

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC143339.7 - phase: 0
(358 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
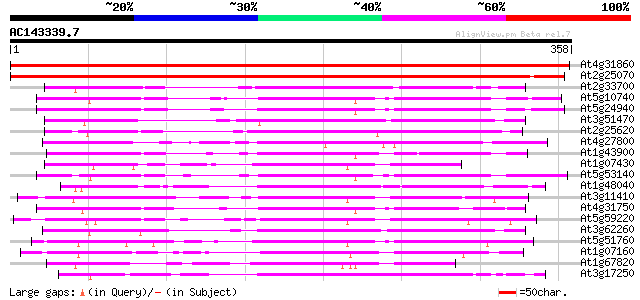
Score E
Sequences producing significant alignments: (bits) Value
At4g31860 protein phosphatase 2C - like protein 536 e-153
At2g25070 putative protein phosphatase 2C 524 e-149
At2g33700 putative protein phosphatase 2C 142 3e-34
At5g10740 protein phosphatase 2C -like protein 137 9e-33
At5g24940 Protein phosphatase 2C-like protein 137 1e-32
At3g51470 protein phosphatase 2C-like protein 135 3e-32
At2g25620 protein phosphatase 2C like protein 135 3e-32
At4g27800 protein phosphatase homolog (PPH1) 134 6e-32
At1g43900 protein phosphatase type 2C like protein 130 1e-30
At1g07430 protein phosphatase 2C, putative 127 1e-29
At5g53140 protein phosphatase 2C-like 124 6e-29
At1g48040 protein phosphatase-2C, putative 123 1e-28
At3g11410 protein phosphatase 2C (PP2C) 122 2e-28
At4g31750 unknown protein 122 4e-28
At5g59220 protein phosphatase 2C - like 121 5e-28
At3g62260 unknown protein 119 2e-27
At5g51760 protein phosphatase-2C; PP2C-like protein 119 3e-27
At1g07160 unknown protein 115 4e-26
At1g67820 protein phosphatase-like protein 111 7e-25
At3g17250 protein phosphatase like 106 2e-23
>At4g31860 protein phosphatase 2C - like protein
Length = 357
Score = 536 bits (1380), Expect = e-153
Identities = 254/357 (71%), Positives = 302/357 (84%)
Query: 1 MGTTLSIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHLDVDSSTSFFGVYDGHG 60
MG LS PKT+KFSEDGEN LRYGLSSMQGWRA+MEDAHAA LD+D +TSF GVYDGHG
Sbjct: 1 MGIYLSTPKTDKFSEDGENHKLRYGLSSMQGWRASMEDAHAAILDLDDNTSFLGVYDGHG 60
Query: 61 GKAVAKFCAKHLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKG 120
GK V+KFCAK+LHQQVL E Y AGDVGTSL KAF RMDEMM+GQRGWRELAVLGDK+
Sbjct: 61 GKVVSKFCAKYLHQQVLSDEAYAAGDVGTSLQKAFFRMDEMMQGQRGWRELAVLGDKINK 120
Query: 121 FNGIIEGLIRSPRSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGD 180
F+G+IEGLI SPRS D+ ++ D WAFE+GPHSDF GPNSGSTACVA++R+ LFVANAGD
Sbjct: 121 FSGMIEGLIWSPRSGDSANKPDAWAFEEGPHSDFAGPNSGSTACVAVVRDKQLFVANAGD 180
Query: 181 SRCVISRNGQAYNLSRDHKPELVIEKERIYKAGGFIHAGRINGSLNLARAIGDVDFKNNR 240
SRCVISR QAYNLSRDHKP+L EKERI KAGGFIHAGR+NGSLNL+RAIGD++FK N+
Sbjct: 181 SRCVISRKNQAYNLSRDHKPDLEAEKERILKAGGFIHAGRVNGSLNLSRAIGDMEFKQNK 240
Query: 241 FLSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEV 300
FL +EKQ+VTA+PD+N V+L D+D+F+V+ACDGIWDC++SQQLVDF+ ++L ETKLS V
Sbjct: 241 FLPSEKQIVTASPDVNTVELCDDDDFLVLACDGIWDCMTSQQLVDFIHEQLNSETKLSVV 300
Query: 301 CERVLDRCLAPSLAVGDGCDNMTMILVQFKKPLQTSAPAQEQSSSNEQDASEPTLEN 357
CE+VLDRCLAP+ + G+GCDNMTMILV+FK P + + ++S E + EP+ N
Sbjct: 301 CEKVLDRCLAPNTSGGEGCDNMTMILVRFKNPTPSETELKPEASQAEGNHDEPSSSN 357
>At2g25070 putative protein phosphatase 2C
Length = 355
Score = 524 bits (1349), Expect = e-149
Identities = 250/354 (70%), Positives = 292/354 (81%), Gaps = 2/354 (0%)
Query: 1 MGTTLSIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHLDVDSSTSFFGVYDGHG 60
MGT LS PKTEK SEDGEND LR+GLSSMQGWRATMEDAHAA LD+D TSFFGVYDGHG
Sbjct: 1 MGTYLSSPKTEKLSEDGENDKLRFGLSSMQGWRATMEDAHAAILDLDDKTSFFGVYDGHG 60
Query: 61 GKAVAKFCAKHLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKG 120
GK VAKFCAK+LHQQV+ +E Y GDV TSL +AF RMD+MM+GQRGWRELAVLGDK+
Sbjct: 61 GKVVAKFCAKYLHQQVISNEAYKTGDVETSLRRAFFRMDDMMQGQRGWRELAVLGDKMNK 120
Query: 121 FNGIIEGLIRSPRSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGD 180
F+G+IEG I SPRS D +Q D W E GPHSDF GP SG TACVA+I++ LFVANAGD
