

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142143.13 - phase: 0
(259 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
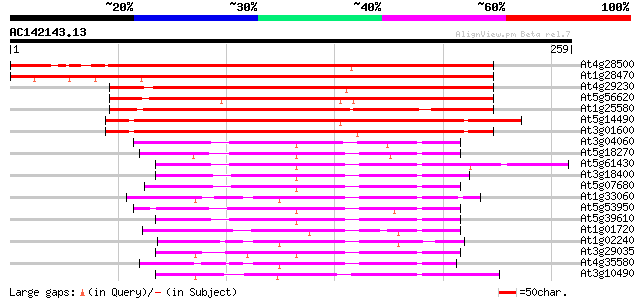
Score E
Sequences producing significant alignments: (bits) Value
At4g28500 predicted protein 323 4e-89
At1g28470 unknown protein 320 6e-88
At4g29230 putative protein 276 6e-75
At5g56620 putative protein 250 6e-67
At1g25580 unknown protein 215 2e-56
At5g14490 putative protein 176 1e-44
At3g01600 hypothetical protein 171 4e-43
At3g04060 NAM-like protein (no apical meristem) 80 8e-16
At5g18270 NAM (no apical meristem)-like protein 71 7e-13
At5g61430 NAM, no apical meristem, - like protein 70 9e-13
At3g18400 organ separation protein, putative 70 1e-12
At5g07680 NAM (no apical meristem)-like protein 69 2e-12
At1g33060 NAC domain containing protein 69 3e-12
At5g53950 CUC2 (dbj|BAA19529.1) 66 2e-11
At5g39610 NAM / CUC2 - like protein 65 3e-11
At1g01720 unknown protein with NAC domain protein 65 4e-11
At1g02240 hypothetical protein 64 1e-10
At3g29035 NAM-like protein (No Apical Meristem) 63 1e-10
At4g35580 NAM / CUC2 -like protein 62 3e-10
At3g10490 unknown protein 62 4e-10
>At4g28500 predicted protein
Length = 305
Score = 323 bits (829), Expect = 4e-89
Identities = 157/224 (70%), Positives = 179/224 (79%), Gaps = 10/224 (4%)
Query: 1 MTWCNDSDDQDRGIELITSTVIPGTSNESPLVTDDETKPTEVRTLTCPSCGHNIEFQDQG 60
MTWCND D +I S PG + ESP+ + P TCPSCGHN +F +Q
Sbjct: 1 MTWCNDRSDVQTVERIIPS---PGAA-ESPVAS----LPVSCHK-TCPSCGHNFKFHEQA 51
Query: 61 GINELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEK 120
GI++LPGLPAGVKFDP DQE+L+HLE KV D KLHPLIDEFI T++GENGICYTHPEK
Sbjct: 52 GIHDLPGLPAGVKFDPTDQEVLEHLEGKVRDDAKKLHPLIDEFIRTIDGENGICYTHPEK 111
Query: 121 LPGVKKDGQIRHFFHRPSKAYTTGTRKRRKVQTDEE-GSETRWHKTGKTRPVIIDGVMKG 179
LPGV KDG +RHFFHRPSKAYTTGTRKRRKV TD + G ETRWHKTGKTRPV+ G ++G
Sbjct: 112 LPGVNKDGTVRHFFHRPSKAYTTGTRKRRKVHTDSDVGGETRWHKTGKTRPVLAGGRVRG 171
Query: 180 FKKILVLYTNYGRQKKPEKTNWVMHQYHLGSNEEEKDGELVVSK 223
+KKILVLYTNYG+QKKPEKTNWVMHQYHLG++EEEK+GELVVSK
Sbjct: 172 YKKILVLYTNYGKQKKPEKTNWVMHQYHLGTSEEEKEGELVVSK 215
>At1g28470 unknown protein
Length = 314
Score = 320 bits (819), Expect = 6e-88
Identities = 158/232 (68%), Positives = 179/232 (77%), Gaps = 9/232 (3%)
Query: 1 MTWCNDSDDQ-DRGIELITSTVIPGTS----NESPLVTDDETK-PTEVRTLTCPSCGHNI 54
M+WC+ SDD D +E +++T P ++S + + ++K E TCPSCGH +
Sbjct: 1 MSWCDGSDDNYDLNLERVSNTDHPSVQLKDQSQSCVTSRPDSKISAETPITTCPSCGHKL 60
