

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
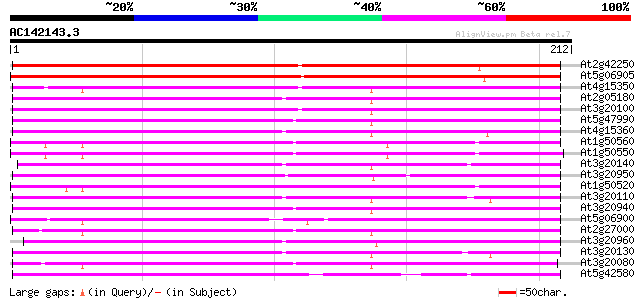
Query= AC142143.3 + phase: 0 /pseudo
(212 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At2g42250 putative cytochrome P450 186 1e-47
At5g06905 cytochrome P450-like protein 175 1e-44
At4g15350 cytochrome P450 like protein 139 8e-34
At2g05180 putative cytochrome P450 135 2e-32
At3g20100 cytochrome P450, putative 134 5e-32
At5g47990 cytochrome P450 133 6e-32
At4g15360 cytochrome P450 like protein 133 8e-32
At1g50560 cytochrome P450, putative 133 8e-32
At1g50550 cytochrome P450, putative 133 8e-32
At3g20140 cytochrome P450, putative 132 1e-31
At3g20950 cytochrome P450, putative 131 3e-31
At1g50520 cytochrome P450 like protein 131 3e-31
At3g20110 cytochrome P450, putative 130 4e-31
At3g20940 cytochrome P450, putative 129 9e-31
At5g06900 cytochrome P450 128 2e-30
At2g27000 putative cytochrome P450 127 4e-30
At3g20960 cytochrome P450 like protein 127 6e-30
At3g20130 cytochrome P450, putative 126 7e-30
At3g20080 cytochrome P450, putative 122 2e-28
At5g42580 cytochrome P450 121 3e-28
>At2g42250 putative cytochrome P450
Length = 514
Score = 186 bits (471), Expect = 1e-47
Identities = 94/208 (45%), Positives = 138/208 (66%), Gaps = 2/208 (0%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
K +LNFS RP+FGS+ + Y+GS F+ A YG YWRFMKK+C+TKLL+ QL +F IRE
Sbjct: 99 KEQELNFSSRPEFGSAEYFKYRGSRFVLAQYGDYWRFMKKLCMTKLLAVPQLEKFADIRE 158
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVRE 121
+E KL+ S+ C E DL F TNN++CRM+M T C N + E I+ LV++
Sbjct: 159 EEKLKLVDSVAKCCREGLPCDLSSQFIKYTNNVICRMAMSTRCSGTDN-EAEEIRELVKK 217
Query: 122 FMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTED-CQGD 180
+ + K+S+G+VLGPL D G GKKL ++ ++D ++E +KE E + +D + D
Sbjct: 218 SLELAGKISVGDVLGPLKVMDFSGNGKKLVAVMEKYDLLVERIMKEREAKAKKKDGTRKD 277
Query: 181 MMDILLQVYRNENAEVRLTRNDIKAFFL 208
++DILL+ YR+ AE+++TRND+K+F L
Sbjct: 278 ILDILLETYRDPTAEMKITRNDMKSFLL 305
>At5g06905 cytochrome P450-like protein
Length = 528
Score = 175 bits (444), Expect = 1e-44
Identities = 95/210 (45%), Positives = 130/210 (61%), Gaps = 3/210 (1%)
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTHD +F+ + FG +YKGS F APYG YWRFMKK+C+TKL + QL RF+ IR
Sbjct: 95 LKTHDPDFASKFVFGPRQFNVYKGSEFFNAPYGSYWRFMKKLCMTKLFAGYQLDRFVDIR 154
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
E+E LL +L+ S + DLGL+FT LT IL +M MG C + NI P+ I+ +V
Sbjct: 155 EEETLALLSTLVERSRNGEACDLGLEFTALTTKILSKMVMGKRCRQNSNI-PKEIRKIVS 213
Query: 121 EFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQ-- 178
+ M + E+ GPL DLFG GKKLR + +D+++E LKE+E + E+ +
Sbjct: 214 DIMACATRFGFMELFGPLRDLDLFGNGKKLRSSIWRYDELVEKILKEYENDKSNEEEEKD 273
Query: 179 GDMMDILLQVYRNENAEVRLTRNDIKAFFL 208
D++DILL Y + AE+RLT N IK F L
Sbjct: 274 KDIVDILLDTYNDPKAELRLTMNQIKFFIL 303