Sbjct: 121 FSGMIEGFIWSPRSGDTNNQPDSWPLEDGPHSDFTGPTSGCTACVALIKDKKLFVANAGD 180
Query: 181 SRCVISRNGQAYNLSRDHKPELVIEKERIYKAGGFIHAGRINGSLNLARAIGDVDFKNNR 240
SRCVISR QAYNLS+DHKP+L +EKERI KAGGFIHAGRINGSLNL RAIGD++FK N+
Sbjct: 181 SRCVISRKSQAYNLSKDHKPDLEVEKERILKAGGFIHAGRINGSLNLTRAIGDMEFKQNK 240
Query: 241 FLSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEV 300
FL +EKQ+VTA+PDIN +DL D+D+F+V+ACDGIWDC+SSQ+LVDF+ ++L ETKLS V
Sbjct: 241 FLPSEKQMVTADPDINTIDLCDDDDFLVVACDGIWDCMSSQELVDFIHEQLKSETKLSTV 300
Query: 301 CERVLDRCLAPSLAVGDGCDNMTMILVQFKKPLQTSAPAQEQSSSNEQDASEPT 354
CE+V+DRCLAP A G+GCDNMT+ILVQFKKP + + + S E EP+
Sbjct: 301 CEKVVDRCLAPDTATGEGCDNMTIILVQFKKP--NPSETEPEDSKPEPSEDEPS 352
>At2g33700 putative protein phosphatase 2C
Length = 380
Score = 142 bits (358), Expect = 3e-34
Identities = 100/317 (31%), Positives = 148/317 (46%), Gaps = 66/317 (20%)
Query: 23 RYGLSSMQGWRATMEDAH----------AAHLDVDSSTSFFGVYDGHGGKAVAKFCAKHL 72
R G + QG + MED H A + S +F+GV+DGHGG A F K++
Sbjct: 84 RSGSCAEQGAKQFMEDEHICIDDLVNHLGAAIQCSSLGAFYGVFDGHGGTDAAHFVRKNI 143
Query: 73 HQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKGFNGIIEGLIRSP 132
+ +++ + V ++ AFL+ D
Sbjct: 144 LRFIVEDSSFPLC-VKKAIKSAFLKAD--------------------------------- 169
Query: 133 RSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGDSRCVISRNGQAY 192
+E S D +SG+TA A I L +ANAGD R V+ R G+A
Sbjct: 170 -------------YEFADDSSLD-ISSGTTALTAFIFGRRLIIANAGDCRAVLGRRGRAI 215
Query: 193 NLSRDHKPELVIEKERIYKAGGFIHAGRINGSLNLARAIGDVDFKNNRFLSAEKQVVTAN 252
LS+DHKP EK RI K GG ++ G +NG L++ARAIGD K + + ++
Sbjct: 216 ELSKDHKPNCTAEKVRIEKLGGVVYDGYLNGQLSVARAIGDWHMKGPKGSACP---LSPE 272
Query: 253 PDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEVCERVLDRCLAPS 312
P++ DL ++DEF+++ CDG+WD +SSQ V R+EL++ E C R L R
Sbjct: 273 PELQETDLSEDDEFLIMGCDGLWDVMSSQCAVTIARKELMIHND-PERCSRELVR----E 327
Query: 313 LAVGDGCDNMTMILVQF 329
+ CDN+T+I+V F
Sbjct: 328 ALKRNTCDNLTVIVVCF 344
>At5g10740 protein phosphatase 2C -like protein
Length = 354
Score = 137 bits (345), Expect = 9e-33
Identities = 107/340 (31%), Positives = 165/340 (48%), Gaps = 72/340 (21%)
Query: 18 ENDNLRYGLSSMQGWRATMEDAHAAHLDVDSS--TSFFGVYDGHGGKAVAKFCAKHLHQQ 75
+N YG +S G R++MED +D + FGV+DGHGG A++ +HL
Sbjct: 28 QNGKFSYGYASSAGKRSSMEDFFETRIDGINGEIVGLFGVFDGHGGARAAEYVKRHLFSN 87
Query: 76 VLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKGFNGIIEGLIRSPRSN 135
++ ++I+ D +++T A+ D L++S S+
Sbjct: 88 LITHPKFIS-DTKSAITDAYNHTDSE--------------------------LLKSENSH 120
Query: 136 DNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGDSRCVISRNGQAYNLS 195
N+D +GSTA AI+ + L VAN GDSR VISR G+A +S
Sbjct: 121 -NRD-------------------AGSTASTAILVGDRLVVANVGDSRAVISRGGKAIAVS 160
Query: 196 RDHKPELVIEKERIYKAGGFI-HAG--RINGSLNLARAIGDVDFKNNRFLSAEKQVVTAN 252
RDHKP+ E+ERI AGGF+ AG R+ G L ++RA GD R L KQ V A+
Sbjct: 161 RDHKPDQSDERERIENAGGFVMWAGTWRVGGVLAVSRAFGD------RLL---KQYVVAD 211
Query: 253 PDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEVCERVLDRCLAPS 312
P+I + D EF+++A DG+WD S++ V V++ E ++ + R
Sbjct: 212 PEIQEEKIDDTLEFLILASDGLWDVFSNEAAVAMVKEVEDPEDSAKKLVGEAIKR----- 266
Query: 313 LAVGDGCDNMTMILVQFKKPLQTSAPAQEQSSSNEQDASE 352
DN+T ++V+F + + SA + SSS+ ++A E
Sbjct: 267 ----GSADNITCVVVRFLE--KKSASSSHISSSSSKEAKE 300
>At5g24940 Protein phosphatase 2C-like protein
Length = 447
Score = 137 bits (344), Expect = 1e-32
Identities = 104/342 (30%), Positives = 162/342 (46%), Gaps = 73/342 (21%)
