGPRAFYRGLCPSLIG 361
Query: 673 EVPFYVAGMGLYA-ESKKGVQK--LLGRELEAWETIAVGA--LSGGLAAVVTTPFDVMKT 727
+P+ AG+ L A E+ K + + L E I +G SG L A P V++T
Sbjct: 362 IIPY--AGIDLAAYETLKDLSRAHFLHDTAEPGPLIQLGCGMTSGALGASCVYPLQVIRT 419
Query: 728 RMMTAQGRSVSMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARK 782
RM A SM LR EG G ++G P FF + P ++++ YE +K
Sbjct: 420 RMQ-ADSSKTSMGQEFLKTLRGEGLKGFYRGIFPNFFKVIPSASISYLVYEAMKK 473
Score = 63.5 bits (153), Expect = 4e-10
Identities = 50/178 (28%), Positives = 87/178 (48%), Gaps = 14/178 (7%)
Query: 526 LAGGLSCALSCALLHPVDSIKTRVQA--SSMSFPEIIAKLPEI----GTRGLYRGSIPAI 579
LAGGL+ A++ ++P+D +KTR+Q S + P++ +I G R YRG P++
Sbjct: 300 LAGGLAGAVAQTAIYPMDLVKTRLQTFVSEVGTPKLWKLTKDIWIQEGPRAFYRGLCPSL 359
Query: 580 LGQFSSHGLRTGIFEASKLV-----LVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVL 634
+G G+ +E K + L + A P +Q+ S LG + P +V+
Sbjct: 360 IGIIPYAGIDLAAYETLKDLSRAHFLHDTAEPGPLIQLG--CGMTSGALGASCVYPLQVI 417
Query: 635 KQRLQAGLFN-NVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGV 691
+ R+QA ++G+ + T + +GLKGF+RG + +P +Y KK +
Sbjct: 418 RTRMQADSSKTSMGQEFLKTLRGEGLKGFYRGIFPNFFKVIPSASISYLVYEAMKKNL 475
Score = 35.4 bits (80), Expect = 0.13
Identities = 21/84 (25%), Positives = 38/84 (45%), Gaps = 1/84 (1%)
Query: 705 IAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFSILRHEGPLGLFKGAVPRFF 764
+ G ++G ++ T P D +K + Q ++ + I R + LG F+G
Sbjct: 208 LLAGGIAGAVSRTATAPLDRLKVALQV-QRTNLGVVPTIKKIWREDKLLGFFRGNGLNVA 266
Query: 765 WIAPLGAMNFAGYELARKAMNKND 788
+AP A+ FA YE+ + + D
Sbjct: 267 KVAPESAIKFAAYEMLKPIIGGAD 290
>At5g51050 calcium-binding transporter-like protein
Length = 487
Score = 104 bits (259), Expect = 2e-22
Identities = 103/447 (23%), Positives = 201/447 (44%), Gaps = 26/447 (5%)
Query: 360 RRFFEELDRDGDGQVTLEDLEIAMRRRKLPR--RYAKEFMSRTRSHLFSRSFGWKQFLSF 417
R F D + G + +E + ++P +YAKE ++ R + +F +
Sbjct: 42 RSLFSFFDSENVGYLDCAQIEKGLCALQIPSGYKYAKELFRVCDANRDGR-VDYHEFRRY 100
Query: 418 MEQKEPTILRAYTSLCLTKSGTLKKSEILESLKNSGLPANEDNAAAMMRFLNADTEESIS 477
M+ KE + R + ++ + +G + + +SL +G+ ++ A + ++ D + I
Sbjct: 101 MDDKELELYRIFQAIDVEHNGCISPEGLWDSLVKAGIEIKDEELARFVEHVDKDNDGIIM 160
Query: 478 YGHFRNFMLLLPSDRLQEDPRSIWFEAATVVAVPPSVEIPAG---SVLRSA--LAGGLSC 532
+ +R+F+LL P + E+ W E +V + IP G + RS +AGG++
Sbjct: 161 FEEWRDFLLLYPHEATIENIYHHW-ERVCLVDIGEQAVIPEGISKHIKRSNYFIAGGIAG 219
Query: 533 ALSCALLHPVDSIKT--RVQASSMSFPEIIAKL-PEIGTRGLYRGSIPAILGQFSSHGLR 589
A S P+D +K ++Q + E I + + G RG +RG+ I+ ++
Sbjct: 220 AASRTATAPLDRLKVLLQIQKTDARIREAIKLIWKQGGVRGFFRGNGLNIVKVAPESAIK 279
Query: 590 TGIFEASKLVL-VNVAPNLPEL--QVQSIASFCSTFLGTAVRIPCEVLKQRLQAGLFNNV 646
+E K + N+ + ++ V+ A + + A P +++K RLQ +
Sbjct: 280 FYAYELFKNAIGENMGEDKADIGTTVRLFAGGMAGAVAQASIYPLDLVKTRLQT-YTSQA 338
Query: 647 GEAL--VGTWQQD-----GLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGREL 699
G A+ +GT +D G + F++G +L +P+ + Y K + + ++
Sbjct: 339 GVAVPRLGTLTKDILVHEGPRAFYKGLFPSLLGIIPYAGIDLAAYETLKDLSRTYILQDA 398
Query: 700 EAWETIAVGA--LSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFSILRHEGPLGLFK 757
E + +G +SG L A P V++TRM + R+ SMS V + EG L+K
Sbjct: 399 EPGPLVQLGCGTISGALGATCVYPLQVVRTRMQAERART-SMSGVFRRTISEEGYRALYK 457
Query: 758 GAVPRFFWIAPLGAMNFAGYELARKAM 784
G +P + P ++ + YE +K++
Sbjct: 458 GLLPNLLKVVPAASITYMVYEAMKKSL 484
Score = 35.4 bits (80), Expect = 0.13
Identities = 29/116 (25%), Positives = 48/116 (41%), Gaps = 8/116 (6%)