>At4g15350 cytochrome P450 like protein
Length = 509
Score = 139 bits (351), Expect = 8e-34
Identities = 79/209 (37%), Positives = 125/209 (59%), Gaps = 4/209 (1%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSY-FITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+ D+N S+R + L+ GSY FI+APYG YW+FM+K+ VTK+L L R R
Sbjct: 94 RDQDVNVSFRHS-PPIEESLFLGSYSFISAPYGDYWKFMRKLMVTKILGPQALERSRRFR 152
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
E EL++ K+LL + + S ++ + L NN +C+M MG +C ++ + E I+ LV
Sbjct: 153 EDELDRFYKTLLDKAMKKESVEIVEEAAKLNNNTICKMIMGRSCSEETG-EAERIRGLVT 211
Query: 121 EFMHVGAKLSMGEVL-GPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQG 179
E M + K+ + + PL K + + K++ + +FD++LE L EHEE++
Sbjct: 212 ESMALTKKIFLATIFHKPLKKLGISLFKKEIMSVSRKFDELLEKILVEHEEKMEEHHQGT 271
Query: 180 DMMDILLQVYRNENAEVRLTRNDIKAFFL 208
DMMD+LL+ YR+ENAE ++TRN IK+ F+
Sbjct: 272 DMMDVLLEAYRDENAEYKITRNHIKSMFV 300
>At2g05180 putative cytochrome P450
Length = 442
Score = 135 bits (340), Expect = 2e-32
Identities = 81/208 (38%), Positives = 115/208 (54%), Gaps = 2/208 (0%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
+ HD+N S R L+ S ITAPYG YW+FMKK+ TKLL L IR
Sbjct: 100 RAHDVNVSSRGVAAIDESLLFGSSGIITAPYGDYWKFMKKLIATKLLRPQSLESSRGIRG 159
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVRE 121
+EL + SLL + S ++ + L NN LCRMSMG + F + N + E I+ LV E
Sbjct: 160 EELTRFYNSLLDKARMTESVEISTEAMKLVNNTLCRMSMGRS-FSEENGEAEKIRGLVGE 218
Query: 122 FMHVGAKLSMGEVL-GPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQGD 180
+ K+ + +L PL K + + K++ + D++LE L EHEE+++ E D
Sbjct: 219 SYALTKKMFLASLLRKPLKKLRISLFEKEIMGVSDRLDELLERILVEHEEKLHEEHQGTD 278
Query: 181 MMDILLQVYRNENAEVRLTRNDIKAFFL 208
MMD+LL +ENAE +TRN IK+FF+
Sbjct: 279 MMDVLLAASGDENAEYNITRNHIKSFFV 306
>At3g20100 cytochrome P450, putative
Length = 513
Score = 134 bits (336), Expect = 5e-32
Identities = 78/208 (37%), Positives = 119/208 (56%), Gaps = 2/208 (0%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
+ HDLN S R L+ + FI+APYG Y++FMKK VTKLL L R IR
Sbjct: 100 RVHDLNVSSRGSPPFEESLLFGSTGFISAPYGDYFKFMKKHLVTKLLGPQALERSRLIRT 159
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVRE 121
ELE+ +LL + + S ++G + L+NN +C+M MG +C ++ + E ++ L+ E
Sbjct: 160 NELERFYINLLDKATKKESVEIGKEAMKLSNNSICKMIMGRSCLEEKG-EAERVRGLIIE 218
Query: 122 FMHVGAKLSMGEVL-GPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQGD 180
++ K + L G L K + + K++ + FD +LE +L+EHEE+ + E D
Sbjct: 219 SFYLTKKFFLAFTLRGLLEKLGISLFKKEIMGVSRRFDDLLERYLREHEEKPDNEHQDTD 278
Query: 181 MMDILLQVYRNENAEVRLTRNDIKAFFL 208
M+D LL YR+E AE ++TRN IKAF +
Sbjct: 279 MIDALLAAYRDEKAEYKITRNQIKAFLV 306
>At5g47990 cytochrome P450
Length = 511
Score = 133 bits (335), Expect = 6e-32
Identities = 76/208 (36%), Positives = 118/208 (56%), Gaps = 2/208 (0%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
