

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141865.9 - phase: 0 /pseudo
(547 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
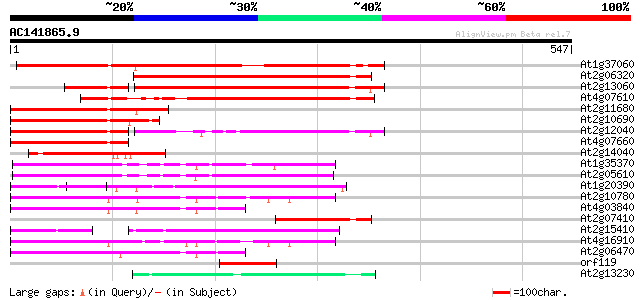
Score E
Sequences producing significant alignments: (bits) Value
At1g37060 Athila retroelment ORF 1, putative 381 e-106
At2g06320 putative retroelement pol polyprotein 279 2e-75
At2g13060 pseudogene 266 3e-71
At4g07610 247 1e-65
At2g11680 putative retroelement pol polyprotein 188 6e-48
At2g10690 putative retroelement pol polyprotein 151 1e-36
At2g12040 T10J7.2 150 1e-36
At4g07660 putative athila transposon protein 141 1e-33
At2g14040 putative retroelement pol polyprotein 115 8e-26
At1g35370 hypothetical protein 110 3e-24
At2g05610 putative retroelement pol polyprotein 107 1e-23
At1g20390 hypothetical protein 106 3e-23
At2g10780 pseudogene 95 1e-19
At4g03840 putative transposon protein 93 3e-19
At2g07410 putative retroelement pol polyprotein 91 1e-18
At2g15410 putative retroelement pol polyprotein 91 2e-18
At4g16910 retrotransposon like protein 88 1e-17
At2g06470 putative retroelement pol polyprotein 86 4e-17
orf119 -mitochondrial genome- 85 9e-17
At2g13230 pseudogene 77 3e-14
>At1g37060 Athila retroelment ORF 1, putative
Length = 1734
Score = 381 bits (979), Expect = e-106
Identities = 197/362 (54%), Positives = 246/362 (67%), Gaps = 34/362 (9%)
Query: 7 TLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAKPRLIQL 66
T+D A V YATTEKELLAVV+A +KFR YLVGSK+T+YTDH+ ++++ KKD KPRL++
Sbjct: 1218 TMDDAQVRYATTEKELLAVVFAFEKFRSYLVGSKVTIYTDHAALRHIYAKKDTKPRLLRW 1277
Query: 67 IFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLAQTNA---PW 123
I LLQEFD+EI DKKG+EN VADHLSR+R EDE+ +DDS P++QL + Q N PW
Sbjct: 1278 ILLLQEFDMEIVDKKGIENGVADHLSRMRI--EDEVLIDDSMPEEQLMAIQQLNEKKLPW 1335
Query: 124 YADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIPENEVSSI 183
YAD VN+L +G PP LS +KKKFF D+ H+Y DEPYL+ D I+R C+ E+E+ I
Sbjct: 1336 YADHVNYLVSGEEPPNLSSYEKKKFFKDINHFYWDEPYLYTLCKDKIYRTCVSEDEIEGI 1395
Query: 184 LTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKRNEMPL 243
L HCH +YGGH +T K KIL +GFWWPS+FKD FISKCD
Sbjct: 1396 LLHCHGFAYGGHFATFKTMSKILQAGFWWPSMFKDAQEFISKCD---------------- 1439
Query: 244 NNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQVVIKLFKKN 303
E FDVW IDF+GPFPSS+ N+YILVA+DYVSKWVE IAS TNDA+VV+KLFK
Sbjct: 1440 ----SFENFDVWGIDFMGPFPSSYGNKYILVAIDYVSKWVEAIASHTNDARVVLKLFKTI 1495
Query: 304 HFSRFGVPRVVISDGGSQNILKNFCKNLE*GTKLQHPITHKLVDKWRSQIDKSKLSLKKL 363
F RFGVPR+VISDGG I K F L+ +H + HK Q++ S +K +
