

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144481.6 - phase: 0
(754 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
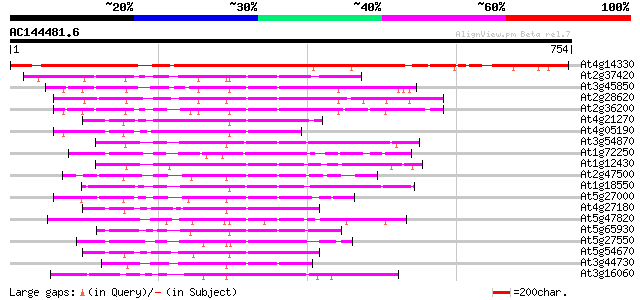
Score E
Sequences producing significant alignments: (bits) Value
At4g14330 kinesin like protein 823 0.0
At2g37420 putative kinesin heavy chain 142 7e-34
At3g45850 kinesin-related protein - like 141 1e-33
At2g28620 putative kinesin-like spindle protein 137 2e-32
At2g36200 putative kinesin-related cytokinesis protein 137 3e-32
At4g21270 kinesin-related protein katA 134 2e-31
At4g05190 kinesin - like protein 130 2e-30
At3g54870 kinesin-like protein 130 2e-30
At1g72250 kinesin, putative 127 2e-29
At1g12430 kinesin-like protein 125 9e-29
At2g47500 kinesin heavy chain like protein 125 1e-28
At1g18550 hypothetical protein 122 7e-28
At5g27000 kinesin-like heavy chain (KATD) 121 1e-27
At4g27180 heavy chain polypeptide of kinesin-like protein (katB) 120 4e-27
At5g47820 kinesin-like protein 116 5e-26
At5g65930 kinesin-like calmodulin-binding protein 115 1e-25
At5g27550 kinesin-like protein 114 2e-25
At5g54670 heavy chain polypeptide of kinesin-like protein (katC) 114 2e-25
At3g44730 kinesin-like protein heavy chain (KP1) 114 2e-25
At3g16060 kinesin like protein 112 6e-25
>At4g14330 kinesin like protein
Length = 959
Score = 823 bits (2126), Expect = 0.0
Identities = 465/781 (59%), Positives = 557/781 (70%), Gaps = 85/781 (10%)
Query: 1 MAPTPSSS-SKQIHTTHLITPRSKHRLNFNGVKPAPTPPHPHPSPHPNFNNKDSPPEHPI 59
MAPTPSSS S Q T + TP++K RLNF+ +P+P+ + SPPEHP+
Sbjct: 1 MAPTPSSSRSNQTQYTLIRTPQTKQRLNFHS-----------KTPNPDGSKDPSPPEHPV 49
Query: 60 EVIARIRDYPDRKDKPLSVLQASSNSRSIRVKADFGYRDFTLDGVSVSEEEELDLFYKKF 119
EVI RIRDYPDRK+K S+LQ +++++++RV+AD GYRDFTLDGVS SE+E L+ FYKKF
Sbjct: 50 EVIGRIRDYPDRKEKSPSILQVNTDNQTVRVRADVGYRDFTLDGVSFSEQEGLEEFYKKF 109
Query: 120 VESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQAGIVYRALRDILGDGDTDSEGSDG 179
+E RI GVK+G+KCTIMMYGPTG+GKSHTMFGC K+ GIVYR+LRDILGD D D
Sbjct: 110 IEERIKGVKVGNKCTIMMYGPTGAGKSHTMFGCGKEPGIVYRSLRDILGDSDQD------ 163
Query: 180 DSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSNASKVKLEVMG 239
GV TFVQVTVLE+YNEEIYDLLSTN G GW K ++KV+LEVMG
Sbjct: 164 --------GV-TFVQVTVLEVYNEEIYDLLSTNSSNN----LGIGWPKGASTKVRLEVMG 210
Query: 240 KKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVILDVPTVGGRLMLV 299
KKAKNA++ISG EAGKISKEI KVEKRRIVKSTLCN+RSSRSHC++ILDVPTVGGRLMLV
Sbjct: 211 KKAKNASFISGTEAGKISKEIVKVEKRRIVKSTLCNERSSRSHCIIILDVPTVGGRLMLV 270
Query: 300 DMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDS 359
DMAGSENI+QAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDS
Sbjct: 271 DMAGSENIDQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDS 330
Query: 360 FEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVRGPHTPVKE----EDSSSTVIL 415
FEDDKSKILMILCASPDPKE+HKT+ TLEYGAKAKCIVRG HTP K+ ++S+S VIL
Sbjct: 331 FEDDKSKILMILCASPDPKEMHKTLCTLEYGAKAKCIVRGSHTPNKDKYGGDESASAVIL 390
Query: 416 GSRIAAMDEFIMKLQMENKLREKERNEAHKKLMKKEEEIAALR---TKVETAPASEEEIN 472
GSRIAAMDEFI+KLQ E K +EKERNEA K+L KKEEE+AALR T+ E +EEEI
Sbjct: 391 GSRIAAMDEFIIKLQSEKKQKEKERNEAQKQLKKKEEEVAALRSLLTQREACATNEEEIK 450
Query: 473 LKVNERTRHLRQELEKKLEECQRMTNEFVELERKRMEERILQQQEEVEILRKRLEEIELQ 532
KVNERT+ L+ EL+KKLEEC+RM EFVE+ER+RMEERI+QQQEE+E++R+RLEEIE+
Sbjct: 451 EKVNERTQLLKSELDKKLEECRRMAEEFVEMERRRMEERIVQQQEELEMMRRRLEEIEV- 509
Query: 533 LCSSSKQERKDENESKEMEPNGFMRKLLSVYKSTDDLGMVKSMDLDMDDQEPFLAREVIV 592
E + N E +GF ++L S+Y S DD GMVKSMDLDM D EP V
Sbjct: 510 -------EFRRSNGGSVDETSGFAKRLRSLY-SDDDPGMVKSMDLDMGDPEPVKQVWGAV 561
Query: 593 GMQG---ISPNQPCPNTLKNGVQEDAYVCAPNFGQKACLSTVYEEEGEEEAEQDHDKVEE 649
Q IS N N L+ E+ + + + CLSTV+EEE E
Sbjct: 562 SHQSSNTISSN--FTNLLQPKPSEN--MLTQMYPDRVCLSTVFEEE------------EV 605
Query: 650 DEEVEKEVIEEKRVCSVVNKSPKIED--------YTGADKENNGSNRLLRIHNIFTLCGN 701
+EE EK ++E+K +C + P + + GAD + + S+R LRI NIFTLCGN
Sbjct: 606 EEEEEKVIVEDKSICLITTPMPSLNSEGLGKENCFNGADDKESASSRRLRIQNIFTLCGN 665
Query: 702 QRELSQYG------TPIPTKKRSDESF---DFKCSPVKSSEKKDSVLRVSNKEN--LEAY 750
QRELSQ+ I + + D F K + E K++ + V +EN L+ Y
Sbjct: 666 QRELSQHSGQEEDQANIASPDKKDNQFFSITNKAEALAVEEAKENNISVDQRENGQLDIY 725
Query: 751 V 751
V
Sbjct: 726 V 726
>At2g37420 putative kinesin heavy chain
Length = 1022
Score = 142 bits (358), Expect = 7e-34
Identities = 136/494 (27%), Positives = 227/494 (45%), Gaps = 77/494 (15%)
Query: 19 TPRSKHRLNFNGVKPAPTP------PHPHPSPHPNFNNKDSPPEHPIEVIARIRDYPDRK 72
TP R + GV P+P P P N ++D+ E ++VI R + + +
Sbjct: 4 TPEVVSRKSGVGVIPSPAPFLTPRLERRRPDSFSNRLDRDNK-EVNVQVILRCKPLSEEE 62
Query: 73 DKPL--SVLQASSNSRSIRVKADFGYRD----FTLDGVSVSEEEELDLFYKKFVESRING 126
K V+ + R + V + F D V + ++ + Y + + ++
Sbjct: 63 QKSSVPRVISCNEMRREVNVLHTIANKQVDRLFNFDKVFGPKSQQRSI-YDQAIAPIVHE 121
Query: 127 VKLGDKCTIMMYGPTGSGKSHTMFGCSK--------QAGIVYRALRDILGDGDTDSEGSD 178
V G CT+ YG TG+GK++TM G + +AG++ RA+R I DT E +
Sbjct: 122 VLEGFSCTVFAYGQTGTGKTYTMEGGMRKKGGDLPAEAGVIPRAVRHIF---DT-LEAQN 177
Query: 179 GDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSNASKVKLEV- 237
D S ++VT LE+YNEE+ DLL+ + S+S+ K + +
Sbjct: 178 ADYS----------MKVTFLELYNEEVTDLLAQDDS-----------SRSSEDKQRKPIS 216