K D+N S RP + S FI PYG Y +FMKK V KLL L R +IR
Sbjct: 101 KAQDVNVSSRPPPPIEESLILGSSSFINTPYGDYSKFMKKFMVQKLLGPQALQRSRNIRA 160
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVRE 121
ELE+ K+LL + + ++ ++ + LTNN +C+M MG +C ++ N + E ++ LV E
Sbjct: 161 DELERFYKTLLDKAMKKQTVEIRNEAMKLTNNTICKMIMGRSCSEE-NGEAETVRGLVTE 219
Query: 122 FMHVGAKLSMGEVL-GPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQGD 180
+ + K +G + PL K + + K+L + FD++LE L EHEE++ D
Sbjct: 220 SIFLTKKHFLGAMFHKPLKKLGISLFAKELMNVSNRFDELLEKILVEHEEKLQEHHQTSD 279
Query: 181 MMDILLQVYRNENAEVRLTRNDIKAFFL 208
M+D+LL+ Y +ENAE ++TR+ IK+ F+
Sbjct: 280 MLDMLLEAYGDENAEYKITRDQIKSLFV 307
>At4g15360 cytochrome P450 like protein
Length = 527
Score = 133 bits (334), Expect = 8e-32
Identities = 74/209 (35%), Positives = 121/209 (57%), Gaps = 3/209 (1%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
K HD+N S R ++ S + APYG YW+FMKK+ TKLL L R +R
Sbjct: 99 KAHDVNVSSRGIIALDESLMFGASGILNAPYGDYWKFMKKLMATKLLRPQVLERSRGVRV 158
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVRE 121
+EL + +S+L + +N S ++G + L NN LC++ MG + F + N + ++ LV E
Sbjct: 159 EELHRFYRSILDKATKNESVEIGKEAMKLMNNTLCKLIMGRS-FSEDNGESNRVRGLVDE 217
Query: 122 FMHVGAKLSMGEVL-GPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQG- 179
+ K+ + +L PL K + + K++ + +FD++LE L+E +E + ++ +G
Sbjct: 218 TYALSEKIFLAAILRRPLAKLRISLFKKEIMGVSNKFDELLERILQERKENLEEKNNEGM 277
Query: 180 DMMDILLQVYRNENAEVRLTRNDIKAFFL 208
DMMD+LL+ Y +ENAE ++T IKAFF+
Sbjct: 278 DMMDVLLEAYGDENAEYKITWKHIKAFFV 306
>At1g50560 cytochrome P450, putative
Length = 519
Score = 133 bits (334), Expect = 8e-32
Identities = 77/212 (36%), Positives = 126/212 (59%), Gaps = 6/212 (2%)
Query: 1 MKTHDLNFSYRP--QFGSSHDYLYKGSY-FITAPYGPYWRFMKKICVTKLLSSSQLGRFM 57
++ DLNF+ R Q L GS+ F++ PYG YWRFMKK+ V KLL S L +
Sbjct: 101 LRIQDLNFASRDSGQTPIMEKSLLFGSFGFVSVPYGDYWRFMKKLLVKKLLGSHSLEQTR 160
Query: 58 HIREQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQC 117
+R +EL+ L + +N + D+G + LTNN +CRM+MG +C ++ N + E ++
Sbjct: 161 LLRGKELQTFRAMLFDKAAKNETVDVGKEMMKLTNNSICRMTMGRSCSEE-NGEAEQVRG 219
Query: 118 LVREFMHVGAKLSMGEVLGPLGKF-DLFGYGKKLRKIVGEFDKILEGFLKEHEERINTED 176
LV + + + K + ++G K + +GK++ ++ +D++LE +KEHEE N +
Sbjct: 220 LVTKSLSLTKKFLIASIVGQFSKLVGISLFGKEIMEVSQRYDELLEKIIKEHEENPNNGE 279
Query: 177 CQGDMMDILLQVYRNENAEVRLTRNDIKAFFL 208
DMMD+LL+V ++NAE +++RN IKA F+
Sbjct: 280 -DRDMMDVLLEVCADDNAEFKISRNQIKALFV 310
>At1g50550 cytochrome P450, putative
Length = 218
Score = 133 bits (334), Expect = 8e-32
Identities = 78/213 (36%), Positives = 127/213 (59%), Gaps = 6/213 (2%)
Query: 1 MKTHDLNFSYRP--QFGSSHDYLYKGSY-FITAPYGPYWRFMKKICVTKLLSSSQLGRFM 57
++ DLNF+ R Q L GS+ F++ PYG YWRFMKK+ V KLL S L +
Sbjct: 6 LRIQDLNFASRDSGQTPIMEKSLLFGSFGFVSVPYGDYWRFMKKLLVKKLLGSHSLEQTR 65
