

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC143339.11 + phase: 0
(678 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
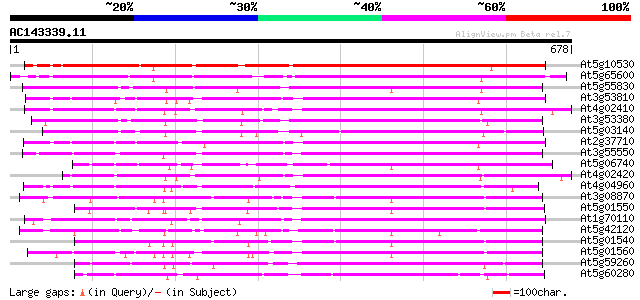
Score E
Sequences producing significant alignments: (bits) Value
At5g10530 lectin-like protein kinase - like 512 e-145
At5g65600 receptor protein kinase-like protein 459 e-129
At5g55830 serine/threonine-specific kinase like protein 346 2e-95
At3g53810 serine/threonine-specific kinase like protein 342 4e-94
At4g02410 unknown protein 335 5e-92
At3g53380 receptor lectin kinase -like protein 335 5e-92
At5g03140 receptor like protein kinase 334 1e-91
At2g37710 putative receptor-like protein kinase 332 3e-91
At3g55550 probable serine/threonine-specific protein kinase 328 8e-90
At5g06740 lectin-like protein kinase 325 5e-89
At4g02420 323 1e-88
At4g04960 unknown protein 305 7e-83
At3g08870 putative serine/threonine protein kinase 297 1e-80
At5g01550 receptor like protein kinase 291 8e-79
At1g70110 hypothetical protein 290 2e-78
At5g42120 receptor lectin kinase-like protein 289 4e-78
At5g01540 receptor like protein kinase 288 5e-78
At5g01560 receptor like protein kinase 288 7e-78
At5g59260 receptor-like protein kinase 284 1e-76
At5g60280 receptor like protein kinase 280 2e-75
>At5g10530 lectin-like protein kinase - like
Length = 651
Score = 512 bits (1318), Expect = e-145
Identities = 289/640 (45%), Positives = 401/640 (62%), Gaps = 32/640 (5%)
Query: 18 NILVLFSLLLVLSYLPKTNSLLFNITNFDDPTVASNMSYQGDGKSTNGSIDLNKVSYLFR 77
N ++LFS +LVL P S+ FNI+ F S ++YQGD ++ NG+++L + Y R
Sbjct: 3 NSILLFSFVLVL---PFVCSVQFNISRFGSDV--SEIAYQGDARA-NGAVELTNIDYTCR 56
Query: 78 VGRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDT--TYGDGFVFYLAPLGYQIPPNS 135
G A Y + + LW+ T+ + F+TRFSF ID N YG GF F+LAP Q+PPNS
Sbjct: 57 AGWATYGKQVPLWNPGTSKPSDFSTRFSFRIDTRNVGYGNYGHGFAFFLAPARIQLPPNS 116
Query: 136 AGGVYGLFNATTNSNFVMNYVVGVEFDTFVGPTDPPM---KHVGIDDNALTSVAFGKFDI 192
AGG GLFN T N + V VEFDTF P P+ HVGI++N+L S + ++
Sbjct: 117 AGGFLGLFNGTNNQSSAFPLVY-VEFDTFTNPEWDPLDVKSHVGINNNSLVSSNYTSWNA 175
Query: 193 DKNLGRVCYVLIDYNSDEKMLEVFWSFKGRFVKGDGSYGNSSISYQIDLMKKLPEFVNIG 252
+ + VLI Y+S + L V W++ NSS+SY IDL K LP V IG
Sbjct: 176 TSHNQDIGRVLIFYDSARRNLSVSWTYD----LTSDPLENSSLSYIIDLSKVLPSEVTIG 231
Query: 253 FSASTGLSTESNVIHSWEFSSNLEDSNSTTSLVEGNDGKGSLKTVIVVVAVIVPVILVFL 312
FSA++G TE N + SWEFSS+LE L++ + K +I+ ++V V+L F
Sbjct: 232 FSATSGGVTEGNRLLSWEFSSSLE-------LIDIKKSQNDKKGMIIGISVSGFVLLTFF 284
Query: 313 IASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFDLDKATIPRRFEYSELVAATNGFDDDRM 372
I S++ ++ K+++K +E + I DL++ PR+F Y +L +A N F DDR
Sbjct: 285 ITSLIVFLKRKQQKKKAEETENLTSI----NEDLERGAGPRKFTYKDLASAANNFADDRK 340
Query: 373 LGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIISRLIHRNLVQFIGWCH 432
LG GG+G VY+G L+ L +VA+K+ + +R F+ EV+IIS L HRNLVQ IGWCH
Sbjct: 341 LGEGGFGAVYRGYLNSLDMMVAIKKFAGGSKQGKREFVTEVKIISSLRHRNLVQLIGWCH 400
Query: 433 EQDELLLVFEYMPNGSLDTHLFGDKKSLAWEVRYKVALGVANALRYLHDDAEQCVLHRDI 492
E+DE L+++E+MPNGSLD HLFG K LAW VR K+ LG+A+AL YLH++ EQCV+HRDI
Sbjct: 401 EKDEFLMIYEFMPNGSLDAHLFGKKPHLAWHVRCKITLGLASALLYLHEEWEQCVVHRDI 460
Query: 493 KSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYGYLAPEYINGGRASKESDMYS 552
K++NV+LD++F+ KLGDFG+A+L+D L Q T + GT+GY+APEYI+ GRASKESD+YS
Sbjct: 461 KASNVMLDSNFNAKLGDFGLARLMDHELGPQTTGLAGTFGYMAPEYISTGRASKESDVYS 520
Query: 553 FGIVALELATGRRFFQDGDFHV-PLMNWV---WGLYVEGNLMCAADEKLNM-EFDVSEMK 607
FG+V LE+ TGR+ V P+ N V W LY +G ++ A DEKL + FD + +
Sbjct: 521 FGVVTLEIVTGRKSVDRRQGRVEPVTNLVEKMWDLYGKGEVITAIDEKLRIGGFDEKQAE 580
Query: 608 SLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPELPLDM 647
L+IVGLWC H + RP + I+VL LE +P LP M
Sbjct: 581 CLMIVGLWCAHPDVNTRPSIKQAIQVLNLEAPVPHLPTKM 620
>At5g65600 receptor protein kinase-like protein
Length = 675
Score = 459 bits (1180), Expect = e-129
Identities = 276/692 (39%), Positives = 405/692 (57%), Gaps = 61/692 (8%)
Query: 2 LNSPSSCSSNIINQSKNILVLFSLLLVLSYLPKTNSLLFNITNF--DDPTVASNMSYQGD 59
L+S SS S++I+ SL L L ++ +SL FN T+F DP ++ Y GD
Sbjct: 10 LSSSSSMSNSIL--------FLSLFLFLPFV--VDSLYFNFTSFRQGDP---GDIFYHGD 56
Query: 60 GK-STNGSIDLNKVSYLFRVGRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTYGD 118
+G+++ N +VG YS+ + +W KT + F+T FSF ID N + G
Sbjct: 57 ATPDEDGTVNFNNAEQTSQVGWITYSKKVPIWSHKTGKASDFSTSFSFKIDARNLSADGH 116
Query: 119 GFVFYLAPLGYQIPPNSAGGVYGLFNATTNSNFVMNY-VVGVEFDTFVGPTDPPM---KH 174
G F+LAP+G Q+P S GG LF T +N+ ++ +V VEFDTF P P H
Sbjct: 117 GICFFLAPMGAQLPAYSVGGFLNLF--TRKNNYSSSFPLVHVEFDTFNNPGWDPNDVGSH 174
Query: 175 VGIDDNALTSVAFGKFDIDKNLGRVCYVLIDYNSDEKMLEVFWSFKGRFVKGDGSYGNSS 234
VGI++N+L S + ++ + +C+ I Y+S K L V W+++ +SS
Sbjct: 175 VGINNNSLVSSNYTSWNASSHSQDICHAKISYDSVTKNLSVTWAYE--LTATSDPKESSS 232
Query: 235 ISYQIDLMKKLPEFVNIGFSASTGLSTESNVIHSWEFSSNLED--SNSTTSLVEGNDGKG 292
+SY IDL K LP V GF A+ G +TE + + SWE SS+L+ ++S LV G G
Sbjct: 233 LSYIIDLAKVLPSDVMFGFIAAAGTNTEEHRLLSWELSSSLDSDKADSRIGLVIGISASG 292
Query: 293 SLKTVIVVVAVIVPVILVFL-IASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFDLDKATI 351
V L F+ I ++V W +RK+K D E I K DL++
Sbjct: 293 F-------------VFLTFMVITTVVVWSRKQRKKKERDI---ENMISINK--DLEREAG 334
Query: 352 PRRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFIN 411
PR+F Y +LV+ATN F R LG GG+G VY+G L + +VAVK++ D + F+N
Sbjct: 335 PRKFSYKDLVSATNRFSSHRKLGEGGFGAVYEGNLKEINTMVAVKKLSGDSRQGKNEFLN 394
Query: 412 EVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKSL-AWEVRYKVAL 470
EV+IIS+L HRNLVQ IGWC+E++E LL++E +PNGSL++HLFG + +L +W++RYK+ L
Sbjct: 395 EVKIISKLRHRNLVQLIGWCNEKNEFLLIYELVPNGSLNSHLFGKRPNLLSWDIRYKIGL 454
Query: 471 GVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGT 530
G+A+AL YLH++ +QCVLHRDIK++N++LD++F+ KLGDFG+A+L++ L + T + GT
Sbjct: 455 GLASALLYLHEEWDQCVLHRDIKASNIMLDSEFNVKLGDFGLARLMNHELGSHTTGLAGT 514
Query: 531 YGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQ---------DGDFHVPLMNWVW 581
+GY+APEY+ G ASKESD+YSFGIV LE+ TGR+ + + D L+ VW
Sbjct: 515 FGYMAPEYVMKGSASKESDIYSFGIVLLEIVTGRKSLERTQEDNSDTESDDEKSLVEKVW 574
Query: 582 GLYVEGNLMCA-ADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMAL 640
LY + L+ + D+KL +FD E + LL++GLWC H + RP + I+V+ E L
Sbjct: 575 ELYGKQELITSCVDDKLGEDFDKKEAECLLVLGLWCAHPDKNSRPSIKQGIQVMNFESPL 634
