

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC143341.5 + phase: 0
(485 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
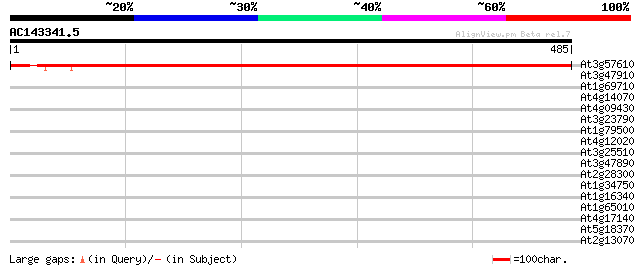
Score E
Sequences producing significant alignments: (bits) Value
At3g57610 adenylosuccinate synthetase 791 0.0
At3g47910 putative protein 32 1.0
At1g69710 putative regulator of chromosome condensation 32 1.0
At4g14070 A6 anther-specific protein (Z97335.16) 31 1.3
At4g09430 putative protein 31 1.3
At3g23790 AMP-binding protein, putative 30 2.3
At1g79500 unknown protein 30 2.3
At4g12020 putative disease resistance protein 30 3.0
At3g25510 unknown protein 30 3.0
At3g47890 putative protein 30 3.9
At2g28300 unknown protein 30 3.9
At1g34750 protein phosphatase type 2C, putative 30 3.9
At1g16340 putative 3-deoxy-D-manno-2-octulosonate-8-phosphate sy... 30 3.9
At1g65010 hypothetical protein 29 5.1
At4g17140 hypothetical protein 29 6.6
At5g18370 disease resistance protein -like 28 8.7
At2g13070 pseudogene 28 8.7
>At3g57610 adenylosuccinate synthetase
Length = 490
Score = 791 bits (2044), Expect = 0.0
Identities = 400/495 (80%), Positives = 437/495 (87%), Gaps = 15/495 (3%)
Query: 1 MNISSLRLDSHA-ICTHQPQRPFAFHNRFP-----RNVVVCAAKPVAPPPTKLAAADS-- 52
M++SSL LDS+ P +H R+P R+ V C+AK A + AADS
Sbjct: 1 MSLSSLTLDSNPRFAVGGP-----YHRRYPPLHHPRSFVSCSAKRPAVSASLSVAADSAA 55
Query: 53 --SLRRIESLSQVSGVLGCQWGDEGKGKLVDILAQHFQIVARCQGGANAGHTIYNAEGKK 110
SL RI SLSQVSGVLGCQWGDEGKGKLVDILAQHF IVARCQGGANAGHTIYN+EGKK
Sbjct: 56 TESLGRIGSLSQVSGVLGCQWGDEGKGKLVDILAQHFDIVARCQGGANAGHTIYNSEGKK 115
Query: 111 FALHLVPSGILNEDTLCVIGNGVVVHLPGLFQEIDNLESNGVSCKGRILISDRAHLLFDF 170
FALHLVPSGILNEDT CVIGNGVVVHLPGLF+EID LESNGVSCKGRIL+SDRAHLLFDF
Sbjct: 116 FALHLVPSGILNEDTTCVIGNGVVVHLPGLFKEIDGLESNGVSCKGRILVSDRAHLLFDF 175
Query: 171 HQTVDGLREAELSKSFIGTTKRGIGPCYSSKANRNGIRVGDLRYMETLPQKLDLLLSDAA 230
HQ VDGLRE+EL+KSFIGTTKRGIGP YSSK RNGIRVGDLR+M+TLPQKLDLLLSDAA
Sbjct: 176 HQEVDGLRESELAKSFIGTTKRGIGPAYSSKVIRNGIRVGDLRHMDTLPQKLDLLLSDAA 235
Query: 231 LRFKDFKYGPDVLREEVEKYKRYAERLEPYIADTVHVMNEAITQKKKVLVEGGQATMLDI 290
RF+ FKY P++LREEVE YKRYA+RLEPYI DTVH +N++I+QKKKVLVEGGQATMLDI
Sbjct: 236 ARFQGFKYTPEMLREEVEAYKRYADRLEPYITDTVHFINDSISQKKKVLVEGGQATMLDI 295
Query: 291 DFGTYPFVTSSSPSAGGICTGLGIAPRVIGDLVGVVKAYTTRVGSGPFPTEILGPGGDLL 350
DFGTYPFVTSSSPSAGGICTGLGIAP V+GDL+GVVKAYTTRVGSGPFPTE LG GGDLL
Sbjct: 296 DFGTYPFVTSSSPSAGGICTGLGIAPSVVGDLIGVVKAYTTRVGSGPFPTENLGTGGDLL 355