Query: 58 HIREQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQC 117
+R +EL+ L + +N + D+G + LTNN +CRM+MG +C ++ N + E ++
Sbjct: 66 LLRGKELQTFRAMLFDKATKNETVDVGKEMMKLTNNSICRMTMGRSCSEE-NDEAEQVRG 124
Query: 118 LVREFMHVGAKLSMGEVLGPLGKF-DLFGYGKKLRKIVGEFDKILEGFLKEHEERINTED 176
LV + + + K + ++G K + +GK++++ FD++LE +KEHE+ N +
Sbjct: 125 LVTKSLSLTKKFLIASIVGHFSKLVGISLFGKEIKEHSQRFDELLEKIIKEHEKDPNNGE 184
Query: 177 CQGDMMDILLQVYRNENAEVRLTRNDIKAFFLV 209
DMMD+LL+V ++NAE +++RN IKA F+V
Sbjct: 185 -DRDMMDVLLEVCADDNAEFKISRNQIKALFVV 216
>At3g20140 cytochrome P450, putative
Length = 510
Score = 132 bits (333), Expect = 1e-31
Identities = 80/206 (38%), Positives = 116/206 (55%), Gaps = 3/206 (1%)
Query: 4 HDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIREQE 63
HD+N S+R L F TAPYG YW+FMKK+ VTKLL L R IR
Sbjct: 103 HDVNISFRGNPPIEESLLVGSFGFFTAPYGDYWKFMKKVMVTKLLGPQALQRSRGIRADA 162
Query: 64 LEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVREFM 123
LE+ +LL + + S ++G + L + +C+M MG F + N + E ++ LV E
Sbjct: 163 LERFYMNLLDKAMKKESVEIGKETMKLIYDSICKMIMGRN-FSEENGEAERVRGLVTEST 221
Query: 124 HVGAKLSMGEVL-GPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQGDMM 182
+ K+ M VL PL K + + K++ + FD++LE FL EHEE++N ED DMM
Sbjct: 222 ALTKKIFMANVLHKPLKKLGISLFKKEIMDVSNSFDELLERFLVEHEEKLN-EDQDMDMM 280
Query: 183 DILLQVYRNENAEVRLTRNDIKAFFL 208
+LL R++NAE ++TRN IK+ F+
Sbjct: 281 GVLLAACRDKNAECKITRNHIKSLFV 306
>At3g20950 cytochrome P450, putative
Length = 526
Score = 131 bits (329), Expect = 3e-31
Identities = 78/208 (37%), Positives = 118/208 (56%), Gaps = 3/208 (1%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
K D+N S R + L+ S F APYG Y++FM+K+ TKLL L R IR
Sbjct: 105 KAQDVNVSSRGHAPAGESLLFGSSSFFFAPYGDYFKFMRKLIATKLLGPQALERSRKIRA 164
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVRE 121
EL++ ++LL + + S D+ + L NNI+C+M MG +C + N + E ++ LV E
Sbjct: 165 DELDRFYRNLLDKAMKKESVDIVEEAAKLNNNIICKMIMGRSC-SEDNGEAERVRGLVIE 223
Query: 122 FMHVGAKLSMGEVLG-PLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQGD 180
+ ++ +G + PL K + + K + K V FD++LE L EHEER+ D
Sbjct: 224 STALTKQIFLGMIFDKPLKKLGISLFQKDI-KSVSRFDELLEKILVEHEERMGKHYKAND 282
Query: 181 MMDILLQVYRNENAEVRLTRNDIKAFFL 208
MMD+LL+ Y +ENAE ++TRN IK+ F+
Sbjct: 283 MMDLLLEAYGDENAEYKITRNHIKSLFV 310
>At1g50520 cytochrome P450 like protein
Length = 533
Score = 131 bits (329), Expect = 3e-31
Identities = 79/210 (37%), Positives = 123/210 (57%), Gaps = 3/210 (1%)
Query: 1 MKTHDLNFSYRPQFGSSHDY-LYKGSY-FITAPYGPYWRFMKKICVTKLLSSSQLGRFMH 58
++T DLNF+ R + S + L GS+ F++APYG YWRFMKK+ VT L S L +
Sbjct: 101 LRTQDLNFATRQREVSIMEKSLLFGSFGFVSAPYGDYWRFMKKLLVTNLFGSHSLEQTRL 160
Query: 59 IREQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCL 118
IRE+EL+ L + + + D+G + LTNN +CRM MG C ++ + +V +