Query: 18 ENDNLRYGLSSMQGWRATMEDAHAAHLD-VDSS-TSFFGVYDGHGGKAVAKFCAKHLHQQ 75
+N YG +S G R++MED +D +D FGV+DGHGG A++ +HL
Sbjct: 28 QNGKFSYGYASSAGKRSSMEDFFETRIDGIDGEIVGLFGVFDGHGGSRAAEYVKRHLFSN 87
Query: 76 VLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKGFNGIIEGLIRSPRSN 135
++ ++I+ D +++ A+ D L++S S+
Sbjct: 88 LITHPKFIS-DTKSAIADAYTHTDSE--------------------------LLKSENSH 120
Query: 136 DNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGDSRCVISRNGQAYNLS 195
++GSTA AI+ + L VAN GDSR VI R G A+ +S
Sbjct: 121 TR--------------------DAGSTASTAILVGDRLLVANVGDSRAVICRGGNAFAVS 160
Query: 196 RDHKPELVIEKERIYKAGGFI-HAG--RINGSLNLARAIGDVDFKNNRFLSAEKQVVTAN 252
RDHKP+ E+ERI AGGF+ AG R+ G L ++RA GD R L KQ V A+
Sbjct: 161 RDHKPDQSDERERIENAGGFVMWAGTWRVGGVLAVSRAFGD------RLL---KQYVVAD 211
Query: 253 PDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEVCERVLDRCLAPS 312
P+I + D EF+++A DG+WD S+++ V V++ E ++ + R
Sbjct: 212 PEIQEEKIDDSLEFLILASDGLWDVFSNEEAVAVVKEVEDPEESTKKLVGEAIKR----- 266
Query: 313 LAVGDGCDNMTMILVQFKKPLQTSAPAQEQSSSNEQDASEPT 354
DN+T ++V+F L++ + SSS+E+ PT
Sbjct: 267 ----GSADNITCVVVRF---LESKSANNNGSSSSEEANQVPT 301
>At3g51470 protein phosphatase 2C-like protein
Length = 361
Score = 135 bits (341), Expect = 3e-32
Identities = 93/316 (29%), Positives = 149/316 (46%), Gaps = 69/316 (21%)
Query: 23 RYGLSSMQGWRATMEDAHAAHLDV-----DSSTSFFGVYDGHGGKAVAKFCAKHLHQQVL 77
R G S +G + +MED D+ S+ +F+GV+DGHGG A F K++ + V+
Sbjct: 72 RSGSWSDKGPKQSMEDEFICVDDLTEYIGSSTGAFYGVFDGHGGVDAASFTKKNIMKLVM 131
Query: 78 KSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKGFNGIIEGLIRSPRSNDN 137
+ + + P S
Sbjct: 132 EDKHF-------------------------------------------------PTSTKK 142
Query: 138 KDQSDDWAFEKGPHSDFDGPN----SGSTACVAIIRNNLLFVANAGDSRCVISRNGQAYN 193
+S AF K H+ D + SG+TA A+I + + +ANAGDSR V+ + G+A
Sbjct: 143 ATRS---AFVKTDHALADASSLDRSSGTTALTALILDKTMLIANAGDSRAVLGKRGRAIE 199
Query: 194 LSRDHKPELVIEKERIYKAGGFIHAGRINGSLNLARAIGDVDFKNNRFLSAEKQVVTANP 253
LS+DHKP E+ RI K GG I+ G +NG L++ARA+GD K + ++ P
Sbjct: 200 LSKDHKPNCTSERLRIEKLGGVIYDGYLNGQLSVARALGDWHIKGTK---GSLCPLSCEP 256
Query: 254 DINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEVCERVLDRCLAPSL 313
++ + L +EDE++++ CDG+WD +SSQ V VR+EL+ + ++ L
Sbjct: 257 ELEEIVLTEEDEYLIMGCDGLWDVMSSQCAVTMVRRELMQHNDPERCSQALVKEALQ--- 313
Query: 314 AVGDGCDNMTMILVQF 329
+ CDN+T+++V F
Sbjct: 314 --RNSCDNLTVVVVCF 327
>At2g25620 protein phosphatase 2C like protein
Length = 392
Score = 135 bits (341), Expect = 3e-32
Identities = 100/321 (31%), Positives = 152/321 (47%), Gaps = 71/321 (22%)
Query: 23 RYGLSSMQGWRATMEDAHAAHLDVDS-------------STSFFGVYDGHGGKAVAKFCA 69
R G S G R++MEDA+ L VD+ ++F+GV+DGHGGK A+F
Sbjct: 89 RSGAWSDIGSRSSMEDAY---LCVDNFMDSFGLLNSEAGPSAFYGVFDGHGGKHAAEFAC 145
Query: 70 KHLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKGFNGIIEGLI 129
H+ + +++ +E+ ++ L+ AFL+
Sbjct: 146 HHIPRYIVEDQEF-PSEINKVLSSAFLQT------------------------------- 173
Query: 130 RSPRSNDNKDQSDDWAFEKGPHSDFDGP-NSGSTACVAIIRNNLLFVANAGDSRCVISRN 188
D AF + DG SG+TA AI+ L VANAGD R V+SR
Sbjct: 174 -------------DTAFLEA--CSLDGSLASGTTALAAILFGRSLVVANAGDCRAVLSRQ 218
Query: 189 GQAYNLSRDHKPELVIEKERIYKAGGFIHAGRINGSLNLARAIGD--VDFKNNRFLSAEK 246
G+A +SRDHKP E+ RI +GG + G +NG LN+ARA+GD ++ + ++
Sbjct: 219 GKAIEMSRDHKPMSSKERRRIEASGGHVFDGYLNGQLNVARALGDFHMEGMKKKKDGSDC 278
Query: 247 QVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEVCERVLD 306
+ A P++ L +EDEF++I CDG+WD SQ VDF R+ L + +++
Sbjct: 279 GPLIAEPELMTTKLTEEDEFLIIGCDGVWDVFMSQNAVDFARRRLQEHNDPVMCSKELVE 338
