

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
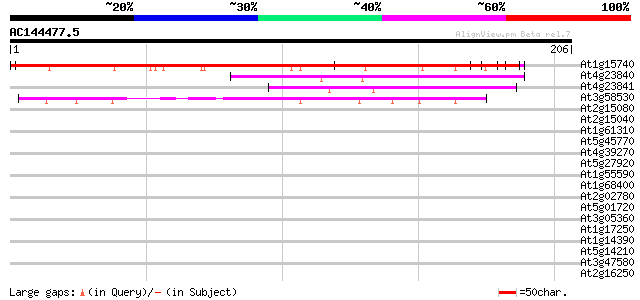
Query= AC144477.5 - phase: 1 /pseudo
(206 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At1g15740 putative leucine rich protein 191 2e-49
At4g23840 leucine rich repeat protein - like 55 3e-08
At4g23841 putative protein 41 5e-04
At3g58530 unknown protein 40 7e-04
At2g15080 putative disease resistance protein 38 0.004
At2g15040 putative disease resistance protein 38 0.004
At1g61310 similar to disease resistance protein 38 0.004
At5g45770 predicted GPI-anchored protein 37 0.006
At4g39270 receptor protein kinase - like protein 37 0.007
At5g27920 unknown protein 37 0.009
At1g55590 unknown protein 36 0.012
At1g68400 receptor kinase like protein 35 0.021
At2g02780 putative receptor-like protein kinase 35 0.028
At5g01720 putative F-box protein family, AtFBL3 35 0.036
At3g05360 putative disease resistance protein 35 0.036
At1g17250 putative receptor protein kinase 35 0.036
At1g14390 hypothetical protein 35 0.036
At5g14210 receptor protein kinase-like protein 34 0.047
At3g47580 receptor kinase-like protein 34 0.062
At2g16250 putative LRR receptor protein kinase 34 0.062
>At1g15740 putative leucine rich protein
Length = 585
Score = 191 bits (486), Expect = 2e-49
Identities = 109/192 (56%), Positives = 144/192 (74%), Gaps = 5/192 (2%)
Query: 1 LQKLSTLNVEGCS-ITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPF-LQV 58
L KL+ LN+EGC +TAAC + ++ALA L LNLNRC SD G EKFS L ++ L +
Sbjct: 284 LNKLNLLNLEGCRHVTAACLDTLTALAGLMYLNLNRCNFSDSGCEKFSDLINLKILNLGM 343
Query: 59 LQA*RG*VLHL---TRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYIS 115
++HL T+L++ +IGDEGLV+L+G+ LKSL LSDTEVG++G+R++S
Sbjct: 344 NNITNSCLVHLKGLTKLESLNLDSCRIGDEGLVHLSGMLELKSLELSDTEVGSNGLRHLS 403
Query: 116 GLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFG 175
GL+ LE +NLSFT VTD+GL++L GLT+L++LNLDAR +TDAGL+ LTSL+GL LDLFG
Sbjct: 404 GLSNLESINLSFTVVTDSGLRKLSGLTSLRTLNLDARHVTDAGLSALTSLTGLTHLDLFG 463
Query: 176 ARITDSGTTYLR 187
ARITDSGT +LR
Sbjct: 464 ARITDSGTNHLR 475
Score = 70.9 bits (172), Expect = 5e-13
Identities = 53/177 (29%), Positives = 86/177 (47%), Gaps = 4/177 (2%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPF-LQVL 59
L L LN+ +IT +C ++ L L LNL+ C + D+G S + + L
Sbjct: 333 LINLKILNLGMNNITNSCLVHLKGLTKLESLNLDSCRIGDEGLVHLSGMLELKSLELSDT 392
Query: 60 QA*RG*VLHLT---RLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISG 116
+ + HL+ L++ + D GL L+GLT L++L L V ++G+ ++
Sbjct: 393 EVGSNGLRHLSGLSNLESINLSFTVVTDSGLRKLSGLTSLRTLNLDARHVTDAGLSALTS 452
Query: 117 LNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDL 173
L L L+L +TD+G L L L+SL + +TD G+ N+ LS L L+L
Sbjct: 453 LTGLTHLDLFGARITDSGTNHLRNLKKLQSLEICGGGLTDTGVKNIKDLSSLTLLNL 509
Score = 60.5 bits (145), Expect = 6e-10
Identities = 52/182 (28%), Positives = 89/182 (48%), Gaps = 5/182 (2%)
Query: 3 KLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHM----FPFLQV 58