Query: 351 RFAGQEFGTTTGRPRRCGWLDIVALSYSCQINGFSSLNLTKLDVLSDLDEIQLGVSYKHA 410
R AGQEFGTTTGRPRRCGWLDIVAL +SCQINGF+SLNLTKLDVLSDL+EIQLGV+YK +
Sbjct: 356 RLAGQEFGTTTGRPRRCGWLDIVALKFSCQINGFASLNLTKLDVLSDLNEIQLGVAYKRS 415
Query: 411 DGTPVQSFPSDLRLLEQLKVEYESLPGWKTDISSIRNYSDLPRAAQLYVERIEELVGVPI 470
DGTPV+SFP DLRLLE+L VEYE LPGWK+DISS+RNYSDLP+AAQ YVERIEELVGVPI
Sbjct: 416 DGTPVKSFPGDLRLLEELHVEYEVLPGWKSDISSVRNYSDLPKAAQQYVERIEELVGVPI 475
Query: 471 HYIGVGPGRDALIFK 485
HYIG+GPGRDALI+K
Sbjct: 476 HYIGIGPGRDALIYK 490
>At3g47910 putative protein
Length = 1291
Score = 31.6 bits (70), Expect = 1.0
Identities = 11/37 (29%), Positives = 23/37 (61%)
Query: 399 DEIQLGVSYKHADGTPVQSFPSDLRLLEQLKVEYESL 435
+E++L +++ DG P+ P +LLE+++ +E L
Sbjct: 500 EEVKLSIAFPPPDGWPISDDPERAKLLEKIRAAFEQL 536
>At1g69710 putative regulator of chromosome condensation
Length = 1028
Score = 31.6 bits (70), Expect = 1.0
Identities = 19/53 (35%), Positives = 24/53 (44%), Gaps = 3/53 (5%)
Query: 314 IAPRVIGDLVGVVKAYTTRVGSGPFPTEILGPGGDLLRFAGQEFGTTTGRPRR 366
I RV GDL G+ Y + V GP+ T ++ G L F FG RR
Sbjct: 383 IPKRVTGDLQGL---YVSDVACGPWHTAVVASSGQLFTFGDGTFGALGHGDRR 432
>At4g14070 A6 anther-specific protein (Z97335.16)
Length = 727
Score = 31.2 bits (69), Expect = 1.3
Identities = 17/55 (30%), Positives = 27/55 (48%)
Query: 228 DAALRFKDFKYGPDVLREEVEKYKRYAERLEPYIADTVHVMNEAITQKKKVLVEG 282
DAALR ++K PD+ R EKY ++PY + + + + Q+ EG
Sbjct: 88 DAALRSSEWKAVPDIWRSSAEKYGDRVALVDPYHDPPLKLTYKQLEQEILDFAEG 142
>At4g09430 putative protein
Length = 1039
Score = 31.2 bits (69), Expect = 1.3
Identities = 21/82 (25%), Positives = 39/82 (46%), Gaps = 3/82 (3%)
Query: 390 TKLDVLSDLDEIQLGVSYKHADGTPVQSFPSDLR--LLEQLKVEYESLPGWKTDISSIRN 447
+KL ++SD+ I G+ H D P+++ P + L ++ + Y +L + D + +
Sbjct: 529 SKLQLISDVSSITHGLKLLHWDAYPLETLPFSFQSSTLVEINLRYSNLKHF-WDETKVYR 587
Query: 448 YSDLPRAAQLYVERIEELVGVP 469
LP +L V LV +P
Sbjct: 588 SKQLPNLRRLDVTGSTSLVELP 609
>At3g23790 AMP-binding protein, putative
Length = 722
Score = 30.4 bits (67), Expect = 2.3
Identities = 28/88 (31%), Positives = 42/88 (46%), Gaps = 7/88 (7%)
Query: 174 VDGLREAELSKSF-IGTTKRGIGPCYSSKANRNGIRVGDLRYMETLPQKLDLLLSDAALR 232
V G+R K+F T++RG SK I+ +LR ++L L +AAL
Sbjct: 23 VSGMRLVFGYKAFGCRTSRRGFRVRCESK-----IQEKELRRCSPFLERLSLP-REAALS 76
Query: 233 FKDFKYGPDVLREEVEKYKRYAERLEPY 260
++K PD+ R VEKY ++PY
Sbjct: 77 SNEWKSVPDIWRSSVEKYGDRVAVVDPY 104
>At1g79500 unknown protein
Length = 290