Query: 238 MGKKAKNATYISGNE------AGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVILDV-- 289
+ + K + + G E A I +++ +R TL N RSSRSH + + V
Sbjct: 217 LMEDGKGSVVLRGLEEEVVYSANDIYALLERGSSKRRTADTLLNKRSSRSHSVFTITVHI 276
Query: 290 --PTVG-------GRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIAN 340
++G G+L LVD+AGSENI ++G A+ + +IN+ + L RV+ ++
Sbjct: 277 KEESMGDEELIKCGKLNLVDLAGSENILRSGARDGRAR-EAGEINKSLLTLGRVINALVE 335
Query: 341 GDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVRGP 400
SHVP+RDSKLT LL+DS K+K +I SP + +T+STL+Y +AK I P
Sbjct: 336 HSSHVPYRDSKLTRLLRDSL-GGKTKTCIIATISPSAHSLEETLSTLDYAYRAKNIKNKP 394
Query: 401 HTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKLR-EKERNEAHKKLMKKEEEIAALRT 459
K L + D ++ +M+ +R +++N + + +E +
Sbjct: 395 EANQK---------LSKAVLLKDLYLELERMKEDVRAARDKNGVYIAHERYTQEEVEKKA 445
Query: 460 KVETAPASEEEINL 473
++E E E+NL
Sbjct: 446 RIERIEQLENELNL 459
>At3g45850 kinesin-related protein - like
Length = 1058
Score = 141 bits (356), Expect = 1e-33
Identities = 154/552 (27%), Positives = 246/552 (43%), Gaps = 82/552 (14%)
Query: 49 NNKDSPPEHPIEVIARIRDYPD---RKDKPLSVLQASSNSRSIRVKADFGY----RDFTL 101
N D ++VI R R + R P+ V+ + N R + R F
Sbjct: 39 NRNDKEKGVNVQVILRCRPLSEDEARIHTPV-VISCNENRREVAATQSIAGKHIDRHFAF 97
Query: 102 DGVSVSEEEELDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQ------ 155
D V ++ DL Y + + + V G CTI YG TG+GK++TM G +++
Sbjct: 98 DKVFGPASQQKDL-YDQAICPIVFEVLEGYNCTIFAYGQTGTGKTYTMEGGARKKNGEFP 156
Query: 156 --AGIVYRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNG 213
AG++ RA++ I + G ++VT LE+YNEEI DLL+
Sbjct: 157 SDAGVIPRAVKQIFDILEAQ--------------GAEYSMKVTFLELYNEEISDLLAPEE 202
Query: 214 GGGGGGGFGFGWSKSNASKVKLEVMGKKAKNATYISGNE------AGKISKEIQKVEKRR 267
F KS S +E K + ++ G E A +I K ++K +R
Sbjct: 203 T------IKFVDEKSKKSIALME----DGKGSVFVRGLEEEIVSTANEIYKILEKGSAKR 252
Query: 268 IVKSTLCNDRSSRSHCMVILDVPTVG-----------GRLMLVDMAGSENIEQAGQTGFE 316
TL N +SSRSH + + + G+L LVD+AGSENI ++G
Sbjct: 253 RTAETLLNKQSSRSHSIFSITIHIKENTPEGEEMIKCGKLNLVDLAGSENISRSGAREGR 312
Query: 317 AKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPD 376
A+ + +IN+ + L RV+ ++ H+P+RDSKLT LL++S K+K +I SP
Sbjct: 313 AR-EAGEINKSLLTLGRVINALVEHSGHIPYRDSKLTRLLRESL-GGKTKTCVIATISPS 370
Query: 377 PKEIHKTISTLEYGAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKLR 436
+ +T+STL+Y +AK I P K S+ L S I + + + + +N +
Sbjct: 371 IHCLEETLSTLDYAHRAKNIKNKPEINQKMMKSAVMKDLYSEIDRLKQEVYAAREKNGIY 430
Query: 437 -EKER---NEAHKKLMKKEEEIAALRTKVETAPASE-EEINLKVNERTRHLRQELEKKLE 491
K+R EA KK M ++ E L+++ + + +E+ T L ++LEK +
Sbjct: 431 IPKDRYIQEEAEKKAMAEKIERLELQSESKDKRVVDLQELYNSQQILTAELSEKLEKTEK 490
Query: 492 ECQRMTNEFVELERKRMEERILQQQEEVEI-----LRKRLEE------IELQLCSS---- 536
+ + + +LE K + +++E I K L E EL+ SS
Sbjct: 491 KLEETEHSLFDLEEKYRQANATIKEKEFVISNLLKSEKSLVERAFQLRTELESASSDVSN 550
Query: 537 --SKQERKDENE 546
SK ERKD+ E
Sbjct: 551 LFSKIERKDKIE 562
>At2g28620 putative kinesin-like spindle protein
Length = 1076
Score = 137 bits (345), Expect = 2e-32
Identities = 154/585 (26%), Positives = 257/585 (43%), Gaps = 91/585 (15%)
Query: 59 IEVIARIRDYPDRKDK--PLSVLQASSNSRSIRVKADFGYRD----FTLDGVSVSEEEEL 112
I+VI R R + + + +VL + + + V + + F D V ++
Sbjct: 51 IQVIVRCRPFNSEETRLQTPAVLTCNDRKKEVAVAQNIAGKQIDKTFLFDKVFGPTSQQK 110
Query: 113 DLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQ--------AGIVYRALR 164
DL+++ V + V G CTI YG TG+GK++TM G +++ AG++ RA++
Sbjct: 111 DLYHQA-VSPIVFEVLDGYNCTIFAYGQTGTGKTYTMEGGARKKNGEIPSDAGVIPRAVK 169
Query: 165 DILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFG 224
I + S ++V+ LE+YNEE+ DLL+
Sbjct: 170 QIFDILEAQSAAEYS-------------LKVSFLELYNEELTDLLAPEETKFA------- 209
Query: 225 WSKSNASKVKLEVMGKKAKNATYISGNE------AGKISKEIQKVEKRRIVKSTLCNDRS 278
+ SK L +M + K ++ G E A +I K ++K +R TL N +S
Sbjct: 210 ---DDKSKKPLALM-EDGKGGVFVRGLEEEIVSTADEIYKVLEKGSAKRRTAETLLNKQS 265
Query: 279 SRSHCMVILDVP-----------TVGGRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQG 327
SRSH + + + G+L LVD+AGSENI ++G A+ + +IN+
Sbjct: 266 SRSHSIFSVTIHIKECTPEGEEIVKSGKLNLVDLAGSENISRSGAREGRAR-EAGEINKS 324
Query: 328 NIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTL 387
+ L RV+ ++ H+P+R+SKLT LL+DS K+K +I SP + +T+STL
Sbjct: 325 LLTLGRVINALVEHSGHIPYRESKLTRLLRDSL-GGKTKTCVIATVSPSVHCLEETLSTL 383
Query: 388 EYGAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKLR-EKER---NEA 443
+Y +AK I P K S+ L S I + + + + +N + KER EA
Sbjct: 384 DYAHRAKHIKNKPEVNQKMMKSAIMKDLYSEIERLKQEVYAAREKNGIYIPKERYTQEEA 443
Query: 444 HKKLMKKEEEIAALRTKVETAPASEEEINLK--------VNERTRHLRQELEKKLEECQR 495
KK M + E + +VE ++ I+L+ V R + EKKL E ++
Sbjct: 444 EKKAMADKIE----QMEVEGEAKDKQIIDLQELYNSEQLVTAGLREKLDKTEKKLYETEQ 499
Query: 496 MTNEFVELER------KRMEERILQQQEEVEILRKRLEEIELQLCSS--------SKQER 541
+ E R K E I + + L R E++ +L ++ +K R
Sbjct: 500 ALLDLEEKHRQAVATIKEKEYLISNLLKSEKTLVDRAVELQAELANAASDVSNLFAKIGR 559
Query: 542 KDE-NESKEMEPNGFMRKLLSVYKSTDD--LGMVKSMDLDMDDQE 583
KD+ +S F +LL + ++ G V + + D E
Sbjct: 560 KDKIEDSNRSLIQDFQSQLLRQLELLNNSVAGSVSQQEKQLQDME 604
>At2g36200 putative kinesin-related cytokinesis protein
Length = 1056
Score = 137 bits (344), Expect = 3e-32