+L +L + + + ++S L+ L +NL+ ++D G K S L + V
Sbjct: 383 ELKSLELSDTEVGSNGLRHLSGLSNLESINLSFTVVTDSGLRKLSGLTSLRTLNLDARHV 442
Query: 59 LQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLN 118
A + LT L F +I D G +L L L+SL + + ++G++ I L+
Sbjct: 443 TDAGLSALTSLTGLTHLDLFGARITDSGTNHLRNLKKLQSLEICGGGLTDTGVKNIKDLS 502
Query: 119 KLEDLNLSFTS-VTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGAR 177
L LNLS S +TD L+ + GLT L SLN+ +++ +GL +L L L +L L +
Sbjct: 503 SLTLLNLSQNSNLTDKTLELISGLTGLVSLNVSNSRVSSSGLRHLKPLKNLRSLTLESCK 562
Query: 178 IT 179
++
Sbjct: 563 LS 564
Score = 56.6 bits (135), Expect = 9e-09
Identities = 57/191 (29%), Positives = 86/191 (44%), Gaps = 22/191 (11%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCG-LSDDGFEKFSVLQHMFPFLQVL 59
L L L++E C ++ AL L LN+ C ++D E SVL ++ LQ+
Sbjct: 211 LVNLKKLDLEKCPGIDGGLVHLRALTKLESLNIKWCNCITDADMEPLSVLTNLRS-LQIC 269
Query: 60 QA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTE-VGNSGIRYISGLN 118
+ KI D G+ L GL L L L V + + ++ L
Sbjct: 270 CS-------------------KITDIGISYLKGLNKLNLLNLEGCRHVTAACLDTLTALA 310
Query: 119 KLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARI 178
L LNL+ + +D+G ++ L NLK LNL IT++ L +L L+ L +L+L RI
Sbjct: 311 GLMYLNLNRCNFSDSGCEKFSDLINLKILNLGMNNITNSCLVHLKGLTKLESLNLDSCRI 370
Query: 179 TDSGTTYLRCM 189
D G +L M
Sbjct: 371 GDEGLVHLSGM 381
Score = 51.2 bits (121), Expect = 4e-07
Identities = 48/176 (27%), Positives = 89/176 (50%), Gaps = 7/176 (3%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQ---HMFPF-L 56
L L ++N+ +T + +S L +L LNL+ ++D G + L H+ F
Sbjct: 405 LSNLESINLSFTVVTDSGLRKLSGLTSLRTLNLDARHVTDAGLSALTSLTGLTHLDLFGA 464
Query: 57 QVLQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLS-DTEVGNSGIRYIS 115
++ + + +L +LQ+ + D G+ N+ L+ L L LS ++ + + + IS
Sbjct: 465 RITDSGTNHLRNLKKLQSLEICGGGLTDTGVKNIKDLSSLTLLNLSQNSNLTDKTLELIS 524
Query: 116 GLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANL--TSLSGLI 169
GL L LN+S + V+ +GL+ L L NL+SL L++ +++ + L T L L+
Sbjct: 525 GLTGLVSLNVSNSRVSSSGLRHLKPLKNLRSLTLESCKLSANDIRKLQATDLPNLV 580
Score = 40.8 bits (94), Expect = 5e-04
Identities = 27/65 (41%), Positives = 39/65 (59%), Gaps = 2/65 (3%)
Query: 120 LEDLNLSFTSVTDNGLKRLLGLTNLKSLNLD-ARQITDAGLANLTSLSGLITLDL-FGAR 177
L ++ S + +TD+GL L G TNL+SLN + QI++ GL +L+ LS L +L A
Sbjct: 140 LLSVDFSGSDITDSGLVSLKGCTNLESLNFNFCDQISNRGLVHLSGLSNLTSLSFRRNAA 199
Query: 178 ITDSG 182
IT G
Sbjct: 200 ITAQG 204
>At4g23840 leucine rich repeat protein - like
Length = 192
Score = 55.1 bits (131), Expect = 3e-08
Identities = 39/110 (35%), Positives = 58/110 (52%), Gaps = 2/110 (1%)
Query: 82 IGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRY-ISGLNKLEDLNLSFT-SVTDNGLKRLL 139
+ E + + G L +L LSD + NS + I+GL L +L+LS VTD G+K L
Sbjct: 74 VNAEWMAYIGGFVNLITLNLSDCQRINSSTLWPITGLTSLTELDLSRCFKVTDAGMKHLQ 133
Query: 140 GLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARITDSGTTYLRCM 189
+ NLK L + +T+ G++ L SL L LDL G +TD L+ +