Query: 641 PELPLDMHDRAPPIVPFRQSNGPSMAPPMTNS 672
P+LPL P+ + S S + P NS
Sbjct: 635 PDLPLKR-----PVAMYYISTTTSSSSPSVNS 661
>At5g55830 serine/threonine-specific kinase like protein
Length = 681
Score = 346 bits (888), Expect = 2e-95
Identities = 228/663 (34%), Positives = 349/663 (52%), Gaps = 51/663 (7%)
Query: 16 SKNILVLFSLLLVLSYLPKTNSLLFNITNFDDPTVA-SNMSYQGDGKSTNGSIDLNKVSY 74
S+ +LV+F + + K + + NF + N+++ GD NG + L +
Sbjct: 4 SRKLLVIFFTWITALSMSKPIFVSSDNMNFTFKSFTIRNLTFLGDSHLRNGVVGLTRELG 63
Query: 75 L--FRVGRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLND--TTYGDGFVFYLAPLGYQ 130
+ G Y+ P+ +D +NT SF+T FSFT+ LN T+ GDG F+L+
Sbjct: 64 VPDTSSGTVIYNNPIRFYDPDSNTTASFSTHFSFTVQNLNPDPTSAGDGLAFFLSHDNDT 123
Query: 131 IPPNSAGGVYGLFNATTNSNFVMNYVVGVEFDTFVGP--TDPPMKHVGIDDNALTSVAFG 188
+ S GG GL N+ S + N V +EFDT + P DP H+G+D ++L S++
Sbjct: 124 L--GSPGGYLGLVNS---SQPMKNRFVAIEFDTKLDPHFNDPNGNHIGLDVDSLNSISTS 178
Query: 189 ----KFDIDKNLGRVCYVLIDYNSDEKMLEVFWSFKGRFVKGDGSYGNSSISYQIDLMKK 244
ID G+ IDY +D ++L VF S+ V +S IDL
Sbjct: 179 DPLLSSQIDLKSGKSITSWIDYKNDLRLLNVFLSYTDP-VTTTKKPEKPLLSVNIDLSPF 237
Query: 245 LPEFVNIGFSASTGLSTESNVIHSWEFSS--------------NLEDSNSTTS--LVEGN 288
L + +GFS ST STE ++I +W F + N+ DS+ +V +
Sbjct: 238 LNGEMYVGFSGSTEGSTEIHLIENWSFKTSGFLPVRSKSNHLHNVSDSSVVNDDPVVIPS 297
Query: 289 DGKGSLKTVIVVVAVIVPVILVFLIASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFDLDK 348
+ + + + + PV L+ L + G+ +K+ + + K+ +
Sbjct: 298 KKRRHRHNLAIGLGISCPV-LICLALFVFGYFTLKKWKS----------VKAEKELKTEL 346
Query: 349 ATIPRRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERV 408
T R F Y EL AT GF R++GRG +G VY+ G I AVKR + +
Sbjct: 347 ITGLREFSYKELYTATKGFHSSRVIGRGAFGNVYRAMFVSSGTISAVKRSRHNSTEGKTE 406
Query: 409 FINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS----LAWEV 464
F+ E+ II+ L H+NLVQ GWC+E+ ELLLV+E+MPNGSLD L+ + ++ L W
Sbjct: 407 FLAELSIIACLRHKNLVQLQGWCNEKGELLLVYEFMPNGSLDKILYQESQTGAVALDWSH 466
Query: 465 RYKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQR 524
R +A+G+A+AL YLH + EQ V+HRDIK++N++LD +F+ +LGDFG+A+L +
Sbjct: 467 RLNIAIGLASALSYLHHECEQQVVHRDIKTSNIMLDINFNARLGDFGLARLTEHDKSPVS 526
Query: 525 TDVVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQ---DGDFHVPLMNWVW 581
T GT GYLAPEY+ G A++++D +S+G+V LE+A GRR + V L++WVW
Sbjct: 527 TLTAGTMGYLAPEYLQYGTATEKTDAFSYGVVILEVACGRRPIDKEPESQKTVNLVDWVW 586
Query: 582 GLYVEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALP 641
L+ EG ++ A DE+L EFD MK LL+VGL C H + ERP V+++L E+
Sbjct: 587 RLHSEGRVLEAVDERLKGEFDEEMMKKLLLVGLKCAHPDSNERPSMRRVLQILNNEIEPS 646
Query: 642 ELP 644
+P
Sbjct: 647 PVP 649
>At3g53810 serine/threonine-specific kinase like protein
Length = 677
Score = 342 bits (877), Expect = 4e-94
Identities = 240/656 (36%), Positives = 355/656 (53%), Gaps = 57/656 (8%)
Query: 20 LVLFSLLLVLSYLPKTNSLLFNITNFDDPTVASNMSYQGDGKST-NGSIDLNKVSYLFRV 78
L+ F LL + + +L F F P +++S QG T NG + L S + +
Sbjct: 7 LIFFFFLLCQIMISSSQNLNFTYNGFHPPL--TDISLQGLATVTPNGLLKLTNTS-VQKT 63
Query: 79 GRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTYGDGFVFYLAP---LGYQIPPNS 135
G AF ++ + D + ++SF+T F F I T G G F +AP L + +P
Sbjct: 64 GHAFCTERIRFKDSQNGNVSSFSTTFVFAIHSQIPTLSGHGIAFVVAPTLGLPFALPSQ- 122
Query: 136 AGGVYGLFNATTNSNFVMNYVVGVEFDTFVGPT--DPPMKHVGIDDNALTSVAFG----K 189
GLFN + N N N++ VEFDT DP HVGID N L S + +
Sbjct: 123 ---YIGLFNISNNGNDT-NHIFAVEFDTIQSSEFGDPNDNHVGIDLNGLRSANYSTAGYR 178
Query: 190 FDIDK--NLGRVC----YVLIDYNSDEKMLEV----FWSFKGRFVKGDGSYGNSSISYQI 239
D DK NL + V IDY++ ++V F S K R +SY
Sbjct: 179 DDHDKFQNLSLISRKRIQVWIDYDNRSHRIDVTVAPFDSDKPR---------KPLVSYVR 229
Query: 240 DLMKKLPEFVNIGFSASTGLSTESNVIHSWEFSSNLEDSNSTTSLVEGNDGKGSLKTVIV 299
DL L E + +GFS++TG + + W F N E + S + + + +
Sbjct: 230 DLSSILLEDMYVGFSSATGSVLSEHFLVGWSFRLNGEAPMLSLSKLPKLP-RFEPRRISE 288
Query: 300 VVAVIVPVILVFLIASIV--GWVIVKRKRKNCDEGLDEYGIPFPKKFDLDKATIPRRFEY 357
+ +P+I + LI SI+ + IV+RK+K +E LD++ F K RF +
Sbjct: 289 FYKIGMPLISLSLIFSIIFLAFYIVRRKKKY-EEELDDWETEFGKN----------RFRF 337
Query: 358 SELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIIS 417
EL AT GF + +LG GG+G+VY+G L VAVKR+ D + + F+ E+ I
Sbjct: 338 KELYHATKGFKEKDLLGSGGFGRVYRGILPTTKLEVAVKRVSHDSKQGMKEFVAEIVSIG 397
Query: 418 RLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS-LAWEVRYKVALGVANAL 476
R+ HRNLV +G+C + ELLLV++YMPNGSLD +L+ + ++ L W+ R + GVA+ L
Sbjct: 398 RMSHRNLVPLLGYCRRRGELLLVYDYMPNGSLDKYLYNNPETTLDWKQRSTIIKGVASGL 457
Query: 477 RYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYGYLAP 536
YLH++ EQ V+HRD+K++NVLLD DF+ +LGDFG+A+L D Q T VVGT GYLAP
Sbjct: 458 FYLHEEWEQVVIHRDVKASNVLLDADFNGRLGDFGLARLYDHGSDPQTTHVVGTLGYLAP 517
Query: 537 EYINGGRASKESDMYSFGIVALELATGRR---FFQDGDFHVPLMNWVWGLYVEGNLMCAA 593
E+ GRA+ +D+Y+FG LE+ +GRR F D L+ WV+ L++ GN+M A
Sbjct: 518 EHSRTGRATTTTDVYAFGAFLLEVVSGRRPIEFHSASDDTFLLVEWVFSLWLRGNIMEAK 577
Query: 594 DEKLNME-FDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPEL-PLDM 647
D KL +D+ E++ +L +GL C+HS+ + RP +V++ L+ +MALPEL PLD+
Sbjct: 578 DPKLGSSGYDLEEVEMVLKLGLLCSHSDPRARPSMRQVLQYLRGDMALPELTPLDL 633
>At4g02410 unknown protein
Length = 674
Score = 335 bits (859), Expect = 5e-92
Identities = 227/693 (32%), Positives = 364/693 (51%), Gaps = 59/693 (8%)
Query: 19 ILVLFSLLLVLSYLPKTNSLLFNITNFDDPTVASNMSYQGDGKSTNGSIDLNKVSYLFRV 78
I F +LL + SL F +F P +N+S QG T+ I +
Sbjct: 8 IFFFFIILLSKPLNSSSQSLNFTYNSFHRPP--TNISIQGIATVTSNGILKLTDKTVIST 65
Query: 79 GRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTYGDGFVFYLAPLGYQIPPNSAGG 138
G AFY++P+ D +T++SF+T F I T G G F++AP + A
Sbjct: 66 GHAFYTEPIRFKDSPNDTVSSFSTTFVIGIYSGIPTISGHGMAFFIAP-NPVLSSAMASQ 124
Query: 139 VYGLFNATTNSNFVMNYVVGVEFDTFVGP--TDPPMKHVGIDDNALTSV---AFGKFDID 193
GLF++T N N N+++ VEFDT + P D HVGI+ N+LTSV G +D
Sbjct: 125 YLGLFSSTNNGNDT-NHILAVEFDTIMNPEFDDTNDNHVGININSLTSVKSSLVGYWDEI 183
Query: 194 KNLGRV-------CYVLIDYNSDEKMLEVFWSFKGRFVKGDGSYGNSSISYQIDLMKKLP 246
+ V +DY+ ++V + G+ + +S DL
Sbjct: 184 NQFNNLTLISRKRMQVWVDYDDRTNQIDVTMA-----PFGEVKPRKALVSVVRDLSSVFL 238
Query: 247 EFVNIGFSASTGLSTESNVIHSWEFSSNLEDSN-------------STTSLVEGNDGKGS 293
+ + +GFSA+TG + + W F + + TSL +
Sbjct: 239 QDMYLGFSAATGYVLSEHFVFGWSFMVKGKTAPPLTLSKVPKFPRVGPTSLQRFYKNRMP 298
Query: 294 LKTVIVVVAVIVPVILVFLIASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFDLDKATIPR 353