Sbjct: 1496 IFPRFGVPRIVISDGGKHFINKGFENLLK-----KHGVKHKT----SGQVEISNREIKAI 1546
Query: 364 FQ 365
+
Sbjct: 1547 LE 1548
>At2g06320 putative retroelement pol polyprotein
Length = 466
Score = 279 bits (714), Expect = 2e-75
Identities = 136/232 (58%), Positives = 166/232 (70%), Gaps = 5/232 (2%)
Query: 121 APWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIPENEV 180
+PWYAD VN+LA G+ PP L+ ++K FF D+ HYY DEPYL+ D I+R + E+EV
Sbjct: 37 SPWYADHVNYLACGIEPPNLTSYERKNFFRDIHHYYWDEPYLYTLCKDKIYRSYVSEDEV 96
Query: 181 SSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKRNE 240
IL HCH S+YGGH +T K KIL +GFWWP++FKD F+SKCD CQR GNI++RNE
Sbjct: 97 EGILLHCHDSAYGGHFATFKTVSKILQAGFWWPTMFKDAQEFVSKCDSCQRKGNISRRNE 156
Query: 241 MPLNNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQVVIKLF 300
MP N ILEV+IFDVW IDF+GPFPSS+ N+YILV VDYVSKWVE IAS TNDA+VV+KLF
Sbjct: 157 MPQNPILEVDIFDVWGIDFMGPFPSSYGNKYILVVVDYVSKWVEAIASPTNDAKVVLKLF 216
Query: 301 KKNHFSRFGVPRVVISDGGSQNILKNFCKNLE*GTKLQHPITHKLVDKWRSQ 352
K F RFGV VVISDGG I K F L+ +H + HK+ + Q
Sbjct: 217 KTIIFPRFGVSWVVISDGGKHFINKVFENLLK-----KHGVKHKVATPYHPQ 263
>At2g13060 pseudogene
Length = 930
Score = 266 bits (679), Expect = 3e-71
Identities = 134/248 (54%), Positives = 167/248 (67%), Gaps = 9/248 (3%)
Query: 122 PWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIPENEVS 181
PWYAD VN+LAA V P + KK+F +++ Y DEPYL+K DGI+RRCI EV
Sbjct: 137 PWYADIVNYLAADVEPDNFTDYNKKRFLREIRRYQWDEPYLYKHSYDGIYRRCIAATEVP 196
Query: 182 SILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKRNEM 241
SIL+HCHSSSYGGH +T K K+L + FWWP++F+D FIS+CD CQR G I+KRNEM
Sbjct: 197 SILSHCHSSSYGGHFATFKTVSKVLQADFWWPTMFRDTQKFISQCDPCQRRGKISKRNEM 256
Query: 242 PLNNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQVVIKLFK 301
P N LEVE+FD W IDF+GPFP S N YILV VDYVSKWVE IASL ND+ VV+KLFK
Sbjct: 257 PPNFRLEVEVFDRWGIDFMGPFPPSNKNLYILVDVDYVSKWVEAIASLKNDSAVVMKLFK 316
Query: 302 KNHFSRFGVPRVVISDGGSQNILKNFCKNLE*GTKLQHPITHKLVDKWR----SQIDKSK 357
F RFGVPR+VISDG I K K L LQ+ + H++ + Q++ S
Sbjct: 317 SIIFPRFGVPRIVISDGDKHFINKILEKLL-----LQYGVQHRVATPYHPQTSGQVEVSN 371
Query: 358 LSLKKLFQ 365
+K++ +
Sbjct: 372 RQIKEILE 379
Score = 78.2 bits (191), Expect = 1e-14
Identities = 35/63 (55%), Positives = 52/63 (81%), Gaps = 2/63 (3%)
Query: 54 LNKKDAKPRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQL 113
+ KKDAKPRL+ I LLQEFD+E++DKKGVEN VADHLSR+R +D++P++D P++ +
Sbjct: 1 MQKKDAKPRLLIWILLLQEFDIEVRDKKGVENGVADHLSRIR--IDDDVPINDFLPEENI 58
Query: 114 FLL 116
+++
Sbjct: 59 YMI 61
>At4g07610
Length = 276