Sbjct: 134 SVVNLKKLWISQTGVTEVGISLLASLKKLSLLDLGGLPVTDQNLISLQVL 183
>At4g23841 putative protein
Length = 364
Score = 40.8 bits (94), Expect = 5e-04
Identities = 36/129 (27%), Positives = 58/129 (44%), Gaps = 38/129 (29%)
Query: 96 LKSLVLSDTEVGNSGIRYISG-LNKLEDLNLSFTSVTD---------------------- 132
LK+L +SDT++ SG+ ++G + +LE L++S T V D
Sbjct: 91 LKNLNVSDTQITPSGVGNLAGHVPQLETLSMSQTFVDDLSILLISTTMPCIKALDLGMNS 150
Query: 133 ---------------NGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGAR 177
L L LT+L++L+L+ + D L+ L+SL+GL L L
Sbjct: 151 TLGFYYLISPQEEKEKSLAALQSLTSLETLSLEHPYLGDKALSGLSSLTGLTHLSLTSTS 210
Query: 178 ITDSGTTYL 186
+TDS +L
Sbjct: 211 LTDSTLHHL 219
Score = 38.9 bits (89), Expect = 0.002
Identities = 26/81 (32%), Positives = 44/81 (54%)
Query: 69 LTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFT 128
LT L+T + +GD+ L L+ LT L L L+ T + +S + ++S L L L +
Sbjct: 174 LTSLETLSLEHPYLGDKALSGLSSLTGLTHLSLTSTSLTDSTLHHLSSLPNLVSLGVRDG 233
Query: 129 SVTDNGLKRLLGLTNLKSLNL 149
+T NGL++ L++L+L
Sbjct: 234 VLTSNGLEKFRPPNRLRTLDL 254
Score = 37.4 bits (85), Expect = 0.006
Identities = 27/91 (29%), Positives = 46/91 (49%), Gaps = 6/91 (6%)
Query: 100 VLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTD---NGLKRLLGLTNLKSLNLDARQITD 156
++S E + + L LE L+L + D +GL L GLT+ L+L + +TD
Sbjct: 157 LISPQEEKEKSLAALQSLTSLETLSLEHPYLGDKALSGLSSLTGLTH---LSLTSTSLTD 213
Query: 157 AGLANLTSLSGLITLDLFGARITDSGTTYLR 187
+ L +L+SL L++L + +T +G R
Sbjct: 214 STLHHLSSLPNLVSLGVRDGVLTSNGLEKFR 244
Score = 33.1 bits (74), Expect = 0.10
Identities = 25/77 (32%), Positives = 42/77 (54%), Gaps = 5/77 (6%)
Query: 88 VNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDN--GLKRLLGLTNLK 145
++ T + + L +S T + N ++ + LE L+LS T+ D+ G +G NLK
Sbjct: 36 LSFTNKSCITYLDVSKTSLKN--FSFLETMFNLEHLDLSSTAFGDDSVGFVACVG-ENLK 92
Query: 146 SLNLDARQITDAGLANL 162
+LN+ QIT +G+ NL
Sbjct: 93 NLNVSDTQITPSGVGNL 109
>At3g58530 unknown protein
Length = 353
Score = 40.4 bits (93), Expect = 7e-04
Identities = 53/181 (29%), Positives = 88/181 (48%), Gaps = 27/181 (14%)
Query: 4 LSTLNVEGC-SITAACFEYIS-ALAALACLNLNRC-GLSDDGFEKFSVLQHMFPFLQVLQ 60
++ LN+ GC S+T + ++ + L LN+ RC ++DDG LQVLQ
Sbjct: 165 ITDLNLSGCKSLTDKSMQLVAESYPDLESLNITRCVKITDDGL------------LQVLQ 212
Query: 61 A*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTE-VGNSGIRYISGLNK 119
L L +A + D+ + ++ L L+ L + + + + GI +I+ NK
Sbjct: 213 K----CFSLQTLNLYA--LSGFTDKAYMKISLLADLRFLDICGAQNISDEGIGHIAKCNK 266
Query: 120 LEDLNLSF-TSVTDNGLKRLL-GLTNLKSLNL-DARQITDAGLANL--TSLSGLITLDLF 174
LE LNL++ +TD G+ + T+L+ L+L +TD L L T + L TLD+
Sbjct: 267 LESLNLTWCVRITDAGVNTIANSCTSLEFLSLFGIVGVTDRCLETLSQTCSTTLTTLDVN 326
Query: 175 G 175
G
Sbjct: 327 G 327
>At2g15080 putative disease resistance protein
Length = 983
Score = 37.7 bits (86), Expect = 0.004
Identities = 44/180 (24%), Positives = 77/180 (42%), Gaps = 12/180 (6%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFP------ 54
L L++ N+ + + I L+ L L L+R + L H+