Query: 681 MGLYAESKKGVQKLLGRELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMS 740
+G A +G+ K + R G ++G + T P D +K ++ Q +
Sbjct: 192 IGEQAVIPEGISKHIKRS----NYFIAGGIAGAASRTATAPLDRLKV-LLQIQKTDARIR 246
Query: 741 IVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAMNKN---DEAKTG 793
I + G G F+G +AP A+ F YEL + A+ +N D+A G
Sbjct: 247 EAIKLIWKQGGVRGFFRGNGLNIVKVAPESAIKFYAYELFKNAIGENMGEDKADIG 302
>At2g30160 putative mitochondrial carrier protein
Length = 331
Score = 103 bits (256), Expect = 5e-22
Identities = 86/300 (28%), Positives = 141/300 (46%), Gaps = 36/300 (12%)
Query: 512 PSVEIPAGSVL----RSALAGGLSCALSCALLHPVDSIKTRVQASS----------MSFP 557
P++ +PA + + +AG ++ ++ + PVD++KT +QA +F
Sbjct: 25 PAIIVPAQNTTLKFWQLMVAGSIAGSVEHMAMFPVDTVKTHMQALRSCPIKPIGIRQAFR 84
Query: 558 EIIAKLPEIGTRGLYRGSIPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIAS 617
II G LYRG LG +H + +E SK L PN +I+
Sbjct: 85 SIIKT---DGPSALYRGIWAMGLGAGPAHAVYFSFYEVSKKFLSGGNPN--NSAAHAISG 139
Query: 618 FCSTFLGTAVRIPCEVLKQRLQAG--LFNNVGEALVGTWQQDGLKGFFRGTGATLCREVP 675
+T AV P +++KQRLQ G + V + + +++G F+ T+ P
Sbjct: 140 VFATISSDAVFTPMDMVKQRLQIGNGTYKGVWDCIKRVTREEGFGAFYASYRTTVLMNAP 199
Query: 676 FYVAGMGLYAESKKGVQKLL------GRELEAWETIAV-GALSGGLAAVVTTPFDVMKTR 728
F Y K+G++++L + E W A GA +GGLAA VTTP DV+KT+
Sbjct: 200 FTAVHFTTYEAVKRGLREMLPEHAVGAEDEEGWLIYATAGAAAGGLAAAVTTPLDVVKTQ 259
Query: 729 MMTAQG-------RSVSMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELAR 781
+ QG +S S+S V +I++ +G GL +G +PR + AP A+ ++ YE +
Sbjct: 260 LQ-CQGVCGCDRFKSSSISDVFRTIVKKDGYRGLARGWLPRMLFHAPAAAICWSTYETVK 318
Score = 42.4 bits (98), Expect = 0.001
Identities = 24/98 (24%), Positives = 49/98 (49%), Gaps = 6/98 (6%)
Query: 699 LEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQG---RSVSMSIVAFSILRHEGPLGL 755
L+ W+ + G+++G + + P D +KT M + + + + SI++ +GP L
Sbjct: 36 LKFWQLMVAGSIAGSVEHMAMFPVDTVKTHMQALRSCPIKPIGIRQAFRSIIKTDGPSAL 95
Query: 756 FKGAVPRFFWIAPLGAMNFAGYELARKAM---NKNDEA 790
++G P A+ F+ YE+++K + N N+ A
Sbjct: 96 YRGIWAMGLGAGPAHAVYFSFYEVSKKFLSGGNPNNSA 133
>At5g07320 peroxisomal Ca-dependent solute carrier-like protein
Length = 479
Score = 100 bits (249), Expect = 3e-21
Identities = 98/457 (21%), Positives = 197/457 (42%), Gaps = 21/457 (4%)
Query: 345 LFSVQDFFRYTESEGRRFFEELDRDGDGQVTLEDLEIAMRRRKLPR--RYAKEFMSRTRS 402
L ++++ E R F+ D G + +E + ++P +YA++ +
Sbjct: 24 LLALRETMDEREIRIRSLFDFFDNSNLGFLDYAQIEKGLASLQIPPEYKYARDLFRVCDA 83
Query: 403 HLFSRSFGWKQFLSFMEQKEPTILRAYTSLCLTKSGTLKKSEILESLKNSGLPANEDNAA 462
+ R +++F +++ KE + R + ++ + +G + E+ E+L +G+ +++ A
Sbjct: 84 NRDGR-VDYQEFRRYIDAKELELYRIFQAIDVEHNGCILPEELWEALVKAGIEIDDEELA 142
Query: 463 AMMRFLNADTEESISYGHFRNFMLLLPSDRLQEDPRSIWFEAATVVAVPPSVEIPAG--- 519
+ ++ D +I++ +R+F+LL P + E+ W E ++ + IP G
Sbjct: 143 RFVEHVDKDNNGTITFEEWRDFLLLYPHEATLENIYHHW-ERVCLIDIGEQAVIPDGISK 201
Query: 520 SVLRSAL--AGGLSCALSCALLHPVDSIKTRVQ---ASSMSFPEIIAKLPEIGTRGLYRG 574
V RS L AGGL+ A+S P+D +K +Q A + P I E G +RG
Sbjct: 202 HVKRSRLLLAGGLAGAVSRTATAPLDRLKVVLQVQRAHAGVLPTIKKIWREDKLMGFFRG 261
Query: 575 SIPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVL 634
+ ++ ++ +E K ++ ++ TA+ P +++
Sbjct: 262 NGLNVMKVAPESAIKFCAYEMLKPMIGGEDGDIGTSGRLMAGGMAGALAQTAI-YPMDLV 320
Query: 635 KQRLQA-----GLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKK 689
K RLQ G + + W ++G + F++G +L VP+ + Y K