Score = 247 bits (630), Expect = 1e-65
Identities = 140/286 (48%), Positives = 178/286 (61%), Gaps = 39/286 (13%)
Query: 70 LQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLAQTNAPWYADFVN 129
++ FD+EI DKK +EN ADHLSR+R E+ +P+DDS ++QL + V
Sbjct: 5 IRGFDIEIVDKKDIENGAADHLSRMRI--EERIPIDDSMLEEQLMV------------VE 50
Query: 130 FLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIPENEVSSILTHCHS 189
F SY K+ F+ L P+ R + E+EV IL HCH
Sbjct: 51 FFGG-------SYSGKE--FHQLNVVEGRSPW-----------RYVSEDEVEGILLHCHC 90
Query: 190 SSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKRNEMPLNNILEV 249
S+YGG+ +T K KIL +GFWWP++FK+ F+ KCD C R GNI+KRNEMP N ILEV
Sbjct: 91 STYGGNFTTFKTVSKILQAGFWWPTMFKEAQEFVLKCDSCHRKGNISKRNEMPQNPILEV 150
Query: 250 EIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQVVIKLFKKNHFSRFG 309
E+FDVW IDFIGPFPSS+ N+YILVAVDYVSKW+E AS TNDA+VV+KLFK FSRFG
Sbjct: 151 ELFDVWGIDFIGPFPSSYGNKYILVAVDYVSKWIEATASPTNDAKVVLKLFKTIIFSRFG 210
Query: 310 VPRVVISDGGSQNILKNFCKNLE*GTKLQHPITHKLVDKWRSQIDK 355
VPRVVISDGG I K F L+ +H + H + +++Q K
Sbjct: 211 VPRVVISDGGKHFINKVFENLLK-----KHGVKHGVYVMFKTQKKK 251
>At2g11680 putative retroelement pol polyprotein
Length = 301
Score = 188 bits (478), Expect = 6e-48
Identities = 96/173 (55%), Positives = 119/173 (68%), Gaps = 20/173 (11%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
IYYAS T+D V YATTEKELLAVV+A +KFR YLVGSK+TVYTDH+ ++++ KKD K
Sbjct: 131 IYYASGTMDDTQVRYATTEKELLAVVFAFEKFRSYLVGSKVTVYTDHAALRHIYAKKDTK 190
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLAQTN 120
PRL++ I LLQEFD+EI DKKG+EN VADHLSR+R EDE+P+DDS P++QL + Q N
Sbjct: 191 PRLLRWILLLQEFDMEIVDKKGIENGVADHLSRMR--IEDEVPIDDSMPEEQLMAIQQLN 248
Query: 121 AP------------------WYADFVNFLAAGVLPPELSYQQKKKFFNDLKHY 155
WYAD V +L G PP LS +KKKFF D+ H+
Sbjct: 249 ESAQINKSLDQVCAVEDKLMWYADHVKYLFGGEEPPNLSSYEKKKFFKDINHF 301
>At2g10690 putative retroelement pol polyprotein
Length = 622
Score = 151 bits (381), Expect = 1e-36
Identities = 81/165 (49%), Positives = 108/165 (65%), Gaps = 23/165 (13%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
IYYAS+T+D A YAT EKELLAVV+A +K R YLVG K+ VY DH+ ++++ KK+ K
Sbjct: 376 IYYASRTMDEAQTRYATIEKELLAVVFAFEKIRSYLVGFKVKVYIDHAALRHIYAKKETK 435
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFL----- 115
RL++ I LLQEFD+EI DKKGVEN VADHLSR+R ED +P+DD+ P++QL
Sbjct: 436 SRLLRWILLLQEFDMEIIDKKGVENGVADHLSRMR--IEDSVPIDDTMPEEQLMFYDLVN 493
Query: 116 --------------LAQTNAPWYADFVNFLAAGVLPPELSYQQKK 146
+ + PWYAD VN+L +G + E+S +Q K
Sbjct: 494 KSFDTKDMPEEADAVEEEKLPWYADLVNYLTSGQV--EVSNRQMK 536