Query: 307 RCLAPSLAVGDGCDNMTMILV 327
L A DN+T ++V
Sbjct: 339 EALKRKSA-----DNVTAVVV 354
>At4g27800 protein phosphatase homolog (PPH1)
Length = 388
Score = 134 bits (338), Expect = 6e-32
Identities = 106/346 (30%), Positives = 163/346 (46%), Gaps = 65/346 (18%)
Query: 22 LRYGLSSMQGWRATMEDAHAAHLDVDSSTSFFGVYDGHGGKAVAKFCAKHLHQQVLKSEE 81
+R+G +S+QG+R MED D S S+ V+DGH G + KF + L+++ + +
Sbjct: 58 IRWGYTSVQGFRDEMEDDIVIRSDAVDSFSYAAVFDGHAGSSSVKFLREELYKECVGA-- 115
Query: 82 YIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKGFNGIIEGLIRSPRSNDNKDQS 141
L+ ++ G GD F I E LI++ S D
Sbjct: 116 --------------LQAGSLLNG----------GD----FAAIKEALIKAFESVDRNLLK 147
Query: 142 DDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGDSRCVISRNGQAYNLSRDHKP- 200
W G D SGSTA V IIRN++ F+A+ GDS V+SR+GQ L+ H+P
Sbjct: 148 --WLEANGDEED----ESGSTATVMIIRNDVSFIAHIGDSCAVLSRSGQIEELTDYHRPY 201
Query: 201 ----ELVIEKERIYKAGGFIHAGRINGSLNLARAIGDVDFK----------------NNR 240
+ E +R+ +AGG+I GRI G + ++RA GD+ FK + +
Sbjct: 202 GSSRAAIQEVKRVKEAGGWIVNGRICGDIAVSRAFGDIRFKTKKNDMLKKGVDEGRWSEK 261
Query: 241 FLSA---EKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKL 297
F+S + +V A PDI V L + EFI++A DG+WD + S +V +VR +L +
Sbjct: 262 FVSRIEFKGDMVVATPDIFQVPLTSDVEFIILASDGLWDYMKSSDVVSYVRDQLRKHGNV 321
Query: 298 SEVCERVLDRCLAPSLAVGDGCDNMTMILVQFKKPLQTSAPAQEQS 343
CE LA DN+++I+ + + PAQ Q+
Sbjct: 322 QLACE-----SLAQVALDRRSQDNISIIIADLGRTEWKNLPAQRQN 362
>At1g43900 protein phosphatase type 2C like protein
Length = 371
Score = 130 bits (326), Expect = 1e-30
Identities = 106/312 (33%), Positives = 149/312 (46%), Gaps = 71/312 (22%)
Query: 24 YGLSSMQGWRATMEDAHAAHL-DVDSS-TSFFGVYDGHGGKAVAKFCAKHLHQQVLKSEE 81
YG SS++G RATMED + DV+ +FFGV+DGHGG A++ +L + ++ ++
Sbjct: 124 YGYSSLKGKRATMEDYFETRISDVNGQMVAFFGVFDGHGGARTAEYLKNNLFKNLVSHDD 183
Query: 82 YIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKGFNGIIEGLIRSPRSNDNKDQS 141
+I+ D ++ + F + DE E LI
Sbjct: 184 FIS-DTKKAIVEVFKQTDE-------------------------EYLIE----------- 206
Query: 142 DDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGDSRCVISRNGQAYNLSRDHKPE 201
E G N+GSTA A + + L VAN GDSR V SRNG A LS DHKP+
Sbjct: 207 -----EAGQPK-----NAGSTAATAFLIGDKLIVANVGDSRVVASRNGSAVPLSDDHKPD 256
Query: 202 LVIEKERIYKAGGF-IHAG--RINGSLNLARAIGDVDFKNNRFLSAEKQVVTANPDINIV 258
E++RI AGGF I AG R+ G L ++RA GD K V A P+I
Sbjct: 257 RSDERQRIEDAGGFIIWAGTWRVGGILAVSRAFGDKQL---------KPYVIAEPEIQEE 307
Query: 259 DLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEVCERVLDRCLAPSLAVGDG 318
D+ EFIV+A DG+W+ LS++ V VR ET ++ + R
Sbjct: 308 DI-STLEFIVVASDGLWNVLSNKDAVAIVRDISDAETAARKLVQEGYAR---------GS 357
Query: 319 CDNMTMILVQFK 330
CDN+T I+V+F+
Sbjct: 358 CDNITCIVVRFE 369
>At1g07430 protein phosphatase 2C, putative
Length = 442
Score = 127 bits (318), Expect = 1e-29
Identities = 97/283 (34%), Positives = 138/283 (48%), Gaps = 64/283 (22%)
Query: 23 RYGLSSMQGWRATMEDAHAAHLD-VDSSTSF-------FGVYDGHGGKAVAKFCAKHLHQ 74
RYG++S+ G R MEDA A H V T F FGVYDGHG VA C + LH+
Sbjct: 120 RYGVASVCGRRRDMEDAVALHPSFVRKQTEFSRTRWHYFGVYDGHGCSHVAARCKERLHE 179
Query: 75 QVL------KSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKGFNGIIEGL 128
V K EE+ + ++F RMD +E+ G+ V N E
Sbjct: 180 LVQEEALSDKKEEW-----KKMMERSFTRMD---------KEVVRWGETVMSANCRCE-- 223
Query: 129 IRSPRSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGDSRCVISRN 188
+++P D GSTA V++I + VAN GDSR V+ RN
Sbjct: 224 LQTP----------------------DCDAVGSTAVVSVITPEKIIVANCGDSRAVLCRN 261