Sbjct: 321 KTRLQTCVSEGGKAPKLWKLTKDIWVREGPRAFYKGLFPSLLGIVPYAGIDLAAYETLKD 380
Query: 690 GVQKLLGRELEAWETIAV--GALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFSIL 747
+ + ++ E I + G SG L A P V++TRM A +M + +
Sbjct: 381 LSRTYILQDTEPGPLIQLSCGMTSGALGASCVYPLQVVRTRMQ-ADSSKTTMKQEFMNTM 439
Query: 748 RHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAM 784
+ EG G ++G +P + P ++ + YE +K M
Sbjct: 440 KGEGLRGFYRGLLPNLLKVVPAASITYIVYEAMKKNM 476
Score = 35.8 bits (81), Expect = 0.097
Identities = 19/94 (20%), Positives = 41/94 (43%), Gaps = 1/94 (1%)
Query: 695 LGRELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFSILRHEGPLG 754
+ + ++ + G L+G ++ T P D +K + + + + + I R + +G
Sbjct: 199 ISKHVKRSRLLLAGGLAGAVSRTATAPLDRLKVVLQVQRAHAGVLPTIK-KIWREDKLMG 257
Query: 755 LFKGAVPRFFWIAPLGAMNFAGYELARKAMNKND 788
F+G +AP A+ F YE+ + + D
Sbjct: 258 FFRGNGLNVMKVAPESAIKFCAYEMLKPMIGGED 291
>At1g07030 unknown protein
Length = 326
Score = 99.4 bits (246), Expect = 7e-21
Identities = 83/276 (30%), Positives = 131/276 (47%), Gaps = 29/276 (10%)
Query: 526 LAGGLSCALSCALLHPVDSIKTRVQASSM----------SFPEIIAKLPEIGTRGLYRGS 575
+AG ++ ++ + PVD+IKT +QA +F II K G LYRG
Sbjct: 41 IAGSIAGSVEHMAMFPVDTIKTHMQALRPCPLKPVGIREAFRSIIQKE---GPSALYRGI 97
Query: 576 IPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVLK 635
LG +H + +E SK L A + +++ +T AV P +++K
Sbjct: 98 WAMGLGAGPAHAVYFSFYEVSKKYLS--AGDQNNSVAHAMSGVFATISSDAVFTPMDMVK 155
Query: 636 QRLQAG--LFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQK 693
QRLQ G + V + + +++G+ F+ T+ PF Y +KKG+ +
Sbjct: 156 QRLQMGEGTYKGVWDCVKRVLREEGIGAFYASYRTTVLMNAPFTAVHFATYEAAKKGLME 215
Query: 694 LLGREL---EAWETIAV-GALSGGLAAVVTTPFDVMKTRMMTAQG-------RSVSMSIV 742
+ E W A GA +GGLAA VTTP DV+KT++ QG S S+S V
Sbjct: 216 FSPDRISDEEGWLVHATAGAAAGGLAAAVTTPLDVVKTQLQ-CQGVCGCDRFTSSSISHV 274
Query: 743 AFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYE 778
+I++ +G GL +G +PR + AP A+ ++ YE
Sbjct: 275 LRTIVKKDGYRGLLRGWLPRMLFHAPAAAICWSTYE 310
Score = 44.7 bits (104), Expect = 2e-04
Identities = 24/94 (25%), Positives = 48/94 (50%), Gaps = 3/94 (3%)
Query: 699 LEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQG---RSVSMSIVAFSILRHEGPLGL 755
L+ W+ + G+++G + + P D +KT M + + V + SI++ EGP L
Sbjct: 34 LKFWQFMIAGSIAGSVEHMAMFPVDTIKTHMQALRPCPLKPVGIREAFRSIIQKEGPSAL 93
Query: 756 FKGAVPRFFWIAPLGAMNFAGYELARKAMNKNDE 789
++G P A+ F+ YE+++K ++ D+
Sbjct: 94 YRGIWAMGLGAGPAHAVYFSFYEVSKKYLSAGDQ 127
Score = 40.4 bits (93), Expect = 0.004
Identities = 34/139 (24%), Positives = 55/139 (39%), Gaps = 2/139 (1%)
Query: 646 VGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELEAWETI 705
+ EA Q++G +RG A P + Y SKK + G + +
Sbjct: 77 IREAFRSIIQKEGPSALYRGIWAMGLGAGPAHAVYFSFYEVSKKYLSA--GDQNNSVAHA 134
Query: 706 AVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFSILRHEGPLGLFKGAVPRFFW 765
G + + V TP D++K R+ +G + +LR EG +
Sbjct: 135 MSGVFATISSDAVFTPMDMVKQRLQMGEGTYKGVWDCVKRVLREEGIGAFYASYRTTVLM 194
Query: 766 IAPLGAMNFAGYELARKAM 784
AP A++FA YE A+K +
Sbjct: 195 NAPFTAVHFATYEAAKKGL 213
Score = 33.1 bits (74), Expect = 0.63
Identities = 35/157 (22%), Positives = 64/157 (40%), Gaps = 16/157 (10%)
Query: 525 ALAGGLSCALSCALLHPVDSIKTRVQASSMSFPE----IIAKLPEIGTRGLYRGSIPAIL 580
A++G + S A+ P+D +K R+Q ++ + L E G Y +L
Sbjct: 134 AMSGVFATISSDAVFTPMDMVKQRLQMGEGTYKGVWDCVKRVLREEGIGAFYASYRTTVL 193
Query: 581 GQFSSHGLRTGIFEASKLVLVNVAPNLPELQ----VQSIASFCSTFLGTAVRIPCEVLKQ 636