Score = 30.4 bits (67), Expect = 2.3
Identities = 18/54 (33%), Positives = 25/54 (45%)
Query: 91 ARCQGGANAGHTIYNAEGKKFALHLVPSGILNEDTLCVIGNGVVVHLPGLFQEI 144
A C A+ H++ GKK V SG L E C+ V V + G+F E+
Sbjct: 188 ANCPVVADITHSLQQPAGKKLDGGGVASGGLRELIPCIARTAVAVGVDGIFMEV 241
>At4g12020 putative disease resistance protein
Length = 1895
Score = 30.0 bits (66), Expect = 3.0
Identities = 23/82 (28%), Positives = 38/82 (46%), Gaps = 4/82 (4%)
Query: 387 LNLTKLDVLSDLDEIQLGVSYKHADGTPVQSFPSDLR---LLEQLKVE-YESLPGWKTDI 442
LNL+ L + EI V + GT +Q PS ++ LLE+L +E L T I
Sbjct: 1333 LNLSGCSKLGNFPEISPNVKELYMGGTMIQEIPSSIKNLVLLEKLDLENSRHLKNLPTSI 1392
Query: 443 SSIRNYSDLPRAAQLYVERIEE 464
+++ L + + +ER +
Sbjct: 1393 YKLKHLETLNLSGCISLERFPD 1414
>At3g25510 unknown protein
Length = 1981
Score = 30.0 bits (66), Expect = 3.0
Identities = 19/69 (27%), Positives = 30/69 (42%), Gaps = 2/69 (2%)
Query: 366 RCGWLDIVALSYSCQINGFSSLNLTKLDVLSDLDEIQLGVSYKHADGTPVQSFPSDLRLL 425
RC L+ AL + + L+LT EI + + DGT V+ PS ++
Sbjct: 982 RCQKLE--ALPSNINLKSLERLDLTDCSQFKSFPEISTNIECLYLDGTAVEEVPSSIKSW 1039
Query: 426 EQLKVEYES 434
+L V + S
Sbjct: 1040 SRLTVLHMS 1048
>At3g47890 putative protein
Length = 1528
Score = 29.6 bits (65), Expect = 3.9
Identities = 12/40 (30%), Positives = 24/40 (60%)
Query: 396 SDLDEIQLGVSYKHADGTPVQSFPSDLRLLEQLKVEYESL 435
+D +E +L +++ DG P+ P +LLE+++ +E L
Sbjct: 457 NDPEEGKLSITFPPPDGWPISDDPERAKLLEKIRAAFELL 496
>At2g28300 unknown protein
Length = 2218
Score = 29.6 bits (65), Expect = 3.9
Identities = 14/52 (26%), Positives = 26/52 (49%)
Query: 426 EQLKVEYESLPGWKTDISSIRNYSDLPRAAQLYVERIEELVGVPIHYIGVGP 477
E+ + E +P D+ ++ +D+P + E ++E+V VP GV P
Sbjct: 1270 EKTNISSEQVPDVSHDLKVSQDQTDIPPVGGIVPENLQEIVDVPASPHGVVP 1321
>At1g34750 protein phosphatase type 2C, putative
Length = 282
Score = 29.6 bits (65), Expect = 3.9
Identities = 29/122 (23%), Positives = 47/122 (37%), Gaps = 17/122 (13%)
Query: 216 ETLPQKLDLLLSDAALRFKDFKYGPDVLREEVEKYKRYAERLEPYIADTVHVMNEAIT-- 273
E +P L L L+ + F+Y P R + Y++ + + + +D + A+T
Sbjct: 76 ERVPAYLQKHLFSNILKEEQFRYDPQ--RSIIAAYEKTDQAILSHSSDLGRGGSTAVTAI 133
Query: 274 -------------QKKKVLVEGGQATMLDIDFGTYPFVTSSSPSAGGICTGLGIAPRVIG 320
+ VL +GGQA + ID + S G + G PRV G
Sbjct: 134 LMNGRRLWVANVGDSRAVLSQGGQAIQMTIDHEPHTERLSIEGKGGFVSNMPGDVPRVNG 193
Query: 321 DL 322
L
Sbjct: 194 QL 195
>At1g16340 putative 3-deoxy-D-manno-2-octulosonate-8-phosphate
synthase
Length = 291
Score = 29.6 bits (65), Expect = 3.9