Query: 189 GQAYNLSRDHKPELVIEKERIYKAGG---FIHAGRINGSLNLARAIGDVDFKNNRFLSAE 245
G+A LS DHKP+ E +RI +AGG + R+ G L ++RAIGD +
Sbjct: 262 GKAVPLSTDHKPDRPDELDRIQEAGGRVIYWDGARVLGVLAMSRAIGD---------NYL 312
Query: 246 KQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVR 288
K VT+ P++ + D +EDEF+++A DG+WD ++++ VR
Sbjct: 313 KPYVTSEPEVTVTDRTEEDEFLILATDGLWDVVTNEAACTMVR 355
>At5g53140 protein phosphatase 2C-like
Length = 420
Score = 124 bits (312), Expect = 6e-29
Identities = 100/348 (28%), Positives = 158/348 (44%), Gaps = 78/348 (22%)
Query: 18 ENDNLRYGLSSMQGWRATMEDAHAAHLDVDSST------SFFGVYDGHGGKAVAKFCAKH 71
++ +L G S +G R+TMED + D+ +ST FG++DGHGG A++ +H
Sbjct: 96 DDGSLSCGYCSFRGKRSTMEDFY----DIKASTIEGQAVCMFGIFDGHGGSRAAEYLKEH 151
Query: 72 LHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKGFNGIIEGLIRS 131
L ++K +++ D +L + + + D +A L
Sbjct: 152 LFNNLMKHPQFLT-DTKLALNETYKQTD-----------VAFLES--------------- 184
Query: 132 PRSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGDSRCVISRNGQA 191
EK + D GSTA A++ N L+VAN GDSR ++S+ G+A
Sbjct: 185 ---------------EKDTYRD-----DGSTASAAVLVGNHLYVANVGDSRTIVSKAGKA 224
Query: 192 YNLSRDHKPELVIEKERIYKAGGFI-HAG--RINGSLNLARAIGDVDFKNNRFLSAEKQV 248
LS DHKP E++RI AGG I AG R+ G L ++RA G NR L KQ
Sbjct: 225 IALSDDHKPNRSDERKRIESAGGVIMWAGTWRVGGVLAMSRAFG------NRML---KQF 275
Query: 249 VTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEVCERVLDRC 308
V A P+I +++ E E +V+A DG+WD + ++ V + E E ++ + R
Sbjct: 276 VVAEPEIQDLEIDHEAELLVLASDGLWDVVPNEDAVALAQSEEEPEAAARKLTDTAFSR- 334
Query: 309 LAPSLAVGDGCDNMTMILVQFKKPLQTSAPAQEQSSSNEQDASEPTLE 356
DN+T I+V+F+ S + + + + PT E
Sbjct: 335 --------GSADNITCIVVKFRHDKTESPKIETNAMAESEPELNPTTE 374
>At1g48040 protein phosphatase-2C, putative
Length = 377
Score = 123 bits (309), Expect = 1e-28
Identities = 100/319 (31%), Positives = 147/319 (45%), Gaps = 55/319 (17%)
Query: 33 RATMEDAH------AAHL---DVDSSTSFFGVYDGHGGKAVAKFCAKHLHQQVLKSEEYI 83
R TMED H +AHL + ++F+GV+DGHGG A F ++L + + +
Sbjct: 82 RETMEDEHICIDDLSAHLGSYNFSVPSAFYGVFDGHGGPEAAIFMKENLTRLFFQDAVF- 140
Query: 84 AGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKGFNGIIEGLIRSPRSNDNKDQSDD 143
S+ AF ++E+ + R+ L D I+ G
Sbjct: 141 --PEMPSIVDAFF-LEEL---ENSHRKAFALADLAMADETIVSG---------------- 178
Query: 144 WAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGDSRCVISRNGQAYNLSRDHKPELV 203
+ G+TA A+I L VANAGD R V+ R G A ++S DH+
Sbjct: 179 --------------SCGTTALTALIIGRHLLVANAGDCRAVLCRRGVAVDMSFDHRSTYE 224
Query: 204 IEKERIYKAGGFIHAGRINGSLNLARAIGDVDFKNNRFLSAEKQVVTANPDINIVDLHDE 263
E+ RI GG+ G +NG L + RAIGD + K N F + ++ ++P+I + L ++
Sbjct: 225 PERRRIEDLGGYFEDGYLNGVLAVTRAIGDWELK-NPFTDSSSPLI-SDPEIGQIILTED 282
Query: 264 DEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEVCERVLDRCLAPSLAVGDGCDNMT 323
DEF+++ACDGIWD LSSQ V VRQ L + L A DNMT
Sbjct: 283 DEFLILACDGIWDVLSSQNAVSNVRQGLRRHGDPRQCAME-----LGKEAARLQSSDNMT 337
Query: 324 MILVQFKKPLQTSAPAQEQ 342
+I++ F S+P Q Q
Sbjct: 338 VIVICFSS--VPSSPKQPQ 354
>At3g11410 protein phosphatase 2C (PP2C)
Length = 399
Score = 122 bits (307), Expect = 2e-28
Identities = 102/340 (30%), Positives = 162/340 (47%), Gaps = 53/340 (15%)
Query: 6 SIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDA---HAAHLDVDSSTS-FFGVYDGHGG 61
S+ + E F D + G +S+ G R MEDA H + L +S F+GV+DGHG
Sbjct: 91 SVTEAESFFSDVP----KIGTTSVCGRRRDMEDAVSIHPSFLQRNSENHHFYGVFDGHGC 146
Query: 62 KAVAKFCAKHLHQQVLKSEEYIAGDVGT-SLTKAFLRMDEMMRGQRGWRELAVLGDKVKG 120
VA+ C + LH V K E +A D T ++ K+F +MD+ + +
