

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144345.11 - phase: 0
(339 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
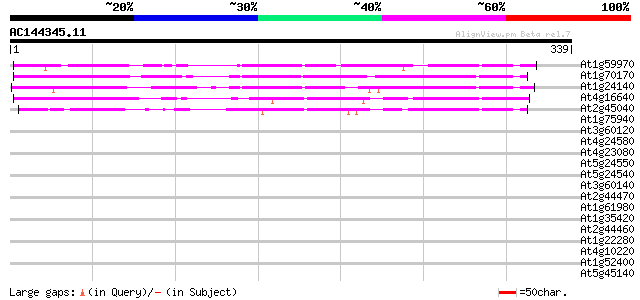
Score E
Sequences producing significant alignments: (bits) Value
At1g59970 predicted GPI-anchored protein 134 9e-32
At1g70170 predicted GPI-anchored protein 128 4e-30
At1g24140 metalloproteinase like protein, predicted GPI-anchor 119 2e-27
At4g16640 proteinase-like protein, predicted GPI-anchored protei... 112 2e-25
At2g45040 metalloproteinase like protein ,predicted GPI-anchored... 111 6e-25
At1g75940 ATA27 35 0.059
At3g60120 beta-glucosidase-like protein 33 0.22
At4g24580 putative protein 32 0.38
At4g23080 putative protein 32 0.50
At5g24550 beta-glucosidase 31 1.1
At5g24540 beta-glucosidase 30 1.4
At3g60140 beta-glucosidase 30 2.5
At2g44470 beta-glucosidase like protein 30 2.5
At1g61980 unknown protein 29 3.2
At1g35420 unknown protein 29 3.2
At2g44460 putative beta-glucosidase 29 4.2
At1g22280 protein phosphatase type 2C like protein 29 4.2
At4g10220 putative protein 28 5.5
At1g52400 beta-glucosidase, putative 28 7.2
At5g45140 DNA-directed RNA polymerase subunit 28 9.3
>At1g59970 predicted GPI-anchored protein
Length = 360
Score = 134 bits (336), Expect = 9e-32
Identities = 111/329 (33%), Positives = 156/329 (46%), Gaps = 75/329 (22%)
Query: 3 IQGLSQIKKHLSTFGYFR---QFLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQ 59
I GLS++K++ FGY FDDVL SAI TYQ+ FNL+VTG L++ TL+
Sbjct: 57 INGLSKLKQYFRRFGYITTTGNCTDDFDDVLQ----SAINTYQKNFNLKVTGKLDSSTLR 112
Query: 60 KISLPRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDM 119
+I PRCG PD+ VS G K + T+K Y F PGK
Sbjct: 113 QIVKPRCGNPDLI--------DGVSEMNGGKIL-RTTEK--YSFFPGKP----------- 150
Query: 120 RYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTTRVL 179
+W PK + LTY F P + ++ F AFTRW++ T L
Sbjct: 151 ------------------RW-PKRKRDLTYAFAPQNNLTDEVKRVFSRAFTRWAEVTP-L 190
Query: 180 NFSETTSYDDADIKIGFYHIYNNDIVDDVVVGDSFISRNLDSNVKSGMIRLERSKFW--- 236
NF+ + S ADI IGF+ + + D + + S+ +GM+ L+ + W
Sbjct: 191 NFTRSESILRADIVIGFF---SGEHGDGEPFDGAMGTLAHASSPPTGMLHLDGDEDWLIS 247
Query: 237 -------VSPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKK 289
+ P TT DLE+V +H+IGHLLGL HSS +++IM+P I ++K
Sbjct: 248 NGEISRRILPVTTVV--------DLESVAVHEIGHLLGLGHSSVEDAIMFPAISG-GDRK 298
Query: 290 VQITDSDNQAIQQLYTNTAKANPNSDHSG 318
V++ D + IQ LY NPN D G
Sbjct: 299 VELAKDDIEGIQHLY----GGNPNGDGGG 323
>At1g70170 predicted GPI-anchored protein
Length = 378
Score = 128 bits (322), Expect = 4e-30
Identities = 105/313 (33%), Positives = 149/313 (47%), Gaps = 42/313 (13%)
Query: 3 IQGLSQIKKHLSTFGYFRQ-FLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQKI 61
+ GL +IKK+ FGY + F F D D+ +A++ YQ FNL VTG L+ T+Q I
Sbjct: 56 VDGLYRIKKYFQRFGYIPETFSGNFTDDFDDILKAAVELYQTNFNLNVTGELDALTIQHI 115