Sbjct: 183 LSHLTSFNLSYNNFSGRVPSSIGNLSYLTTLRLSRNSFFGELPSSLGSLFHLTDLILDTN 242
Query: 55 -FLQVLQA*RG*VLHLTRLQTHA*-FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIR 112
F+ + + G + HLT + H F+ +I +L L+ L S +LSD +
Sbjct: 243 HFVGKIPSSLGNLSHLTSIDLHKNNFVGEIP----FSLGNLSCLTSFILSDNNIVGEIPS 298
Query: 113 YISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLD 172
LN+L+ LN+ ++ + LL L L +L+L ++T +N++SLS L D
Sbjct: 299 SFGNLNQLDILNVKSNKLSGSFPIALLNLRKLSTLSLFNNRLTGTLPSNMSSLSNLKLFD 358
>At2g15040 putative disease resistance protein
Length = 1011
Score = 37.7 bits (86), Expect = 0.004
Identities = 26/77 (33%), Positives = 36/77 (45%)
Query: 93 LTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDAR 152
L+ L +L LS + + N+L L + T N L LL LTNL L+L
Sbjct: 151 LSHLTNLDLSQNNLVGEIPSFFGSFNQLVSLAVEENEFTGNFLLILLNLTNLSDLSLSRN 210
Query: 153 QITDAGLANLTSLSGLI 169
Q T N++SLS L+
Sbjct: 211 QFTGTLPPNMSSLSNLV 227
Score = 28.9 bits (63), Expect = 2.0
Identities = 21/78 (26%), Positives = 33/78 (41%)
Query: 96 LKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQIT 155
L SL + + E + + + L L DL+LS T + L+NL DA T
Sbjct: 178 LVSLAVEENEFTGNFLLILLNLTNLSDLSLSRNQFTGTLPPNMSSLSNLVLFYADANAFT 237
Query: 156 DAGLANLTSLSGLITLDL 173
++L ++ L DL
Sbjct: 238 GTIPSSLLNIPSLSCFDL 255
Score = 27.3 bits (59), Expect = 5.8
Identities = 18/54 (33%), Positives = 28/54 (51%), Gaps = 2/54 (3%)
Query: 122 DLNLSFTSVTD--NGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDL 173
+LNLS + + N + L NL+ L+L + L++L + S L TLDL
Sbjct: 10 ELNLSSSCLHGLLNSKSNIFSLQNLRFLDLSNNHFSGQILSSLGNFSSLTTLDL 63
>At1g61310 similar to disease resistance protein
Length = 925
Score = 37.7 bits (86), Expect = 0.004
Identities = 40/122 (32%), Positives = 56/122 (45%), Gaps = 11/122 (9%)
Query: 87 LVNLTG-----LTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTD--NGLKRLL 139
L NL+G + L L LSD N ISGL L+ L+LSFT + GLK L
Sbjct: 558 LKNLSGEFIRYMQKLVVLDLSDNRDFNELPEQISGLVSLQYLDLSFTRIEQLPVGLKELK 617
Query: 140 GLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARITDSGTTYLRCMYIYQLQAIVV 199
LT L L AR + +G++ L SL L L G+++ + + LQ + +
Sbjct: 618 KLTFL-DLAYTARLCSISGISRLLSLR---VLSLLGSKVHGDASVLKELQQLENLQDLAI 673
Query: 200 FL 201
L
Sbjct: 674 TL 675
>At5g45770 predicted GPI-anchored protein
Length = 425
Score = 37.4 bits (85), Expect = 0.006
Identities = 30/91 (32%), Positives = 50/91 (53%), Gaps = 6/91 (6%)
Query: 89 NLTGL------TLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLT 142
NLTGL + L+ + LS+ + S I+ L L+ LNLS S++ ++ LT
Sbjct: 182 NLTGLIPKSFHSNLRYIDLSNNSLKGSIRISITRLKNLKSLNLSHNSLSGQIPNKIKSLT 241
Query: 143 NLKSLNLDARQITDAGLANLTSLSGLITLDL 173
LK+L+L + +++ +L+S+S L LDL
Sbjct: 242 FLKNLSLASNKLSGTIPNSLSSISELTHLDL 272
Score = 28.5 bits (62), Expect = 2.6
Identities = 17/60 (28%), Positives = 30/60 (49%)
Query: 90 LTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNL 149
+ LT LK+L L+ ++ + +S +++L L+LS + + NLK LNL