Sbjct: 161 IREKELKTFRTMLFDKAAKKGTVDVGKEMMKLTNNSICRMIMGRRCSEENSEAEKVEDLV 220
Query: 119 VREFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQ 178
++ F V L V L KF + + K++ ++ +D++LE +KEHEE N ++
Sbjct: 221 IKSFSLVKKVLIANTVGRLLKKFGISLFEKEIMEVSQRYDELLEKIIKEHEEDPNKKE-D 279
Query: 179 GDMMDILLQVYRNENAEVRLTRNDIKAFFL 208
DMMD+LL+V ++ AEV++TRN IKA +
Sbjct: 280 RDMMDVLLEVCADDKAEVKITRNQIKALIV 309
>At3g20110 cytochrome P450, putative
Length = 510
Score = 130 bits (328), Expect = 4e-31
Identities = 80/209 (38%), Positives = 122/209 (58%), Gaps = 5/209 (2%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
K HDLN S R + L S F+ APYG YW+FMKK+ VTKLL L R IR
Sbjct: 98 KAHDLNISSRDNPPINESLLVGSSVFVGAPYGDYWKFMKKLLVTKLLGPQALERSRSIRA 157
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVRE 121
ELE+ +SLL + + S ++G + T L+ N +CRMSMG + F + + + E ++ LV E
Sbjct: 158 DELERFYRSLLDKAMKKESVEIGKEATKLSINSICRMSMGRS-FSEESGEAERVRGLVTE 216
Query: 122 FMHVGAKLSMGEVL-GPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQGD 180
+ K+ + +L PL K + + K++ + FD++LE + E E++ N + QG
Sbjct: 217 LDGLTKKVLLVNILRWPLEKLRISLFKKEIMYVSNSFDELLERIIVEREKKPN--EHQGT 274
Query: 181 -MMDILLQVYRNENAEVRLTRNDIKAFFL 208
+MD+LL+ Y +E AE ++TRN IK+ F+
Sbjct: 275 YLMDVLLEAYEDEKAEHKITRNHIKSLFV 303
>At3g20940 cytochrome P450, putative
Length = 563
Score = 129 bits (325), Expect = 9e-31
Identities = 76/208 (36%), Positives = 116/208 (55%), Gaps = 2/208 (0%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
K D+N S R + S F APYG Y++FM+K+ TKLL L R IR
Sbjct: 101 KAQDVNVSSRGHAPVGESLWFGSSSFFFAPYGDYFKFMRKLIATKLLGPQALERSRKIRA 160
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVRE 121
EL++ K+LL + + S ++G + L NNI+C+M MG +C ++ N + E + LV E
Sbjct: 161 DELDRFYKTLLDKAMKKESVEIGEEAAKLNNNIICKMIMGRSCSEE-NGEAEKFRHLVIE 219
Query: 122 FMHVGAKLSMGEVL-GPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQGD 180
M + ++ G + PL K + + K + + +FD++LE L EHEE+ + D
Sbjct: 220 SMALTKQIFFGMIFHKPLKKLGISLFQKDILSLSRKFDELLEKILFEHEEKKAEHNQAND 279
Query: 181 MMDILLQVYRNENAEVRLTRNDIKAFFL 208
MMD LL+ Y +ENAE ++TRN IK+ F+
Sbjct: 280 MMDFLLEAYGDENAEYKITRNHIKSLFV 307
Score = 28.9 bits (63), Expect = 2.1
Identities = 13/40 (32%), Positives = 20/40 (49%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKK 41
+ HD+N S R ++ S + APYG +F+KK
Sbjct: 523 RAHDMNVSSRGAAAIDESLVFGSSGVVYAPYGDNLKFVKK 562
>At5g06900 cytochrome P450
Length = 507
Score = 128 bits (322), Expect = 2e-30
Identities = 79/216 (36%), Positives = 124/216 (56%), Gaps = 15/216 (6%)
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSY-FITAPYGPYWRFMKKICVTKLLSSSQLGRFMHI 59
+K+++LNF RP + DYL GS F +APYG +W+FMK+IC+ +L SS L F+ +
Sbjct: 91 LKSNELNFLNRPTM-QNVDYLTYGSADFFSAPYGLHWKFMKRICMVELFSSRALDSFVSV 149
Query: 60 REQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNID-------P 112