+ +EA+K L+ +P+ + V + A + L AV P +V+K
Sbjct: 194 MNAPFTAVHFATYEAAKKGLMEFSPDRISDEEGWLVHATAGAAAGGLAAAVTTPLDVVKT 253
Query: 637 RLQAG--------LFNNVGEALVGTWQQDGLKGFFRG 665
+LQ +++ L ++DG +G RG
Sbjct: 254 QLQCQGVCGCDRFTSSSISHVLRTIVKKDGYRGLLRG 290
Score = 29.6 bits (65), Expect = 6.9
Identities = 20/66 (30%), Positives = 34/66 (51%), Gaps = 10/66 (15%)
Query: 525 ALAGGLSCALSCALLHPVDSIKTRVQA---------SSMSFPEIIAKL-PEIGTRGLYRG 574
A AG + L+ A+ P+D +KT++Q +S S ++ + + G RGL RG
Sbjct: 231 ATAGAAAGGLAAAVTTPLDVVKTQLQCQGVCGCDRFTSSSISHVLRTIVKKDGYRGLLRG 290
Query: 575 SIPAIL 580
+P +L
Sbjct: 291 WLPRML 296
>At4g32400 adenylate translocator (brittle-1) - like protein
Length = 392
Score = 93.2 bits (230), Expect = 5e-19
Identities = 72/284 (25%), Positives = 137/284 (47%), Gaps = 12/284 (4%)
Query: 518 AGSVLRSALAGGLSCALSCALLHPVDSIKTRVQASS--MSFPEIIAKLPEI-GTRGLYRG 574
A LR L+G ++ A+S ++ P+++I+T + S S E+ + + + G GL+RG
Sbjct: 107 ANPSLRRLLSGAVAGAVSRTVVAPLETIRTHLMVGSGGNSSTEVFSDIMKHEGWTGLFRG 166
Query: 575 SIPAILGQFSSHGLRTGIFEA--SKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCE 632
++ ++ + + +FE KL + + + +A C+ T + P E
Sbjct: 167 NLVNVIRVAPARAVELFVFETVNKKLSPPHGQESKIPIPASLLAGACAGVSQTLLTYPLE 226
Query: 633 VLKQRL--QAGLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKG 690
++K RL Q G++ + +A + +++G +RG +L VP+ Y +K
Sbjct: 227 LVKTRLTIQRGVYKGIFDAFLKIIREEGPTELYRGLAPSLIGVVPYAATNYFAYDSLRKA 286
Query: 691 VQKLLGRE-LEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTA--QGRSV--SMSIVAFS 745
+ +E + ET+ +G+L+G L++ T P +V + M GR V +M +
Sbjct: 287 YRSFSKQEKIGNIETLLIGSLAGALSSTATFPLEVARKHMQVGAVSGRVVYKNMLHALVT 346
Query: 746 ILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAMNKNDE 789
IL HEG LG +KG P + P ++F YE +K + +N++
Sbjct: 347 ILEHEGILGWYKGLGPSCLKLVPAAGISFMCYEACKKILIENNQ 390
Score = 66.6 bits (161), Expect = 5e-11
Identities = 50/196 (25%), Positives = 84/196 (42%), Gaps = 8/196 (4%)
Query: 605 PNLPELQVQSIASFCSTFLGTAVRIPCEVLKQRLQAGLFNNVG-EALVGTWQQDGLKGFF 663
P+L L ++A S V P E ++ L G N E + +G G F
Sbjct: 109 PSLRRLLSGAVAGAVSR----TVVAPLETIRTHLMVGSGGNSSTEVFSDIMKHEGWTGLF 164
Query: 664 RGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRE--LEAWETIAVGALSGGLAAVVTTP 721
RG + R P + ++ K + G+E + ++ GA +G ++T P
Sbjct: 165 RGNLVNVIRVAPARAVELFVFETVNKKLSPPHGQESKIPIPASLLAGACAGVSQTLLTYP 224
Query: 722 FDVMKTRMMTAQGRSVSMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELAR 781
+++KTR+ +G + I+R EGP L++G P + P A N+ Y+ R
Sbjct: 225 LELVKTRLTIQRGVYKGIFDAFLKIIREEGPTELYRGLAPSLIGVVPYAATNYFAYDSLR 284
Query: 782 KAMNK-NDEAKTGNLE 796
KA + + K GN+E
Sbjct: 285 KAYRSFSKQEKIGNIE 300
Score = 51.6 bits (122), Expect = 2e-06
Identities = 50/206 (24%), Positives = 83/206 (40%), Gaps = 25/206 (12%)
Query: 502 FEAATVVAVPPSVEIPAGSVLRSALAGGLSCALSCALLHPVDSIKTRVQAS----SMSFP 557
FE PP + + S LAG + L +P++ +KTR+ F
Sbjct: 185 FETVNKKLSPPHGQESKIPIPASLLAGACAGVSQTLLTYPLELVKTRLTIQRGVYKGIFD 244
Query: 558 EIIAKLPEIGTRGLYRGSIPAILG--------QFSSHGLRTGIFEASKLVLVNVAPNLPE 609
+ + E G LYRG P+++G F+ LR SK + N+
Sbjct: 245 AFLKIIREEGPTELYRGLAPSLIGVVPYAATNYFAYDSLRKAYRSFSKQEKIG---NIET 301
Query: 610 LQVQSIASFCSTFLGTAVRIPCEVLKQRLQAG------LFNNVGEALVGTWQQDGLKGFF 663
L + S+A L + P EV ++ +Q G ++ N+ ALV + +G+ G++
Sbjct: 302 LLIGSLAG----ALSSTATFPLEVARKHMQVGAVSGRVVYKNMLHALVTILEHEGILGWY 357
Query: 664 RGTGATLCREVPFYVAGMGLYAESKK 689