Sbjct: 147 SHVAEKCRERLHDIVKKEVEVMASDEWTETMVKSFQKMDKEVSQRE-------------- 192
Query: 121 FNGIIEGLIRSPRSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGD 180
N ++ G RS +++ + + P D GSTA V+++ + V+N GD
Sbjct: 193 CNLVVNGATRSMKNSCRCEL-------QSPQCDA----VGSTAVVSVVTPEKIIVSNCGD 241
Query: 181 SRCVISRNGQAYNLSRDHKPELVIEKERIYKAGG---FIHAGRINGSLNLARAIGDVDFK 237
SR V+ RNG A LS DHKP+ E RI +AGG + R+ G L ++RAIGD
Sbjct: 242 SRAVLCRNGVAIPLSVDHKPDRPDELIRIQQAGGRVIYWDGARVLGVLAMSRAIGD---- 297
Query: 238 NNRFLSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKL 297
+ K V +P++ + D DEDE +++A DG+WD + ++ R L
Sbjct: 298 -----NYLKPYVIPDPEVTVTDRTDEDECLILASDGLWDVVPNETACGVARM-CLRGAGA 351
Query: 298 SEVCERVLDRC-----LAPSLAVG-DGCDNMTMILVQFKK 331
+ + + C L LA+ DN+++++V +K
Sbjct: 352 GDDSDAAHNACSDAALLLTKLALARQSSDNVSVVVVDLRK 391
>At4g31750 unknown protein
Length = 311
Score = 122 bits (305), Expect = 4e-28
Identities = 95/318 (29%), Positives = 151/318 (46%), Gaps = 72/318 (22%)
Query: 18 ENDNLRYGLSSMQGWRATMEDAHAAHLD--VDSSTSFFGVYDGHGGKAVAKFCAKHLHQQ 75
+N YG +S G R++MED + +D FGV+DGHGG A++ ++L
Sbjct: 28 QNGKFSYGYASSPGKRSSMEDFYETRIDGVEGEIVGLFGVFDGHGGARAAEYVKQNLFSN 87
Query: 76 VLKSEEYIAGDVGTSLTKAFLRMD-EMMRGQRGWRELAVLGDKVKGFNGIIEGLIRSPRS 134
+++ ++I+ D ++ A+ + D E ++ + +
Sbjct: 88 LIRHPKFIS-DTTAAIADAYNQTDSEFLKSE----------------------------N 118
Query: 135 NDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGDSRCVISRNGQAYNL 194
+ N+D +GSTA AI+ + L VAN GDSR VI R G A +
Sbjct: 119 SQNRD-------------------AGSTASTAILVGDRLLVANVGDSRAVICRGGNAIAV 159
Query: 195 SRDHKPELVIEKERIYKAGGFI-HAG--RINGSLNLARAIGDVDFKNNRFLSAEKQVVTA 251
SRDHKP+ E++RI AGGF+ AG R+ G L ++RA GD R L KQ V A
Sbjct: 160 SRDHKPDQSDERQRIEDAGGFVMWAGTWRVGGVLAVSRAFGD------RLL---KQYVVA 210
Query: 252 NPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEVCERVLDRCLAP 311
+P+I + EF+++A DG+WD +S+++ V + K E E R +
Sbjct: 211 DPEIQEEKVDSSLEFLILASDGLWDVVSNEEAVGMI--------KAIEDPEEGAKRLMME 262
Query: 312 SLAVGDGCDNMTMILVQF 329
+ G DN+T ++V+F
Sbjct: 263 AYQRG-SADNITCVVVRF 279
>At5g59220 protein phosphatase 2C - like
Length = 413
Score = 121 bits (304), Expect = 5e-28
Identities = 109/353 (30%), Positives = 163/353 (45%), Gaps = 71/353 (20%)
Query: 3 TTLSIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHLDVD------SSTSFF--G 54
T +++P E + +YG++S+ G R MEDA A H SST F G
Sbjct: 99 TVVALPDPEAYP--------KYGVASVCGRRREMEDAVAVHPFFSRHQTEYSSTGFHYCG 150
Query: 55 VYDGHGGKAVAKFCAKHLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVL 114
VYDGHG VA C + LH+ V + E A D S+ ++F RMD E+ L
Sbjct: 151 VYDGHGCSHVAMKCRERLHELVREEFEADA-DWEKSMARSFTRMD---------MEVVAL 200
Query: 115 GDKVKGFNGIIEGLIRSPRSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLF 174
N D + E D D GSTA V+++ +
Sbjct: 201 ----------------------NADGAAKCRCEL-QRPDCDAV--GSTAVVSVLTPEKII 235
Query: 175 VANAGDSRCVISRNGQAYNLSRDHKPELVIEKERIYKAGG---FIHAGRINGSLNLARAI 231
VAN GDSR V+ RNG+A LS DHKP+ E +RI AGG + R+ G L ++RAI
Sbjct: 236 VANCGDSRAVLCRNGKAIALSSDHKPDRPDELDRIQAAGGRVIYWDGPRVLGVLAMSRAI 295
Query: 232 GDVDFKNNRFLSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQEL 291
GD + K V + P++ + D + D+F+++A DG+WD +S++ VR L
Sbjct: 296 GD---------NYLKPYVISRPEVTVTDRANGDDFLILASDGLWDVVSNETACSVVRMCL 346
Query: 292 --LLETKLSEVCERVLDRCLAPSLAVGDG------CDNMTMILVQFKKPLQTS 336
+ ++S ER + A ++ VG G C+ +++L + Q+S
Sbjct: 347 RGKVNGQVSSSPEREMTGVGAGNVVVGGGDLPDKACEEASLLLTRLALARQSS 399
>At3g62260 unknown protein
Length = 383