Identities = 155/578 (26%), Positives = 260/578 (44%), Gaps = 96/578 (16%)
Query: 59 IEVIARIRDYPD---RKDKPLSVLQASSNSRSIRVKADFGY----RDFTLDGVSVSEEEE 111
++V+ R R + D R + P VL + R + V + R FT D V ++
Sbjct: 13 VQVLLRCRPFSDDELRSNAP-QVLTCNDLQREVAVSQNIAGKHIDRVFTFDKVFGPSAQQ 71
Query: 112 LDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFG-CSK-----------QAGIV 159
DL Y + V +N V G CTI YG TG+GK++TM G C + +AG++
Sbjct: 72 KDL-YDQAVVPIVNEVLEGFNCTIFAYGQTGTGKTYTMEGECRRSKSAPCGGLPAEAGVI 130
Query: 160 YRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGG 219
RA++ I DT EG + S V+VT LE+YNEEI DLL+
Sbjct: 131 PRAVKQIF---DT-LEGQQAEYS----------VKVTFLELYNEEITDLLAPE------- 169
Query: 220 GFGFGWSKSNASKVKLEVMGKK-------AKNATYISGNE------AGKISKEIQKVEKR 266
+ S+V E KK K + G E A +I +++ +
Sbjct: 170 ---------DLSRVAAEEKQKKPLPLMEDGKGGVLVRGLEEEIVTSANEIFTLLERGSSK 220
Query: 267 RIVKSTLCNDRSSRSHCMVILDVPTVG-----------GRLMLVDMAGSENIEQAGQTGF 315
R T N +SSRSH + + + G+L LVD+AGSENI ++G
Sbjct: 221 RRTAETFLNKQSSRSHSLFSITIHIKEATPEGEELIKCGKLNLVDLAGSENISRSGARDG 280
Query: 316 EAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASP 375
A+ + +IN+ + L RV+ ++ HVP+RDSKLT LL+DS ++K +I SP
Sbjct: 281 RAR-EAGEINKSLLTLGRVISALVEHLGHVPYRDSKLTRLLRDSL-GGRTKTCIIATVSP 338
Query: 376 DPKEIHKTISTLEYGAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKL 435
+ +T+STL+Y +AK I P K S+ L I + + + +N +
Sbjct: 339 AVHCLEETLSTLDYAHRAKNIRNKPEVNQKMMKSTLIKDLYGEIERLKAEVYASREKNGV 398
Query: 436 -REKER---NEAHKKLMKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQELEKKLE 491
KER E+ +K+M E+I + ++E EE+ K + R +L KL+
Sbjct: 399 YMPKERYYQEESERKVM--AEQIEQMGGQIENYQKQLEELQDKYVGQVREC-SDLTTKLD 455
Query: 492 ECQRMTNEFVEL-----ERKRMEERILQQQEEVEILRKRLEEIELQLCS--SSKQERKDE 544
++ ++ ++ E + + +++++ + +K+ E + +Q S E+ +
Sbjct: 456 ITEKNLSQTCKVLASTNEELKKSQYAMKEKDFIISEQKKSENVLVQQACILQSNLEKATK 515
Query: 545 NESKEMEPNGFMRKLLSVYKSTDDLGMVKSMDLDMDDQ 582
+ S + G KL S D+ +V + +++ +Q
Sbjct: 516 DNSSLHQKIGREDKL-----SADNRKVVDNYQVELSEQ 548
>At4g21270 kinesin-related protein katA
Length = 793
Score = 134 bits (337), Expect = 2e-31
Identities = 112/337 (33%), Positives = 168/337 (49%), Gaps = 38/337 (11%)
Query: 99 FTLDGVSVSEEEELDLFYK--KFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFG---CS 153
FT D V E + ++F++ + V+S ++G K+ I YG TGSGK++TM G
Sbjct: 478 FTFDKVFNHEASQEEVFFEISQLVQSALDGYKV----CIFAYGQTGSGKTYTMMGRPEAP 533
Query: 154 KQAGIVYRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNG 213
Q G++ R+L I + S G+ G K +QV++LEIYNE I DLLSTN
Sbjct: 534 DQKGLIPRSLEQIFQA--SQSLGAQGWKYK---------MQVSMLEIYNETIRDLLSTNR 582
Query: 214 GGGGGGGFGFGWSKSNASKVKLEVMGKK-AKNATYISGNEAGKISKEIQKVEKRRIVKST 272
+ + +V G + T GKIS +Q+ + R V T
Sbjct: 583 TTSMDLVRADSGTSGKQYTITHDVNGHTHVSDLTIFDVCSVGKISSLLQQAAQSRSVGKT 642
Query: 273 LCNDRSSRSHCMVILDVPTVG--------GRLMLVDMAGSENIEQAGQTGFEAKMQTAKI 324
N++SSRSH + + + V G L L+D+AGSE + ++G TG K +T I
Sbjct: 643 QMNEQSSRSHFVFTMRISGVNESTEQQVQGVLNLIDLAGSERLSKSGATGDRLK-ETQAI 701
Query: 325 NQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTI 384
N+ AL V+ ++A + HVPFR+SKLT LLQ D SK LM + SPDP +++
Sbjct: 702 NKSLSALSDVIFALAKKEDHVPFRNSKLTYLLQPCLGGD-SKTLMFVNISPDPTSAGESL 760
Query: 385 STLEYGAKAK-CIVRGPHTPVKEEDSSSTVILGSRIA 420
+L + A+ C + P +ST +L SR++
Sbjct: 761 CSLRFAARVNACEIGIPRR------QTSTKLLDSRLS 791
Score = 42.0 bits (97), Expect = 0.001
Identities = 52/243 (21%), Positives = 105/243 (42%), Gaps = 26/243 (10%)
Query: 323 KINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHK 382
K+ + +A K+ V S+ + + ++ L D ++ + A + I +
Sbjct: 203 KVKEEKMAAKQKVTSLEDMYKRLQEYNTSLQQYNSKLQTDLETVRAALTRAEKEKSSILE 262
Query: 383 TISTLEYGAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKLREKERNE 442
+STL RG ++++ SSS V+ I D + ++ ++ R++
Sbjct: 263 NLSTL----------RGHSKSLQDQLSSSRVLQDDAIKQKDSLLSEVTNLRNELQQVRDD 312
Query: 443 AHKKLMKKEEEIAALRTKVETAPASEEEINLKVN-----ERTRHLRQE----LEKKL--- 490
+++++ ++ +R E S +E+++ E T L++E LE++L
Sbjct: 313 RDRQVVQSQKLSEEIRKYQENVGKSSQELDILTAKSGSLEETCSLQKERLNMLEQQLAIA 372
Query: 491 EECQRMTNEFVELERKRMEERILQQQEEVEILRKRLEEIELQLCSSSKQERKDENESKEM 550
E Q+M + V L R EE Q+ + L+ RL ++E QLC +K N E+
Sbjct: 373 NERQKMADASVSLTRTEFEE----QKHLLCELQDRLADMEHQLCEGELLRKKLHNTILEL 428
Query: 551 EPN 553
+ N
Sbjct: 429 KGN 431
Score = 29.6 bits (65), Expect = 6.5
Identities = 33/115 (28%), Positives = 54/115 (46%), Gaps = 11/115 (9%)
Query: 428 KLQMENKLREKERNEAHKKL--MKKEEEIAALRTKVETAPASEEEINLKVNERTRHL--- 482
K ++ L E+ HK+L KEEE+ A +K+E S E K T+
Sbjct: 120 KENLKVSLESSEQKYNHKELEARTKEEELQATISKLEENVVSLHEKLAKEESSTQDAIEC 179
Query: 483 -RQELEKKL--EECQRMTNEFVELERKRMEERILQQQ-EEVEILRKRLEEIELQL 533
R+E E ++ E+ Q E EL++ + E+ +Q+ +E + KRL+E L
Sbjct: 180 HRREKEARVAAEKVQASLGE--ELDKVKEEKMAAKQKVTSLEDMYKRLQEYNTSL 232
>At4g05190 kinesin - like protein
Length = 777
Score = 130 bits (328), Expect = 2e-30
Identities = 114/354 (32%), Positives = 176/354 (49%), Gaps = 37/354 (10%)
Query: 59 IEVIARIRDY-PD---RKDKPLSVLQASSNS--RSIRVKADFGYRDFTLDGVSVSEEEEL 112
I V R+R PD R++ + S+ S R I V FT D V +
Sbjct: 416 IRVFCRVRPLLPDDGGRQEASVIAYPTSTESLGRGIDVVQSGNKHPFTFDKVFDHGASQE 475