+G G + + VP Y KK
Sbjct: 358 KGLGPSCLKLVPAAGISFMCYEACKK 383
>At3g54110 uncoupling protein (ucp/PUMP)
Length = 306
Score = 82.8 bits (203), Expect = 7e-16
Identities = 70/275 (25%), Positives = 122/275 (43%), Gaps = 25/275 (9%)
Query: 541 PVDSIKTRVQ------ASSMSFPEIIAKLPEIGT-------RGLYRGSIPAILGQFSSHG 587
P+D+ K R+Q A ++ P+ L +GT R L++G +P + Q G
Sbjct: 31 PLDTAKVRLQLQKSALAGDVTLPKYRGLLGTVGTIAREEGLRSLWKGVVPGLHRQCLFGG 90
Query: 588 LRTGIFEASKLVLV--NVAPNLPELQVQSIASFCSTFLGTAVRIPCEVLKQRLQAG---- 641
LR G++E K + V + ++P L + +A + LG V P +++K RLQA
Sbjct: 91 LRIGMYEPVKNLYVGKDFVGDVP-LSKKILAGLTTGALGIMVANPTDLVKVRLQAEGKLA 149
Query: 642 -----LFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLG 696
++ A +Q+G++ + G G + R A + Y + K+ + K+ G
Sbjct: 150 AGAPRRYSGALNAYSTIVRQEGVRALWTGLGPNVARNAIINAAELASYDQVKETILKIPG 209
Query: 697 RELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFSILRHEGPLGLF 756
I G +G A + +P DV+K+RMM G L+ +GP+ +
Sbjct: 210 FTDNVVTHILSGLGAGFFAVCIGSPVDVVKSRMMGDSGAYKGTIDCFVKTLKSDGPMAFY 269
Query: 757 KGAVPRFFWIAPLGAMNFAGYELARKAMNKNDEAK 791
KG +P F + + F E A+K + + D +K
Sbjct: 270 KGFIPNFGRLGSWNVIMFLTLEQAKKYVRELDASK 304
Score = 36.6 bits (83), Expect = 0.057
Identities = 30/156 (19%), Positives = 61/156 (38%), Gaps = 13/156 (8%)
Query: 523 RSALAGGLSCALSCALLHPVDSIKTRVQAS-----------SMSFPEIIAKLPEIGTRGL 571
+ LAG + AL + +P D +K R+QA S + + + G R L
Sbjct: 116 KKILAGLTTGALGIMVANPTDLVKVRLQAEGKLAAGAPRRYSGALNAYSTIVRQEGVRAL 175
Query: 572 YRGSIPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPC 631
+ G P + + ++ K ++ + + ++ + F + P
Sbjct: 176 WTGLGPNVARNAIINAAELASYDQVKETILKIPGFTDNVVTHILSGLGAGFFAVCIGSPV 235
Query: 632 EVLKQRL--QAGLFNNVGEALVGTWQQDGLKGFFRG 665
+V+K R+ +G + + V T + DG F++G
Sbjct: 236 DVVKSRMMGDSGAYKGTIDCFVKTLKSDGPMAFYKG 271
>At3g20240 mitochondrial carrier protein, putative
Length = 348
Score = 82.0 bits (201), Expect = 1e-15
Identities = 67/284 (23%), Positives = 132/284 (45%), Gaps = 28/284 (9%)
Query: 523 RSALAGGLSCALSCALLHPVDSIKTR--VQASSMSFP-EIIAKLPEIGTRGLYRGSIPAI 579
R L+G L+ A++ A+L P+++I+TR V S S P + + + G +GL+ G+ +
Sbjct: 50 REFLSGALAGAMTKAVLAPLETIRTRMIVGVGSRSIPGSFLEVVQKQGWQGLWAGNEINM 109
Query: 580 LGQFSSHGLRTGIFEASKLVLVNV-------------------APNLPELQVQSIASFCS 620
+ + + G FE K + + +P++ + ++A +
Sbjct: 110 IRIIPTQAIELGTFEWVKRAMTSAQVKLKKIEDAKIEIGDFSFSPSISWISPVAVAGASA 169
Query: 621 TFLGTAVRIPCEVLKQRLQAG--LFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYV 678
T V P EVLK RL ++ ++ A+ ++ DG++GF+ G G TL +P+
Sbjct: 170 GIASTLVCHPLEVLKDRLTVSPEIYPSLSLAIPRIFRADGIRGFYAGLGPTLVGMLPYST 229
Query: 679 AGMGLYAESKKGVQKLLGRE-LEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRS- 736
+Y + K K ++ L E + +GAL+G A+ ++ P +V + R+M +
Sbjct: 230 CYYFMYDKMKTSYCKSKNKKALSRPEMLVLGALAGLTASTISFPLEVARKRLMVGALKGE 289
Query: 737 --VSMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYE 778
+M+ +++ EG +GL++G + P + + YE
Sbjct: 290 CPPNMAAAIAEVVKKEGVMGLYRGWGASCLKVMPSSGITWVFYE 333
Score = 37.7 bits (86), Expect = 0.025
Identities = 23/78 (29%), Positives = 40/78 (50%), Gaps = 3/78 (3%)
Query: 708 GALSGGLAAVVTTPFDVMKTRMMTAQG-RSVSMSIVAFSILRHEGPLGLFKGAVPRFFWI 766
GAL+G + V P + ++TRM+ G RS+ S + +++ +G GL+ G I
Sbjct: 55 GALAGAMTKAVLAPLETIRTRMIVGVGSRSIPGSFL--EVVQKQGWQGLWAGNEINMIRI 112
Query: 767 APLGAMNFAGYELARKAM 784
P A+ +E ++AM
Sbjct: 113 IPTQAIELGTFEWVKRAM 130