R +EL+KLL +L + S +LG LT+NI+ RM F+K D
Sbjct: 150 RSEELKKLLIRVLKKAEAEESVNLGEQLKELTSNIITRM-----MFRKMQSDSDGGEKSE 204
Query: 113 EVIQCLVREFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERI 172
EVI+ +V E + ++ E L + DL G K+L+ ++D I+E ++EHE
Sbjct: 205 EVIKMVV-ELNELAGFFNVSETFWFLKRLDLQGLKKRLKNARDKYDVIIERIMEEHESSK 263
Query: 173 NTEDCQGDMMDILLQVYRNENAEVRLTRNDIKAFFL 208
+ +M+D+LL +Y ++NAE++LTR +IKAF +
Sbjct: 264 KNATGERNMLDVLLDIYEDKNAEMKLTRENIKAFIM 299
>At2g27000 putative cytochrome P450
Length = 514
Score = 127 bits (319), Expect = 4e-30
Identities = 79/209 (37%), Positives = 122/209 (57%), Gaps = 4/209 (1%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSY-FITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+ D+N S R F ++ L+ GS+ FITAPYG YW+FMKK+ VTKLL L R IR
Sbjct: 98 RAQDVNVSTR-DFPTNEGSLFLGSFSFITAPYGEYWKFMKKLIVTKLLGPQALERSQRIR 156
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
E+E+ +LL + + S ++ + L NNI+C+M MG TC ++ N + E I+ LV
Sbjct: 157 ANEVERFYSNLLDKAMKKESVEIADEAMKLVNNIICKMIMGRTCSEE-NGEAERIRGLVT 215
Query: 121 EFMHVGAKLSMGEVL-GPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQG 179
+ + K + +L PL K + + K I +FD++LE L E+EER+
Sbjct: 216 KSDALLKKFLLAAILRKPLKKIGITLFKKVFMDISLKFDEVLEKILVENEERLEENQQGT 275
Query: 180 DMMDILLQVYRNENAEVRLTRNDIKAFFL 208
D+MD LL+VY ++ +E ++TR+ IK+ F+
Sbjct: 276 DIMDKLLEVYGDKTSEYKITRDHIKSLFV 304
>At3g20960 cytochrome P450 like protein
Length = 418
Score = 127 bits (318), Expect = 6e-30
Identities = 75/204 (36%), Positives = 114/204 (55%), Gaps = 2/204 (0%)
Query: 6 LNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIREQELE 65
+N S R ++ S + APYG Y +F+KKI TKLL L R +R +EL+
Sbjct: 1 MNISSRGAAAIDESLVFGSSGVVYAPYGDYLKFVKKIIATKLLRPQVLERSRGLRAEELQ 60
Query: 66 KLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVREFMHV 125
+L +L + N + ++G + T L NNI CRMSMG + F + N + E ++ LV E +
Sbjct: 61 RLYNRILDKAKMNENVEIGKEATMLMNNIFCRMSMGRS-FSEENGEAERVRGLVAESYAL 119
Query: 126 GAKLSMGEVLGP-LGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQGDMMDI 184
K+ VL L K + + K + + F+++LE L EH E+++ E DMMD+
Sbjct: 120 AKKIFFASVLRRLLKKLRIPLFKKDIMDVSNRFNELLEKILVEHNEKLDEEHKDTDMMDV 179
Query: 185 LLQVYRNENAEVRLTRNDIKAFFL 208
LL Y +ENAE ++TRN IK+FF+
Sbjct: 180 LLAAYADENAEYKITRNHIKSFFV 203
>At3g20130 cytochrome P450, putative
Length = 515
Score = 126 bits (317), Expect = 7e-30
Identities = 76/209 (36%), Positives = 120/209 (57%), Gaps = 5/209 (2%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
+T D+N S R ++ S F+TAPYG YW+FMKK+ V KLL + IR
Sbjct: 101 RTQDVNISSRGVTAVDESLVFGSSSFVTAPYGDYWKFMKKLTVMKLLGPQAQEQSRDIRA 160
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVRE 121
++++ ++LL + + S ++G + L NNILC+MSMG + F + N + E ++ LV E
Sbjct: 161 DDIKRFCRNLLDKARKKESVEIGKEAMNLMNNILCKMSMGRS-FSEENGETEKLRGLVTE 219