Score = 119 bits (299), Expect = 2e-27
Identities = 92/328 (28%), Positives = 148/328 (45%), Gaps = 75/328 (22%)
Query: 22 LRYGLSSMQGWRATMEDAHAAHLDVDSS----------TSFFGVYDGHGGKAVAKFCAKH 71
+R G + G + MED H D+ S ++F+ V+DGHGG A + ++
Sbjct: 76 IRSGSFADIGPKRNMEDEHIRIDDLSSQVGSLFELPKPSAFYAVFDGHGGPEAAAYVREN 135
Query: 72 LHQQVLKSEEY---------IAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKGFN 122
+ + E++ +V TSL AFL+ D +
Sbjct: 136 AIRFFFEDEQFPQTSEVSSVYVEEVETSLRNAFLQADLAL-------------------- 175
Query: 123 GIIEGLIRSPRSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGDSR 182
++D + + G+TA A+I LL VANAGD R
Sbjct: 176 ------------------AEDCSISD---------SCGTTALTALICGRLLMVANAGDCR 208
Query: 183 CVISRNGQAYNLSRDHKPELVIEKERIYKAGGFI-HAGRINGSLNLARAIGDVDFKNNRF 241
V+ R G+A ++S DHKP ++E+ R+ ++GGFI + G +N L + RA+GD D K
Sbjct: 209 AVLCRKGRAIDMSEDHKPINLLERRRVEESGGFITNDGYLNEVLAVTRALGDWDLK---L 265
Query: 242 LSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEVC 301
+ + + P+I + L ++DEF+VI CDGIWD L+SQ+ V VR+ L +
Sbjct: 266 PHGSQSPLISEPEIKQITLTEDDEFLVIGCDGIWDVLTSQEAVSIVRRGLNRHNDPTRCA 325
Query: 302 ERVLDRCLAPSLAVGDGCDNMTMILVQF 329
++ L + DN+T ++V F
Sbjct: 326 RELVMEALG-----RNSFDNLTAVVVCF 348
>At5g51760 protein phosphatase-2C; PP2C-like protein
Length = 416
Score = 119 bits (298), Expect = 3e-27
Identities = 111/360 (30%), Positives = 154/360 (41%), Gaps = 84/360 (23%)
Query: 15 EDGENDNLRYGLSSMQGWRATMEDAHAA-------HLDVDSSTSFFGVYDGHGGKAVAKF 67
E+ E++ L YG+ S+ G MED+ ++ FF VYDGHGG V+
Sbjct: 101 EETEDEPL-YGIVSVMGRSRKMEDSVTVKPNLCKPEVNRQRPVHFFAVYDGHGGSQVSTL 159
Query: 68 CAKHLH--------QQVLKSEEYIAGDVGTS-----LTKAFLRMDEMMRGQRGWRELAVL 114
C+ +H Q + + EE DV + ++F RMDEM V
Sbjct: 160 CSTTMHTFVKEELEQNLEEEEEGSENDVVERKWRGVMKRSFKRMDEMATST------CVC 213
Query: 115 GDKVKGFNGIIEGLIRSPRSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLF 174
G V N PR + SGSTA A++ ++ +
Sbjct: 214 GTSVPLCNC-------DPR---------------------EAAISGSTAVTAVLTHDHII 245
Query: 175 VANAGDSRCVISRNGQAYNLSRDHKPELVIEKERIYKAGG---FIHAGRINGSLNLARAI 231
VAN GDSR V+ RNG A LS DHKP+ E+ RI AGG + R+ G L +RAI
Sbjct: 246 VANTGDSRAVLCRNGMAIPLSNDHKPDRPDERARIEAAGGRVLVVDGARVEGILATSRAI 305
Query: 232 GDVDFKNNRFLSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQEL 291
GD R+L K +V P++ + DE +V+A DG+WD LSSQ D R L
Sbjct: 306 GD------RYL---KPMVAWEPEVTFMRRESGDECLVLASDGLWDVLSSQLACDIARFCL 356
Query: 292 LLETKLSEVCER----------------VLDRCLAPSLAVG-DGCDNMTMILVQFKKPLQ 334
ET S R VL L LA+G DN++++++ K Q
Sbjct: 357 REETPSSLDLNRMAQEDDNDGEQNPSRSVLAATLLTRLALGRQSSDNISVVVIDLKNSSQ 416
>At1g07160 unknown protein
Length = 380
Score = 115 bits (288), Expect = 4e-26
Identities = 99/331 (29%), Positives = 158/331 (46%), Gaps = 74/331 (22%)
Query: 8 PKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAA--HLDVDSSTSFFGVYDGHGGKAVA 65
P+ E + + E D Y + +G R MED +A +L D + FGVYDGHGG A
Sbjct: 109 PREESRAVEREGDG--YSVYCKRGKREAMEDRFSAITNLQGDPKQAIFGVYDGHGGPTAA 166
Query: 66 KFCAKHLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKGFNGII 125
+F AK+L +L E + G + + +A +RG+ LA + +K N +
Sbjct: 167 EFAAKNLCSNILG--EIVGGRNESKIEEAV---------KRGY--LATDSEFLKEKN--V 211
Query: 126 EGLIRSPRSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGDSRCVI 185
+G GS A+I + L VANAGD R V+
Sbjct: 212 KG--------------------------------GSCCVTALISDGNLVVANAGDCRAVL 239
Query: 186 SRNGQAYNLSRDHKPELVIEKERIYKAGGFI----HAGRINGSLNLARAIGDVDFKNNRF 241
S G A L+ DH+P E+ RI +GG++ RI GSL ++R IGD