Sbjct: 237 IKSLTFLKNLSLASNKLSGTIPNSLSSISELTHLDLSMNQLNGTVPSFFSEMKNLKHLNL 296
Score = 28.5 bits (62), Expect = 2.6
Identities = 33/118 (27%), Positives = 50/118 (41%), Gaps = 26/118 (22%)
Query: 90 LTGLTLLKSLVLSDTEVGNSGIRYI--SGLNKLEDL-----------------NLSFTSV 130
L L LK+L +S T + S Y+ ++KL L NL + +
Sbjct: 141 LARLKNLKTLYISSTPIQTSRRLYVILGNMHKLTSLTISNSNLTGLIPKSFHSNLRYIDL 200
Query: 131 TDNGLK-----RLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARITDSGT 183
++N LK + L NLKSLNL ++ + SL+ L L L ++ SGT
Sbjct: 201 SNNSLKGSIRISITRLKNLKSLNLSHNSLSGQIPNKIKSLTFLKNLSLASNKL--SGT 256
>At4g39270 receptor protein kinase - like protein
Length = 864
Score = 37.0 bits (84), Expect = 0.007
Identities = 46/159 (28%), Positives = 72/159 (44%), Gaps = 20/159 (12%)
Query: 23 SALAALACLNLNRCGLSDDGFEKFSVLQHMFPFLQVLQA*RG*VLHLTRLQTHA*FI*KI 82
S+L L L+L+ C ++ E + L H+ VL L++ +
Sbjct: 123 SSLLTLEVLDLSSCSITGTIPESLTRLSHLK------------VLDLSKNAIN------- 163
Query: 83 GDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLT 142
GD L +LT L L L LS V S I L+KL+ LNLS ++T + L L+
Sbjct: 164 GDIPL-SLTSLQNLSILDLSSNSVFGSIPANIGALSKLQRLNLSRNTLTSSIPPSLGDLS 222
Query: 143 NLKSLNLDARQITDAGLANLTSLSGLITLDLFGARITDS 181
L L+L ++ + ++L L L TL + G R++ S
Sbjct: 223 VLIDLDLSFNGMSGSVPSDLKGLRNLQTLVIAGNRLSGS 261
Score = 36.6 bits (83), Expect = 0.009
Identities = 46/179 (25%), Positives = 84/179 (46%), Gaps = 25/179 (13%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPFLQVLQ 60
L L L++ CSIT E ++ L+ L L+L++ ++ D + LQ+
Sbjct: 125 LLTLEVLDLSSCSITGTIPESLTRLSHLKVLDLSKNAINGDIPLSLTSLQN--------- 175
Query: 61 A*RG*VLHLTRLQTHA*FI*KIGDEGLV--NLTGLTLLKSLVLSDTEVGNSGIRYISGLN 118
L + L +++ F G + N+ L+ L+ L LS + +S + L+
Sbjct: 176 ------LSILDLSSNSVF-------GSIPANIGALSKLQRLNLSRNTLTSSIPPSLGDLS 222
Query: 119 KLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTS-LSGLITLDLFGA 176
L DL+LSF ++ + L GL NL++L + +++ + +L S LS L +D G+
Sbjct: 223 VLIDLDLSFNGMSGSVPSDLKGLRNLQTLVIAGNRLSGSLPPDLFSLLSKLQIIDFRGS 281
>At5g27920 unknown protein
Length = 642
Score = 36.6 bits (83), Expect = 0.009
Identities = 45/174 (25%), Positives = 79/174 (44%), Gaps = 33/174 (18%)
Query: 4 LSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPFLQVLQA*R 63
L +L+V IT I+ L L L++ C L DDG +F L++ P LQ +
Sbjct: 200 LKSLDVSYLKITNDSIRSIALLVKLEVLDMVSCPLIDDGGLQF--LENGSPSLQEVD--- 254
Query: 64 G*VLHLTRLQTHA*FI*KIGDEGLVNLT----GLTLLKSLVLSDTEVGNSGIRYISGLNK 119
+TR ++ GL+++ + LLK+ +EV S ++YI GL
Sbjct: 255 -----VTRCD-------RVSLSGLISIVRGHPDIQLLKASHCV-SEVSGSFLKYIKGLKH 301
Query: 120 LEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLS--GLITL 171
L+ + + V+D ++L SL+ R + + GL+ ++ G+I+L
Sbjct: 302 LKTIWIDGAHVSD---------SSLVSLSSSCRSLMEIGLSRCVDVTDIGMISL 346
Score = 31.2 bits (69), Expect = 0.40
Identities = 29/99 (29%), Positives = 51/99 (51%), Gaps = 5/99 (5%)
Query: 87 LVNLTGLTLLK-SLVLSDTEVGNSGIRYISGLNKLEDLNLSFT-SVTDNGLKRLLGLTN- 143
L + TGL LK LS ++VG + R + G + L ++L + ++D G+ L +