Query: 55 EFQDQ---GGINELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGEN 111
G I +LP LPAGVKFDP+D+EIL HLEAKV D KLHPLIDEFIPTLEGEN
Sbjct: 61 HHHQDDQVGSIKDLPSLPAGVKFDPSDKEILMHLEAKVSSDKRKLHPLIDEFIPTLEGEN 120
Query: 112 GICYTHPEKLPGVKKDGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPV 171
GICYTHPEKLPGV KDGQ+RHFFHRPSKAYTTGTRKRRKV TDEEG ETRWHKTGKTRPV
Sbjct: 121 GICYTHPEKLPGVSKDGQVRHFFHRPSKAYTTGTRKRRKVSTDEEGHETRWHKTGKTRPV 180
Query: 172 IIDGVMKGFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSNEEEKDGELVVSK 223
+ GFKKILVLYTNYGRQKKPEKTNWVMHQYHLGS+E+EKDGE V+SK
Sbjct: 181 LSQSGETGFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSSEDEKDGEPVLSK 232
>At4g29230 putative protein
Length = 498
Score = 276 bits (707), Expect = 6e-75
Identities = 131/185 (70%), Positives = 151/185 (80%), Gaps = 12/185 (6%)
Query: 47 CPSCGHNIEFQDQGGINELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPT 106
CPSCGH +E + Q + GLPAGVKFDP DQE+++HLEAKVL K HPLIDEFIPT
Sbjct: 31 CPSCGHKLEGKPQDWV----GLPAGVKFDPTDQELIEHLEAKVLAKDFKSHPLIDEFIPT 86
Query: 107 LEGENGICYTHPEKLPGVKKDGQIRHFFHRPSKAYTTGTRKRRKVQTD--------EEGS 158
+EGE+GICYTHPEKLPGV +DG RHFFHRPSKAYTTGTRKRRK+QT+
Sbjct: 87 IEGEDGICYTHPEKLPGVTRDGLSRHFFHRPSKAYTTGTRKRRKIQTECDNNLQGSSSSG 146
Query: 159 ETRWHKTGKTRPVIIDGVMKGFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSNEEEKDGE 218
ETRWHKTGKTRPV+++G KG KKILVLYTN+G+ +KPEKTNWVMHQYHLG++EEEK+GE
Sbjct: 147 ETRWHKTGKTRPVMVNGKQKGCKKILVLYTNFGKNRKPEKTNWVMHQYHLGTHEEEKEGE 206
Query: 219 LVVSK 223
LVVSK
Sbjct: 207 LVVSK 211
>At5g56620 putative protein
Length = 386
Score = 250 bits (638), Expect = 6e-67
Identities = 122/188 (64%), Positives = 146/188 (76%), Gaps = 14/188 (7%)
Query: 47 CPSCGHNIEFQDQGGINELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKL-----HPLID 101
CP CG I+ + GLPAGVKFDP DQE+++HLEAKV HPLID
Sbjct: 25 CPGCGRMIQAATKPN---WVGLPAGVKFDPTDQELIEHLEAKVKGKEENKKWSSSHPLID 81
Query: 102 EFIPTLEGENGICYTHPEKLPGVKKDGQIRHFFHRPSKAYTTGTRKRRKV-QTDEEG--- 157
EFIPT++GE+GICYTHP+KLPGV +DG +HFFH+PS+AYTTGTRKRRK+ QTD +
Sbjct: 82 EFIPTIDGEDGICYTHPQKLPGVTRDGLSKHFFHKPSRAYTTGTRKRRKIIQTDHDSELT 141
Query: 158 --SETRWHKTGKTRPVIIDGVMKGFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSNEEEK 215
SETRWHKTGKTRPV+I+G +G KKILVLYTN+G+ ++PEKTNWVMHQYHLG NEEE+
Sbjct: 142 GSSETRWHKTGKTRPVMINGQQRGCKKILVLYTNFGKNRRPEKTNWVMHQYHLGINEEER 201
Query: 216 DGELVVSK 223
+GELVVSK
Sbjct: 202 EGELVVSK 209
>At1g25580 unknown protein
Length = 449
