

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144345.5 - phase: 0
(131 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
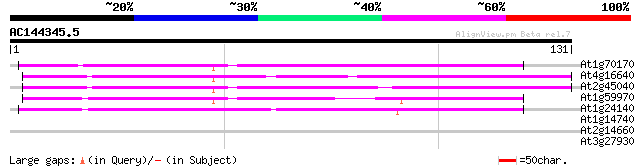
At1g70170 predicted GPI-anchored protein 75 1e-14
At4g16640 proteinase-like protein, predicted GPI-anchored protei... 73 4e-14
At2g45040 metalloproteinase like protein ,predicted GPI-anchored... 71 1e-13
At1g59970 predicted GPI-anchored protein 66 6e-12
At1g24140 metalloproteinase like protein, predicted GPI-anchor 59 5e-10
At1g14740 unknown protein 26 5.1
At2g14660 hypothetical protein 26 6.6
At3g27930 unknown protein 25 8.7
>At1g70170 predicted GPI-anchored protein
Length = 378
Score = 75.1 bits (183), Expect = 1e-14
Identities = 48/119 (40%), Positives = 67/119 (55%), Gaps = 4/119 (3%)
Query: 3 NVFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVD-DVVVGDTVIKLGSNV 61
+VF AF RWS T LNF+ + S+ ++I IGFY ++ DG D V+G + +
Sbjct: 187 SVFSRAFGRWSDVT-ALNFTLSESFSTSDITIGFYTGDHGDGEPFDGVLG--TLAHAFSP 243
Query: 62 NSGFIRLIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYPAI 120
SG L A + WV+ D + DLE+ A+H+IGHLLGL HS +E +MYP I
Sbjct: 244 PSGKFHLDADENWVVSGDLDSFLSVTAAVDLESVAVHEIGHLLGLGHSSVEESIMYPTI 302
>At4g16640 proteinase-like protein, predicted GPI-anchored protein
(by homology)
Length = 364
Score = 73.2 bits (178), Expect = 4e-14
Identities = 46/129 (35%), Positives = 71/129 (54%), Gaps = 6/129 (4%)
Query: 4 VFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVD-DVVVGDTVIKLGSNVN 62
VFR AF++WS V +F E + A++KIGFY ++ DG+ D V+G
Sbjct: 185 VFRRAFSQWSSVIPV-SFEEVDDFTTADLKIGFYAGDHGDGLPFDGVLGTLAHAFAPE-- 241
Query: 63 SGFIRLIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYPAILP 122
+G + L A++ W++ D + DLE+ A H+IGHLLGL HS + VMYP++ P
Sbjct: 242 NGRLHLDAAETWIVDDD--LKGSSEVAVDLESVATHEIGHLLGLGHSSQESAVMYPSLRP 299
Query: 123 LQQQRKLQI 131
++ L +
Sbjct: 300 RTKKVDLTV 308
>At2g45040 metalloproteinase like protein ,predicted GPI-anchored
protein (by homology)
Length = 342
Score = 71.2 bits (173), Expect = 1e-13
Identities = 48/129 (37%), Positives = 72/129 (55%), Gaps = 7/129 (5%)
Query: 4 VFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVD-DVVVGDTVIKLGSNVN 62
VFR AF +W+ V +F ET Y A+IKIGF+N ++ DG D V+G V+ +
Sbjct: 163 VFRRAFGKWASVIPV-SFIETEDYVIADIKIGFFNGDHGDGEPFDGVLG--VLAHTFSPE 219
Query: 63 SGFIRLIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYPAILP 122
+G + L ++ W + D S DLE+ A+H+IGH+LGL HS K+ MYP + P
Sbjct: 220 NGRLHLDKAETWAVDFDEEKSSVA---VDLESVAVHEIGHVLGLGHSSVKDAAMYPTLKP 276
Query: 123 LQQQRKLQI 131
++ L +
Sbjct: 277 RSKKVNLNM 285
>At1g59970 predicted GPI-anchored protein
Length = 360
Score = 65.9 bits (159), Expect = 6e-12
Identities = 46/129 (35%), Positives = 69/129 (52%), Gaps = 24/129 (18%)
Query: 4 VFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVD-DVVVGDTVIKLGSNVN 62
VF AFTRW++ T LNF+ + S A+I IGF++ + DG D +G + S+
Sbjct: 176 VFSRAFTRWAEVTP-LNFTRSESILRADIVIGFFSGEHGDGEPFDGAMG--TLAHASSPP 232
Query: 63 SGFIRLIASKYWVLPTDNYMWSWQNGEF-----------DLETAAMHQIGHLLGLDHSFD 111
+G + L + W++ NGE DLE+ A+H+IGHLLGL HS
Sbjct: 233 TGMLHLDGDEDWLI---------SNGEISRRILPVTTVVDLESVAVHEIGHLLGLGHSSV 283
Query: 112 KEYVMYPAI 120
++ +M+PAI
Sbjct: 284 EDAIMFPAI 292
>At1g24140 metalloproteinase like protein, predicted GPI-anchor
Length = 384
Score = 59.3 bits (142), Expect = 5e-10
Identities = 41/119 (34%), Positives = 60/119 (49%), Gaps = 3/119 (2%)
Query: 3 NVFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVDDVVVGDTVIKLGSNVN 62
+VF AFTRW + T L F+ + ++I IGFY+ + DG T+ S
Sbjct: 191 SVFSRAFTRWEEVTP-LTFTRVERFSTSDISIGFYSGEHGDGEPFDGPMRTLAHAFSPP- 248
Query: 63 SGFIRLIASKYWVLPTDNYMWSWQNGE-FDLETAAMHQIGHLLGLDHSFDKEYVMYPAI 120
+G L + W++ + E DLE+ A+H+IGHLLGL HS + +MYP I
Sbjct: 249 TGHFHLDGEENWIVSGEGGDGFISVSEAVDLESVAVHEIGHLLGLGHSSVEGSIMYPTI 307
>At1g14740 unknown protein
Length = 733
Score = 26.2 bits (56), Expect = 5.1
Identities = 10/22 (45%), Positives = 16/22 (72%)
Query: 18 VLNFSETTSYDDANIKIGFYNI 39
V+ FSE +SYDD ++ F+N+
Sbjct: 93 VVTFSENSSYDDKWVERDFFNL 114
>At2g14660 hypothetical protein
Length = 158
Score = 25.8 bits (55), Expect = 6.6
Identities = 8/18 (44%), Positives = 12/18 (66%)
Query: 72 KYWVLPTDNYMWSWQNGE 89
+YW+L T+ WSW + E
Sbjct: 6 RYWLLKTEPNEWSWSDQE 23
>At3g27930 unknown protein
Length = 425
Score = 25.4 bits (54), Expect = 8.7
Identities = 10/29 (34%), Positives = 16/29 (54%)
Query: 8 AFTRWSQTTRVLNFSETTSYDDANIKIGF 36
AF W + + N S TT++ N++ GF
Sbjct: 331 AFKSWWKPSFAFNISATTNHRTGNVQCGF 359
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.138 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,055,731
Number of Sequences: 26719
Number of extensions: 123958
Number of successful extensions: 245
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 229
Number of HSP's gapped (non-prelim): 8
length of query: 131
length of database: 11,318,596
effective HSP length: 88
effective length of query: 43
effective length of database: 8,967,324
effective search space: 385594932
effective search space used: 385594932
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 54 (25.4 bits)
Medicago: description of AC144345.5