Sbjct: 142 LSSATGLRELKMDKCLSLSDVGLA--RIVVGCSNLNKISLKWCMEISDLGIDLLCKICKG 199
Query: 144 LKSLNLDARQITDAGLANLTSLSGLITLDLFGARITDSG 182
LKSL++ +IT+ + ++ L L LD+ + D G
Sbjct: 200 LKSLDVSYLKITNDSIRSIALLVKLEVLDMVSCPLIDDG 238
>At1g55590 unknown protein
Length = 607
Score = 36.2 bits (82), Expect = 0.012
Identities = 28/84 (33%), Positives = 44/84 (52%), Gaps = 3/84 (3%)
Query: 106 VGNSGIRYISGLNKLEDLNL-SFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTS 164
+ + ++ + LE L+L S S++D+ L + L L SLNL +TD+G+ L
Sbjct: 376 ITSEAVKKLGLCGNLEVLDLGSCKSISDSCLNSVSALRKLTSLNLAGADVTDSGMLALGK 435
Query: 165 LSGLIT-LDLFGA-RITDSGTTYL 186
IT L L G R++D G +YL
Sbjct: 436 SDVPITQLSLRGCRRVSDRGISYL 459
Score = 34.3 bits (77), Expect = 0.047
Identities = 25/76 (32%), Positives = 41/76 (53%), Gaps = 9/76 (11%)
Query: 106 VGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLD---ARQITDAGLANL 162
+ +S + +S L KL LNL+ VTD+G+ LG +++ L R+++D G++ L
Sbjct: 401 ISDSCLNSVSALRKLTSLNLAGADVTDSGM-LALGKSDVPITQLSLRGCRRVSDRGISYL 459
Query: 163 TSLSGLI-----TLDL 173
+ G I TLDL
Sbjct: 460 LNNEGTISKTLSTLDL 475
Score = 34.3 bits (77), Expect = 0.047
Identities = 23/65 (35%), Positives = 42/65 (64%), Gaps = 2/65 (3%)
Query: 120 LEDLNLSFTS-VTDNGLKRLLGLTNLKSLNLDA-RQITDAGLANLTSLSGLITLDLFGAR 177
L+++ LS +T +K+L NL+ L+L + + I+D+ L ++++L L +L+L GA
Sbjct: 365 LQEVRLSTCPLITSEAVKKLGLCGNLEVLDLGSCKSISDSCLNSVSALRKLTSLNLAGAD 424
Query: 178 ITDSG 182
+TDSG
Sbjct: 425 VTDSG 429
Score = 27.3 bits (59), Expect = 5.8
Identities = 15/40 (37%), Positives = 22/40 (54%), Gaps = 1/40 (2%)
Query: 4 LSTLNVEGC-SITAACFEYISALAALACLNLNRCGLSDDG 42
L L++ C SI+ +C +SAL L LNL ++D G
Sbjct: 390 LEVLDLGSCKSISDSCLNSVSALRKLTSLNLAGADVTDSG 429
>At1g68400 receptor kinase like protein
Length = 670
Score = 35.4 bits (80), Expect = 0.021
Identities = 26/78 (33%), Positives = 42/78 (53%), Gaps = 4/78 (5%)
Query: 89 NLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLN 148
NL+ LT LK L LS+ + + I+ L +L L+LSF + + L LT+L +L
Sbjct: 109 NLSNLTALKLLFLSNNQFSGNFPTSITSLTRLYRLDLSFNNFSGQIPPDLTDLTHLLTLR 168
Query: 149 LDAR----QITDAGLANL 162
L++ QI + L++L
Sbjct: 169 LESNRFSGQIPNINLSDL 186
Score = 28.5 bits (62), Expect = 2.6
Identities = 25/75 (33%), Positives = 37/75 (49%), Gaps = 2/75 (2%)
Query: 99 LVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAG 158
LVL D + S I ++ L L L+L +++ + L LT LK L L Q +
Sbjct: 73 LVLEDINLTGS-ISSLTSLTSLRVLSLKHNNLS-GPIPNLSNLTALKLLFLSNNQFSGNF 130
Query: 159 LANLTSLSGLITLDL 173
++TSL+ L LDL
Sbjct: 131 PTSITSLTRLYRLDL 145
>At2g02780 putative receptor-like protein kinase
Length = 735
Score = 35.0 bits (79), Expect = 0.028
Identities = 31/91 (34%), Positives = 47/91 (51%), Gaps = 2/91 (2%)
Query: 90 LTGLTLLKSLVLSDTEV-GNSGIRYISGLN-KLEDLNLSFTSVTDNGLKRLLGLTNLKSL 147
LT L+ LK+L L+ + G+ + I+ L+ LE LNLS ++ + ++ L NLKSL
Sbjct: 101 LTQLSSLKTLSLTSLGISGSLSPKIITKLSPSLESLNLSSNFISGKIPEEIVSLKNLKSL 160
Query: 148 NLDARQITDAGLANLTSLSGLITLDLFGARI 178
L +L LS L LDL G ++
Sbjct: 161 VLRDNMFWGFVSDDLRGLSNLQELDLGGNKL 191
