

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144481.4 + phase: 0
(152 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
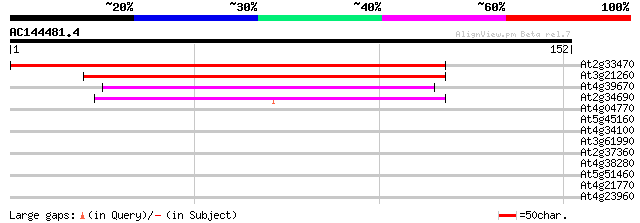
Score E
Sequences producing significant alignments: (bits) Value
At2g33470 unknown protein 187 2e-48
At3g21260 unknown protein 79 7e-16
At4g39670 unknown protein 48 2e-06
At2g34690 Accelerated-Cell-Death11 (ACD11) 42 2e-04
At4g04770 ABC transporter like protein 30 0.66
At5g45160 GTP-binding protein-like; root hair defective 3 protei... 28 1.5
At4g34100 unknown protein 28 1.9
At3g61990 unknown protein 27 3.3
At2g37360 ABC transporter like protein 27 4.3
At4g38280 putative protein 27 5.6
At5g51460 trehalose-6-phosphate phosphatase (AtTPPA) 26 7.3
At4g21770 unknown protein (At4g21770) 26 7.3
At4g23960 putative protein 26 9.5
>At2g33470 unknown protein
Length = 202
Score = 187 bits (475), Expect = 2e-48
Identities = 87/118 (73%), Positives = 100/118 (84%)
Query: 1 MAIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMD 60
M +VKSDIGGNI+RLE YLS+P KF LY+ VQVE+E+K AK SSSCT GLLWLTRAMD
Sbjct: 46 MTLVKSDIGGNITRLEKNYLSDPDKFKYLYTFVQVEIESKIAKGSSSCTNGLLWLTRAMD 105
Query: 61 FLVAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVV 118
FLV +FRNL+ H DWSM QAC DSY KTLKKWHGWLASST ++A+ LAPDRKKFM+V+
Sbjct: 106 FLVELFRNLVAHQDWSMPQACADSYQKTLKKWHGWLASSTFSMALKLAPDRKKFMDVI 163
>At3g21260 unknown protein
Length = 144
Score = 79.3 bits (194), Expect = 7e-16
Identities = 34/98 (34%), Positives = 60/98 (60%)
Query: 21 SNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMDFLVAVFRNLIEHADWSMSQA 80
S+P ++ L +++ E + +++ SC+ LWLTRAMDF +A+ + L++ +M QA
Sbjct: 4 SDPLVYSNLVEILRKEAKEGSSRKPKSCSRAALWLTRAMDFTLALLQRLVKDMSQNMEQA 63
Query: 81 CTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVV 118
+ Y T+K WHGW++S+ VA+ L P+ F+ V+
Sbjct: 64 IEECYNLTIKPWHGWISSAAFKVALKLVPNNNTFINVL 101
>At4g39670 unknown protein
Length = 229
Score = 47.8 bits (112), Expect = 2e-06
Identities = 22/90 (24%), Positives = 44/90 (48%)
Query: 26 FNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMDFLVAVFRNLIEHADWSMSQACTDSY 85
F L++++ ++VE +T K S + L + + +D + A+F + D+S+ A T +Y
Sbjct: 99 FETLHNILDLDVEKETVKTPGSHSRNLRRVRQGLDLIRAIFEQFLIADDYSLKDAATTAY 158
Query: 86 YKTLKKWHGWLASSTVTVAMMLAPDRKKFM 115
+ +H W + V M P R + +
Sbjct: 159 TEVCAPFHTWAVRTAVYAGMYTLPTRDQLL 188
>At2g34690 Accelerated-Cell-Death11 (ACD11)
Length = 206
Score = 41.6 bits (96), Expect = 2e-04
Identities = 22/96 (22%), Positives = 42/96 (42%), Gaps = 1/96 (1%)
Query: 24 AKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMDFLVAVFRNLI-EHADWSMSQACT 82
+ + L ++ ++E + + S T LL + R +D + +F +I D S+ T
Sbjct: 73 SSISTLVVMMDKDIEADCVRKAGSHTRNLLRVKRGLDMVKVLFEQIIASEGDNSLKDPAT 132
Query: 83 DSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVV 118
SY + HGW V++ M P R + ++
Sbjct: 133 KSYAQVFAPHHGWAIRKAVSLGMYALPTRAHLLNML 168
>At4g04770 ABC transporter like protein
Length = 557
Score = 29.6 bits (65), Expect = 0.66
Identities = 16/41 (39%), Positives = 25/41 (60%)
Query: 13 SRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLL 53
SR+ SK +S NC LVQV+ + + AK +S+C + L+
Sbjct: 432 SRIISKGISAGHSRNCYRGLVQVQSKAEGAKNTSTCDSMLI 472
>At5g45160 GTP-binding protein-like; root hair defective 3