>At5g15640 putative mitochondrial carrier protein
Length = 323
Score = 76.6 bits (187), Expect = 5e-14
Identities = 74/276 (26%), Positives = 116/276 (41%), Gaps = 32/276 (11%)
Query: 539 LHPVDSIKTRVQASSMSFPE------IIAKLPEIGTRGLYRGSIPAILGQFSSH-----G 587
L+PV +KTR+Q +S E + L G GLYRG I G +
Sbjct: 42 LYPVSVVKTRLQVASKEIAERSAFSVVKGILKNDGVPGLYRGFGTVITGAVPARIIFLTA 101
Query: 588 LRTGIFEASKLVL-VNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVLKQRLQ----AGL 642
L T A KLV + ++ IA ++ AV +P +V+ Q+L +G
Sbjct: 102 LETTKISAFKLVAPLELSEPTQAAIANGIAGMTASLFSQAVFVPIDVVSQKLMVQGYSGH 161
Query: 643 FNNVGEALVGTW--QQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELE 700
G V T + G++G +RG G ++ P A Y S++ + + LG +
Sbjct: 162 ATYTGGIDVATKIIKSYGVRGLYRGFGLSVMTYSPSSAAWWASYGSSQRVIWRFLGYGGD 221
Query: 701 AWETIAV------------GALSGGLAAVVTTPFDVMKTRM--MTAQGRSVSMSIVAFSI 746
+ T A G ++G A+ +TTP D +KTR+ M Q S V +
Sbjct: 222 SDATAAPSKSKIVMVQAAGGIIAGATASSITTPLDTIKTRLQVMGHQENRPSAKQVVKKL 281
Query: 747 LRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARK 782
L +G G ++G PRFF ++ G YE ++
Sbjct: 282 LAEDGWKGFYRGLGPRFFSMSAWGTSMILTYEYLKR 317
Score = 38.5 bits (88), Expect = 0.015
Identities = 40/188 (21%), Positives = 77/188 (40%), Gaps = 24/188 (12%)
Query: 504 AATVVAVPPSVEIPAGSVLRSALAGGLSCALSCALLHPVDSIKTRVQASSMSFP------ 557
+A + P + P + + + +AG + S A+ P+D + ++ S
Sbjct: 108 SAFKLVAPLELSEPTQAAIANGIAGMTASLFSQAVFVPIDVVSQKLMVQGYSGHATYTGG 167
Query: 558 -EIIAK-LPEIGTRGLYRGSIPAILGQFSSHGLRTGIFEASKLVL-----------VNVA 604
++ K + G RGLYRG +++ S + +S+ V+ A
Sbjct: 168 IDVATKIIKSYGVRGLYRGFGLSVMTYSPSSAAWWASYGSSQRVIWRFLGYGGDSDATAA 227
Query: 605 PNLPEL-QVQSIASFCSTFLGTAVRIPCEVLKQRLQA-GLFNN---VGEALVGTWQQDGL 659
P+ ++ VQ+ + +++ P + +K RLQ G N + + +DG
Sbjct: 228 PSKSKIVMVQAAGGIIAGATASSITTPLDTIKTRLQVMGHQENRPSAKQVVKKLLAEDGW 287
Query: 660 KGFFRGTG 667
KGF+RG G
Sbjct: 288 KGFYRGLG 295
Score = 31.6 bits (70), Expect = 1.8
Identities = 21/63 (33%), Positives = 35/63 (55%), Gaps = 6/63 (9%)
Query: 521 VLRSALAGGLSCALSCALLHPVDSIKTRVQA-----SSMSFPEIIAK-LPEIGTRGLYRG 574
V+ A G ++ A + ++ P+D+IKTR+Q + S +++ K L E G +G YRG
Sbjct: 234 VMVQAAGGIIAGATASSITTPLDTIKTRLQVMGHQENRPSAKQVVKKLLAEDGWKGFYRG 293
Query: 575 SIP 577
P
Sbjct: 294 LGP 296
>At2g33820 mitochondrial basic amino acid carrier (BAC1)
Length = 311
Score = 75.5 bits (184), Expect = 1e-13
Identities = 76/290 (26%), Positives = 125/290 (42%), Gaps = 33/290 (11%)
Query: 526 LAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAK---------LPEIGTRGLYRGSI 576
+AG ++ + A+ HP D++K ++Q + + K L G +GLYRG+
Sbjct: 19 VAGMMAGLATVAVGHPFDTVKVKLQKHNTDVQGLRYKNGLHCASRILQTEGVKGLYRGAT 78
Query: 577 PAILGQFSSHGLRTGIFEASKLVLVNVAPN---LPELQVQSIASFCSTFLGTAVRIPCEV 633
+ +G L GI+ +KL L P+ PE+ V S A F + + V P E+
Sbjct: 79 SSFMGMAFESSLMFGIYSQAKLFLRGTLPDDGPRPEIIVPS-AMFGGAII-SFVLCPTEL 136
Query: 634 LKQRLQA----------GLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGL 683
+K R+Q +N+ + V T + DG+ G FRG ATL RE +
Sbjct: 137 VKCRMQIQGTDSLVPNFRRYNSPLDCAVQTVKNDGVTGIFRGGSATLLRECTGNAVFFTV 196
Query: 684 YAESKKGV-QKLLGRELEAWETI--AVGALSGGLAAV----VTTPFDVMKTRMMTAQGRS 736
Y + + +L +L+ + +G L+GGL + PFDV KT + T+ ++
Sbjct: 197 YEYLRYHIHSRLEDSKLKDGYLVDMGIGVLTGGLGGIACWSAVLPFDVAKTIIQTSSEKA 256
Query: 737 VSMS--IVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAM 784
+ V SI + G G + G P P A +E + K +