L +++++ + V V L+FL+ IV R+R+ E +++ F K
Sbjct: 299 LFSLLLIPVLFV-VSLIFLVRFIV------RRRRKFAEEFEDWETEFGK----------N 341
Query: 354 RFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEV 413
R + +L AT GF D +LG GG+G+VY+G + K +AVKR+ + + F+ E+
Sbjct: 342 RLRFKDLYYATKGFKDKDLLGSGGFGRVYRGVMPTTKKEIAVKRVSNESRQGLKEFVAEI 401
Query: 414 RIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFG-DKKSLAWEVRYKVALGV 472
I R+ HRNLV +G+C +DELLLV++YMPNGSLD +L+ + +L W+ R+ V +GV
Sbjct: 402 VSIGRMSHRNLVPLLGYCRRRDELLLVYDYMPNGSLDKYLYDCPEVTLDWKQRFNVIIGV 461
Query: 473 ANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYG 532
A+ L YLH++ EQ V+HRDIK++NVLLD +++ +LGDFG+A+L D Q T VVGT+G
Sbjct: 462 ASGLFYLHEEWEQVVIHRDIKASNVLLDAEYNGRLGDFGLARLCDHGSDPQTTRVVGTWG 521
Query: 533 YLAPEYINGGRASKESDMYSFGIVALELATGRRFFQ---DGDFHVPLMNWVWGLYVEGNL 589
YLAP+++ GRA+ +D+++FG++ LE+A GRR + + D V L++ V+G ++EGN+
Sbjct: 522 YLAPDHVRTGRATTATDVFAFGVLLLEVACGRRPIEIEIESDESVLLVDSVFGFWIEGNI 581
Query: 590 MCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPEL-PLDMH 648
+ A D L +D E++++L +GL C+HS+ + RP +V++ L+ + LP+L PLD
Sbjct: 582 LDATDPNLGSVYDQREVETVLKLGLLCSHSDPQVRPTMRQVLQYLRGDATLPDLSPLDFR 641
Query: 649 DRAPPI---VPFRQSNGPSMAPPMTNSLITSGR 678
+ F +S S + S+++ GR
Sbjct: 642 GSGKMLGMNHRFSESCTFSSGSSIAYSIVSGGR 674
>At3g53380 receptor lectin kinase -like protein
Length = 715
Score = 335 bits (859), Expect = 5e-92
Identities = 232/684 (33%), Positives = 339/684 (48%), Gaps = 91/684 (13%)
Query: 27 LVLSYLPKTNSLLFN---ITNFDDPTVA-SNMSYQGDGKSTNGSIDLNKVSYLFR--VGR 80
L LS+ FN T FD T+A SN+ GD + +NG + L + + G+
Sbjct: 3 LFLSFFISILLCFFNGATTTQFDFSTLAISNLKLLGDARLSNGIVGLTRDLSVPNSGAGK 62
Query: 81 AFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTYGDGFVFYLAPLGYQIPPNSAGGVY 140
YS P+ T+ TSF++ FSF+I +N ++ G G F ++P I AGG
Sbjct: 63 VLYSNPIRFRQPGTHFPTSFSSFFSFSITNVNPSSIGGGLAFVISPDANSI--GIAGGSL 120
Query: 141 GLFNATTNSNFVMNYVVGVEFDTF--VGPTDPPMKHVGIDDNALTSVA---FGKFDIDKN 195
GL T N + V VEFDT V D HVG D N + S G +ID
Sbjct: 121 GL----TGPNGSGSKFVAVEFDTLMDVDFKDINSNHVGFDVNGVVSSVSGDLGTVNIDLK 176
Query: 196 LGRVCYVLIDYNSDEKMLEVFWSFKGRFVKGDGSYGNSSISYQIDLMKKLPEFVNIGFSA 255
G I+Y+ ++ V S+ K +S+ +DL + + +F+ +GFS
Sbjct: 177 SGNTINSWIEYDGLTRVFNVSVSYSNLKPKVP------ILSFPLDLDRYVNDFMFVGFSG 230
Query: 256 STGLSTESNVIHSWEFSSNLEDS------------------------------------- 278
ST STE + I W FSS+ S
Sbjct: 231 STQGSTEIHSIEWWSFSSSFGSSLGSGSGSPPPRANLMNPKANSVKSPPPLASQPSSSAI 290
Query: 279 ---------NSTTSLVEGNDGKGSLKTVIVVVAVIVPVILVFLIASIVGWVIVKRKRKNC 329
S++S K + T+ VV + L A + WV K+ ++
Sbjct: 291 PISSNTQLKTSSSSSCHSRFCKENPGTIAGVVTA--GAFFLALFAGALFWVYSKKFKR-- 346
Query: 330 DEGLDEYGIPFPKKFDLDKATIPRRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYL 389
+ F + P+ F Y EL A T F++ R++G G +G VY+G L
Sbjct: 347 --------VERSDSFASEIIKAPKEFSYKELKAGTKNFNESRIIGHGAFGVVYRGILPET 398
Query: 390 GKIVAVKRIFADFENSERVFINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSL 449
G IVAVKR ++ + F++E+ II L HRNLV+ GWCHE+ E+LLV++ MPNGSL
Sbjct: 399 GDIVAVKRCSHSSQDKKNEFLSELSIIGSLRHRNLVRLQGWCHEKGEILLVYDLMPNGSL 458
Query: 450 DTHLFGDKKSLAWEVRYKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGD 509
D LF + +L W+ R K+ LGVA+AL YLH + E V+HRD+KS+N++LD F+ KLGD
Sbjct: 459 DKALFESRFTLPWDHRKKILLGVASALAYLHRECENQVIHRDVKSSNIMLDESFNAKLGD 518
Query: 510 FGMAKLVDPMLRTQRTDVVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQD 569
FG+A+ ++ + T GT GYLAPEY+ GRAS+++D++S+G V LE+ +GRR +
Sbjct: 519 FGLARQIEHDKSPEATVAAGTMGYLAPEYLLTGRASEKTDVFSYGAVVLEVVSGRRPIEK 578
Query: 570 GDFHVP---------LMNWVWGLYVEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSN 620
D +V L+ WVWGLY EG + AAD +L +FD EM +L+VGL C+H +
Sbjct: 579 -DLNVQRHNVGVNPNLVEWVWGLYKEGKVSAAADSRLEGKFDEGEMWRVLVVGLACSHPD 637
Query: 621 DKERPKAYEVIKVLQLEMALPELP 644
RP V+++L E +P +P
Sbjct: 638 PAFRPTMRSVVQMLIGEADVPVVP 661
>At5g03140 receptor like protein kinase
Length = 711
Score = 334 bits (856), Expect = 1e-91
Identities = 224/655 (34%), Positives = 336/655 (51%), Gaps = 77/655 (11%)
Query: 40 FNITNFDDPTVA-SNMSYQGDGKSTNGSIDLNKVSYL--FRVGRAFYSQPLHLWDKKTNT 96
F T FD T+ S++ GD NG+I L + + G+A Y +P+ +T +
Sbjct: 32 FPATRFDLGTLTLSSLKLLGDAHLNNGTIKLTRELSVPTSTAGKALYGKPVKFRHPETKS 91
Query: 97 LTSFTTRFSFTIDKLNDTTYGDGFVFYLAPLGYQIPPNSAGGVYGLFNAT-TNSNFVMNY 155
SFTT FSF++ LN ++ G G F ++P + S GG GL T + S FV
Sbjct: 92 PASFTTYFSFSVTNLNPSSIGGGLAFVISPDEDYL--GSTGGFLGLTEETGSGSGFV--- 146
Query: 156 VVGVEFDTFVGPT--DPPMKHVGIDDNALTSVA---FGKFDIDKNLGRVCYVLIDYNSDE 210
VEFDT + D HVG+D NA+ S A G DID G I Y+
Sbjct: 147 --AVEFDTLMDVQFKDVNGNHVGLDLNAVVSAAVADLGNVDIDLKSGNAVNSWITYDGSG 204
Query: 211 KMLEVFWSFKGRFVKGDGSYGNSSISYQIDLMKKLPEFVNIGFSASTGLSTESNVIHSWE 270
++L V+ S+ K + +S +DL + + + + +GFS ST STE + + W
Sbjct: 205 RVLTVYVSYSNLKPK------SPILSVPLDLDRYVSDSMFVGFSGSTQGSTEIHSVDWWS 258
Query: 271 FSSNLEDS--------NSTTSLVEGNDGKGSLKTV---------------------IVVV 301
FSS+ E+S NS + SL TV V
Sbjct: 259 FSSSFEESSESPPPMPNSPPPSSPSSSITPSLSTVRRKTADPSSSCRNKLCKKSPAAVAG 318
Query: 302 AVIVPVILVFLIASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFDLDKATI---PRRFEYS 358
V + L A ++ WV K+ I + +K + + I PR F Y
Sbjct: 319 VVTAGAFFLALFAGVIIWVYSKK-------------IKYTRKSESLASEIMKSPREFTYK 365
Query: 359 ELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIISR 418
EL AT+ F R++G G +G VYKG L G+I+A+KR + F++E+ +I
Sbjct: 366 ELKLATDCFSSSRVIGNGAFGTVYKGILQDSGEIIAIKRC-SHISQGNTEFLSELSLIGT 424
Query: 419 LIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKSLAWEVRYKVALGVANALRY 478
L HRNL++ G+C E+ E+LL+++ MPNGSLD L+ +L W R K+ LGVA+AL Y
Sbjct: 425 LRHRNLLRLQGYCREKGEILLIYDLMPNGSLDKALYESPTTLPWPHRRKILLGVASALAY 484
Query: 479 LHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYGYLAPEY 538
LH + E ++HRD+K++N++LD +F+ KLGDFG+A+ + T GT GYLAPEY
Sbjct: 485 LHQECENQIIHRDVKTSNIMLDANFNPKLGDFGLARQTEHDKSPDATAAAGTMGYLAPEY 544
Query: 539 INGGRASKESDMYSFGIVALELATGRRFFQDGD--------FHVPLMNWVWGLYVEGNLM 590
+ GRA++++D++S+G V LE+ TGRR + L++WVWGLY EG L+
Sbjct: 545 LLTGRATEKTDVFSYGAVVLEVCTGRRPITRPEPEPGLRPGLRSSLVDWVWGLYREGKLL 604
Query: 591 CAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPELPL 645
A DE+L+ EF+ EM +++VGL C+ + RP V+++L E +PE+P+
Sbjct: 605 TAVDERLS-EFNPEEMSRVMMVGLACSQPDPVTRPTMRSVVQILVGEADVPEVPI 658
>At2g37710 putative receptor-like protein kinase