>At2g12040 T10J7.2
Length = 976
Score = 150 bits (380), Expect = 1e-36
Identities = 70/116 (60%), Positives = 95/116 (81%), Gaps = 2/116 (1%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
IYYAS+TLD A NYATTEKE LAVV+A +KFR YLVG K+ V+TDH+ +KYL+ KDAK
Sbjct: 267 IYYASRTLDDAKRNYATTEKEFLAVVFAFEKFRSYLVGLKVIVHTDHAALKYLMQNKDAK 326
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLL 116
PRL++ I LLQEFD+E++DKKGVE VADHLSR+R +D++P++D P++ ++++
Sbjct: 327 PRLLRWILLLQEFDIEVRDKKGVEKCVADHLSRIR--IDDDVPINDFLPEENIYMI 380
Score = 148 bits (374), Expect = 7e-36
Identities = 98/254 (38%), Positives = 134/254 (52%), Gaps = 41/254 (16%)
Query: 122 PWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIPENEVS 181
PWYAD V +L A V KK+F +++ YY D+PYL +S
Sbjct: 459 PWYADIVIYLGADVELDNFIDYNKKRFLREIRRYYWDKPYL----------------PIS 502
Query: 182 SILT------HCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNI 235
+L HC + S Q +S ++L + + S ++H+ D CQR G I
Sbjct: 503 IVLMEFIGEMHCCNRD-----SIQSSSSRLLVA--YNVSGCSEIHI---ASDPCQRRGKI 552
Query: 236 TKRNEMPLNNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQV 295
+ NEMP ILEV++FD W I+F+GP P S N YILV VDYVSKWVE IAS ND+ V
Sbjct: 553 SNCNEMPQKFILEVKVFDCWGINFMGPIPPSNKNLYILVVVDYVSKWVEAIASPKNDSAV 612
Query: 296 VIKLFKKNHFSRFGVPRVVISDGGSQNILKNFCKNLE*GTKLQHPITHKLVDKWR----S 351
V+KLFK F FGVPR+VISDGG I K K L LQ+ + H++ +
Sbjct: 613 VMKLFKCIIFPHFGVPRIVISDGGKHFINKILTKLL-----LQYGVQHRVATPYHPQTSG 667
Query: 352 QIDKSKLSLKKLFQ 365
Q++ S +K++ +
Sbjct: 668 QVEVSNRQIKEILE 681
>At4g07660 putative athila transposon protein
Length = 724
Score = 141 bits (355), Expect = 1e-33
Identities = 70/116 (60%), Positives = 91/116 (78%), Gaps = 2/116 (1%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
IYYAS+TLD A YATTEKELL VV+A KFR YLV SK+TVYTDH+ ++++ KKD K
Sbjct: 598 IYYASRTLDEAQGRYATTEKELLVVVFAFKKFRSYLVESKVTVYTDHAALRHMYAKKDTK 657
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLL 116
PRL++ I LLQEFD+EI +KKG+EN ADHLSR+R E+ + +DDS P++QL ++
Sbjct: 658 PRLLRGILLLQEFDMEIVEKKGIENGAADHLSRMR--IEEPILIDDSMPEEQLMVV 711
>At2g14040 putative retroelement pol polyprotein
Length = 841
Score = 115 bits (287), Expect = 8e-26
Identities = 68/155 (43%), Positives = 98/155 (62%), Gaps = 25/155 (16%)
Query: 19 EKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAKPRLIQLIFLLQEFDLEIK 78
+K+L A+ YA + L G+++ V+T H+ ++YLL+KKDAKPRL++ I LLQ+FDLEIK
Sbjct: 681 DKKLHAIYYA----SRTLDGAQVIVHTVHAALRYLLSKKDAKPRLLRWILLLQDFDLEIK 736
Query: 79 DKKGVENVVADHLSRLRETNE----DELP------LDDSFPDD---------QLFLL--A 117