Score = 27.7 bits (60), Expect = 4.4
Identities = 18/60 (30%), Positives = 33/60 (55%)
Query: 114 ISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDL 173
I LN L+ L+LS T + + L + +L+ L+LD ++ + + + S +ITLD+
Sbjct: 220 IKKLNNLQSLDLSSNEFTGSIPEFLFSIPSLQILSLDQNLLSGSLPNSSCTSSKIITLDV 279
>At5g01720 putative F-box protein family, AtFBL3
Length = 665
Score = 34.7 bits (78), Expect = 0.036
Identities = 30/86 (34%), Positives = 44/86 (50%), Gaps = 4/86 (4%)
Query: 120 LEDLNLSFTSVTDNGLKRLLGLTNLKSLNLD-ARQITDAGLANL-TSLSGLITLDLF-GA 176
LE+L+L+ + D GLK + +L SL L ITD GL+ + S L LDL+
Sbjct: 409 LEELDLTDNEIDDEGLKSISSCLSLSSLKLGICLNITDKGLSYIGMGCSNLRELDLYRSV 468
Query: 177 RITDSGTTYLRCMYIYQLQAIVVFLC 202
ITD G + + I+ L+ I + C
Sbjct: 469 GITDVGISTIAQGCIH-LETINISYC 493
Score = 33.5 bits (75), Expect = 0.080
Identities = 49/174 (28%), Positives = 75/174 (42%), Gaps = 19/174 (10%)
Query: 4 LSTLNVEGCSITAACFEYISALAALACLNLNRC-GLSDDGFEKFSVLQHMFPFLQVLQA* 62
+ TL++ IT C I L L L L C G+ DD + L+H L+ L A
Sbjct: 204 IRTLDLSYLPITGKCLHDILKLQHLEELLLEGCFGVDDDSLKS---LRHDCKSLKKLDA- 259
Query: 63 RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLED 122
+ TH + G + L+ S++ D S ++ +S L +
Sbjct: 260 -----SSCQNLTHRGLTSLLSGAGYLQRLDLSHCSSVISLDFA---SSLKKVSAL---QS 308
Query: 123 LNLSFTSVTDNGLKRLLGLTN-LKSLNLD-ARQITDAGLANLT-SLSGLITLDL 173
+ L SVT +GLK + L N LK ++L +TD GL++L L L LD+
Sbjct: 309 IRLDGCSVTPDGLKAIGTLCNSLKEVSLSKCVSVTDEGLSSLVMKLKDLRKLDI 362
>At3g05360 putative disease resistance protein
Length = 786
Score = 34.7 bits (78), Expect = 0.036
Identities = 23/56 (41%), Positives = 33/56 (58%)
Query: 117 LNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLD 172
L +L LNLS S T N + L LTNL++L+L Q++ +L SLS L T++
Sbjct: 621 LKELRLLNLSGNSFTSNIPQSLANLTNLETLDLSRNQLSGHIPRDLGSLSFLSTMN 676
>At1g17250 putative receptor protein kinase
Length = 756
Score = 34.7 bits (78), Expect = 0.036
Identities = 25/69 (36%), Positives = 33/69 (47%), Gaps = 4/69 (5%)
Query: 81 KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLG 140
KI D+ +T LT LKSL L +G I L++L+ L L ++T L
Sbjct: 290 KINDD----ITHLTKLKSLELYSNHLGGEIPMDIGQLSRLQSLQLHINNITGTVPPSLAN 345
Query: 141 LTNLKSLNL 149
TNL LNL
Sbjct: 346 CTNLVKLNL 354
>At1g14390 hypothetical protein
Length = 747
Score = 34.7 bits (78), Expect = 0.036
Identities = 32/92 (34%), Positives = 49/92 (52%), Gaps = 6/92 (6%)
Query: 90 LTGLTLLKSLVLSDTEVGNSG---IRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKS 146
LT L+ LK+L L +G SG + I + L+ LNLS ++ N K + L NL+S
Sbjct: 103 LTKLSNLKTLSL--VSLGISGPLPSQIIRLSSSLQSLNLSSNFISGNIPKEISSLKNLRS 160
Query: 147 LNLDARQITDAGLANLTSLSGLITLDLFGARI 178
L L A + + + +L LS L L+L G ++
Sbjct: 161 LVL-ANNLFNGSVPDLRGLSNLQELNLGGNKL 191
Score = 33.1 bits (74), Expect = 0.10
Identities = 28/89 (31%), Positives = 47/89 (52%), Gaps = 7/89 (7%)
Query: 96 LKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQIT 155
L+SL LS + + + IS L L L L+ ++ + + L GL+NL+ LNL ++