Length = 675
Score = 332 bits (852), Expect = 3e-91
Identities = 229/650 (35%), Positives = 347/650 (53%), Gaps = 40/650 (6%)
Query: 17 KNILVLFSLLLVLSYLPKTNSLLFNITN-FDDPTVASNMSYQGDGKST-NGSIDLNKVSY 74
K + + F L + + SL F N F+ PT ++S QG T NG + L +
Sbjct: 4 KLLTIFFFFFFNLIFQSSSQSLNFAYNNGFNPPT---DLSIQGITTVTPNGLLKLTNTT- 59
Query: 75 LFRVGRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTYGDGFVFYLAPLGYQIPPN 134
+ + G AFY++P+ D T++SF+T F F I G G F +AP +P
Sbjct: 60 VQKTGHAFYTKPIRFKDSPNGTVSSFSTSFVFAIHSQIAILSGHGIAFVVAP-NASLPYG 118
Query: 135 SAGGVYGLFNATTNSNFVMNYVVGVEFDTFVGP--TDPPMKHVGIDDNALTSVAFGKFDI 192
+ GLFN N N N+V VE DT + D HVGID N+L SV
Sbjct: 119 NPSQYIGLFNLANNGNET-NHVFAVELDTILSTEFNDTNDNHVGIDINSLKSVQSSPAGY 177
Query: 193 DKNLGRVCYVLIDYNSDEKMLEVFWSFKGRFVKGDGSYGNSS--------ISYQIDLMKK 244
G+ + + K ++V+ + GR K D + + ++ DL
Sbjct: 178 WDEKGQFKNLTL---ISRKPMQVWVDYDGRTNKIDVTMAPFNEDKPTRPLVTAVRDLSSV 234
Query: 245 LPEFVNIGFSASTGLSTESNVIHSWEFSSNLEDSNSTTSLVEGNDGKGSLKTVIVVVAVI 304
L + + +GFS++TG + I W F N + S + + K + +
Sbjct: 235 LLQDMYVGFSSATGSVLSEHYILGWSFGLNEKAPPLALSRLPKLP-RFEPKRISEFYKIG 293
Query: 305 VPVILVFLIASIVGWV--IVKRKRKNCDEGLDEYGIPFPKKFDLDKATIPRRFEYSELVA 362
+P+I +FLI S + V IV+R+RK +E L+E+ F K RF + +L
Sbjct: 294 MPLISLFLIFSFIFLVCYIVRRRRKFAEE-LEEWEKEFGKN----------RFRFKDLYY 342
Query: 363 ATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIISRLIHR 422
AT GF + +LG GG+G VYKG + +AVKR+ + + F+ E+ I R+ HR
Sbjct: 343 ATKGFKEKGLLGTGGFGSVYKGVMPGTKLEIAVKRVSHESRQGMKEFVAEIVSIGRMSHR 402
Query: 423 NLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKK-SLAWEVRYKVALGVANALRYLHD 481
NLV +G+C + ELLLV++YMPNGSLD +L+ + +L W+ R KV LGVA+ L YLH+
Sbjct: 403 NLVPLLGYCRRRGELLLVYDYMPNGSLDKYLYNTPEVTLNWKQRIKVILGVASGLFYLHE 462
Query: 482 DAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYGYLAPEYING 541
+ EQ V+HRD+K++NVLLD + + +LGDFG+A+L D Q T VVGT GYLAPE+
Sbjct: 463 EWEQVVIHRDVKASNVLLDGELNGRLGDFGLARLYDHGSDPQTTHVVGTLGYLAPEHTRT 522
Query: 542 GRASKESDMYSFGIVALELATGRR---FFQDGDFHVPLMNWVWGLYVEGNLMCAADEKLN 598
GRA+ +D+++FG LE+A GRR F Q+ D L++WV+GL+ +G+++ A D +
Sbjct: 523 GRATMATDVFAFGAFLLEVACGRRPIEFQQETDETFLLVDWVFGLWNKGDILAAKDPNMG 582
Query: 599 MEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPEL-PLDM 647
E D E++ +L +GL C+HS+ + RP +V+ L+ + LPEL PLD+
Sbjct: 583 SECDEKEVEMVLKLGLLCSHSDPRARPSMRQVLHYLRGDAKLPELSPLDL 632
>At3g55550 probable serine/threonine-specific protein kinase
Length = 684
Score = 328 bits (840), Expect = 8e-90
Identities = 226/650 (34%), Positives = 345/650 (52%), Gaps = 47/650 (7%)
Query: 16 SKNILVLFSLLLVLSYLPKTNSLLFNITNFDDPTVASNMSYQGDGK-STNGSIDLNKVSY 74
S+ V+ LL+ L++L +SL+ + + + N++ G + + G+I L +
Sbjct: 2 SQTFAVILLLLIFLTHL--VSSLIQDFSFIGFKKASPNLTLNGVAEIAPTGAIRLTTETQ 59
Query: 75 LFRVGRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTYGDGFVFYLAPLGYQIPPN 134
+G AFYS P+ N SF+T F+ + T G G F + P P+
Sbjct: 60 RV-IGHAFYSLPIRFKPIGVNRALSFSTSFAIAMVPEFVTLGGHGLAFAITPT-----PD 113
Query: 135 SAGGVYGLFNATTNSNFV--MNYVVGVEFDTF--VGPTDPPMKHVGIDDNALTS------ 184
G + + NS+ V ++ VEFDT + D HVGID N++ S
Sbjct: 114 LRGSLPSQYLGLLNSSRVNFSSHFFAVEFDTVRDLEFEDINDNHVGIDINSMESSISTPA 173
Query: 185 ----VAFGKFDIDKNLGRVCYVLIDYNSDEKMLEVFWSFKGRFVKGDGSYGNSSISYQID 240
K ++ + GRV IDY+S++K L+V S K S +SY +D
Sbjct: 174 GYFLANSTKKELFLDGGRVIQAWIDYDSNKKRLDVKLSPFSEKPK------LSLLSYDVD 227
Query: 241 LMKKLPEFVNIGFSASTGLSTESNVIHSWEFSSNLED-SNSTTSL--VEGNDGKGSLKTV 297
L L + + +GFSASTGL S+ I W F+ + E S S SL + + K K
Sbjct: 228 LSSVLGDEMYVGFSASTGLLASSHYILGWNFNMSGEAFSLSLPSLPRIPSSIKKRKKKRQ 287
Query: 298 IVVVAVIVPVILVFLIASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFDLDKATIPRRFEY 357
+++ V + L+ + + V RK K+ D ++E+ + F P RF Y
Sbjct: 288 SLILGVSLLCSLLIFAVLVAASLFVVRKVKDEDR-VEEWELDFG----------PHRFSY 336
Query: 358 SELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIIS 417
EL ATNGF D +LG GG+G+VYKG L + VAVKRI + R F++EV I
Sbjct: 337 RELKKATNGFGDKELLGSGGFGKVYKGKLPGSDEFVAVKRISHESRQGVREFMSEVSSIG 396
Query: 418 RLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS--LAWEVRYKVALGVANA 475
L HRNLVQ +GWC +D+LLLV+++MPNGSLD +LF + L W+ R+K+ GVA+
Sbjct: 397 HLRHRNLVQLLGWCRRRDDLLLVYDFMPNGSLDMYLFDENPEVILTWKQRFKIIKGVASG 456
Query: 476 LRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYGYLA 535
L YLH+ EQ V+HRDIK+ANVLLD++ + ++GDFG+AKL + T VVGT+GYLA
Sbjct: 457 LLYLHEGWEQTVIHRDIKAANVLLDSEMNGRVGDFGLAKLYEHGSDPGATRVVGTFGYLA 516
Query: 536 PEYINGGRASKESDMYSFGIVALELATGRRFFQDGDF--HVPLMNWVWGLYVEGNLMCAA 593
PE G+ + +D+Y+FG V LE+A GRR + + +++WVW + G++
Sbjct: 517 PELTKSGKLTTSTDVYAFGAVLLEVACGRRPIETSALPEELVMVDWVWSRWQSGDIRDVV 576
Query: 594 DEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPEL 643
D +LN EFD E+ ++ +GL C++++ + RP +V+ L+ + PE+
Sbjct: 577 DRRLNGEFDEEEVVMVIKLGLLCSNNSPEVRPTMRQVVMYLEKQFPSPEV 626
>At5g06740 lectin-like protein kinase
Length = 652
Score = 325 bits (833), Expect = 5e-89
Identities = 214/603 (35%), Positives = 319/603 (52%), Gaps = 66/603 (10%)
Query: 77 RVGRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTYGDGFVFYLAPLGYQIPPNSA 136
+ GRA Y +P LW K + +F T F I D G+G F L P P NS+
Sbjct: 69 QAGRALYKKPFRLWSKHKSA--TFNTTFVINISNKTDPG-GEGLAFVLTP-EETAPQNSS 124
Query: 137 GGVYGLFNATTNSNFVMNYVVGVEFDTFVGPTDP-PMKHVGIDDNALTSVAFGKFD---- 191
G G+ N TN N +V VEFDT +D HV ++ N + SV
Sbjct: 125 GMWLGMVNERTNRNNESR-IVSVEFDTRKSHSDDLDGNHVALNVNNINSVVQESLSGRGI 183
Query: 192 -IDKNLGRVCYVLIDYNSDEKMLEVFWS-----FKGRFVKGDGSYGNSSISYQIDLMKKL 245
ID L +V D K L V+ S F+ R N S IDL L
Sbjct: 184 KIDSGLDLTAHV----RYDGKNLSVYVSRNLDVFEQR---------NLVFSRAIDLSAYL 230
Query: 246 PEFVNIGFSASTGLSTESNVIHSWEFSSNLEDSNSTTSLVEGNDGKGSLKTVIVVVAVIV 305
PE V +GF+AST TE N + SW F D +GN ++ + + +
Sbjct: 231 PETVYVGFTASTSNFTELNCVRSWSFEGLKIDG-------DGN---------MLWLWITI 274
Query: 306 PVILVFLIASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFDLDK-ATIPRRFEYSELVAAT 364
P++ + I + +G + ++ + K + D + +LD A P++F+ EL AT
Sbjct: 275 PIVFIVGIGAFLGALYLRSRSKAGETNPDI-------EAELDNCAANPQKFKLRELKRAT 327
Query: 365 NGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIISRLIHRNL 424
F + LG+GG+G V+KG + G+ +AVKR+ ++ FI E+ I L HRNL
Sbjct: 328 GNFGAENKLGQGGFGMVFKG--KWQGRDIAVKRVSEKSHQGKQEFIAEITTIGNLNHRNL 385
Query: 425 VQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS---LAWEVRYKVALGVANALRYLHD 481