Sbjct: 257 TERNPFKVLSSIHKRAGLKGCYAGLGPTIVRAFPANAAAIVAWEFSMKML 306
>At5g01500 unknown protein
Length = 415
Score = 69.7 bits (169), Expect = 6e-12
Identities = 43/145 (29%), Positives = 76/145 (51%), Gaps = 2/145 (1%)
Query: 648 EALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELEAWETIAV 707
EA+ +++G+KG+++G + R VP+ + Y KK + G +L +
Sbjct: 163 EAITLIGKEEGIKGYWKGNLPQVIRIVPYSAVQLFAYETYKKLFRGKDG-QLSVLGRLGA 221
Query: 708 GALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFSILRHEGPLGLFKGAVPRFFWIA 767
GA +G + ++T P DV++ R+ G +MS VA ++LR EG + G P IA
Sbjct: 222 GACAGMTSTLITYPLDVLRLRLAVEPGYR-TMSQVALNMLREEGVASFYNGLGPSLLSIA 280
Query: 768 PLGAMNFAGYELARKAMNKNDEAKT 792
P A+NF ++L +K++ + + KT
Sbjct: 281 PYIAINFCVFDLVKKSLPEKYQQKT 305
Score = 62.4 bits (150), Expect = 1e-09
Identities = 55/277 (19%), Positives = 121/277 (42%), Gaps = 18/277 (6%)
Query: 527 AGGLSCALSCALLHPVDSIKTRVQA-----------SSMSFPEIIAKL-PEIGTRGLYRG 574
AG + A + ++ P+D IK +Q ++ F E I + E G +G ++G
Sbjct: 121 AGAFAGAAAKSVTAPLDRIKLLMQTHGVRAGQQSAKKAIGFIEAITLIGKEEGIKGYWKG 180
Query: 575 SIPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVL 634
++P ++ ++ +E K + L L A C+ T + P +VL
Sbjct: 181 NLPQVIRIVPYSAVQLFAYETYKKLFRGKDGQLSVLGRLG-AGACAGMTSTLITYPLDVL 239
Query: 635 KQRLQAGL-FNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQK 693
+ RL + + + + +++G+ F+ G G +L P+ ++ KK + +
Sbjct: 240 RLRLAVEPGYRTMSQVALNMLREEGVASFYNGLGPSLLSIAPYIAINFCVFDLVKKSLPE 299
Query: 694 LLGRELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFS-ILRHEGP 752
++ ++ ++ ++ +A P D ++ R M +G + AFS I+ EG
Sbjct: 300 KYQQKTQS--SLLTAVVAAAIATGTCYPLDTIR-RQMQLKGTPYKSVLDAFSGIIAREGV 356
Query: 753 LGLFKGAVPRFFWIAPLGAMNFAGYELARKAMNKNDE 789
+GL++G VP P ++ +++ +K + +++
Sbjct: 357 VGLYRGFVPNALKSMPNSSIKLTTFDIVKKLIAASEK 393
>At3g21390 unknown protein
Length = 335
Score = 69.7 bits (169), Expect = 6e-12
Identities = 76/308 (24%), Positives = 119/308 (37%), Gaps = 52/308 (16%)
Query: 525 ALAGGLSCALSCALLHPVDSIKTRVQAS-------SMSFPEIIAK-----------LPEI 566
A AGG++ A+S + P+D IK R Q ++ ++ K E
Sbjct: 19 ASAGGVAGAISRMVTSPLDVIKIRFQVQLEPTATWALKDSQLKPKYNGLFRTTKDIFREE 78
Query: 567 GTRGLYRGSIPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLG-- 624
G G +RG++PA+L ++ + K + Q+ S+ S L
Sbjct: 79 GLSGFWRGNVPALLMVVPYTSIQFAVLHKVKSFAAGSSKAENHAQLSPYLSYISGALAGC 138
Query: 625 --TAVRIPCEVLKQRL----QAGLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYV 678
T P ++L+ L + ++ N+ A + Q G+KG + G TL +P+
Sbjct: 139 AATVGSYPFDLLRTVLASQGEPKVYPNMRSAFLSIVQTRGIKGLYAGLSPTLIEIIPYAG 198
Query: 679 AGMGLYAESKKGVQKLLGR-------------ELEAWETIAVGALSGGLAAVVTTPFDVM 725
G Y K+ R L +++ G SG ++ +V P DV+
Sbjct: 199 LQFGTYDTFKRWSMVYNKRYRSSSSSSTNPSDSLSSFQLFLCGLASGTVSKLVCHPLDVV 258
Query: 726 KTRMMTA-------QGRSVSMSIVAF------SILRHEGPLGLFKGAVPRFFWIAPLGAM 772
K R G V ++ ILR EG GL+KG VP AP GA+
Sbjct: 259 KKRFQVEGLQRHPKYGARVELNAYKNMFDGLGQILRSEGWHGLYKGIVPSTIKAAPAGAV 318
Query: 773 NFAGYELA 780
F YELA
Sbjct: 319 TFVAYELA 326
>At2g17270 putative mitochondrial phosphate translocator protein
Length = 309
Score = 69.7 bits (169), Expect = 6e-12
Identities = 60/268 (22%), Positives = 118/268 (43%), Gaps = 21/268 (7%)
Query: 526 LAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAK----LPEIGTRGLYRGSIPAILG 581
+ G LS + + P+D +K +Q + + + I + L E G L+RG +LG
Sbjct: 23 MGGMLSAGTTHLAITPLDVLKVNMQVNPVKYNSIPSGFSTLLREHGHSYLWRGWSGKLLG 82