Query: 113 DLFYK--KFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFG---CSKQAGIVYRALRDIL 167
++F++ + V+S ++G K+ I YG TGSGK++TM G +Q G++ R+L I
Sbjct: 476 EVFFEISQLVQSALDGYKV----CIFAYGQTGSGKTYTMMGRPETPEQKGLIPRSLEQIF 531
Query: 168 GDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFGWSK 227
+ S S++G+ + +QV++LEIYNE I DLLST+ +
Sbjct: 532 -------KTSQSLSTQGW----KYKMQVSMLEIYNESIRDLLSTSRTIAIESVRADSSTS 580
Query: 228 SNASKVKLEVMGKK-AKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVI 286
+ +V G + T + G+IS +Q+ + R V T N++SSRSH +
Sbjct: 581 GRQYTITHDVNGNTHVSDLTIVDVCSIGQISSLLQQAAQSRSVGKTHMNEQSSRSHFVFT 640
Query: 287 LDVPTVG--------GRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESI 338
L + V G L L+D+AGSE + ++G TG K +T IN+ AL V+ ++
Sbjct: 641 LRISGVNESTEQQVQGVLNLIDLAGSERLSRSGATGDRLK-ETQAINKSLSALSDVIFAL 699
Query: 339 ANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAK 392
A + HVPFR+SKLT LLQ D SK LM + SPDP +++ +L + A+
Sbjct: 700 AKKEDHVPFRNSKLTYLLQPCLGGD-SKTLMFVNISPDPSSTGESLCSLRFAAR 752
>At3g54870 kinesin-like protein
Length = 1070
Score = 130 bits (328), Expect = 2e-30
Identities = 124/475 (26%), Positives = 215/475 (45%), Gaps = 77/475 (16%)
Query: 116 YKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSK----QAGIVYRALRDILGDGD 171
Y+ + + GV G TIM YG TG+GK++T+ K + GI+ RAL DIL +
Sbjct: 185 YEGVAKPVVEGVLSGYNGTIMAYGQTGTGKTYTVGKIGKDDAAERGIMVRALEDILLNAS 244
Query: 172 TDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSNAS 231
+ S V+++ L++Y E I DLL+ K+N S
Sbjct: 245 SASIS----------------VEISYLQLYMETIQDLLAPE--------------KNNIS 274
Query: 232 KVKLEVMGK-KAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVILDV- 289
+ G+ AT ++ + + +Q E R +T N SSRSH ++ + V
Sbjct: 275 INEDAKTGEVSVPGATVVNIQDLDHFLQVLQVGETNRHAANTKMNTESSRSHAILTVYVR 334
Query: 290 --------------------PTVG-GRLMLVDMAGSENIEQAGQTGFEAKMQTAK-INQG 327
P V +L++VD+AGSE I ++G G ++ AK IN
Sbjct: 335 RAMNEKTEKAKPESLGDKAIPRVRKSKLLIVDLAGSERINKSGTDGH--MIEEAKFINLS 392
Query: 328 NIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTL 387
+L + + ++A G SH+P RDSKLT LL+DSF ++ +I+ P + +T ST+
Sbjct: 393 LTSLGKCINALAEGSSHIPTRDSKLTRLLRDSF-GGSARTSLIITIGPSARYHAETTSTI 451
Query: 388 EYGAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKLREKERNEAHKKL 447
+G +A IV +KEE ++ +D +++ +NKLR E++E K+L
Sbjct: 452 MFGQRAMKIVN--MVKLKEEFDYESLCRKLE-TQVDHLTAEVERQNKLRNSEKHELEKRL 508
Query: 448 MKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQELEKKLEECQRMTNEFVELERKR 507
+ E A T E+ N ++ + L ++L+ + ++C M ++ ++LE K
Sbjct: 509 RECENSFAEAEKNAVTRSKFLEKENTRLELSMKELLKDLQLQKDQCDLMHDKAIQLEMKL 568
Query: 508 MEERILQQQEE-----------VEILRKRLEEIELQLCSSSKQERKDENESKEME 551
+ QQQ E ++ K++ E+ ++ + E++ EM+
Sbjct: 569 KNTK--QQQLENSAYEAKLADTSQVYEKKIAELVQRVEDEQARSTNAEHQLTEMK 621
>At1g72250 kinesin, putative
Length = 1195
Score = 127 bits (320), Expect = 2e-29
Identities = 128/478 (26%), Positives = 214/478 (43%), Gaps = 76/478 (15%)
Query: 79 LQASSNSRSIRVKADFGYRDFTLDGVSVSEEEELDLFYKK--FVESRINGVKLGDKCTIM 136
++++ N I + F + F D V + D+F F S I+G + I
Sbjct: 516 VESTKNGEVIVMSNGFPKKSFKFDSVFGPNASQADVFEDTAPFATSVIDGYNV----CIF 571
Query: 137 MYGPTGSGKSHTMFGCSKQAGIVYRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVT 196
YG TG+GK+ TM G G+ YR L ++ + + + V+
Sbjct: 572 AYGQTGTGKTFTMEGTQHDRGVNYRTLENLFRIIKAREHRYNYE------------ISVS 619
Query: 197 VLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSNASKVKLEVMGKKAKNATYISGNEAGKI 256
VLE+YNE+I DLL + +AS K + + ++ ++ G +
Sbjct: 620 VLEVYNEQIRDLLVP--------------ASQSASAPKRFEIRQLSEGNHHVPGLVEAPV 665
Query: 257 SKEIQKV-------EKRRIVKSTLCNDRSSRSHCM--------VILDVPTVGGRLMLVDM 301
K I++V R V T N+ SSRSHC+ +L+ +L LVD+
Sbjct: 666 -KSIEEVWDVLKTGSNARAVGKTTANEHSSRSHCIHCVMVKGENLLNGECTKSKLWLVDL 724
Query: 302 AGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFE 361
AGSE + + G E +T IN+ AL V+ ++AN SH+PFR+SKLT LLQDS
Sbjct: 725 AGSERVAKTEVQG-ERLKETQNINKSLSALGDVIFALANKSSHIPFRNSKLTHLLQDSLG 783
Query: 362 DDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVRGPHTPVKEEDSSSTVILGSRIAA 421
D SK LM + SP+ + +T+ +L + ++ + I GP K+ D++
Sbjct: 784 GD-SKTLMFVQISPNENDQSETLCSLNFASRVRGIELGP--AKKQLDNT----------- 829
Query: 422 MDEFIMKLQMENKLREKERNEAHKKLMKKEEEIAALRTKVETAPASEEEINLKVNERTRH 481
E + QM K ++ + + +++ K EE + L K++ + + KV
Sbjct: 830 --ELLKYKQMVEKWKQDMKGK-DEQIRKMEETMYGLEAKIKERDTKNKTLQDKV------ 880
Query: 482 LRQELEKKLEECQRMTNEFVELERKRMEERILQQQEEVEILRKRLEEIELQLCSSSKQ 539
+ELE +L +++ + V + K E++ QQ E+ KR + L S+SK+
Sbjct: 881 --KELESQLLVERKLARQHV--DTKIAEQQTKQQTEDENNTSKRPPLTNILLGSASKE 934
>At1g12430 kinesin-like protein
Length = 919
Score = 125 bits (314), Expect = 9e-29
Identities = 126/475 (26%), Positives = 214/475 (44%), Gaps = 70/475 (14%)
Query: 116 YKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQ----AGIVYRALRDILGDGD 171
Y+ + + GV G TIM YG TG+GK++T+ ++ GI+ RA+ DIL +
Sbjct: 132 YEVVAKPVVEGVLDGYNGTIMAYGQTGTGKTYTLGQLGEEDVADRGIMVRAMEDILAEVS 191
Query: 172 TDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNG-----------GGGGGGG 220
+++ + V+ L++Y E + DLL + G G
Sbjct: 192 LETDS----------------ISVSYLQLYMETVQDLLDPSNDNIAIVEDPKNGDVSLPG 235
Query: 221 FGFGWSKSNASKVKLEVMGKKAKNATYISGNEAGKISKEIQKVEKRRIVKST--LCNDRS 278