V+ +GWC+E+ E LLV+EYMPNGSLD +LF + KS L WE R + G++ AL YLH+
Sbjct: 386 VKLLGWCYERKEYLLVYEYMPNGSLDKYLFLEDKSRSNLTWETRKNIITGLSQALEYLHN 445
Query: 482 DAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRT--DVVGTYGYLAPEYI 539
E+ +LHRDIK++NV+LD+DF+ KLGDFG+A+++ T + ++ GT GY+APE
Sbjct: 446 GCEKRILHRDIKASNVMLDSDFNAKLGDFGLARMIQQSEMTHHSTKEIAGTPGYMAPETF 505
Query: 540 NGGRASKESDMYSFGIVALELATGRR----FFQD--GDFHVPLMNWVWGLYVEGNLMCAA 593
GRA+ E+D+Y+FG++ LE+ +G++ +D +++ ++NW+W LY G + AA
Sbjct: 506 LNGRATVETDVYAFGVLMLEVVSGKKPSYVLVKDNQNNYNNSIVNWLWELYRNGTITDAA 565
Query: 594 DEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPELPLDMHDRAPP 653
D + FD EMKS+L++GL C H N +RP V+KVL E + P++P + P
Sbjct: 566 DPGMGNLFDKEEMKSVLLLGLACCHPNPNQRPSMKTVLKVLTGETSPPDVPTERPAFVWP 625
Query: 654 IVP 656
+P
Sbjct: 626 AMP 628
>At4g02420
Length = 669
Score = 323 bits (829), Expect = 1e-88
Identities = 223/642 (34%), Positives = 343/642 (52%), Gaps = 50/642 (7%)
Query: 64 NGSIDLNKVSYLFRVGRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTYGDGFVFY 123
NG + L + + G AFY++P+ D T++SF+T F F I +G FV
Sbjct: 51 NGLLKLTNTT-MQSTGHAFYTKPIRFKDSPNGTVSSFSTTFVFAIHSQIPIAHGMAFVIA 109
Query: 124 LAPLGYQIPPNSAGGVYGLFNATTNSNFVMNYVVGVEFDTFVGP--TDPPMKHVGIDDNA 181
P ++P S GLFN T N N V N+V VE DT + D HVGID N+
Sbjct: 110 PNP---RLPFGSPLQYLGLFNVTNNGN-VRNHVFAVELDTIMNIEFNDTNNNHVGIDINS 165
Query: 182 LTSVA------FGKFDIDKNLGRVC----YVLIDYNSDEKMLEVFWSFKGRFVKGDGSYG 231
L SV + + D NL + V +D++ +++V + G+
Sbjct: 166 LNSVKSSPAGYWDENDQFHNLTLISSKRMQVWVDFDGPTHLIDVTMA-----PFGEVKPR 220
Query: 232 NSSISYQIDLMKKLPEFVNIGFSASTGLSTESNVIHSWEFSSNLEDSNSTTSLVEGNDGK 291
+S DL L + + +GFS++TG + W F N E S +
Sbjct: 221 KPLVSIVRDLSSVLLQDMFVGFSSATGNIVSEIFVLGWSFGVNGEAQPLALSKLPRLP-V 279
Query: 292 GSLKTVIVV------VAVIVPVILVFLIASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFD 345
LK V V +I +++ FL+ + I+KR+RK +E ++++ F K
Sbjct: 280 WDLKPTRVYRFYKNWVPLISLLLIPFLLIIFLVRFIMKRRRKFAEE-VEDWETEFGKN-- 336
Query: 346 LDKATIPRRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENS 405
R + +L AT GF D +LG GG+G VYKG + K +AVKR+ +
Sbjct: 337 --------RLRFKDLYYATKGFKDKNILGSGGFGSVYKGIMPKTKKEIAVKRVSNESRQG 388
Query: 406 ERVFINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKK-SLAWEV 464
+ F+ E+ I ++ HRNLV +G+C +DELLLV++YMPNGSLD +L+ + +L W+
Sbjct: 389 LKEFVAEIVSIGQMSHRNLVPLVGYCRRRDELLLVYDYMPNGSLDKYLYNSPEVTLDWKQ 448
Query: 465 RYKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQR 524
R+KV GVA+AL YLH++ EQ V+HRD+K++NVLLD + + +LGDFG+A+L D Q
Sbjct: 449 RFKVINGVASALFYLHEEWEQVVIHRDVKASNVLLDAELNGRLGDFGLAQLCDHGSDPQT 508
Query: 525 TDVVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFF----QDGDFHVPLMNWV 580
T VVGT+GYLAP++I GRA+ +D+++FG++ LE+A GRR Q G+ V L++WV
Sbjct: 509 TRVVGTWGYLAPDHIRTGRATTTTDVFAFGVLLLEVACGRRPIEINNQSGE-RVVLVDWV 567
Query: 581 WGLYVEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMAL 640
+ ++E N++ A D L E+D E++ +L +GL C+HS+ RP +V++ L+ + L
Sbjct: 568 FRFWMEANILDAKDPNLGSEYDQKEVEMVLKLGLLCSHSDPLARPTMRQVLQYLRGDAML 627
Query: 641 PEL-PLDMHDRAPPIVPFRQSNGPSM---APPMTNSLITSGR 678
P+L PLD+ + SN M + SL++SGR
Sbjct: 628 PDLSPLDLRGSGIMLGTHNGSNESGMFTSGSSVAYSLLSSGR 669
>At4g04960 unknown protein
Length = 686
Score = 305 bits (780), Expect = 7e-83
Identities = 230/648 (35%), Positives = 342/648 (52%), Gaps = 49/648 (7%)
Query: 17 KNILVLFSLLLVLSYLPKTNSLLFNITNFDDPTVASNMSYQGDGKSTNGSIDL-NKVSYL 75
K +L L +L L+L +FN F+D + SN+S G + + L N+ S
Sbjct: 2 KALLFLLTLFLILPNPISAIDFIFN--GFNDSS--SNVSLFGIATIESKILTLTNQTS-- 55
Query: 76 FRVGRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTYGDGFVFYLAPLGYQIPPNS 135
F GRA Y++ + D T+++ F+T F FT+ +T G G VF AP I +S
Sbjct: 56 FATGRALYNRTIRTKDPITSSVLPFSTSFIFTMAPYKNTLPGHGIVFLFAP-STGINGSS 114
Query: 136 AGGVYGLFNATTNSNFVMNYVVGVEFDTFVGP--TDPPMKHVGIDDNALTSV---AFGKF 190
+ GLFN T N N N++ GVEFD F +D HVGID N+L SV G +
Sbjct: 115 SAQHLGLFNLTNNGN-PSNHIFGVEFDVFANQEFSDIDANHVGIDVNSLHSVYSNTSGYW 173
Query: 191 DIDK--------NLGRVCYVLIDYNSDEKMLEVFWSFKGRFVKGDGSYGNSSISYQIDLM 242
D N GR V IDY + ++ V G+ +S ++L
Sbjct: 174 SDDGVVFKPLKLNDGRNYQVWIDYR--DFVVNVTMQVAGKIRPKI-----PLLSTSLNLS 226
Query: 243 KKLPEFVNIGFSASTGLSTESNVIHSWEFS-SNLEDSNS--TTSLVEGNDGKGSLKTVIV 299
+ + + +GF+A+TG +S+ I +W FS SN SNS TT L K S+ V
Sbjct: 227 DVVEDEMFVGFTAATGRLVQSHKILAWSFSNSNFSLSNSLITTGLPSFVLPKDSI--VKA 284
Query: 300 VVAVIVPVILVFLIASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFDLDKATIPRRFEYSE 359
V V V++ FL+ ++VG V+ RK + + D + P R Y E
Sbjct: 285 KWFVFVLVLICFLVVALVGLVLFAVVRKRLERARKRALME-----DWEMEYWPHRIPYEE 339
Query: 360 LVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIISRL 419
+ + T GFD+ ++G GG G+VYKG L VAVKRI + + R F+ E+ + RL
Sbjct: 340 IESGTKGFDEKNVIGIGGNGKVYKGLLQGGVVEVAVKRISQESSDGMREFVAEISSLGRL 399
Query: 420 IHRNLVQFIGWCHEQ-DELLLVFEYMPNGSLDTHLF-GDKK--SLAWEVRYKVALGVANA 475
HRNLV GWC ++ +LV++YM NGSLD +F D+K +L+ E R ++ GVA+
Sbjct: 400 KHRNLVSLRGWCKKEVGSFMLVYDYMENGSLDRWIFENDEKITTLSCEERIRILKGVASG 459
Query: 476 LRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYGYLA 535
+ YLH+ E VLHRDIK++NVLLD D +L DFG+A++ + T VVGT GYLA
Sbjct: 460 ILYLHEGWESKVLHRDIKASNVLLDRDMIPRLSDFGLARVHGHEQPVRTTRVVGTAGYLA 519
Query: 536 PEYINGGRASKESDMYSFGIVALELATGRRFFQDGDFHVPLMNWVWGLYVEGNLMCAADE 595
PE + GRAS ++D++++GI+ LE+ GRR ++G PLM+WVWGL G ++ D
Sbjct: 520 PEVVKTGRASTQTDVFAYGILVLEVMCGRRPIEEG--KKPLMDWVWGLMERGEILNGLDP 577
Query: 596 KLNMEFDVSEM----KSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMA 639
++ M V+E+ + +L +GL C H + +RP +V++V + + A
Sbjct: 578 QMMMTQGVTEVIDEAERVLQLGLLCAHPDPAKRPSMRQVVQVFEGDKA 625
>At3g08870 putative serine/threonine protein kinase
Length = 693
Score = 297 bits (760), Expect = 1e-80
Identities = 215/668 (32%), Positives = 340/668 (50%), Gaps = 62/668 (9%)
Query: 13 INQSKNILVLFSLLLVLSYLPKTNSLL------FNITNFDDPTVASNMSYQGDGKST-NG 65
I +S N + F L++LS K++ L F F + + Q +G ST
Sbjct: 3 IARSINSFMFFFFLMILSNASKSSVLAEATTAKFTFIGFKE----NQTDIQTEGASTIQH 58
Query: 66 SIDLNKVSYLFR--VGRAFYSQPLHLWDKKTNT---LTSFTTRFSFTIDKLNDTTYGDGF 120
DL +++ + G AFY +P+ L + ++ + SF+T F F I + G GF
Sbjct: 59 DNDLLRLTNRKQNVTGTAFYRKPIRLRELTNSSDIKVCSFSTSFVFVILPSSPGNGGFGF 118
Query: 121 VFYLAPLGYQIPPNSAGGVYGLFNATTNSNFVMNYVVGVEFDTFVGPTDPPMK---HVGI 177
F L+P + P + GL N T N N N+V VEFDT G D + H+G+