Score = 215 bits (547), Expect = 2e-56
Identities = 103/177 (58%), Positives = 128/177 (72%), Gaps = 8/177 (4%)
Query: 47 CPSCGHNIEFQDQGGINELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPT 106
CP C H I+ D +++ PGLP GVKFDP+D EI+ HL AK HP IDEFIPT
Sbjct: 39 CPKCQHVIDNSDV--VDDWPGLPRGVKFDPSDPEIIWHLLAKSGLSGLSSHPFIDEFIPT 96
Query: 107 LEGENGICYTHPEKLPGVKKDGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTG 166
+ ++GICYTHP+ LPGVK DG + HFFH+ KAY+TGTRKRRK+ D+ G + RWHKTG
Sbjct: 97 VNQDDGICYTHPKNLPGVKSDGTVSHFFHKAIKAYSTGTRKRRKIHDDDFG-DVRWHKTG 155
Query: 167 KTRPVIIDGVMKGFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSNEEEKDGELVVSK 223
+T+PV++DGV +G KKI+VLY K KTNWVMHQYHLG E+EK+G+ VVSK
Sbjct: 156 RTKPVVLDGVQRGCKKIMVLYGG-----KAVKTNWVMHQYHLGIEEDEKEGDYVVSK 207
>At5g14490 putative protein
Length = 350
Score = 176 bits (446), Expect = 1e-44
Identities = 89/193 (46%), Positives = 127/193 (65%), Gaps = 4/193 (2%)
Query: 45 LTCPSCGHNIEFQDQGGINELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFI 104
+ C +C + I+ + + PGLP GVKF+P D+E+++HLEAK D K H LI +FI
Sbjct: 37 INCFNCSYRID--NSNVLTPWPGLPKGVKFEPTDEEVIEHLEAKCGIDGLKPHLLIQDFI 94
Query: 105 PTLEGENGICYTHPEKLPGVKKDGQIRHFFHRPSKAYTTGTRKRRKV-QTDEEGSETRWH 163
++ + GI YTHP+ LPGV KDG FF++ + AY G RKRR++ T + RWH
Sbjct: 95 CSVTQDVGINYTHPQNLPGVSKDGTSVFFFNKTAHAYQNGQRKRRRITPTSLKDDTVRWH 154
Query: 164 KTGKTRPVIIDGVMKGFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSNEEEKDGELVVSK 223
KTG+T+PV+++G+ KG KKI+VLY + + KPEK+NWV+HQYHLG+ EE + GE VVSK
Sbjct: 155 KTGQTKPVMLNGIQKGCKKIMVLYKSARKGFKPEKSNWVLHQYHLGT-EEGEIGEYVVSK 213
Query: 224 KILEYLSSSERRL 236
+ E+ +
Sbjct: 214 ITYQQPKQQEKTI 226
>At3g01600 hypothetical protein
Length = 370
Score = 171 bits (433), Expect = 4e-43
Identities = 87/180 (48%), Positives = 120/180 (66%), Gaps = 4/180 (2%)
Query: 45 LTCPSCGHNIEFQDQGGINELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFI 104
+ CP+C + I+ + + PGLP GVKF+P D++I++ LEAK + H LI+EFI
Sbjct: 34 IKCPNCTYRID--NSNVLIPWPGLPKGVKFEPTDEDIIEFLEAKCGIGGSEPHVLIEEFI 91
Query: 105 PTLEGENGICYTHPEKLPGVKKDGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSE-TRWH 163
+ + GI YTHP+ LPG KDG FFH+ +AY TG RKRRK+ E RWH
Sbjct: 92 RPVTEDVGINYTHPQNLPGANKDGVSVFFFHKTVQAYGTGQRKRRKITPTLVNDEPVRWH 151
Query: 164 KTGKTRPVIIDGVMKGFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSNEEEKDGELVVSK 223
KTG+T+PV++ GV +G KKI+VLY + + KPEK+NWV+HQYHLG+ E ++ G+ VVSK
Sbjct: 152 KTGRTKPVMLSGVQRGCKKIMVLYKSARKGTKPEKSNWVLHQYHLGT-EGKEIGDYVVSK 210