+ S ++L +G+ + A N S I V RR +K+ L ++ +
Sbjct: 236 ATLVEIRDQQSFLELLQLGEAHRFAANTKLNTESSRSHAILMVNVRRSMKTRDGLSSESN 295
Query: 279 SRSHCMVILDVPTVG-GRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVES 337
SH L P V G+L++VD+AGSE I ++G G + + IN AL + + +
Sbjct: 296 GNSHMTKSLKPPVVRKGKLVVVDLAGSERINKSGSEGHTLE-EAKSINLSLSALGKCINA 354
Query: 338 IANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIV 397
+A SHVPFRDSKLT LL+DSF ++ +++ P P+ +T ST+ +G +A +
Sbjct: 355 LAENSSHVPFRDSKLTRLLRDSF-GGTARTSLVITIGPSPRHRGETTSTIMFGQRAMKV- 412
Query: 398 RGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKLREKERNEAHKKLMKKEEEIAAL 457
VK ++ L R +++Q++N + E ER + K E E +
Sbjct: 413 ---ENMVKIKEEFDYKSLSRR--------LEVQLDNLIEENERQQ---KAFVDEIERITV 458
Query: 458 RTKVETAPASEEEINLKVNERTRHLRQELE--KKLEECQRMTNEFVELERKRMEER---- 511
+ + A + N +E+ R+ +E KKLEE + + ER + E+
Sbjct: 459 EAHNQISEAEKRYANALEDEKLRYQNDYMESIKKLEENWSKNQKKLAAERLALGEKNGLD 518
Query: 512 --------ILQQQEEVEILRKRLEEIELQLCSSSKQE----RKDENESKEMEPNG 554
I EEV L+K L++ E Q ++++E + NE K++E +G
Sbjct: 519 ITSNGNRSIAPALEEVSELKKLLQK-EAQSKMAAEEEVNRLKHQLNEFKKVEASG 572
>At2g47500 kinesin heavy chain like protein
Length = 983
Score = 125 bits (313), Expect = 1e-28
Identities = 123/442 (27%), Positives = 192/442 (42%), Gaps = 87/442 (19%)
Query: 72 KDKPLSVLQASSNSRSIRVKADFGYRDFTLDGVSVSEEEELDLFYKKFVESRINGVKLGD 131
+D + + AS + +S++ FT + V + ++F ++ I V G
Sbjct: 424 EDDTIGINTASRHGKSLK--------SFTFNKVFGPSATQEEVFSD--MQPLIRSVLDGY 473
Query: 132 KCTIMMYGPTGSGKSHTMFG----CSKQAGIVYRALRDI--LGDGDTDSEGSDGDSSKGF 185
I YG TGSGK+ TM G K G+ YRAL D+ L + D+ D
Sbjct: 474 NVCIFAYGQTGSGKTFTMSGPRDLTEKSQGVNYRALGDLFLLAEQRKDTFRYD------- 526
Query: 186 CLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSNASKVKLEVMGKKAK-- 243
+ V ++EIYNE++ DLL T+G S +LE+ K
Sbjct: 527 -------IAVQMIEIYNEQVRDLLVTDG-----------------SNKRLEIRNSSQKGL 562
Query: 244 ---NATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVILDVP--------TV 292
+A+ + + + ++ K R V ST NDRSSRSH + + V +
Sbjct: 563 SVPDASLVPVSSTFDVIDLMKTGHKNRAVGSTALNDRSSRSHSCLTVHVQGRDLTSGAVL 622
Query: 293 GGRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKL 352
G + LVD+AGSE ++++ TG K + IN+ AL V+ S+A+ + HVP+R+SKL
Sbjct: 623 RGCMHLVDLAGSERVDKSEVTGDRLK-EAQHINRSLSALGDVIASLAHKNPHVPYRNSKL 681
Query: 353 TMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVRGPHTPVKEEDSSST 412
T LLQDS ++K LM + SP+ + +TISTL++ + + G D+S
Sbjct: 682 TQLLQDSL-GGQAKTLMFVHISPEADAVGETISTLKFAERVATVELG--AARVNNDTSDV 738
Query: 413 VILGSRIAAMDEFIMKLQMENKLREKERNEAHKKLMKKEEEIAALRTKVETAPASEEEIN 472
L +IA +K + K E ++N K P E+
Sbjct: 739 KELKEQIAT-----LKAALARKEAESQQNNILK------------------TPGGSEKHK 775
Query: 473 LKVNERTRHLRQELEKKLEECQ 494
K E H + KK E C+
Sbjct: 776 AKTGEVEIHNNNIMTKKSESCE 797
>At1g18550 hypothetical protein
Length = 799
Score = 122 bits (306), Expect = 7e-28
Identities = 120/464 (25%), Positives = 200/464 (42%), Gaps = 56/464 (12%)
Query: 97 RDFTLDGVSVSEEEELDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQA 156
R FT D S E Y + V G ++ YG TG+GK++TM G +
Sbjct: 202 RHFTFDS-SFPETTTQQEVYSTTTGDLVEAVLEGRNGSVFCYGATGAGKTYTMLGTMENP 260
Query: 157 GIVYRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGG 216
G++ A++D+ DG+ V ++ LE+YNE + DLLS
Sbjct: 261 GVMVLAIKDLFAK--VRQRSLDGNH----------VVHLSYLEVYNETVRDLLSPG---- 304
Query: 217 GGGGFGFGWSKSNASKVKLEVMGKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCND 276
++ + G A T ++ +Q+ + R + T CN+
Sbjct: 305 ------------RPLILREDKQGIVAAGLTQYRAYSTDEVMALLQRGNQNRTTEPTRCNE 352
Query: 277 RSSRSHCM--VILDVPTVG---------GRLMLVDMAGSENIEQAGQTGFEAKMQTAKIN 325
SSRSH + VI++ T G+L L+D+AGSE Q + ++ A IN
Sbjct: 353 TSSRSHAILQVIVEYKTRDASMNIISRVGKLSLIDLAGSERALATDQRTLRS-LEGANIN 411
Query: 326 QGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTIS 385
+ +AL + ++ G H+P+R+SKLT LL+DS +MI SP + +T +
Sbjct: 412 RSLLALSSCINALVEGKKHIPYRNSKLTQLLKDSL-GGSCNTVMIANISPSSQSFGETQN 470
Query: 386 TLEYGAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDE--FIMKLQMEN-KLREKERNE 442
TL + +AK I VKE + + V+ D+ +++LQ EN +LR + +
Sbjct: 471 TLHWADRAKEI------RVKECEVNEEVVQVGEEEGADQAKLLLELQKENSELRVQLAKQ 524
Query: 443 AHKKLMKKEEEIAAL--RTKVETAPASEEEINLKVNERTRHLRQELEKKLEECQRMTNEF 500
K L + E IAA + P S + + T +++ L T E
Sbjct: 525 QQKLLTLQAENIAAANNNNNISLTPPSISSLMTPPSALTAQQKKKPRHSLLSGTCFTPE- 583
Query: 501 VELERKRMEERILQQQEEVEILRKRLEEIELQLCSSSKQERKDE 544
L+R + EE + + Q V+ L+ +E ++ + K++ KDE
Sbjct: 584 -SLKRTKAEEAVKELQLTVKALKMEMERMKREHGLQMKKQ-KDE 625
>At5g27000 kinesin-like heavy chain (KATD)
Length = 987
Score = 121 bits (304), Expect = 1e-27
Identities = 119/427 (27%), Positives = 192/427 (44%), Gaps = 65/427 (15%)
Query: 59 IEVIARIRDY-PDRKDKPLSVLQ-ASSNSRSIRVKADFG---YRDFTLDGVSVSEEEELD 113
I V R+R + P ++ LS ++ + +IRV + +G + F + V + +
Sbjct: 395 IRVYCRVRPFLPGQESGGLSAVEDIDEGTITIRVPSKYGKAGQKPFMFNKVFGPSATQEE 454
Query: 114 LFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFG----CSKQAGIVYRALRDILGD 169
+F ++ + V G I YG TGSGK+ TM G + G+ YRAL D+
Sbjct: 455 VFSD--MQPLVRSVLDGYNVCIFAYGQTGSGKTFTMTGPKELTEESLGVNYRALADLFL- 511
Query: 170 GDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSN 229