Sbjct: 119 TFTLSPTPNR-PGAESAQYLGLLNRTNNGN-PSNHVFAVEFDTVQGFKDGADRRGNHIGL 176
Query: 178 DDNALTS----------VAFGKFDIDKNLGRVCYVLIDYNSDEKMLEVFWSFKGRFVKGD 227
+ N L+S K D G VLIDY+ + L V K
Sbjct: 177 NFNNLSSNVQEPLIYYDTEDRKEDFQLESGEPIRVLIDYDGSSETLNVTIYPTRLEFKPK 236
Query: 228 GSYGNSSISYQIDLMKKLPEFVNIGFSASTGLSTES-NVIHSWEFSSNLEDSNSTTSLVE 286
+ +S +++K + + +GF+A+TG S + + W FSS E+ + +
Sbjct: 237 KPLISRRVSELSEIVK---DEMYVGFTAATGKDQSSAHYVMGWSFSSCGENPMADWLEIS 293
Query: 287 G-------NDGKGSLKTVIVVVAVIVPVILVFLIASIVGWVIVKRKRKNCDEGLDEYGIP 339
++ KG VIV++ + V LV L+ + +V+ KR+ + ++ L+++ I
Sbjct: 294 RLPPPPRLSNKKGYNSQVIVLIVALSIVTLVLLVLLFI-FVMYKRRIQE-EDTLEDWEID 351
Query: 340 FPKKFDLDKATIPRRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIF 399
+P RF Y +L AT F + ++G GG+G VY+G LS G I AVK+I
Sbjct: 352 YP-----------HRFRYRDLYLATKKFKESEIIGTGGFGIVYRGNLSSSGPI-AVKKIT 399
Query: 400 ADFENSERVFINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS 459
++ R F+ E+ + RL H+NLV GWC ++ELLL+++Y+PNGSLD+ L+ +
Sbjct: 400 SNSLQGVREFMAEIESLGRLGHKNLVNLQGWCKHKNELLLIYDYIPNGSLDSLLYQTPRR 459
Query: 460 ----LAWEVRYKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKL 515
L W+VR+++ G+A+ L YLH++ EQ V+HRD+K +NVL+D D + KLGDFG+A+L
Sbjct: 460 NGIVLPWDVRFEIIKGIASGLLYLHEEWEQIVVHRDVKPSNVLIDEDMNAKLGDFGLARL 519
Query: 516 VDPMLRTQRTDVVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDGDFHVP 575
+ TQ T +VGT GY+APE G+ S SD+++FG++ LE+ G + +F
Sbjct: 520 YERGTLTQTTKIVGTLGYMAPELTRNGKGSTASDVFAFGVLLLEIVCGNKPTNAENFF-- 577
Query: 576 LMNWVWGLYVEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQ 635
L +WV + G ++C D+ L F+ E K L+VGL C H K RP V++ L
Sbjct: 578 LADWVMEFHTNGGILCVVDQNLGSSFNGREAKLALVVGLLCCHQKPKFRPSMRMVLRYLN 637
Query: 636 LEMALPEL 643
E +P++
Sbjct: 638 GEENVPQI 645
>At5g01550 receptor like protein kinase
Length = 688
Score = 291 bits (745), Expect = 8e-79
Identities = 199/604 (32%), Positives = 319/604 (51%), Gaps = 59/604 (9%)
Query: 79 GRAFYSQPLHLWDKKTN----TLTSFTTRFSFTIDKLNDTTYGDGFVFYLAPLGYQIPPN 134
G +FY +P+ L + T+ T+ SF+T F F I + + G GF F L+P +
Sbjct: 63 GTSFYHKPVRLLETNTSSTNSTIRSFSTSFVFVIIPTSSSNGGFGFTFTLSPTPDRTGAE 122
Query: 135 SAGGVYGLFNATTNSNFVMNYVVGVEFDTFVG---PTDPPMKHVGIDDNALTS-----VA 186
SA + GL N + N N+V VEFDT G D H+G++ N+LTS V
Sbjct: 123 SAQYL-GLLNKANDGNST-NHVFAVEFDTVQGFKDGADRTGNHIGLNFNSLTSDVQEPVV 180
Query: 187 F-------GKFDIDKNLGRVCYVLIDYNSDEKMLEVFW---SFKGRFVKGDGSYGNSSIS 236
+ K D G ++DY+ + L + + K R V+ IS
Sbjct: 181 YYDNEDPNRKEDFPLQSGDPIRAILDYDGPTQTLNLTVYPANLKSRPVR-------PLIS 233
Query: 237 YQIDLMKKL-PEFVNIGFSASTGLSTES-NVIHSWEFSSNLED-SNSTTSLVE------G 287
+ + ++ E + +GF+A+TG S + + W FSS + + T L+E
Sbjct: 234 RPVPKLSQIVQEEMYVGFTAATGRDQSSAHYVMGWSFSSGGDLLTEDTLDLLELPRPPPN 293
Query: 288 NDGKGSLKTVIVVVAVIVPVILVFLIASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFDLD 347
K + ++ + V + + V L+A + +V+ K KR E L+++ I P
Sbjct: 294 TAKKRGYNSQVLALIVALSGVTVILLALLFFFVMYK-KRLQQGEVLEDWEINHP------ 346
Query: 348 KATIPRRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKI-VAVKRIFADFENSE 406
R Y +L AAT+GF ++R++G GG+G V++G LS +AVK+I +
Sbjct: 347 -----HRLRYKDLYAATDGFKENRIVGTGGFGTVFRGNLSSPSSDQIAVKKITPNSMQGV 401
Query: 407 RVFINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS----LAW 462
R FI E+ + RL H+NLV GWC ++++LLL+++Y+PNGSLD+ L+ + L+W
Sbjct: 402 REFIAEIESLGRLRHKNLVNLQGWCKQKNDLLLIYDYIPNGSLDSLLYSRPRQSGVVLSW 461
Query: 463 EVRYKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRT 522
R+K+A G+A+ L YLH++ E+ V+HRDIK +NVL++ D + +LGDFG+A+L + ++
Sbjct: 462 NARFKIAKGIASGLLYLHEEWEKVVIHRDIKPSNVLIEDDMNPRLGDFGLARLYERGSQS 521
Query: 523 QRTDVVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDGDFHVPLMNWVWG 582
T VVGT GY+APE G++S SD+++FG++ LE+ +GRR G F L +WV
Sbjct: 522 NTTVVVGTIGYMAPELARNGKSSSASDVFAFGVLLLEIVSGRRPTDSGTFF--LADWVME 579
Query: 583 LYVEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPE 642
L+ G ++ A D +L +D E + L+VGL C H RP V++ L + +PE
Sbjct: 580 LHARGEILHAVDPRLGFGYDGVEARLALVVGLLCCHQRPTSRPSMRTVLRYLNGDDDVPE 639
Query: 643 LPLD 646
+ D
Sbjct: 640 IDND 643
>At1g70110 hypothetical protein
Length = 666
Score = 290 bits (741), Expect = 2e-78
Identities = 211/653 (32%), Positives = 318/653 (48%), Gaps = 51/653 (7%)
Query: 19 ILVLFSLLL-----VLSYLPKTNSLLFNITNFDDPTVASNMSYQGDGKSTNGSIDLNKVS 73
+L+LF +L V+ P N + FN + NM G N + S
Sbjct: 2 VLLLFLVLFFVPESVVCQRPNPNGVEFN--------TSGNMYTSGSAYINNNGLIRLTNS 53
Query: 74 YLFRVGRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTYGDGFVFYLAPLGYQIPP 133
G+ FY+ L + T++SF+T F F+I+ N G G F + P + P
Sbjct: 54 TPQTTGQVFYNDQLRFKNSVNGTVSSFSTTFVFSIEFHNGIYGGYGIAFVICPTR-DLSP 112
Query: 134 NSAGGVYGLFNATTNSNFVMNYVVGVEFDTFVGPT--DPPMKHVGIDDNALTS--VAFGK 189
GLFN + N N++V VE DT V D HVGID N L S VA
Sbjct: 113 TFPTTYLGLFNRS-NMGDPKNHIVAVELDTKVDQQFEDKDANHVGIDINTLVSDTVALAG 171
Query: 190 FDIDK--------NLGRVCYVLIDYNSDEKMLEVFWSFKGRFVKGDGSYGNSSISYQIDL 241
+ +D N G+ + I+Y+S +K + V + +V +S + DL
Sbjct: 172 YYMDNGTFRSLLLNSGQPMQIWIEYDSKQKQINV--TLHPLYVPKPKI---PLLSLEKDL 226
Query: 242 MKKLPEFVNIGFSASTGLSTESNVIHSWEFSSNLE----DSNSTTSLVEGNDGKGSLKTV 297
L E + +GF+++TG T S+ I W F N D + + N
Sbjct: 227 SPYLLELMYVGFTSTTGDLTASHYILGWTFKMNGTTPDIDPSRLPKIPRYNQPWIQSPNG 286
Query: 298 IVVVAVIVPVILVFLIASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFDLDKATIPRRFEY 357
I+ +++ V +++ +I S+ W+ +KRK+ E L+++ + F P RF +
Sbjct: 287 ILTISLTVSGVIILIILSLSLWLFLKRKKLL--EVLEDWEVQFG----------PHRFAF 334
Query: 358 SELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIIS 417
+L AT GF D +LG+GG+G+VYKG L +AVK + D R FI E+ I
Sbjct: 335 KDLHIATKGFKDTEVLGKGGFGKVYKGTLPVSNVEIAVKMVSHDSRQGMREFIAEIATIG 394
Query: 418 RLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKK-SLAWEVRYKVALGVANAL 476
RL H NLV+ G+C + EL LV++ M GSLD L+ + +L W R+K+ VA+ L
Sbjct: 395 RLRHPNLVRLQGYCRHKGELYLVYDCMAKGSLDKFLYHQQTGNLDWSQRFKIIKDVASGL 454
Query: 477 RYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYGYLAP 536
YLH Q ++HRDIK AN+LLD + + KLGDFG+AKL D Q + V GT GY++P
Sbjct: 455 YYLHQQWVQVIIHRDIKPANILLDANMNAKLGDFGLAKLCDHGTDPQTSHVAGTLGYISP 514
Query: 537 EYINGGRASKESDMYSFGIVALELATGRR--FFQDGDFHVPLMNWVWGLYVEGNLMCAAD 594
E G+AS SD+++FGIV LE+A GR+ + + L +WV + ++M D
Sbjct: 515 ELSRTGKASTRSDVFAFGIVMLEIACGRKPILPRASQREMVLTDWVLECWENEDIMQVLD 574