Query: 122 FMHVGAKLSMGEVL-GPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQGD 180
+ + K+ + +L L K + + K + + +FD +LE L EH E+ E QG
Sbjct: 220 SIGLMKKMFLAVLLRRQLQKLGISLFKKDIMGVSNKFDVLLEKVLVEHREK--PEKDQGT 277
Query: 181 -MMDILLQVYRNENAEVRLTRNDIKAFFL 208
M+D+LL Y +ENAE ++T+N IKAFF+
Sbjct: 278 VMLDVLLAAYGDENAEYKITKNHIKAFFV 306
>At3g20080 cytochrome P450, putative
Length = 523
Score = 122 bits (305), Expect = 2e-28
Identities = 71/208 (34%), Positives = 120/208 (57%), Gaps = 4/208 (1%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSY-FITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+ HD+N S R F + D L+ GS+ F +APYG YW+FMKK+ VT LL L R R
Sbjct: 101 REHDVNISSRG-FPPTDDSLFAGSFSFTSAPYGDYWKFMKKLLVTNLLGPQALERSRGFR 159
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
EL+ ++LL + + S D+ ++ L+NN +C+M MG +C ++ N + E ++ L
Sbjct: 160 ADELDLFYENLLDKAMKKESVDICVEALKLSNNSICKMIMGRSCSEE-NGEAERVRALAT 218
Query: 121 EFMHVGAKLSMGEVL-GPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQG 179
+ + K+ + +L K + + K++ + FD++LE L EHE++++
Sbjct: 219 QLDGLTKKILLANMLRAGFKKLVVSLFRKEMMDVSSRFDELLERILVEHEDKLDMHHQGT 278
Query: 180 DMMDILLQVYRNENAEVRLTRNDIKAFF 207
D++D LL R++NAE +++RN IK+FF
Sbjct: 279 DLVDALLAACRDKNAEYKISRNHIKSFF 306
>At5g42580 cytochrome P450
Length = 499
Score = 121 bits (303), Expect = 3e-28
Identities = 75/207 (36%), Positives = 109/207 (52%), Gaps = 13/207 (6%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
+T+D+N SYR + ++ S F+TAPYG YW+FMKK+ TKLL L R
Sbjct: 98 RTNDVNVSYRFVPVNKDSLVFGSSGFVTAPYGDYWKFMKKLISTKLLRPHALELSKGNRA 157
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVRE 121
+EL + L + + S ++G LTNNI+CRMSMG +C +K + RE
Sbjct: 158 EELRRFCLDLQGKARKKESVEIGKVALKLTNNIICRMSMGRSCSEKNGVAER-----ARE 212
Query: 122 FMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQGDM 181
++ LS+ + + D+ G + EFD+ LE L EHEE + D DM
Sbjct: 213 LVNKSFALSVKLFFSNMFRKDIMGVSR-------EFDEFLERILVEHEENLE-GDQDRDM 264
Query: 182 MDILLQVYRNENAEVRLTRNDIKAFFL 208
+D LL+ YRNE AE ++TR IK+ +
Sbjct: 265 IDHLLEAYRNEEAEYKITRKQIKSLIV 291
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.325 0.142 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,783,746
Number of Sequences: 26719
Number of extensions: 199121
Number of successful extensions: 876
Number of sequences better than 10.0: 148
Number of HSP's better than 10.0 without gapping: 131
Number of HSP's successfully gapped in prelim test: 17
Number of HSP's that attempted gapping in prelim test: 655
Number of HSP's gapped (non-prelim): 152
length of query: 212
length of database: 11,318,596
effective HSP length: 95
effective length of query: 117
effective length of database: 8,780,291
effective search space: 1027294047
effective search space used: 1027294047
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC142143.3