++ D S + + V +LEIYNE++ DLL+ +G
Sbjct: 512 --LSNQRKDTTSYE---------ISVQMLEIYNEQVRDLLAQDG---------------- 544
Query: 230 ASKVKLEVM-----GKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSRSHCM 284
+LE+ G A+ + + + + + R V ST NDRSSRSH
Sbjct: 545 -QTKRLEIRNNSHNGINVPEASLVPVSSTDDVIQLMDLGHMNRAVSSTAMNDRSSRSHSC 603
Query: 285 VILDVP--------TVGGRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVE 336
V + V + G + LVD+AGSE ++++ TG K + IN+ AL V+
Sbjct: 604 VTVHVQGRDLTSGSILHGSMHLVDLAGSERVDKSEVTGDRLK-EAQHINKSLSALGDVIS 662
Query: 337 SIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCI 396
S++ SHVP+R+SKLT LLQDS +K LM + SP+P + +TISTL++ + +
Sbjct: 663 SLSQKTSHVPYRNSKLTQLLQDSL-GGSAKTLMFVHISPEPDTLGETISTLKFAERVGSV 721
Query: 397 VRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKLREKERNEAHKKLMKKEEEIAA 456
G K+ S + + E I L+M +R+ N+ + E +
Sbjct: 722 ELGAARVNKD---------NSEVKELKEQIANLKMA-LVRKGNGNDVQPTAIPINRERIS 771
Query: 457 LRTKVET 463
R +ET
Sbjct: 772 RRRSLET 778
>At4g27180 heavy chain polypeptide of kinesin-like protein (katB)
Length = 745
Score = 120 bits (300), Expect = 4e-27
Identities = 104/333 (31%), Positives = 161/333 (48%), Gaps = 36/333 (10%)
Query: 99 FTLDGVSVSEEEELDLFYK--KFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCS--- 153
FT D V V + D+F + + V+S ++G K+ I YG TGSGK++TM G
Sbjct: 434 FTFDKVFVPSASQEDVFVEISQLVQSALDGYKV----CIFAYGQTGSGKTYTMMGRPGNP 489
Query: 154 KQAGIVYRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNG 213
+ G++ R L I + S+G+ + +QV++LEIYNE I DLLSTN
Sbjct: 490 DEKGLIPRCLEQIF-------QTRQSLRSQGW----KYELQVSMLEIYNETIRDLLSTNK 538
Query: 214 GGGGGGGFGFGWSKSNASKVKLEVMGKK-AKNATYISGNEAGKISKEIQKVEKRRIVKST 272
G S + +K + G T + + ++S + + R V T
Sbjct: 539 EAVRADN---GVSPQKYA-IKHDASGNTHVVELTVVDVRSSKQVSFLLDHAARNRSVGKT 594
Query: 273 LCNDRSSRSHCMVILDVP--------TVGGRLMLVDMAGSENIEQAGQTGFEAKMQTAKI 324
N++SSRSH + L + V G L L+D+AGSE + ++G TG K +T I
Sbjct: 595 AMNEQSSRSHFVFTLKISGFNESTEQQVQGVLNLIDLAGSERLSKSGSTGDRLK-ETQAI 653
Query: 325 NQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTI 384
N+ +L V+ ++A + HVPFR+SKLT LLQ D SK LM + +P+P +++
Sbjct: 654 NKSLSSLGDVIFALAKKEDHVPFRNSKLTYLLQPCLGGD-SKTLMFVNITPEPSSTGESL 712
Query: 385 STLEYGAKAK-CIVRGPHTPVKEEDSSSTVILG 416
+L + A+ C + H V + LG
Sbjct: 713 CSLRFAARVNACEIGTAHRHVNARPLDYRLSLG 745
Score = 37.7 bits (86), Expect = 0.024
Identities = 40/188 (21%), Positives = 82/188 (43%), Gaps = 25/188 (13%)
Query: 389 YGAKAKCIVRGPHTPVKEEDSSSTVILGS------RIAAMDEFIMKLQMENKLREKERNE 442
Y +K + + H +K + T I+ S + A+ + + ++ K+++E
Sbjct: 202 YNSKLQGDLDEAHENIKRGEKERTGIVESIGNLKGQFKALQDQLAASKVSQDDVMKQKDE 261
Query: 443 AHKKLMKKEEEIAALR-------TKVETAPASEEEINLKVNERTRHLRQELEKKLEECQR 495
+++ + EI ++ T++ET A + N + L + + +E +
Sbjct: 262 LVNEIVSLKVEIQQVKDDRDRHITEIETLQAEATKQN-DFKDTINELESKCSVQNKEIEE 320
Query: 496 MTNEFVELERK----------RMEERILQQQEEVEILRKRLEEIELQLCSSSKQERKDEN 545
+ ++ V ERK +M E +Q+E + L+ RLEE EL+L K +K N
Sbjct: 321 LQDQLVASERKLQVADLSTFEKMNE-FEEQKESIMELKGRLEEAELKLIEGEKLRKKLHN 379
Query: 546 ESKEMEPN 553
+E++ N
Sbjct: 380 TIQELKGN 387
>At5g47820 kinesin-like protein
Length = 1035
Score = 116 bits (290), Expect = 5e-26
Identities = 138/554 (24%), Positives = 233/554 (41%), Gaps = 101/554 (18%)
Query: 51 KDSPPEHPIEVIARIRDYPDRKDKPLSVLQASSNSRSIRVKADFGYRDFTLDGVSVSEEE 110
+ +PP V + P D+ + Q + + + G FT D V S
Sbjct: 2 ESTPPPDDCSVKVAVHIRPLIGDERIQGCQDCVTVVTGKPQVQIGSHSFTFDHVYGSSGS 61
Query: 111 ELDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTM-FGC--SKQAGIVYRALRDIL 167
Y++ ++G+ G T++ YG TGSGK++TM GC S Q GI+ + + +
Sbjct: 62 PSTEMYEECAAPLVDGLFQGYNATVLAYGQTGSGKTYTMGTGCGDSSQTGIIPQVMNALF 121
Query: 168 GDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLL--------STNGGGGGGG 219
+T + + + V+ +EI+ EE+ DLL TN G G
Sbjct: 122 TKIETLKQ------------QIEFQIHVSFIEIHKEEVQDLLDPCTVNKSDTNNTGHVG- 168
Query: 220 GFGFGWSKSNASKVKLEVMGKKAKN-------ATYISGNEAGKISKEIQKVEKRRIVKST 272
K K + ++ N +T +S + +++ + + R ST
Sbjct: 169 -------KVAHVPGKPPIQIRETSNGVITLAGSTEVSVSTLKEMAACLDQGSVSRATGST 221
Query: 273 LCNDRSSRSHCMVIL----------DVPTVG------------GRLMLVDMAGSENIEQA 310
N++SSRSH + + D P G +L LVD+AGSE ++
Sbjct: 222 NMNNQSSRSHAIFTITVEQMRKINTDSPENGAYNGSLKEEYLCAKLHLVDLAGSERAKRT 281
Query: 311 GQTGFEAKMQTAKINQGNIALKRVVESIAN-----GDSHVPFRDSKLTMLLQDSFEDDKS 365
G G K + IN+G +AL V+ ++ + +HVP+RDSKLT LLQDS S
Sbjct: 282 GSDGLRFK-EGVHINKGLLALGNVISALGDEKKRKDGAHVPYRDSKLTRLLQDSL-GGNS 339
Query: 366 KILMILCASPDPKEIHKTISTLEYGAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEF 425
+ +MI C SP +T++TL+Y +A+ I + PV D S S + M +
Sbjct: 340 RTVMIACISPADINAEETLNTLKYANRARNI---RNKPVVNRDPVS-----SEMLKMRQQ 391
Query: 426 IMKLQMENKLREKERNEAHKKLMKKE------------EEIAALRTKVETAPASEEEI-N 472
+ LQ E LR + A + +K+ E+ R++ SE++ +
Sbjct: 392 VEYLQAELSLRTGGSSCAEVQALKERIVWLETANEELCRELHEYRSRCPGVEHSEKDFKD 451
Query: 473 LKVNERTRHLRQE-LEKKLEECQRMTNEFVEL----------ERKRMEERILQQQEEVEI 521
++ ++ +R + L++ L + VE E K E ++LQ + E+
Sbjct: 452 IRADDIVGSVRPDGLKRSLHSIESSNYPMVEATTGDSREIDEEAKEWEHKLLQNSMDKEL 511
Query: 522 --LRKRLEEIELQL 533
L +RLEE E ++
Sbjct: 512 YELNRRLEEKESEM 525