Query: 595 EKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPELPLDM 647
K+ E+ + +L +GL+C+H RP VI++L LP LD+
Sbjct: 575 HKIGQEYVEEQAALVLKLGLFCSHPVAAIRPNMSSVIQLLDSVAQLPHNLLDI 627
>At5g42120 receptor lectin kinase-like protein
Length = 691
Score = 289 bits (739), Expect = 4e-78
Identities = 218/691 (31%), Positives = 344/691 (49%), Gaps = 96/691 (13%)
Query: 13 INQSKNILVLFSLLLVLSYLPKTNSLLFNITNFDDPTVASNMSYQGDGKSTNGSIDLNKV 72
+N LV+F L+L LS T S F+ N++ GD + +I L +
Sbjct: 1 MNHHHYSLVIFHLILFLSLDFPTLSHRFS-------PPLQNLTLYGDAFFRDRTISLTQQ 53
Query: 73 SYLFR-------------VGRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTYGDG 119
F +GRA Y P+ + TNT SF+ RFSF+I +GDG
Sbjct: 54 QPCFPSVTTPPSKPSSSGIGRALYVYPIKFLEPSTNTTASFSCRFSFSIIASPSCPFGDG 113
Query: 120 FVFYLAPLGYQIPPNSAGGVYGLFNATTNSNFVMNYVVGVEFDTFVGPT--DPPMKHVGI 177
F F + + G GL N + + VEFDT P D HVGI
Sbjct: 114 FAFLITSNADSFV--FSNGFLGLPNPDDS-------FIAVEFDTRFDPVHGDINDNHVGI 164
Query: 178 DDNALTSV----AFGKFDIDKNLGRVCYVLIDYNSDEKMLEVFWSFKGRFVKGDGSYGNS 233
D +++ SV A K D G+ I+Y+ K++ V+ + VK
Sbjct: 165 DVSSIFSVSSVDAISK-GFDLKSGKKMMAWIEYSDVLKLIRVWVGYSR--VKPTSPV--- 218
Query: 234 SISYQIDLMKKLPEFVNIGFSAST-GLSTESNVIHSWEFSS---------NLEDSNSTTS 283
+S QIDL K+ E++++GFSAS G+ + +++ W+F + E+
Sbjct: 219 -LSTQIDLSGKVKEYMHVGFSASNAGIGSALHIVERWKFRTFGSHSDAIQEEEEEKDEEC 277
Query: 284 LVEGNDGKGSLKTV--------IVVVAVIVPV--------ILVFLIASIVGWVIVKRKRK 327
LV + + K + + VV + +PV +V L+A IV +I +KR
Sbjct: 278 LVCSGEVSENPKEIHRKGFNFRVTVVGLKIPVWSLLPGLAAIVILVAFIVFSLICGKKRI 337
Query: 328 NCDEGLDEYGIPFPKKFDLDKATIPRRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALS 387
+ +E G+ +P R +E+ +AT+GF+++ ++G+G VY+G++
Sbjct: 338 S-EEADSNSGL----------VRMPGRLSLAEIKSATSGFNENAIVGQGASATVYRGSIP 386
Query: 388 YLGKIVAVKRIFADF--ENSERVFINEVRIISRLI-HRNLVQFIGWCHEQDELLLVFEYM 444
+G VAVKR + + + F E ++ + H+NLVQF GWC E E LVFEY+
Sbjct: 387 SIGS-VAVKRFDREHWPQCNRNPFTTEFTTMTGYLRHKNLVQFQGWCSEGTETALVFEYL 445
Query: 445 PNGSLDTHLFGDKKS--------LAWEVRYKVALGVANALRYLHDDAEQCVLHRDIKSAN 496
PNGSL L S L+W+ R + LGVA+AL YLH++ E+ ++HRD+K+ N
Sbjct: 446 PNGSLSEFLHKKPSSDPSEEIIVLSWKQRVNIILGVASALTYLHEECERQIIHRDVKTCN 505
Query: 497 VLLDTDFSTKLGDFGMAKLVDP---MLRTQRTDVVGTYGYLAPEYINGGRASKESDMYSF 553
++LD +F+ KLGDFG+A++ + + T GT GYLAPEY+ G S+++D+YSF
Sbjct: 506 IMLDAEFNAKLGDFGLAEIYEHSALLAGRAATLPAGTMGYLAPEYVYTGVPSEKTDVYSF 565
Query: 554 GIVALELATGRRFFQDGDFHVPLMNWVWGLYVEGNLMCAADEKLNMEFDVSEMKSLLIVG 613
G+V LE+ TGRR GD L++ +W + G ++ AD L EFD EM+ +L+VG
Sbjct: 566 GVVVLEVCTGRR--PVGDDGAVLVDLMWSHWETGKVLDGADIMLREEFDAEEMERVLMVG 623
Query: 614 LWCTHSNDKERPKAYEVIKVLQLEMALPELP 644
+ C H + ++RP+ + +++++ E LP LP
Sbjct: 624 MVCAHPDSEKRPRVKDAVRIIRGEAPLPVLP 654
>At5g01540 receptor like protein kinase
Length = 682
Score = 288 bits (738), Expect = 5e-78
Identities = 193/596 (32%), Positives = 313/596 (52%), Gaps = 52/596 (8%)
Query: 79 GRAFYSQPLHLWDKKTNTLT--SFTTRFSFTIDKLNDTTYGDGFVFYLAPLGYQIPPNSA 136
G AFY +P+ L ++ + +T SF+T F F I + + G GF F L+P Y++ SA
Sbjct: 70 GTAFYHKPVRLLNRNSTNVTIRSFSTSFVFVIIPSSSSNKGFGFTFTLSPTPYRLNAGSA 129
Query: 137 GGVYGLFNATTNSNFVMNYVVGVEFDTFVGP----TDPPMKHVGIDDNALTS------VA 186
+ G+FN N + N+V VEFDT G TD +G++ N+ TS V
Sbjct: 130 QYL-GVFNKENNGD-PRNHVFAVEFDTVQGSRDDNTDRIGNDIGLNYNSRTSDLQEPVVY 187
Query: 187 FGKFDIDKN------LGRVCYVLIDYNSDEKMLEVFWSFKGRFVKGDGSYGNSSISYQID 240
+ D +K G L++Y+ +ML V K + + ++
Sbjct: 188 YNNDDHNKKEDFQLESGNPIQALLEYDGATQMLNVTVYPARLGFKPTKPLISQHVPKLLE 247
Query: 241 LMKKLPEFVNIGFSASTGLSTES-NVIHSWEFSSNLEDSNSTTSLVEG------NDGK-- 291
+++ E + +GF+ASTG S + + W FSS E + ++ N K
Sbjct: 248 IVQ---EEMYVGFTASTGKGQSSAHYVMGWSFSSGGERPIADVLILSELPPPPPNKAKKE 304
Query: 292 GSLKTVIVVVAVIVPVILVFLIASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFDLDKATI 351
G VIV++ + V+LV L+ ++ + ++ +KR +E L+++ I P
Sbjct: 305 GLNSQVIVMIVALSAVMLVMLV--LLFFFVMYKKRLGQEETLEDWEIDHP---------- 352
Query: 352 PRRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFIN 411
RR Y +L AT+GF ++G GG+G V+KG L I AVK+I R F+
Sbjct: 353 -RRLRYRDLYVATDGFKKTGIIGTGGFGTVFKGKLPNSDPI-AVKKIIPSSRQGVREFVA 410
Query: 412 EVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS----LAWEVRYK 467
E+ + +L H+NLV GWC +++LLL+++Y+PNGSLD+ L+ + L+W R++
Sbjct: 411 EIESLGKLRHKNLVNLQGWCKHKNDLLLIYDYIPNGSLDSLLYTVPRRSGAVLSWNARFQ 470
Query: 468 VALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDV 527
+A G+A+ L YLH++ E+ V+HRD+K +NVL+D+ + +LGDFG+A+L + ++ T +
Sbjct: 471 IAKGIASGLLYLHEEWEKIVIHRDVKPSNVLIDSKMNPRLGDFGLARLYERGTLSETTAL 530
Query: 528 VGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDGDFHVPLMNWVWGLYVEG 587
VGT GY+APE G S SD+++FG++ LE+ GR+ G F L++WV L+ G
Sbjct: 531 VGTIGYMAPELSRNGNPSSASDVFAFGVLLLEIVCGRKPTDSGTFF--LVDWVMELHANG 588
Query: 588 NLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPEL 643
++ A D +L +D E + L VGL C H RP V++ L E +PE+
Sbjct: 589 EILSAIDPRLGSGYDGGEARLALAVGLLCCHQKPASRPSMRIVLRYLNGEENVPEI 644
>At5g01560 receptor like protein kinase
Length = 685
Score = 288 bits (737), Expect = 7e-78
Identities = 210/669 (31%), Positives = 333/669 (49%), Gaps = 81/669 (12%)
Query: 22 LFSLLLVLSYLPKTNSLLFNITNFDDPTVASNMS---YQGDGKST-NGSIDLNKVSYLFR 77
+ SLLLVL + + T F N S QGD T NG + L +
Sbjct: 1 MVSLLLVLFLVRAHVATTETTTEFIFHGFKGNQSEIHMQGDSTITSNGLLRLTDRNSDV- 59
Query: 78 VGRAFYSQPLHLWDKKTN--TLTSFTTRFSFTIDKLNDTTYGDGFVFYLAPLGYQIPPNS 135
VG AFY +P+ L D + T+ SF+T F F I + + G GF F L+P PN
Sbjct: 60 VGTAFYHKPVRLLDSNSTNTTVRSFSTSFIFIIPSSSTSNGGFGFTFTLSPT-----PNR 114
Query: 136 AGG----VYGLFNATTNSNFVMNYVVGVEFDTFVGPTDPPMK---HVGIDDNALTS---- 184
GL N + N N+V VEFDT G D + H+G++ N+L+S
Sbjct: 115 TDADPEQYMGLLNERNDGNS-SNHVFAVEFDTVQGFKDGTNRIGNHIGLNFNSLSSDVQE 173
Query: 185 --VAFGKFDIDKN-----LGRVCYVLIDYNSDEKMLEVF-----WSFKGRFVKGDGSYGN 232
F D K G V +DY+ K L + +K R
Sbjct: 174 PVAYFNNNDSQKEEFQLVSGEPIQVFLDYHGPTKTLNLTVYPTRLGYKPRI--------- 224
Query: 233 SSISYQIDLMKKLP-EFVNIGFSASTGLSTESNV--IHSWEFSSNLEDSNSTTSLVEG-- 287
IS ++ + + + + +GF+A+TG +S+ + W F+S E + +
Sbjct: 225 PLISREVPKLSDIVVDEMFVGFTAATGRHGQSSAHYVMGWSFASGGEHPLAAMLDISQLP 284
Query: 288 ----NDGK-----GSLKTVIVVVAVIVPVILVFLIASIVGWVIVKRKRKNCDEGLDEYGI 338
N K G + +IV ++ ++ ++LV L ++ +KR +E L+++ I
Sbjct: 285 PPPPNKAKKRGYNGKVIALIVALSTVISIMLVLLFL-----FMMYKKRMQQEEILEDWEI 339
Query: 339 PFPKKFDLDKATIPRRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRI 398