Query: 582 QFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVLKQRLQAG 641
G R G++E K + +V PN + ++S + P E +K R+Q
Sbjct: 83 YGVQGGCRFGLYEYFKTLYSDVLPNHNRTSIYFLSSASAQIFADMALCPFEAIKVRVQTQ 142
Query: 642 LFNNVG--EALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGV-QKLL--- 695
G + ++ +GL GF RG CR +PF + + +S + + QK++
Sbjct: 143 PMFAKGLLDGFPRVYRSEGLAGFHRGLFPLWCRNLPFSMVMFSTFEQSVEFIYQKIIQKR 202
Query: 696 ----GRELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFSILRHEG 751
+ + T G +G + +++ P DV+ + + + ++V ++ R+ G
Sbjct: 203 KQDCSKAQQLGVTCLAGYTAGAVGTIISNPADVVLSSLYNNKAKNVLQAV------RNIG 256
Query: 752 PLGLFKGAVP-RFFWIAPLGAMNFAGYE 778
+GLF ++P R + P+ + + Y+
Sbjct: 257 FVGLFTRSLPVRITIVGPVITLQWFFYD 284
>At4g24570 putative mitochondrial uncoupling protein
Length = 313
Score = 68.9 bits (167), Expect = 1e-11
Identities = 72/309 (23%), Positives = 127/309 (40%), Gaps = 47/309 (15%)
Query: 522 LRSALAGGLSCALSCALLHPVDSIKTRVQ----------------------ASSMSFPEI 559
++S + GG++ ++ HP+D IK R+Q +S +F E
Sbjct: 3 VKSFVEGGIASVIAGCSTHPLDLIKVRLQLHGEAPSTTTVTLLRPALAFPNSSPAAFLET 62
Query: 560 IAKLPEIG-------------TRGLYRGSIPAILGQFSSHGLRTGIFEASKLVLVNVAPN 606
+ +P++G L+ G +L Q R G++E K +
Sbjct: 63 TSSVPKVGPISLGINIVKSEGAAALFSGVSATLLRQTLYSTTRMGLYEVLKNKWTDPESG 122
Query: 607 LPELQVQSIASFCSTFLGTAVRIPCEVLKQRLQAG---------LFNNVGEALVGTWQQD 657
L + A + +G AV P +V R+QA + VG+A+ + +
Sbjct: 123 KLNLSRKIGAGLVAGGIGAAVGNPADVAMVRMQADGRLPLAQRRNYAGVGDAIRSMVKGE 182
Query: 658 GLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELEAWETIAVGALSGG-LAA 716
G+ +RG+ T+ R + A + Y + K+G+ + G + T V + + G +A+
Sbjct: 183 GVTSLWRGSALTINRAMIVTAAQLASYDQFKEGILE-NGVMNDGLGTHVVASFAAGFVAS 241
Query: 717 VVTTPFDVMKTRMMTAQ-GRSVSMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFA 775
V + P DV+KTR+M + G A ++ EG + L+KG VP P + F
Sbjct: 242 VASNPVDVIKTRVMNMKVGAYDGAWDCAVKTVKAEGAMALYKGFVPTVCRQGPFTVVLFV 301
Query: 776 GYELARKAM 784
E RK +
Sbjct: 302 TLEQVRKLL 310
Score = 45.1 bits (105), Expect = 2e-04
Identities = 37/146 (25%), Positives = 59/146 (40%), Gaps = 7/146 (4%)
Query: 655 QQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELEAWETIAVGALSGGL 714
+ +G F G ATL R+ + MGLY K +L I G ++GG+
Sbjct: 80 KSEGAAALFSGVSATLLRQTLYSTTRMGLYEVLKNKWTDPESGKLNLSRKIGAGLVAGGI 139
Query: 715 AAVVTTPFDVMKTRMMT------AQGRS-VSMSIVAFSILRHEGPLGLFKGAVPRFFWIA 767
A V P DV RM AQ R+ + S+++ EG L++G+
Sbjct: 140 GAAVGNPADVAMVRMQADGRLPLAQRRNYAGVGDAIRSMVKGEGVTSLWRGSALTINRAM 199
Query: 768 PLGAMNFAGYELARKAMNKNDEAKTG 793
+ A A Y+ ++ + +N G
Sbjct: 200 IVTAAQLASYDQFKEGILENGVMNDG 225
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.134 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,461,001
Number of Sequences: 26719
Number of extensions: 758638
Number of successful extensions: 2715
Number of sequences better than 10.0: 114
Number of HSP's better than 10.0 without gapping: 51
Number of HSP's successfully gapped in prelim test: 63
Number of HSP's that attempted gapping in prelim test: 2244
Number of HSP's gapped (non-prelim): 290
length of query: 796
length of database: 11,318,596
effective HSP length: 107
effective length of query: 689
effective length of database: 8,459,663
effective search space: 5828707807
effective search space used: 5828707807
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 64 (29.3 bits)
Medicago: description of AC144484.11