>At3g04060 NAM-like protein (no apical meristem)
Length = 338
Score = 80.5 bits (197), Expect = 8e-16
Identities = 53/153 (34%), Positives = 74/153 (47%), Gaps = 18/153 (11%)
Query: 58 DQGGINELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTH 117
+QGG E+ LP G +F P D+EI+ H + +F++ F G+ +
Sbjct: 10 NQGGDQEVVDLPPGFRFHPTDEEIITHYLKEKVFNI--------RFTAAAIGQADLNKNE 61
Query: 118 PEKLPGVKKDGQIR-HFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVII-DG 175
P LP + K G+ +FF + + Y TG R R + W TGK + + G
Sbjct: 62 PWDLPKIAKMGEKEFYFFCQRDRKYPTGMRTNRATVSG------YWKATGKDKEIFRGKG 115
Query: 176 VMKGFKKILVLYTNYGRQKKPEKTNWVMHQYHL 208
+ G KK LV YT GR K EKTNWVMH+Y L
Sbjct: 116 CLVGMKKTLVFYT--GRAPKGEKTNWVMHEYRL 146
>At5g18270 NAM (no apical meristem)-like protein
Length = 335
Score = 70.9 bits (172), Expect = 7e-13
Identities = 54/151 (35%), Positives = 70/151 (45%), Gaps = 20/151 (13%)
Query: 61 GINELPGLPAGVKFDPNDQEILQ-HLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPE 119
G EL LP G +F P D+EI+ +L+ KVL F GE + P
Sbjct: 14 GGEELVDLPPGFRFHPTDEEIITCYLKEKVLNS---------RFTAVAMGEADLNKCEPW 64
Query: 120 KLPGVKKDGQIR-HFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIID-GVM 177
LP K G+ +FF + + Y TG R R ++ W TGK + + G +
Sbjct: 65 DLPKRAKMGEKEFYFFCQRDRKYPTGMRTNRATESGY------WKATGKDKEIFKGKGCL 118
Query: 178 KGFKKILVLYTNYGRQKKPEKTNWVMHQYHL 208
G KK LV Y GR K EKTNWVMH+Y L
Sbjct: 119 VGMKKTLVFYR--GRAPKGEKTNWVMHEYRL 147
>At5g61430 NAM, no apical meristem, - like protein
Length = 336
Score = 70.5 bits (171), Expect = 9e-13
Identities = 57/198 (28%), Positives = 87/198 (43%), Gaps = 25/198 (12%)
Query: 68 LPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVKKD 127
LP G +F P D+E++ H K + D F GE + + P +LP + K
Sbjct: 16 LPPGFRFHPTDEELITHYLHKKVLDT--------SFSAKAIGEVDLNKSEPWELPWMAKM 67
Query: 128 GQIR-HFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMKGFKKILVL 186
G+ +FF + Y TG R R + W TGK + + + G KK LV
Sbjct: 68 GEKEWYFFCVRDRKYPTGLRTNRATEAGY------WKATGKDKEIYRGKSLVGMKKTLVF 121
Query: 187 YTNYGRQKKPEKTNWVMHQYHLGSN------EEEKDGELVVSKKILEYLSSSERRLACNS 240
Y GR K +KTNWVMH+Y L + E V+ + + S+ +++ +S
Sbjct: 122 YR--GRAPKGQKTNWVMHEYRLEGKFSAHNLPKTAKNEWVICRVFQK--SAGGKKIPISS 177
Query: 241 VIRFMYSSVNFFLSLMAS 258
+IR +F SL+ S
Sbjct: 178 LIRIGSLGTDFNPSLLPS 195
>At3g18400 organ separation protein, putative
Length = 314
Score = 70.1 bits (170), Expect = 1e-12
Identities = 46/146 (31%), Positives = 66/146 (44%), Gaps = 17/146 (11%)