P RF Y +L AT GF ++R++G GG+G VY+G + +AVK+I
Sbjct: 340 DHP-----------HRFRYRDLYKATEGFKENRVVGTGGFGIVYRGNIRSSSDQIAVKKI 388
Query: 399 FADFENSERVFINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKK 458
+ R F+ E+ + RL H+NLV GWC +++LLL+++Y+PNGSLD+ L+ +
Sbjct: 389 TPNSMQGVREFVAEIESLGRLRHKNLVNLQGWCKHRNDLLLIYDYIPNGSLDSLLYSKPR 448
Query: 459 S----LAWEVRYKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAK 514
L+W R+++A G+A+ L YLH++ EQ V+HRD+K +NVL+D+D + +LGDFG+A+
Sbjct: 449 RSGAVLSWNARFQIAKGIASGLLYLHEEWEQIVIHRDVKPSNVLIDSDMNPRLGDFGLAR 508
Query: 515 LVDPMLRTQRTDVVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDGDFHV 574
L + ++ T VVGT GY+APE G +S SD+++FG++ LE+ +GR+ G F +
Sbjct: 509 LYERGSQSCTTVVVGTIGYMAPELARNGNSSSASDVFAFGVLLLEIVSGRKPTDSGTFFI 568
Query: 575 PLMNWVWGLYVEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVL 634
+WV L G ++ A D +L +D E + L VGL C H + RP V++ L
Sbjct: 569 --ADWVMELQASGEILSAIDPRLGSGYDEGEARLALAVGLLCCHHKPESRPLMRMVLRYL 626
Query: 635 QLEMALPEL 643
+ +PE+
Sbjct: 627 NRDEDVPEI 635
>At5g59260 receptor-like protein kinase
Length = 674
Score = 284 bits (726), Expect = 1e-76
Identities = 198/590 (33%), Positives = 299/590 (50%), Gaps = 45/590 (7%)
Query: 79 GRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTYGDGFVFYLAPLGYQIPPNSAGG 138
G AFY+ P+ ++ SF+T F F I L +TYG G F ++P SA
Sbjct: 64 GHAFYNIPIKFTASSLSSF-SFSTEFVFAIFPLQKSTYGHGMAFVVSPTKDLRSNGSANS 122
Query: 139 VYGLFNATTNSNFVMNYVVGVEFDTFVGPT--DPPMKHVGIDDNALTSV-----AFGKFD 191
G+FN N N ++ VE DT D VGID N++ SV ++
Sbjct: 123 NLGIFNRA-NDNKTATHIFAVELDTNQNSESFDKGGNDVGIDINSIVSVESADASYFNAR 181
Query: 192 IDKNL------GRVCYVLIDYNSDEKMLEVFWSFKGRFVKGDGSYGNSSISYQIDLMKK- 244
KN+ G+ V IDY+ EK+L V + + K D Y +S I ++ L+ +
Sbjct: 182 KGKNISLPLASGKSILVWIDYDGIEKVLNVTLA-PVQTPKPDSPYFSSFIKPKVPLLSRS 240
Query: 245 ------LPEFVNIGFSASTGLSTESNVIHSWEFSSNLEDSNSTTSLVEGNDGKGSLKTVI 298
E + +GFS STG + I W F + + S + +
Sbjct: 241 INLSEIFTETMYVGFSGSTGSIKSNQYILGWSFKQGGKAESLDISRLSNPPPSPKRFPLK 300
Query: 299 VVVAVIVPVILVFLIASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFDLDKATIPRRFEYS 358
V+ + I + G ++ K+K E L+++ +K P+R+ +
Sbjct: 301 EVLGATISTIAFLTL----GGIVYLYKKKKYAEVLEQW----------EKEYSPQRYSFR 346
Query: 359 ELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIISR 418
L AT GF ++++LG GG+G+VYKG L G +AVKR++ D E + ++ E+ + R
Sbjct: 347 ILYKATKGFRENQLLGAGGFGKVYKGILPS-GTQIAVKRVYHDAEQGMKQYVAEIASMGR 405
Query: 419 LIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKK--SLAWEVRYKVALGVANAL 476
L H+NLV +G+C + ELLLV++YMPNGSLD +LF K L W R + GVA+AL
Sbjct: 406 LRHKNLVHLLGYCRRKGELLLVYDYMPNGSLDDYLFHKNKLKDLTWSQRVNIIKGVASAL 465
Query: 477 RYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYGYLAP 536
YLH++ EQ VLHRDIK++N+LLD D + KLGDFG+A+ D + + T VVGT GY+AP
Sbjct: 466 LYLHEEWEQVVLHRDIKASNILLDADLNGKLGDFGLARFHDRGVNLEATRVVGTIGYMAP 525
Query: 537 EYINGGRASKESDMYSFGIVALELATGRRFFQDGDF---HVPLMNWVWGLYVEGNLMCAA 593
E G + +D+Y+FG LE+ GRR D D V L+ WV L
Sbjct: 526 ELTAMGVTTTCTDVYAFGAFILEVVCGRRPV-DPDAPREQVILVKWVASCGKRDALTDTV 584
Query: 594 DEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPEL 643
D KL ++F V E K LL +G+ C+ N + RP ++++ L+ +++P +
Sbjct: 585 DSKL-IDFKVEEAKLLLKLGMLCSQINPENRPSMRQILQYLEGNVSVPAI 633
>At5g60280 receptor like protein kinase
Length = 657
Score = 280 bits (715), Expect = 2e-75
Identities = 198/587 (33%), Positives = 305/587 (51%), Gaps = 51/587 (8%)
Query: 79 GRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTYGDGFVFYLAPLGYQIPPNSAGG 138
G AF+ QPL ++ SF+T F + + T G+G F+L+P + A
Sbjct: 63 GHAFFRQPLVF---NSSEPLSFSTHFVCAMVRKPGVTGGNGIAFFLSP-SMDLTNADATQ 118
Query: 139 VYGLFNATTNSNFVMNYVVGVEFDTFVGPT--DPPMKHVGIDDNALTSVAFGK---FDID 193
GLFN TTN + +++ +E DT D HVGID N+LTSV F
Sbjct: 119 YLGLFNTTTNRS-PSSHIFAIELDTVQSAEFDDIDNNHVGIDVNSLTSVESAPASYFSDK 177
Query: 194 KNLGRVCYVLIDYNSDEKMLEVFWSFKGRFVK------GDGSYGNSSISYQIDLMKKLPE 247
K L + +L ++V+ F G + G S IS ++L + + +
Sbjct: 178 KGLNKSISLL-----SGDSIQVWVDFDGTVLNVSLAPLGIRKPSQSLISRSMNLSEVIQD 232
Query: 248 FVNIGFSASTGLSTESNVIHSWEFSSNLEDSNSTTSLVEGNDGKGSLKTVIVVVAVIVPV 307
+ +GFSA+TG ++ I W FS + S +KT ++++ +++ V
Sbjct: 233 RMFVGFSAATGQLANNHYILGWSFSRSKASLQSLDISKLPQVPHPKMKTSLLLILLLI-V 291
Query: 308 ILVFLIASIVGWVIVKRKR--KNCDEGLDEYGIPFPKKFDLDKATIPRRFEYSELVAATN 365
+ + L+ +VG + +R + + +E EYG P R+ Y L AT
Sbjct: 292 LGIILLVLLVGAYLYRRNKYAEVREEWEKEYG--------------PHRYSYKSLYKATK 337
Query: 366 GFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIISRLIHRNLV 425
GF D LG+GG+G+VYKG L + +AVKR E + F+ E+ + L HRNLV
Sbjct: 338 GFHKDGFLGKGGFGEVYKGTLPQ--EDIAVKRFSHHGERGMKQFVAEIASMGCLDHRNLV 395
Query: 426 QFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKK-SLAWEVRYKVALGVANALRYLHDDAE 484
G+C + E LLV +YMPNGSLD LF +++ SL W R + G+A+AL+YLH +A
Sbjct: 396 PLFGYCRRKGEFLLVSKYMPNGSLDQFLFHNREPSLTWSKRLGILKGIASALKYLHTEAT 455
Query: 485 QCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYGYLAPEYINGGRA 544
Q VLHRDIK++NV+LDTDF+ KLGDFGMA+ D T VGT GY+ PE + G A
Sbjct: 456 QVVLHRDIKASNVMLDTDFTGKLGDFGMARFHDHGANPTTTGAVGTVGYMGPELTSMG-A 514
Query: 545 SKESDMYSFGIVALELATGRRFFQDGDFHVP-----LMNWVWGLYVEGNLMCAADEKLNM 599
S ++D+Y+FG + LE+ GRR + ++P L+ WV + +L+ A D KL+
Sbjct: 515 STKTDVYAFGALILEVTCGRRPVEP---NLPIEKQLLVKWVCDCWKRKDLISARDPKLSG 571
Query: 600 EFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPELPLD 646
E + +++ +L +GL CT+ + RP +V++ L +++LP+ D
Sbjct: 572 EL-IPQIEMVLKLGLLCTNLVPESRPDMVKVVQYLDRQVSLPDFSPD 617
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.139 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,175,721
Number of Sequences: 26719
Number of extensions: 750119
Number of successful extensions: 5412
Number of sequences better than 10.0: 1012
Number of HSP's better than 10.0 without gapping: 894
Number of HSP's successfully gapped in prelim test: 118
Number of HSP's that attempted gapping in prelim test: 1627
Number of HSP's gapped (non-prelim): 1185
length of query: 678
length of database: 11,318,596
effective HSP length: 106
effective length of query: 572
effective length of database: 8,486,382
effective search space: 4854210504
effective search space used: 4854210504
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 63 (28.9 bits)
Medicago: description of AC143339.11