DKKG+EN VAD+LSRLR E D LP ++D + D+ +L L
Sbjct: 737 DKKGIENEVADNLSRLRVQEEVLMRDILPGENLASIEDCYMDEVGRLRVSTLELMTLHTG 796
Query: 118 QTNAPWYADFVNFLAAGVLPPELSYQQKKKFFNDL 152
++N PWYADF N+L+ V PP+ + KKK ++
Sbjct: 797 ESNLPWYADFANYLSCEVPPPDFTGYFKKKLLKEV 831
>At1g35370 hypothetical protein
Length = 1447
Score = 110 bits (274), Expect = 3e-24
Identities = 98/324 (30%), Positives = 155/324 (47%), Gaps = 33/324 (10%)
Query: 3 YASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAKPR 62
Y S+ L G ++ + EKELLA ++A+ K+R YL+ S + TD ++KYLL ++ P
Sbjct: 891 YISRQLKGKQLHLSIYEKELLAFIFAVRKWRHYLLPSHFIIKTDQRSLKYLLEQRLNTPV 950
Query: 63 LIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLAQTNAP 122
Q + L EFD EI+ ++G EN+VAD LSR+ + + L D L +
Sbjct: 951 QQQWLPKLLEFDYEIQYRQGKENLVADALSRVEGSEVLHMALSIVECD----FLKEIQVA 1006
Query: 123 WYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIPENEV-- 180
+ +D GVL +S Q+ D K +YS + +R + + N+V
Sbjct: 1007 YESD-------GVLKDIISALQQHP---DAKKHYSWSQDILRRK-----SKIVVPNDVEI 1051
Query: 181 -SSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKRN 239
+ +L H S GG S + AS + + S F+W + KD+ FI C CQ+ K +
Sbjct: 1052 TNKLLQWLHCSGMGGR-SGRDASHQRVKSLFYWKGMVKDIQAFIRSCGTCQQ----CKSD 1106
Query: 240 EMPLNNILE-VEIFD-VWV---IDFIGPFPSSFVNQYILVAVDYVSKWVEVIA-SLTNDA 293
+L+ + I D +W +DFI P+S I+V VD +SK +A + A
Sbjct: 1107 NAAYPGLLQPLPIPDKIWCDVSMDFIEGLPNSGGKSVIMVVVDRLSKAAHFVALAHPYSA 1166
Query: 294 QVVIKLFKKNHFSRFGVPRVVISD 317
V + F N + G P ++SD
Sbjct: 1167 LTVAQAFLDNVYKHHGCPTSIVSD 1190
>At2g05610 putative retroelement pol polyprotein
Length = 780
Score = 107 bits (268), Expect = 1e-23
Identities = 92/317 (29%), Positives = 150/317 (47%), Gaps = 23/317 (7%)
Query: 3 YASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAKPR 62
+ S+ L G ++ + EKELLAV++ + K+R YL+ S + TD ++KYLL ++ P
Sbjct: 311 FISRQLKGKQLHLSIYEKELLAVIFVVRKWRHYLISSHFIIKTDQRSLKYLLEQRLNTPI 370
Query: 63 LIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLAQTNAP 122
Q + L EFD EI+ K+G EN+VAD LSR+ + + L D LL + A
Sbjct: 371 QQQWLPKLLEFDYEIQYKQGKENLVADALSRVEGSEVLHMALSVVECD----LLKEIQAG 426
Query: 123 WYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIPENE--V 180
+ D G + ++ Q++ + KHY + L ++ + +P N
Sbjct: 427 YVTD-------GDIQGIITILQQQA--DSKKHYTWSQGVLRRKN-----KIVVPNNSGIK 472
Query: 181 SSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKRNE 240
+IL H S GGH S ++ + + + F+W S+ KD+ FI C CQ+ + +
Sbjct: 473 DTILRWLHCSGMGGH-SGKEVTHQRVKGLFYWKSMVKDIQAFIRSCGTCQQCKSDNAASP 531
Query: 241 MPLNNI-LEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIA-SLTNDAQVVIK 298