>At5g65930 kinesin-like calmodulin-binding protein
Length = 1260
Score = 115 bits (287), Expect = 1e-25
Identities = 92/342 (26%), Positives = 162/342 (46%), Gaps = 50/342 (14%)
Query: 117 KKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQAGIVYRALRDILGDGDTDSEG 176
K V+S ++G + I YG TGSGK+ T++G G+ RA +++ DS
Sbjct: 951 KYLVQSAVDGYNV----CIFAYGQTGSGKTFTIYGHESNPGLTPRATKELFNILKRDS-- 1004
Query: 177 SDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSNASKVKLE 236
K F ++ ++ +E+Y + + DLL +A ++KLE
Sbjct: 1005 ------KRFSFSLKAYM----VELYQDTLVDLLLPK----------------SARRLKLE 1038
Query: 237 VMGKK-----AKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVILDVPT 291
+ +N T I + ++ +++ +RR V T N+ SSRSH ++ + + +
Sbjct: 1039 IKKDSKGMVFVENVTTIPISTLEELRMILERGSERRHVSGTNMNEESSRSHLILSVVIES 1098
Query: 292 VG--------GRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDS 343
+ G+L VD+AGSE ++++G G + K + IN+ AL V+ ++++G+
Sbjct: 1099 IDLQTQSAARGKLSFVDLAGSERVKKSGSAGCQLK-EAQSINKSLSALGDVIGALSSGNQ 1157
Query: 344 HVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVRGPHTP 403
H+P+R+ KLTML+ DS +K LM + SP + +T ++L Y ++ + IV + P
Sbjct: 1158 HIPYRNHKLTMLMSDSL-GGNAKTLMFVNVSPAESNLDETYNSLLYASRVRTIV---NDP 1213
Query: 404 VKEEDSSSTVILGSRIAAMDEFIMKLQMENKLREKERNEAHK 445
K S V L +A E K E L + E + K
Sbjct: 1214 SKHISSKEMVRLKKLVAYWKEQAGKKGEEEDLVDIEEDRTRK 1255
Score = 36.6 bits (83), Expect = 0.053
Identities = 53/229 (23%), Positives = 97/229 (42%), Gaps = 45/229 (19%)
Query: 325 NQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTI 384
NQ + L+ +E+I NG + + LL+ + + DK L LC I +
Sbjct: 649 NQQEVTLREELEAIHNG------LELERRKLLEVTLDRDK---LRSLCDEKGTT-IQSLM 698
Query: 385 STLEYGAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKLREKERNEA- 443
S L G +A+ + + +T +E S + ++I + K+Q E ++R KE + A
Sbjct: 699 SELR-GMEAR-LAKSGNTKSSKETKSELAEMNNQI------LYKIQKELEVRNKELHVAV 750
Query: 444 -HKKLMKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQELEKKLEECQRMTNEFVE 502
+ K + E +I +E E EI+ K R E EKK+
Sbjct: 751 DNSKRLLSENKILEQNLNIEKKKKEEVEIHQK--------RYEQEKKV------------ 790
Query: 503 LERKRMEERILQQQEEVEILRKRLEEIELQLCSSSKQERKDENESKEME 551
++ R+ + + ++E+L + L+ E + S + +N KE+E
Sbjct: 791 -----LKLRVSELENKLEVLAQDLDSAESTIESKNSDMLLLQNNLKELE 834
Score = 35.4 bits (80), Expect = 0.12
Identities = 36/162 (22%), Positives = 81/162 (49%), Gaps = 23/162 (14%)
Query: 408 DSSSTVILGSRIAAMDEFIMKLQMENKLREKERNEAHKKLMK-----KEEEIAALRTKVE 462
D+S ++ ++I + I K + E ++R E KK++K E ++ L ++
Sbjct: 751 DNSKRLLSENKILEQNLNIEKKKKEEVEIHQKRYEQEKKVLKLRVSELENKLEVLAQDLD 810
Query: 463 TAPASEE---------EINLKVNERTRHLRQELEKKLEECQRMTNEFVELERKRMEERIL 513
+A ++ E + NLK E R +++++++K E+ T ++++ ++ E +
Sbjct: 811 SAESTIESKNSDMLLLQNNLKELEELREMKEDIDRKNEQ----TAAILKMQGAQLAELEI 866
Query: 514 QQQEEVEILRKR----LEEIELQLCSSSKQERKDENESKEME 551
+EE ++LRKR +E+++ ++ + +E ES E E
Sbjct: 867 LYKEE-QVLRKRYYNTIEDMKGKIRVYCRIRPLNEKESSERE 907
>At5g27550 kinesin-like protein
Length = 425
Score = 114 bits (286), Expect = 2e-25
Identities = 106/388 (27%), Positives = 181/388 (46%), Gaps = 70/388 (18%)
Query: 90 VKADFGYRDFTLDGVSVSEEEELDLFY--KKFVESRINGVKLGDKCTIMMYGPTGSGKSH 147
+ +D + F D V ++ + +F K V S ++G + I YG TG+GK+
Sbjct: 75 LSSDSSKKHFKFDHVFKPDDGQETVFAQTKPIVTSVLDGYNV----CIFAYGQTGTGKTF 130
Query: 148 TMFGCSKQAGIVYRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYD 207
TM G + G+ YR L ++ ++ S + S V++LE+YNE+I D
Sbjct: 131 TMEGTPENRGVNYRTLEELFRCSESKSHLMKFELS------------VSMLEVYNEKIRD 178
Query: 208 LLSTNGGGGGGGGFGFGWSKSNASKVKLEVMGKKAKNATYISGNEAGKISKE------IQ 261
LL N SN KLEV + A+ + G ++ ++
Sbjct: 179 LLVDN---------------SNQPPKKLEVK-QSAEGTQEVPGLVEAQVYNTDGVWDLLK 222
Query: 262 KVEKRRIVKSTLCNDRSSRSHCMVILDVP---TVGGR-----LMLVDMAGSENIEQAGQT 313
K R V ST N++SSRSHC++ + V + G+ L LVD+AGSE + +
Sbjct: 223 KGYAVRSVGSTAANEQSSRSHCLLRVTVKGENLINGQRTRSHLWLVDLAGSERVGKVEVE 282
Query: 314 GFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCA 373
G K ++ IN+ AL V+ ++A+ SH+P+R+SKLT +LQ+S D K LM +
Sbjct: 283 GERLK-ESQFINKSLSALGDVISALASKTSHIPYRNSKLTHMLQNSLGGD-CKTLMFVQI 340
Query: 374 SPDPKEIHKTISTLEYGAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMEN 433
SP ++ +T+ +L + ++ + I GP + A + E + QM
Sbjct: 341 SPSSADLGETLCSLNFASRVRGIESGP---------------ARKQADVSELLKSKQMAE 385
Query: 434 KLREKERNEAHKKLMKKEEEIAALRTKV 461
KL+ +E K+ K ++ + +L+ ++
Sbjct: 386 KLKHEE-----KETKKLQDNVQSLQLRL 408
>At5g54670 heavy chain polypeptide of kinesin-like protein (katC)
Length = 754
Score = 114 bits (285), Expect = 2e-25
Identities = 101/337 (29%), Positives = 161/337 (46%), Gaps = 44/337 (13%)
Query: 99 FTLDGVSVSEEEELDLFYK--KFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCS--- 153
FT D V + D+F + + V+S ++G K+ I YG TGSGK++TM G
Sbjct: 443 FTFDKVFAPTASQEDVFTEISQLVQSALDGYKV----CIFAYGQTGSGKTYTMMGRPGNV 498
Query: 154 KQAGIVYRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNG 213
++ G++ R L I E S+G+ + +QV++LEIYNE I DLLSTN
Sbjct: 499 EEKGLIPRCLEQIF-------ETRQSLRSQGW----KYELQVSMLEIYNETIRDLLSTNK 547
Query: 214 GGGGGGGFGFGWSKSNASKVKLEVMGKKAKNA-----TYISGNEAGKISKEIQKVEKRRI 268
+ S S K + + N T + + ++S + + R
Sbjct: 548 EAVR--------TDSGVSPQKHAIKHDASGNTHVAELTILDVKSSREVSFLLDHAARNRS 599