Query: 62 SLPRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMRY 121
+PRCG PD+ G+ + K F + + K+++L
Sbjct: 116 VIPRCGNPDVVN------GTSLMHGGRRKTFEVNFSRTHLHAV--KRYTL---------- 157
Query: 122 EYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTTRVLNF 181
FP +W P+ + LTY F P + ++ F AF RWS T LNF
Sbjct: 158 -----------FPGEPRW-PRNRRDLTYAFDPKNPLTEEVKSVFSRAFGRWSDVT-ALNF 204
Query: 182 SETTSYDDADIKIGFYHIYNNDIVD-DVVVGDSFISRNLDSNVKSGMIRLERSKFWVSPT 240
+ + S+ +DI IGFY + D D V+G + + SG L+ + WV
Sbjct: 205 TLSESFSTSDITIGFYTGDHGDGEPFDGVLGTLAHA----FSPPSGKFHLDADENWVVSG 260
Query: 241 TTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITDSDNQAI 300
DLE+V +H+IGHLLGL HSS +ESIMYP I ++KV +T+ D + I
Sbjct: 261 DLDSFLSVTAAVDLESVAVHEIGHLLGLGHSSVEESIMYPTI-TTGKRKVDLTNDDVEGI 319
Query: 301 QQLYTNTAKANPN 313
Q LY ANPN
Sbjct: 320 QYLY----GANPN 328
>At1g24140 metalloproteinase like protein, predicted GPI-anchor
Length = 384
Score = 119 bits (299), Expect = 2e-27
Identities = 102/324 (31%), Positives = 147/324 (44%), Gaps = 51/324 (15%)
Query: 2 KIQGLSQIKKHLSTFGYFRQFLLK--FDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQ 59
K GL +K++ FGY + L F D D+ +A++ YQ+ F L VTG L+ TL+
Sbjct: 57 KYDGLYMLKQYFQHFGYITETNLSGNFTDDFDDILKNAVEMYQRNFQLNVTGVLDELTLK 116
Query: 60 KISLPRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLP-GKKFSLLRCGIPD 118
+ +PRCG PD+ G G K F G++F ++
Sbjct: 117 HVVIPRCGNPDV--------------VNGTSTMHSGRKTFEVSFAGRGQRFHAVK----- 157
Query: 119 MRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTTRV 178
Y F FP +W P+ + LTY F P + ++ F AFTRW + T
Sbjct: 158 ---HYSF-------FPGEPRW-PRNRRDLTYAFDPRNALTEEVKSVFSRAFTRWEEVTP- 205
Query: 179 LNFSETTSYDDADIKIGFYHIYNNDIVDDVVVGDSFIS--RNLDS--NVKSGMIRLERSK 234
L F+ + +DI IGFY + D G+ F R L + +G L+ +
Sbjct: 206 LTFTRVERFSTSDISIGFYSGEHGD-------GEPFDGPMRTLAHAFSPPTGHFHLDGEE 258
Query: 235 FWVSPTTTYFKKWELQE-FDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQIT 293
W+ + E DLE+V +H+IGHLLGL HSS + SIMYP I +KV +T
Sbjct: 259 NWIVSGEGGDGFISVSEAVDLESVAVHEIGHLLGLGHSSVEGSIMYPTI-RTGRRKVDLT 317
Query: 294 DSDNQAIQQLYTNTAKANPNSDHS 317
D + +Q LY ANPN + S
Sbjct: 318 TDDVEGVQYLY----GANPNFNGS 337
>At4g16640 proteinase-like protein, predicted GPI-anchored protein
(by homology)
Length = 364
Score = 112 bits (281), Expect = 2e-25
Identities = 99/320 (30%), Positives = 138/320 (42%), Gaps = 66/320 (20%)
Query: 3 IQGLSQIKKHLSTFGYFRQFLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQKIS 62
+ G+S++K++L FGY F DV D SAI YQ+ L +TG L+T T+ +S
Sbjct: 67 VSGVSELKRYLHRFGYVNDGSEIFSDVFDGPLESAISLYQENLGLPITGRLDTSTVTLMS 126
Query: 63 LPRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMRYE 122
LPRCG+ D D F T TY GK
Sbjct: 127 LPRCGVSDTHMTINND-------------FLHTTAHYTY--FNGK--------------- 156
Query: 123 YGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKF----SLDMIEGFRNAFTRWSQTTRV 178