Sbjct: 240 SVGGFAEALTSDHRPSRDDERNRIESSGGYVDTFNSVWRIQGSLAVSRGIGDAHL----- 294
Query: 242 LSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVR---QELLLETKLS 298
KQ + + P+INI+ ++ + EF+++A DG+WD +S+Q+ VD R + + K
Sbjct: 295 ----KQWIISEPEINILRINPQHEFLILASDGLWDKVSNQEAVDIARPFCKGTDQKRKPL 350
Query: 299 EVCERVLDRCLAPSLAVGDG-CDNMTMILVQ 328
C++++D L+V G D+++++L+Q
Sbjct: 351 LACKKLVD------LSVSRGSLDDISVMLIQ 375
>At1g67820 protein phosphatase-like protein
Length = 447
Score = 111 bits (277), Expect = 7e-25
Identities = 89/278 (32%), Positives = 127/278 (45%), Gaps = 66/278 (23%)
Query: 24 YGLSSMQGWRATMEDAH--AAHLDVDSSTSFFGVYDGHGGKAVAKFCAKHLHQQVLKSEE 81
+G+ S G + MED H L +S SFFGVYDGHGG A+F A++LH+ V+
Sbjct: 104 FGVVSRNGKKKFMEDTHRIVPCLVGNSKKSFFGVYDGHGGAKAAEFVAENLHKYVV---- 159
Query: 82 YIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKGFNGIIEGLIRSPRSNDNKDQS 141
EMM +G E KV+ F R K+QS
Sbjct: 160 ------------------EMMENCKGKEE------KVEAFKAAFLRTDRDFLEKVIKEQS 195
Query: 142 DDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGDSRCVISRNGQAYNLSRDHKPE 201
G SG+ A+I++ + V+N GD R V+ R G A L+ DHKP
Sbjct: 196 ------------LKGVVSGACCVTAVIQDQEMIVSNLGDCRAVLCRAGVAEALTDDHKPG 243
Query: 202 LVIEKERIYK-----------AGGFI--HAG--RINGSLNLARAIGDVDFKNNRFLSAEK 246
EKERI GG++ H G R+ G L ++R+IGD K
Sbjct: 244 RDDEKERIESQSLIPFMTFGLQGGYVDNHQGAWRVQGILAVSRSIGDAHL---------K 294
Query: 247 QVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLV 284
+ V A P+ +++L + EF+V+A DG+WD +S+Q+ V
Sbjct: 295 KWVVAEPETRVLELEQDMEFLVLASDGLWDVVSNQEAV 332
>At3g17250 protein phosphatase like
Length = 384
Score = 106 bits (265), Expect = 2e-23
Identities = 88/322 (27%), Positives = 140/322 (43%), Gaps = 58/322 (18%)
Query: 32 WRATMEDAHAAHLDVDSST-----------SFFGVYDGHGGKAVAKFCAKHLHQQVLKSE 80
+R MED H D+ +F+GV+DGHGG +++ ++ L E
Sbjct: 89 YREYMEDEHICIDDLSDHLGSSFYRFPVPMAFYGVFDGHGGSDASQYIKENAMS--LFFE 146
Query: 81 EYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKGFNGIIEGLIRSPRSNDNKDQ 140
+ + + + FL+ E RE L D I+
Sbjct: 147 DAVFRQSPSVVDSLFLKELETSH-----REAYRLADLAMEDERIVSS------------- 188
Query: 141 SDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGDSRCVISRNGQAYNLSRDHKP 200
+ G+TA A++ L VAN GD R V+ R G+A ++S DHK
Sbjct: 189 -----------------SCGTTALTALVIGRHLMVANVGDCRAVLCRKGKAVDMSFDHKS 231
Query: 201 ELVIEKERIYKAGGFIHAGRINGSLNLARAIGDVDFKNNRFLSAEKQVVTANPDINIVDL 260
E+ R+ GG+ + G L + RA+GD K L + ++PDI + L
Sbjct: 232 TFEPERRRVEDLGGYFEGEYLYGDLAVTRALGDWSIKRFSPLGESLSPLISDPDIQQMIL 291
Query: 261 HDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEVCERVLDRCLAPSLAVGDGCD 320
+EDEF+++ CDG+WD ++SQ V FVRQ L C L R +L + D D
Sbjct: 292 TEEDEFLIMGCDGVWDVMTSQYAVTFVRQGLRRHGD-PRRCAMELGR---EALRL-DSSD 346
Query: 321 NMTMILVQFKKPLQTSAPAQEQ 342
N+T++++ F +S+PA ++
Sbjct: 347 NVTVVVICF-----SSSPAPQR 363
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.134 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,356,327
Number of Sequences: 26719
Number of extensions: 364517
Number of successful extensions: 1163
Number of sequences better than 10.0: 87
Number of HSP's better than 10.0 without gapping: 75
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 896
Number of HSP's gapped (non-prelim): 171
length of query: 358
length of database: 11,318,596
effective HSP length: 100
effective length of query: 258
effective length of database: 8,646,696
effective search space: 2230847568
effective search space used: 2230847568
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC143339.7