Query: 68 LPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVKKD 127
LP G +F P D+E++ H + + D+ F + + P LP
Sbjct: 5 LPPGFRFHPTDEELITHYLCRKVSDIG--------FTGKAVVDVDLNKCEPWDLPAKASM 56
Query: 128 GQIR-HFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMKGFKKILVL 186
G+ +FF + + Y TG R R + W TGK + + GV+ G KK LV
Sbjct: 57 GEKEWYFFSQRDRKYPTGLRTNRATEAGY------WKTTGKDKEIYRSGVLVGMKKTLVF 110
Query: 187 YTNYGRQKKPEKTNWVMHQYHLGSNE 212
Y GR K EK+NWVMH+Y L S +
Sbjct: 111 YK--GRAPKGEKSNWVMHEYRLESKQ 134
>At5g07680 NAM (no apical meristem)-like protein
Length = 329
Score = 69.3 bits (168), Expect = 2e-12
Identities = 49/147 (33%), Positives = 66/147 (44%), Gaps = 17/147 (11%)
Query: 63 NELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLP 122
+E LP G +F P D+E++ H K + D+ F GE + P +LP
Sbjct: 12 DEQMDLPPGFRFHPTDEELITHYLHKKVLDLG--------FSAKAIGEVDLNKAEPWELP 63
Query: 123 GVKKDGQIR-HFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMKGFK 181
K G+ +FF + Y TG R R Q W TGK + + + G K
Sbjct: 64 YKAKIGEKEWYFFCVRDRKYPTGLRTNRATQAGY------WKATGKDKEIFRGKSLVGMK 117
Query: 182 KILVLYTNYGRQKKPEKTNWVMHQYHL 208
K LV Y GR K +KTNWVMH+Y L
Sbjct: 118 KTLVFYR--GRAPKGQKTNWVMHEYRL 142
>At1g33060 NAC domain containing protein
Length = 648
Score = 68.6 bits (166), Expect = 3e-12
Identities = 51/167 (30%), Positives = 75/167 (44%), Gaps = 23/167 (13%)
Query: 55 EFQDQGGINELPGLPAGVKFDPNDQEILQH-LEAKVLFDVPKLHPLIDEFIPTLEGENGI 113
E + + + LP G +F P D+E++ H L K+ L IP ++ +
Sbjct: 11 EMTTEQALLSMEALPLGFRFRPTDEELINHYLRLKI-----NGRDLEVRVIPEID----V 61
Query: 114 CYTHPEKLPG---VKKDGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRP 170
C P LPG +K D Q FF + Y +G R R W TGK R
Sbjct: 62 CKWEPWDLPGLSVIKTDDQEWFFFCPRDRKYPSGHRSNRATDIGY------WKATGKDRT 115
Query: 171 VIIDGVMKGFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSNEEEKDG 217
+ ++ G KK LV Y GR + E+TNW+MH+Y + ++E DG
Sbjct: 116 IKSKKMIIGMKKTLVFYR--GRAPRGERTNWIMHEYR--ATDKELDG 158
>At5g53950 CUC2 (dbj|BAA19529.1)
Length = 375
Score = 65.9 bits (159), Expect = 2e-11
Identities = 50/154 (32%), Positives = 68/154 (43%), Gaps = 21/154 (13%)
Query: 58 DQGGINELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTH 117
D GG ++ LP G +F P D+E++ H + + D F E +
Sbjct: 9 DHGGDSQY--LPPGFRFHPTDEELITHYLLRKVLD--------GCFSSRAIAEVDLNKCE 58
Query: 118 PEKLPGVKKDGQIR-HFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGV 176
P +LPG K G+ +FF + Y TG R R + W TGK R +