PK N+ LTY K S D+ FR AF++WS V
Sbjct: 157 -----------PKWNR------DTLTYAISKTHKLDYLTSEDVKTVFRRAFSQWSSVIPV 199
Query: 179 LNFSETTSYDDADIKIGFYHIYNND-IVDDVVVGD---SFISRNLDSNVKSGMIRLERSK 234
+F E + AD+KIGFY + D + D V+G +F N G + L+ ++
Sbjct: 200 -SFEEVDDFTTADLKIGFYAGDHGDGLPFDGVLGTLAHAFAPEN-------GRLHLDAAE 251
Query: 235 FWVSPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITD 294
W+ K DLE+V H+IGHLLGL HSS + ++MYP + P + KKV +T
Sbjct: 252 TWIVDDD--LKGSSEVAVDLESVATHEIGHLLGLGHSSQESAVMYPSLRP-RTKKVDLTV 308
Query: 295 SDNQAIQQLYTNTAKANPNS 314
D + +LY K +S
Sbjct: 309 DDVAGVLKLYGPNPKLRLDS 328
>At2g45040 metalloproteinase like protein ,predicted GPI-anchored
protein (by homology)
Length = 342
Score = 111 bits (277), Expect = 6e-25
Identities = 102/316 (32%), Positives = 144/316 (45%), Gaps = 73/316 (23%)
Query: 6 LSQIKKHLSTFGYFRQFLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQKISLPR 65
+ +IK+HL +GY Q + DDV E+ A+ YQ+ L +TG +++TL +I LPR
Sbjct: 50 IPEIKRHLQQYGYLPQNK-ESDDVSFEQ---ALVRYQKNLGLPITGKPDSDTLSQILLPR 105
Query: 66 CGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMRYEYGF 125
CG PD DV PK T F GKK+
Sbjct: 106 CGFPD-----------DVE--------PK-----TAPFHTGKKY---------------- 125
Query: 126 DVGSDVSFPKGNKWFPKGTKKLTYGF----LPGKKFSLDMIEGFRNAFTRWSQTTRVLNF 181
V FP +W KLTY F L D+ FR AF +W+ V +F
Sbjct: 126 -----VYFPGRPRWTRDVPLKLTYAFSQENLTPYLAPTDIRRVFRRAFGKWASVIPV-SF 179
Query: 182 SETTSYDDADIKIGFYHIYNND--IVDDV--VVGDSFISRNLDSNVKSGMIRLERSKFWV 237
ET Y ADIKIGF++ + D D V V+ +F N G + L++++ W
Sbjct: 180 IETEDYVIADIKIGFFNGDHGDGEPFDGVLGVLAHTFSPEN-------GRLHLDKAETWA 232
Query: 238 SPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITDSDN 297
+ ++ DLE+V +H+IGH+LGL HSS K++ MYP + P + KKV + D
Sbjct: 233 ---VDFDEEKSSVAVDLESVAVHEIGHVLGLGHSSVKDAAMYPTLKP-RSKKVNLNMDDV 288
Query: 298 QAIQQLYTNTAKANPN 313
+Q LY NPN
Sbjct: 289 VGVQSLY----GTNPN 300
>At1g75940 ATA27
Length = 535
Score = 35.0 bits (79), Expect = 0.059
Identities = 25/80 (31%), Positives = 36/80 (44%), Gaps = 15/80 (18%)
Query: 129 SDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFTR----WSQTTRVLNFSET 184
+D+ F + N FPKG F+ G + +EG N R W T+ F
Sbjct: 33 TDIHFTRAN--FPKG-------FIFGTATAAFQVEGAVNEGCRGPSMWDVYTK--KFPHK 81
Query: 185 TSYDDADIKIGFYHIYNNDI 204
+Y +AD+ + FYH Y DI
Sbjct: 82 CNYHNADVAVDFYHRYKEDI 101
>At3g60120 beta-glucosidase-like protein
Length = 534
Score = 33.1 bits (74), Expect = 0.22
Identities = 23/64 (35%), Positives = 27/64 (41%), Gaps = 2/64 (3%)
Query: 143 GTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQT--TRVLNFSETTSYDDADIKIGFYHIY 200
G GFL G S EG RN R T V + E Y +AD I FY+ Y
Sbjct: 9 GRSDFPEGFLFGTASSAYQYEGARNEAPRGESVWDTFVRKYPERNCYSNADQAIEFYNHY 68
Query: 201 NNDI 204
+DI
Sbjct: 69 KDDI 72
>At4g24580 putative protein
Length = 790
Score = 32.3 bits (72), Expect = 0.38