L + + I+ +DFI P S I+V VD +SK IA + A V +
Sbjct: 532 GLLQPLPIPDRIWSDVSMDFIDGLPLSNGKTVIMVVVDRLSKAAHFIALAHPYSAMTVAQ 591
Query: 299 LFKKNHFSRFGVPRVVI 315
+ N F G P ++
Sbjct: 592 AYLDNVFKLHGCPSSIV 608
>At1g20390 hypothetical protein
Length = 1791
Score = 106 bits (265), Expect = 3e-23
Identities = 81/296 (27%), Positives = 136/296 (45%), Gaps = 27/296 (9%)
Query: 56 KKDAKPRLIQLIFL-LQEFDLEIKDKKGVENVVADHLSRLRETNEDEL------------ 102
K DA +++QL+ + F L K +G +N AD L+ L T++ +L
Sbjct: 1295 KMDAYLKIVQLMTKDFENFKLS-KIPRG-DNAPADALAALALTSDSDLRRIIPVESIDKP 1352
Query: 103 PLDDSFPDDQLFLLAQTNAP-------WYADFVNFLAAGVLPPELSYQQKKKFFNDLKHY 155
+D + + + + +NAP W + ++L+ G LP + + ++ K+
Sbjct: 1353 SIDSTDAVEIVNTIRSSNAPDPADPTDWRVEIRDYLSDGTLPSD-KWTARRLRIKAAKYT 1411
Query: 156 YSDEPYLFKRG*DGIFRRCIPENEVSSILTHCHSSSYGGHASTQKASFKILHSGFWWPSL 215
E +L K G C+ E++ I+ H + G H+ + + K+ GF+WP++
Sbjct: 1412 LMKE-HLLKVSAFGAMLNCLHGTEINEIMKETHEGAAGNHSGGRALALKLKKLGFYWPTM 1470
Query: 216 FKDVHLFISKCDKCQRTGNITKRNEMPLNNILEVEIFDVWVIDFIGPFPSSFVNQYILVA 275
D F +KC++CQR + L + F W +D +GP P+S ++ILV
Sbjct: 1471 ISDCKTFTAKCEQCQRHAPTIHQPTELLRAGVAPYPFMRWAMDIVGPMPASRQKRFILVM 1530
Query: 276 VDYVSKWVEVIASLTNDAQVVIKLFKKNHFSRFGVPRVVISDGGSQNI---LKNFC 328
DY +KWVE + T A V K R G+P +I+D GSQ I +NFC
Sbjct: 1531 TDYFTKWVEAESYATIRANDVQNFVWKFIICRHGLPYEIITDNGSQFISLSFENFC 1586
Score = 44.7 bits (104), Expect = 1e-04
Identities = 30/95 (31%), Positives = 47/95 (48%), Gaps = 2/95 (2%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
I+Y SK+L A Y EK LAVV + K R Y I V TD ++ L+
Sbjct: 1100 IFYTSKSLVEAETRYPVIEKAALAVVTSARKLRPYFQSHTIAVLTD-QPLRVALHSPSQS 1158
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVEN-VVADHLSRL 94
R+ + L E+D++ + + +++ V+AD L L
Sbjct: 1159 GRMTKWAVELSEYDIDFRPRPAMKSQVLADFLIEL 1193
>At2g10780 pseudogene
Length = 1611
Score = 94.7 bits (234), Expect = 1e-19
Identities = 85/333 (25%), Positives = 154/333 (45%), Gaps = 25/333 (7%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
I YAS+ L NY T + E+ AVV+ + +R YL G+K+ +YTDH ++KY+ + +
Sbjct: 977 IAYASRQLRKHEKNYPTHDLEMAAVVFFLKIWRSYLYGAKVQIYTDHKSLKYIFTQPELN 1036
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLR---ETNEDELPLDDSFPDDQLFLLA 117
R + + L+ +++L+I G N VAD LSR R E ++ L + + L+
Sbjct: 1037 LRQRRWMELVADYNLDIAYHPGKANQVADALSRRRSEVEAERSQVDLVNMMGTLHVNALS 1096
Query: 118 QTNAPW---YADFVNFLAAGVLPPELSYQQKKKFFNDLKHYY-SDEPYLFKRG*DGIFRR 173
+ P AD + L+ L E + K N+ Y S+ + G R
Sbjct: 1097 KEVEPLGLGAADQADLLSRIRLAQERDEEIKGWAQNNKTEYQTSNNGTIVVNG-----RV 1151