Sbjct: 59 PWQLPGRAKMGEKEWYFFSLRDRKYPTGLRTNRATEAGY------WKATGKDREIFSSKT 112
Query: 177 --MKGFKKILVLYTNYGRQKKPEKTNWVMHQYHL 208
+ G KK LV Y GR K EK+NWVMH+Y L
Sbjct: 113 CALVGMKKTLVFYK--GRAPKGEKSNWVMHEYRL 144
>At5g39610 NAM / CUC2 - like protein
Length = 285
Score = 65.5 bits (158), Expect = 3e-11
Identities = 47/142 (33%), Positives = 62/142 (43%), Gaps = 17/142 (11%)
Query: 68 LPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVKKD 127
LP G +F P D+E++ H +F+ F T GE + P LP K
Sbjct: 20 LPPGFRFHPTDEELITHYLKPKVFNT--------FFSATAIGEVDLNKIEPWDLPWKAKM 71
Query: 128 GQIR-HFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMKGFKKILVL 186
G+ +FF + Y TG R R + W TGK + + + G KK LV
Sbjct: 72 GEKEWYFFCVRDRKYPTGLRTNRATEAGY------WKATGKDKEIFKGKSLVGMKKTLVF 125
Query: 187 YTNYGRQKKPEKTNWVMHQYHL 208
Y GR K KTNWVMH+Y L
Sbjct: 126 YK--GRAPKGVKTNWVMHEYRL 145
>At1g01720 unknown protein with NAC domain protein
Length = 289
Score = 65.1 bits (157), Expect = 4e-11
Identities = 47/150 (31%), Positives = 69/150 (45%), Gaps = 22/150 (14%)
Query: 62 INELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKL 121
++EL LP G +F P D+E++ H + P+I E + P +L
Sbjct: 1 MSELLQLPPGFRFHPTDEELVMHYLCRKCASQSIAVPIIAEI--------DLYKYDPWEL 52
Query: 122 PGVKKDGQIRHFFHRP-SKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMK-- 178
PG+ G+ +F P + Y G+R R + W TG +P+ G+ K
Sbjct: 53 PGLALYGEKEWYFFSPRDRKYPNGSRPNRSAGSGY------WKATGADKPI---GLPKPV 103
Query: 179 GFKKILVLYTNYGRQKKPEKTNWVMHQYHL 208
G KK LV Y G+ K EKTNW+MH+Y L
Sbjct: 104 GIKKALVFYA--GKAPKGEKTNWIMHEYRL 131
>At1g02240 hypothetical protein
Length = 359
Score = 63.5 bits (153), Expect = 1e-10
Identities = 48/148 (32%), Positives = 67/148 (44%), Gaps = 23/148 (15%)
Query: 69 PAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPG---VK 125
P G +F PND+EI+ H D H +DE I T++ IC P LP +K
Sbjct: 4 PVGFRFRPNDEEIVDHYLRPKNLDSDTSH--VDEVISTVD----ICSFEPWDLPSKSMIK 57
Query: 126 KDGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMK---GFKK 182
+ +FF Y G ++RR+ + W KTGKT V+ + G K+
Sbjct: 58 SRDGVWYFFSVKEMKYNRGDQQRRRTNSGF------WKKTGKTMTVMRKRGNREKIGEKR 111
Query: 183 ILVLYTNYGRQKKPEKTNWVMHQYHLGS 210
+LV G KT+WVMH+YH S
Sbjct: 112 VLVFKNRDG-----SKTDWVMHEYHATS 134
>At3g29035 NAM-like protein (No Apical Meristem)
Length = 318
Score = 63.2 bits (152), Expect = 1e-10
Identities = 48/144 (33%), Positives = 64/144 (44%), Gaps = 21/144 (14%)
Query: 68 LPAGVKFDPNDQEILQH-LEAKVLFDVPKLHPLIDEFIPTLE-GENGICYTHPEKLPGVK 125
LP G +F P D+E++ H L KV ++ F + GE + P LP