Query: 269 VKSTLCNDRSSRSHCMVILDVPTVG--------GRLMLVDMAGSENIEQAGQTGFEAKMQ 320
V T N++SSRSH + L + V G L L+D+AGSE + ++G TG K +
Sbjct: 600 VGKTQMNEQSSRSHFVFTLRISGVNESTEQQVQGVLNLIDLAGSERLSKSGSTGDRLK-E 658
Query: 321 TAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEI 380
T IN+ +L V+ ++A + HVPFR+SKLT LLQ D +K LM + +P+
Sbjct: 659 TQAINKSLSSLGDVIFALAKKEDHVPFRNSKLTYLLQPCLGGD-AKTLMFVNIAPESSST 717
Query: 381 HKTISTLEYGAKAK-CIVRGPHTPVKEEDSSSTVILG 416
+++ +L + A+ C + P + + + LG
Sbjct: 718 GESLCSLRFAARVNACEIGTPRRQTNIKPLENRLSLG 754
Score = 33.5 bits (75), Expect = 0.45
Identities = 39/196 (19%), Positives = 81/196 (40%), Gaps = 64/196 (32%)
Query: 422 MDEFIMKLQMENKLREKERNEAHKKLMKKEEE--------------IAALRTKVETAPAS 467
+ E+ LQ+ N + + +EAH+ + + E+E +AL+ ++ + AS
Sbjct: 201 LQEYNSSLQLYNSKLQGDLDEAHETIKRGEKERTAIIENIGNLKGQFSALQEQLAASKAS 260
Query: 468 EEEI------------NLKV------NERTRHLRQ--ELEKKLEECQRMTNEFVELER-- 505
+E+I +LKV ++R RHL + L+ + + + ELE
Sbjct: 261 QEDIMKQKGELVNEIASLKVELQQVKDDRDRHLVEVKTLQTEATKYNDFKDAITELETTC 320
Query: 506 -------KRMEERILQQQEEVEI---------------------LRKRLEEIELQLCSSS 537
+++++R++ + +++ L+ R+EE EL+L
Sbjct: 321 SSQSTQIRQLQDRLVNSERRLQVSDLSTFEKMNEYEDQKQSIIDLKSRVEEAELKLVEGE 380
Query: 538 KQERKDENESKEMEPN 553
K +K N E++ N
Sbjct: 381 KLRKKLHNTILELKGN 396
>At3g44730 kinesin-like protein heavy chain (KP1)
Length = 1087
Score = 114 bits (285), Expect = 2e-25
Identities = 93/296 (31%), Positives = 145/296 (48%), Gaps = 39/296 (13%)
Query: 124 INGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQA----GIVYRALRDILGDGDTDSEGSDG 179
I V G I YG TGSGK++TM G G+ YRALRD+ + +
Sbjct: 446 IRSVLDGFNVCIFAYGQTGSGKTYTMSGPDLMTETTWGVNYRALRDLFQLSNARTH---- 501
Query: 180 DSSKGFCLGVRTF-VQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSNASKVKLEVM 238
V T+ + V ++EIYNE++ DLL ++G S+ + ++
Sbjct: 502 ---------VVTYEIGVQMIEIYNEQVRDLLVSDGS-----------SRRLDIRNNSQLN 541
Query: 239 GKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVILDVP-------- 290
G +A I + + ++ +K R V +T N+RSSRSH ++ + V
Sbjct: 542 GLNVPDANLIPVSNTRDVLDLMRIGQKNRAVGATALNERSSRSHSVLTVHVQGKELASGS 601
Query: 291 TVGGRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDS 350
+ G L LVD+AGSE +E++ G K + IN+ AL V+ ++A SHVP+R+S
Sbjct: 602 ILRGCLHLVDLAGSERVEKSEAVGERLK-EAQHINKSLSALGDVIYALAQKSSHVPYRNS 660
Query: 351 KLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVRGPHTPVKE 406
KLT +LQDS ++K LM + +P+ + +TISTL++ + I G KE
Sbjct: 661 KLTQVLQDSL-GGQAKTLMFVHINPEVNAVGETISTLKFAQRVASIELGAARSNKE 715
>At3g16060 kinesin like protein
Length = 684
Score = 112 bits (281), Expect = 6e-25
Identities = 123/498 (24%), Positives = 205/498 (40%), Gaps = 74/498 (14%)
Query: 56 EHPIEVIARIRDYPDRKDKPLSVLQASSNSRSIRVKADFGYRDFTLDGVSVSEEEELDLF 115
+ P+ ++ D D + L + + A +F D V + EE D
Sbjct: 176 KRPLNKKESTKNEEDIVDTHANCLTVHETKLKVDLTAYVEKHEFVFDAV-LDEEVSNDEV 234
Query: 116 YKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQAGIVYRALRDILGDGDTDSE 175
Y++ VE + + K T YG TGSGK++TM + +A RDIL
Sbjct: 235 YRETVEPVVPLIFQRIKATCFAYGQTGSGKTYTM------KPLPLKASRDILRLMHHTYR 288
Query: 176 GSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSNASKVKL 235
++GF L V F EIY ++YDLLS K
Sbjct: 289 ------NQGFQLFVSFF------EIYGGKLYDLLSER---------------------KK 315
Query: 236 EVMGKKAKNATYISGNEAGKISKE------IQKVEKRRIVKSTLCNDRSSRSHCMVILDV 289
M + K I G + ++S I++ R +T N+ SSRSH ++ L +
Sbjct: 316 LCMREDGKQQVCIVGLQEYRVSDTDAIMELIERGSATRSTGTTGANEESSRSHAILQLAI 375
Query: 290 -----------PTVGGRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESI 338
P + G+L +D+AGSE + +++ A+IN+ +ALK + ++
Sbjct: 376 KKSVEGNQSKPPRLVGKLSFIDLAGSERGADTTDNDKQTRLEGAEINKSLLALKECIRAL 435
Query: 339 ANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVR 398
N H+PFR SKLT +L+DSF + S+ +MI C SP T++TL Y + K + +
Sbjct: 436 DNDQGHIPFRGSKLTEVLRDSFMGN-SRTVMISCISPSSGSCEHTLNTLRYADRVKSLSK 494
Query: 399 GPHTPVKEEDSSST--------VILGSRIAAMDEFIMKLQ----MENKLREKERNEAHKK 446
G K++ SSST + L S + F + EN + E K+
Sbjct: 495 G--NASKKDVSSSTMNLRESTKIPLSSALPTPSNFDDDVNEMWTEENDEFDASDYEQDKQ 552
Query: 447 LMKKEEEIAALRTKVETAPASEEEINLKVNERTR--HLRQELEKKLEECQRMTNEFVELE 504
+ KK ++ + + I +K + R + + L + + V
Sbjct: 553 MWKKNGKLEPSYNGMAQERIPKPTIQMKSRDMPRPDMKKSNSDDNLNALLQEEEDLVNAH 612
Query: 505 RKRMEERILQQQEEVEIL 522
RK++E+ + +EE+ +L
Sbjct: 613 RKQVEDTMNIVKEEMNLL 630
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.312 0.131 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,341,859
Number of Sequences: 26719
Number of extensions: 933772
Number of successful extensions: 12041
Number of sequences better than 10.0: 575
Number of HSP's better than 10.0 without gapping: 182
Number of HSP's successfully gapped in prelim test: 402
Number of HSP's that attempted gapping in prelim test: 8738
Number of HSP's gapped (non-prelim): 2136
length of query: 754
length of database: 11,318,596
effective HSP length: 107
effective length of query: 647
effective length of database: 8,459,663
effective search space: 5473401961
effective search space used: 5473401961
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 64 (29.3 bits)
Medicago: description of AC144481.6