Query: 174 CIPENEV--SSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQR 231
C+P + IL H S + H + K ++ L + W + KDV +++KC CQ
Sbjct: 1152 CVPNDRALKEEILREAHQSKFSIHPGSNKM-YRDLKRYYHWVGMKKDVARWVAKCPTCQL 1210
Query: 232 TGNITKRNEMPLNNILEVEI----FDVWVIDFIGPFPSSFVNQY--ILVAVDYVSKWVEV 285
+ +++P + + I +D +DF+ P+ +++ + V VD ++K
Sbjct: 1211 ---VKAEHQVPSGLLQNLPIPEWKWDHITMDFVTGLPTGIKSKHNAVWVVVDRLTKSAHF 1267
Query: 286 IASLTND-AQVVIKLFKKNHFSRFGVPRVVISD 317
+A D A+++ + + G+P ++SD
Sbjct: 1268 MAISDKDGAEIIAEKYIDEIVRLHGIPVSIVSD 1300
>At4g03840 putative transposon protein
Length = 973
Score = 93.2 bits (230), Expect = 3e-19
Identities = 72/239 (30%), Positives = 113/239 (47%), Gaps = 15/239 (6%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
I YAS+ L NY T + E+ AVV+A+ +R YL G+K+ +YTDH ++KY+ + +
Sbjct: 427 IAYASRQLRKHEKNYPTHDLEMAAVVFALKIWRSYLYGAKVQIYTDHKSLKYIFTQPELN 486
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLR---ETNEDELPLDDSFPDDQLFLLA 117
R + + L+ ++DL+I G N VAD LSR R E ++ L + + L+
Sbjct: 487 LRQRRWMELVADYDLDIAYHAGKANQVADALSRRRSEVEAERSQVDLVNMMGTLHVNALS 546
Query: 118 QTNAP---WYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYY-SDEPYLFKRG*DGIFRR 173
+ P AD N L+ L E + K N+ Y S+ + G R
Sbjct: 547 KEVEPLGLGAADQANLLSRIRLAQERDEEIKGWALNNKTEYQTSNNGTIVVNG-----RV 601
Query: 174 CIPENEV--SSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQ 230
C+P N IL H S + H + K ++ L + W + KDV +++KC CQ
Sbjct: 602 CVPNNRALKEEILREAHQSKFSIHPGSNKI-YRDLKRYYHWVGMKKDVARWVAKCPTCQ 659
>At2g07410 putative retroelement pol polyprotein
Length = 411
Score = 91.3 bits (225), Expect = 1e-18
Identities = 51/93 (54%), Positives = 62/93 (65%), Gaps = 5/93 (5%)
Query: 260 IGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQVVIKLFKKNHFSRFGVPRVVISDGG 319
+G FPSS+ N+YILVAVDYVSKWVE I S TNDA+VV+KLFK F+RFGV VVISDG
Sbjct: 1 MGSFPSSYGNKYILVAVDYVSKWVEAIVSPTNDAKVVLKLFKTIIFTRFGVHMVVISDGR 60
Query: 320 SQNILKNFCKNLE*GTKLQHPITHKLVDKWRSQ 352
I K F L+ +H + HK+ + Q
Sbjct: 61 KHFINKAFENLLK-----KHGVKHKVATSYHPQ 88
>At2g15410 putative retroelement pol polyprotein
Length = 1787
Score = 90.9 bits (224), Expect = 2e-18
Identities = 51/205 (24%), Positives = 93/205 (44%), Gaps = 4/205 (1%)
Query: 117 AQTNAPWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIP 176
A TN W + +++A ++P + ++ + + Y +L + G+ C+
Sbjct: 1372 ASTN--WIEEIRSYIADRIVPKDKWAARRLRARS--AQYTLPHEHLLRWSATGVLLSCLD 1427
Query: 177 ENEVSSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNIT 236
+ E ++ H + G H+ + + KI G +WP++ D F+++C+KCQ
Sbjct: 1428 DEEAQQVMREIHEGAGGNHSGGRALALKIRKHGQYWPTMNADCEKFVARCEKCQGHAPFI 1487