Sbjct: 24 LPPGFRFHPTDEELITHYLRPKV----------VNSFFSAIAIGEVDLNKVEPWDLPWKA 73
Query: 126 KDGQIR-HFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMKGFKKIL 184
K G+ +FF + Y TG R R + W TGK + + + G KK L
Sbjct: 74 KLGEKEWYFFCVRDRKYPTGLRTNRATKAGY------WKATGKDKEIFKGKSLVGMKKTL 127
Query: 185 VLYTNYGRQKKPEKTNWVMHQYHL 208
V Y GR K KTNWVMH+Y L
Sbjct: 128 VFYK--GRAPKGVKTNWVMHEYRL 149
>At4g35580 NAM / CUC2 -like protein
Length = 512
Score = 62.0 bits (149), Expect = 3e-10
Identities = 45/149 (30%), Positives = 64/149 (42%), Gaps = 19/149 (12%)
Query: 61 GINELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEK 120
G + LP G +F P D+E++ H + + H + IP ++ +C P
Sbjct: 2 GAVSMESLPLGFRFRPTDEELVNHY---LRLKINGRHSDV-RVIPDID----VCKWEPWD 53
Query: 121 LPG---VKKDGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVM 177
LP +K D FF + Y G R R + W TGK R + +
Sbjct: 54 LPALSVIKTDDPEWFFFCPRDRKYPNGHRSNRATDSGY------WKATGKDRSIKSKKTL 107
Query: 178 KGFKKILVLYTNYGRQKKPEKTNWVMHQY 206
G KK LV Y GR K E+TNW+MH+Y
Sbjct: 108 IGMKKTLVFYR--GRAPKGERTNWIMHEY 134
>At3g10490 unknown protein
Length = 451
Score = 61.6 bits (148), Expect = 4e-10
Identities = 51/163 (31%), Positives = 72/163 (43%), Gaps = 21/163 (12%)
Query: 68 LPAGVKFDPNDQEILQH-LEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLP---G 123
L G +F P D+E++ + L+ KVL + + GE I P L
Sbjct: 27 LAPGFRFHPTDEELVSYYLKRKVLGQPVRFDAI---------GEVDIYKHEPWDLAVFSR 77
Query: 124 VKKDGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMKGFKKI 183
+K Q +F+ K Y G R R + W TGK R + D ++ G KK
Sbjct: 78 LKTRDQEWYFYSALDKKYGNGARMNRAT------NRGYWKATGKDREIRRDILLLGMKKT 131
Query: 184 LVLYTNYGRQKKPEKTNWVMHQYHLGSNEEEKDGELVVSKKIL 226
LV ++ GR +TNWVMH+Y L E EK+G LV +L
Sbjct: 132 LVFHS--GRAPDGLRTNWVMHEYRLVEYETEKNGNLVQDAYVL 172
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,542,572
Number of Sequences: 26719
Number of extensions: 308696
Number of successful extensions: 1028
Number of sequences better than 10.0: 109
Number of HSP's better than 10.0 without gapping: 59
Number of HSP's successfully gapped in prelim test: 50
Number of HSP's that attempted gapping in prelim test: 847
Number of HSP's gapped (non-prelim): 112
length of query: 259
length of database: 11,318,596
effective HSP length: 97
effective length of query: 162
effective length of database: 8,726,853
effective search space: 1413750186
effective search space used: 1413750186
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC142143.13