Identities = 43/189 (22%), Positives = 77/189 (39%), Gaps = 25/189 (13%)
Query: 126 DVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTTRVLNFSETT 185
DV SD P P G+K+ +G PGKK + + ++FS
Sbjct: 481 DVASDTQKPSKLSDAPGGSKR-HWGRTPGKK----------------NLSMESIDFSVEV 523
Query: 186 SYDDADI-KIGFYHIYNNDIVDDVVVGDSFISRNLDSNVKSGMIR-------LERSKFWV 237
D+ADI ++ + + + V ++ + +L+ K+ R L++ V
Sbjct: 524 DEDNADIERLESTKLELQSRITEEVKSNAVLQASLERRKKALYGRRQALEQDLKKDLQEV 583
Query: 238 SPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITDSDN 297
+ K E + DLE + H G G HS+ KES P + ++K + T++ +
Sbjct: 584 AQAEADIAKLEHKVDDLENRLGHHDGKASGSTHSASKESRKLPEHNAKMKEKQKDTEAAS 643
Query: 298 QAIQQLYTN 306
I + T+
Sbjct: 644 THISERSTS 652
>At4g23080 putative protein
Length = 244
Score = 32.0 bits (71), Expect = 0.50
Identities = 32/113 (28%), Positives = 42/113 (36%), Gaps = 24/113 (21%)
Query: 88 GNKWFPKGTKKL--TYGFLPGKKFSLLRCGIPDMRYEYGFDVGS--DVSFPKGNKWFPKG 143
GN W GT L G P K+F E+G +V S S P GN +PKG
Sbjct: 127 GNWWLLMGTGILWEKIGVWPAKRFKESF----GTEIEWGGEVHSPSSPSPPMGNSHYPKG 182
Query: 144 TKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTTRVLNFSETTS-YDDADIKIG 195
T +L + + T W + NF + T Y D K+G
Sbjct: 183 TPRL---------------DSYVRLITTWDENYNADNFVKNTERYSDGCYKVG 220
>At5g24550 beta-glucosidase
Length = 531
Score = 30.8 bits (68), Expect = 1.1
Identities = 13/33 (39%), Positives = 18/33 (54%), Gaps = 2/33 (6%)
Query: 172 WSQTTRVLNFSETTSYDDADIKIGFYHIYNNDI 204
W T F E T+ D+ D+ + FYH Y +DI
Sbjct: 66 WDNFTHA--FPERTNMDNGDVAVDFYHRYKDDI 96
>At5g24540 beta-glucosidase
Length = 534
Score = 30.4 bits (67), Expect = 1.4
Identities = 13/33 (39%), Positives = 17/33 (51%), Gaps = 2/33 (6%)
Query: 172 WSQTTRVLNFSETTSYDDADIKIGFYHIYNNDI 204
W T F E T+ D+ D+ + FYH Y DI
Sbjct: 66 WDNFTHA--FPERTNMDNGDVAVDFYHRYKEDI 96
>At3g60140 beta-glucosidase
Length = 577
Score = 29.6 bits (65), Expect = 2.5
Identities = 12/26 (46%), Positives = 16/26 (61%)
Query: 179 LNFSETTSYDDADIKIGFYHIYNNDI 204
L + E T +AD+ I FYH Y +DI
Sbjct: 65 LTYPERTKMHNADVAIDFYHRYKDDI 90
>At2g44470 beta-glucosidase like protein
Length = 451
Score = 29.6 bits (65), Expect = 2.5
Identities = 11/24 (45%), Positives = 16/24 (65%)
Query: 181 FSETTSYDDADIKIGFYHIYNNDI 204
F E T+ +AD+ + FYH Y +DI
Sbjct: 70 FPERTNMQNADVAVDFYHRYKDDI 93
>At1g61980 unknown protein
Length = 418
Score = 29.3 bits (64), Expect = 3.2
Identities = 18/64 (28%), Positives = 30/64 (46%)
Query: 215 ISRNLDSNVKSGMIRLERSKFWVSPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDK 274
+ R D N++ + +R F V T FK+W E +++ I LGL S D+
Sbjct: 249 VQRLSDKNIEDKVNAYKRLGFDVEYVWTVFKRWPNFLTHSEKKILNTIETFLGLGFSRDE 308
Query: 275 ESIM 278
S++
Sbjct: 309 FSVL 312
>At1g35420 unknown protein
Length = 310
Score = 29.3 bits (64), Expect = 3.2
Identities = 23/102 (22%), Positives = 45/102 (43%), Gaps = 5/102 (4%)