protein-like
Length = 834
Score = 28.5 bits (62), Expect = 1.5
Identities = 22/83 (26%), Positives = 43/83 (51%), Gaps = 7/83 (8%)
Query: 64 AVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVVISMLT 123
+++R +++++QA S + K+ + WL + V +M+ +FM ++ + L
Sbjct: 653 SLWRQFKSETEYTVTQAI--SAQEAHKRNNNWLPPAWAIV-LMIVLGFNEFMMLLKNPLY 709
Query: 124 LSNFVLAFLLSLKRITSFWLDLD 146
L F +AFLLS + W+ LD
Sbjct: 710 LLGFFVAFLLS----KALWVQLD 728
>At4g34100 unknown protein
Length = 1051
Score = 28.1 bits (61), Expect = 1.9
Identities = 14/36 (38%), Positives = 18/36 (49%)
Query: 108 APDRKKFMEVVISMLTLSNFVLAFLLSLKRITSFWL 143
AP R F E V+ + + VL F L L + S WL
Sbjct: 128 APSRLPFQEFVVGIAMKACHVLQFFLRLSFVLSVWL 163
>At3g61990 unknown protein
Length = 290
Score = 27.3 bits (59), Expect = 3.3
Identities = 16/40 (40%), Positives = 23/40 (57%), Gaps = 5/40 (12%)
Query: 92 WHGWLASSTVT--VAMMLAPDRKKFME---VVISMLTLSN 126
WHGW+A STV + L KK M+ V ISM+++ +
Sbjct: 243 WHGWVADSTVNDERTISLRNFNKKLMDDQRVSISMVSIGD 282
>At2g37360 ABC transporter like protein
Length = 755
Score = 26.9 bits (58), Expect = 4.3
Identities = 15/49 (30%), Positives = 22/49 (44%)
Query: 95 WLASSTVTVAMMLAPDRKKFMEVVISMLTLSNFVLAFLLSLKRITSFWL 143
W SS VT + P+ VV+++L F +S RI +WL
Sbjct: 591 WAGSSFVTFLSGVIPNVMLGFTVVVAILAYFLLFSGFFISRDRIPVYWL 639
>At4g38280 putative protein
Length = 159
Score = 26.6 bits (57), Expect = 5.6
Identities = 16/42 (38%), Positives = 22/42 (52%)
Query: 6 SDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSS 47
S IG LES +N A +Y +VEV+T A AS++
Sbjct: 21 SSIGAKKPPLESPATTNAASGRLVYVRRRVEVDTSKAAASTT 62
>At5g51460 trehalose-6-phosphate phosphatase (AtTPPA)
Length = 385
Score = 26.2 bits (56), Expect = 7.3
Identities = 13/66 (19%), Positives = 29/66 (43%)
Query: 61 FLVAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVVIS 120
F ++V +E +W++ C D +T K + + ++ D+ K + ++
Sbjct: 252 FCISVHYRNVEEKNWTLVAQCVDDVIRTYPKLRLTHGRKVLEIRPVIDWDKGKAVTFLLE 311
Query: 121 MLTLSN 126
L L+N
Sbjct: 312 SLGLNN 317
>At4g21770 unknown protein (At4g21770)
Length = 472
Score = 26.2 bits (56), Expect = 7.3
Identities = 11/30 (36%), Positives = 22/30 (72%), Gaps = 1/30 (3%)
Query: 18 KYLSNPAKFNCLYSLVQVEVETKTAKASSS 47
+++S+P +F CL SL++ + + +T +SSS
Sbjct: 41 RFISSPKRFTCL-SLLKTDSQNQTTLSSSS 69
>At4g23960 putative protein
Length = 122
Score = 25.8 bits (55), Expect = 9.5
Identities = 18/56 (32%), Positives = 26/56 (46%), Gaps = 14/56 (25%)
Query: 93 HGWLASSTVTVAMMLAPDRKKFMEVVISMLTLSNFVLAFLLSLKRITSFWLDLDWM 148
+GW A VT +M D KF ++ S SL R+TS WL+ D++
Sbjct: 61 NGWKAFFAVTKQVMTVND--KFFSILDSR------------SLPRMTSLWLNSDYV 102
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.327 0.134 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,980,281
Number of Sequences: 26719
Number of extensions: 94300
Number of successful extensions: 343
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 335
Number of HSP's gapped (non-prelim): 13
length of query: 152
length of database: 11,318,596
effective HSP length: 90
effective length of query: 62
effective length of database: 8,913,886
effective search space: 552660932
effective search space used: 552660932
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC144481.4