Identities = 18/54 (33%), Positives = 25/54 (45%)
Query: 91 ARCQGGANAGHTIYNAEGKKFALHLVPSGILNEDTLCVIGNGVVVHLPGLFQEI 144
A C A+ H++ GKK V SG L E C+ V V + G+F E+
Sbjct: 189 ADCPVVADITHSLQQPAGKKLDGGGVASGGLRELIPCIARTAVAVGVDGIFMEV 242
>At1g65010 hypothetical protein
Length = 1318
Score = 29.3 bits (64), Expect = 5.1
Identities = 18/45 (40%), Positives = 26/45 (57%), Gaps = 1/45 (2%)
Query: 422 LRLLEQLKVEYESLPGWKTDISSI-RNYSDLPRAAQLYVERIEEL 465
L+ +E+L V ESL TD+ SI + DL Y+++IEEL
Sbjct: 692 LKKIEELSVANESLADNVTDLQSIVQESKDLKEREVAYLKKIEEL 736
>At4g17140 hypothetical protein
Length = 1322
Score = 28.9 bits (63), Expect = 6.6
Identities = 20/58 (34%), Positives = 28/58 (47%), Gaps = 3/58 (5%)
Query: 380 QINGFSSLNLTKLDVLSDLDEIQLGVSYKHADGTPVQSFPSDLRLLEQLKVEYESLPG 437
Q +GF NL + V LDE+++ SY H D SF + L E E+ +L G
Sbjct: 789 QKDGFDLSNLESVYVTGVLDELKICFSYGHQDDA---SFMAVLLARESKLFEFRALGG 843
>At5g18370 disease resistance protein -like
Length = 1210
Score = 28.5 bits (62), Expect = 8.7
Identities = 25/124 (20%), Positives = 45/124 (36%), Gaps = 29/124 (23%)
Query: 374 ALSYSCQINGFSSLNLTKLDVLSDLDEIQLGVSYKHADGTPVQSFPSDLRLLEQL----- 428
AL + S L L L EI + + GT ++ PS +RL +L
Sbjct: 776 ALPGDSNMRSLSKLLLNGSSRLKTFPEISTNIQELNLSGTAIEEVPSSIRLWSRLDKLDM 835
Query: 429 ----------------------KVEYESLPGWKTDISSIRNYSDL--PRAAQLYVERIEE 464
+ E E +P W ++S +R++ + + + + RI +
Sbjct: 836 SRCKNLKMFPPVPDGISVLNLSETEIEDIPPWVENLSQLRHFVMIRCKKLDNISLSRISK 895
Query: 465 LVGV 468
+ GV
Sbjct: 896 MEGV 899
>At2g13070 pseudogene
Length = 239
Score = 28.5 bits (62), Expect = 8.7
Identities = 19/56 (33%), Positives = 26/56 (45%), Gaps = 2/56 (3%)
Query: 360 TTGRPRRCGWLDIVALSYSCQINGFSSLNLTKLDVLSDLDEIQLGVSYKHADGTPV 415
T RPR LD L C+ S + L+KLD+L L+ +G GTP+
Sbjct: 153 TVERPRLQAELD--GLEAQCKSKEVSDVTLSKLDILPVLEMSVVGPMDVDKQGTPI 206
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.140 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,316,978
Number of Sequences: 26719
Number of extensions: 512799
Number of successful extensions: 1126
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 1114
Number of HSP's gapped (non-prelim): 20
length of query: 485
length of database: 11,318,596
effective HSP length: 103
effective length of query: 382
effective length of database: 8,566,539
effective search space: 3272417898
effective search space used: 3272417898
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC143341.5