Query: 154 GKKFSLDMIEGFRNAFTRWSQTTRVLNFSETTSYDDADIKIGFYHIYNND---IVDDVVV 210
G + S+D +EG+ + + T +L S+ + D+ + Y + N +V D+
Sbjct: 87 GTEVSVDGVEGYLLTAVKNNNGTGLLLLSDVFGFQDSATRDFAYRVACNGYNVLVPDLFR 146
Query: 211 GDSFISRNLDSNVKSGMIRLERSKFWVSPTTTYFKKWELQEF 252
GD + S + + ++ + TT F KW ++EF
Sbjct: 147 GDPWSKNRPKSEYEEWRRGHDPNR--IRQDTTTFTKWMVEEF 186
>At2g44460 putative beta-glucosidase
Length = 577
Score = 28.9 bits (63), Expect = 4.2
Identities = 11/24 (45%), Positives = 15/24 (61%)
Query: 181 FSETTSYDDADIKIGFYHIYNNDI 204
F E T +AD+ + FYH Y +DI
Sbjct: 70 FPERTRMQNADVAVDFYHRYKDDI 93
>At1g22280 protein phosphatase type 2C like protein
Length = 281
Score = 28.9 bits (63), Expect = 4.2
Identities = 16/58 (27%), Positives = 30/58 (51%), Gaps = 9/58 (15%)
Query: 191 DIKIGFYHIYNNDIVDDVVVGDSFISRNLDSNVKSGMIRLERSKFWVSPTTTYFKKWE 248
D ++G + IY+ + D V +++ + L SN+ L+ +FWV P + K +E
Sbjct: 60 DHELGLFAIYDGHMGDSV---PAYLQKRLFSNI------LKEGEFWVDPRRSIAKAYE 108
>At4g10220 putative protein
Length = 246
Score = 28.5 bits (62), Expect = 5.5
Identities = 20/62 (32%), Positives = 27/62 (43%), Gaps = 6/62 (9%)
Query: 88 GNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMRYEYGFDV--GSDVSFPKGNKWFPKGTK 145
GN W GT GF P +F E+G +V S + P GN +PKG+
Sbjct: 133 GNWWLLMGTSWEEVGFWPSSRFK----ESSGTMVEWGGEVYSPSPPNPPMGNSHYPKGSP 188
Query: 146 KL 147
K+
Sbjct: 189 KV 190
>At1g52400 beta-glucosidase, putative
Length = 528
Score = 28.1 bits (61), Expect = 7.2
Identities = 18/59 (30%), Positives = 25/59 (41%), Gaps = 6/59 (10%)
Query: 150 GFLPGKKFSLDMIEGFRNAFTR----WSQTTRVLNFSETTSYDDADIKIGFYHIYNNDI 204
GF+ G + +EG N R W T+ F +AD+ + FYH Y DI
Sbjct: 47 GFIWGTATAAFQVEGAVNEGCRGPSMWDTFTK--KFPHRCENHNADVAVDFYHRYKEDI 103
>At5g45140 DNA-directed RNA polymerase subunit
Length = 1194
Score = 27.7 bits (60), Expect = 9.3
Identities = 20/77 (25%), Positives = 37/77 (47%), Gaps = 6/77 (7%)
Query: 1 MKIQGLSQIKKHLSTFGYFRQFLLKFDDVLDEETISAI--KTYQQFFNLQV--TGNLNTE 56
+K++GL +K+HL +F YF + + S + Y +F ++V +N
Sbjct: 61 LKVRGL--VKQHLDSFNYFINVGIHKIVKANSRITSTVDPSIYLRFKKVRVGEPSIINVN 118
Query: 57 TLQKISLPRCGIPDMRY 73
T++ I+ C + DM Y
Sbjct: 119 TVENINPHMCRLADMTY 135
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.140 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,009,381
Number of Sequences: 26719
Number of extensions: 362880
Number of successful extensions: 819
Number of sequences better than 10.0: 23
Number of HSP's better than 10.0 without gapping: 12
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 783
Number of HSP's gapped (non-prelim): 31
length of query: 339
length of database: 11,318,596
effective HSP length: 100
effective length of query: 239
effective length of database: 8,646,696
effective search space: 2066560344
effective search space used: 2066560344
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC144345.11