Sbjct: 134 LQSLNLSSNFISGNIPKEISSLKNLRSLVLA-NNLFNGSVPDLRGLSNLQELNLGGNKLG 192
Query: 156 DAGLANLTSLSGLITLDL----FGARITD 180
+ +L S LIT+ L FG++I +
Sbjct: 193 PEVVPSLA--SNLITISLKNNSFGSKIPE 219
>At5g14210 receptor protein kinase-like protein
Length = 812
Score = 34.3 bits (77), Expect = 0.047
Identities = 24/83 (28%), Positives = 41/83 (48%)
Query: 96 LKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQIT 155
L +++LS R GL++L+ L+LSF +T + L L N+ L+L + +++
Sbjct: 259 LVTVLLSKNSFSGEIPRRFGGLSQLQHLDLSFNHLTGTPSRFLFSLPNISYLDLASNKLS 318
Query: 156 DAGLANLTSLSGLITLDLFGARI 178
NLT L +DL R+
Sbjct: 319 GKLPLNLTCGGKLGFVDLSNNRL 341
>At3g47580 receptor kinase-like protein
Length = 1011
Score = 33.9 bits (76), Expect = 0.062
Identities = 25/85 (29%), Positives = 42/85 (49%)
Query: 89 NLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLN 148
++ ++ L SL LSD G R + L +LE L ++F S+ L + L +L+
Sbjct: 85 SIGNVSFLISLDLSDNAFGGIIPREVGNLFRLEHLYMAFNSLEGGIPATLSNCSRLLNLD 144
Query: 149 LDARQITDAGLANLTSLSGLITLDL 173
L + + + L SL+ L+ LDL
Sbjct: 145 LYSNPLRQGVPSELGSLTKLVILDL 169
Score = 30.8 bits (68), Expect = 0.52
Identities = 16/48 (33%), Positives = 27/48 (55%)
Query: 84 DEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVT 131
D + N+ GL ++ + LS+ ++ S Y + +KLE LNLS + T
Sbjct: 543 DGAIPNIRGLMGVRRVDLSNNDLSGSIPEYFANFSKLEYLNLSINNFT 590
>At2g16250 putative LRR receptor protein kinase
Length = 925
Score = 33.9 bits (76), Expect = 0.062
Identities = 28/87 (32%), Positives = 37/87 (42%)
Query: 93 LTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDAR 152
L L+ L LS V + L L LNLS S+T L L NL L+L
Sbjct: 127 LLALEVLDLSSCSVNGVVPFTLGNLTSLRTLNLSQNSLTSLVPSSLGQLLNLSQLDLSRN 186
Query: 153 QITDAGLANLTSLSGLITLDLFGARIT 179
T + +SL L+TLD+ +T
Sbjct: 187 SFTGVLPQSFSSLKNLLTLDVSSNYLT 213
Score = 26.9 bits (58), Expect = 7.5
Identities = 26/86 (30%), Positives = 39/86 (45%), Gaps = 3/86 (3%)
Query: 87 LVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKS 146
L NLT L L S T + S + + L++L+ SFT V L NL +
Sbjct: 148 LGNLTSLRTLNLSQNSLTSLVPSSLGQLLNLSQLDLSRNSFTGVLPQSFS---SLKNLLT 204
Query: 147 LNLDARQITDAGLANLTSLSGLITLD 172
L++ + +T L +LS LI L+
Sbjct: 205 LDVSSNYLTGPIPPGLGALSKLIHLN 230
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.335 0.148 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,753,578
Number of Sequences: 26719
Number of extensions: 132096
Number of successful extensions: 1138
Number of sequences better than 10.0: 146
Number of HSP's better than 10.0 without gapping: 91
Number of HSP's successfully gapped in prelim test: 56
Number of HSP's that attempted gapping in prelim test: 799
Number of HSP's gapped (non-prelim): 349
length of query: 206
length of database: 11,318,596
effective HSP length: 95
effective length of query: 111
effective length of database: 8,780,291
effective search space: 974612301
effective search space used: 974612301
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC144477.5