Query: 237 KRNEMPLNNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQVV 296
L + F W +D I PFP+S +YILV DY +KW E + ++ V
Sbjct: 1488 HIPSEVLQTAMPPYPFMRWAMDIIRPFPASRQKKYILVMTDYFTKWFEAESYARIQSKEV 1547
Query: 297 IKLFKKNHFSRFGVPRVVISDGGSQ 321
KN G+P +++D G+Q
Sbjct: 1548 QNFVWKNIICHHGLPYEIVTDNGTQ 1572
Score = 48.1 bits (113), Expect = 1e-05
Identities = 28/80 (35%), Positives = 42/80 (52%), Gaps = 1/80 (1%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
I+Y SKTLDGA + Y T EK AVV + K R Y + V T + ++ +L+
Sbjct: 1085 IFYVSKTLDGAELRYPTLEKLAFAVVISARKLRPYFKSYTVEVLT-NQPLRTILHSPSQS 1143
Query: 61 PRLIQLIFLLQEFDLEIKDK 80
RL + L E+D+ K++
Sbjct: 1144 GRLAKWAVELSEYDIAYKNR 1163
>At4g16910 retrotransposon like protein
Length = 687
Score = 88.2 bits (217), Expect = 1e-17
Identities = 82/333 (24%), Positives = 155/333 (45%), Gaps = 33/333 (9%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
I YAS+ L NY T + E+ AVV+A+ +R YL G+K+ ++TDH ++KY+ + +
Sbjct: 69 IAYASRQLRKHEKNYPTNDLEMAAVVFALKIWRSYLYGAKVQIFTDHKSLKYIFTQPELN 128
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLR---ETNEDELPLDDSFPDDQLFLLA 117
R + I L+ ++DL I G N V D LSRLR E +++ L + L L+
Sbjct: 129 LRQRRWIKLVADYDLNIAYHPGKANQVVDALSRLRTVVEAERNQVNLVNMMGTLHLNALS 188
Query: 118 QTNAPWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIF----RR 173
+ P N A +L S Q++ + ++K + + ++ +G R
Sbjct: 189 KEVEPLGLRAAN--QADLLSRIRSAQERDE---EIKGWAQNNKTEYQSSNNGTIVVNGRV 243
Query: 174 CIPENEV--SSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQR 231
C P ++ IL H S + H + K ++ L + W + KDV +++K
Sbjct: 244 CGPNDKALKEEILKEAHQSKFSIHPGSNK-MYRDLKRYYHWVGMKKDVARWVAK------ 296
Query: 232 TGNITKRNEMPLNNILEVEI----FDVWVIDFIGPFPSSFVNQY--ILVAVDYVSKWVEV 285
+ +++P + + I +D ++DF+ P+ +++ + V VD ++K
Sbjct: 297 -----EEHQVPSGMLQNLPIPEWKWDHIMMDFVTGLPTGIKSKHNAVWVVVDRLTKSAHF 351
Query: 286 IASLTND-AQVVIKLFKKNHFSRFGVPRVVISD 317
+A D A+++ + + G+P ++SD
Sbjct: 352 MAISDKDAAEIIAEKYIDEIVRLHGIPVSIVSD 384
>At2g06470 putative retroelement pol polyprotein
Length = 899
Score = 86.3 bits (212), Expect = 4e-17
Identities = 70/239 (29%), Positives = 111/239 (46%), Gaps = 15/239 (6%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
I YAS+ L NY T + E+ AVV+A+ +R YL G+K+ +YTDH ++KY+ + +
Sbjct: 427 IAYASRQLWKHEKNYPTHDLEMAAVVFALKIWRSYLYGAKVQIYTDHKSLKYIFIQPELN 486
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDD------SFPDDQLF 114
R + + L+ ++DL+I G N VAD LSR R E E D + + L
Sbjct: 487 LRQRRWMELVADYDLDIAYHPGKANQVADALSRRRSEVEAERSQVDLVNMMGTLHVNALS 546
Query: 115 LLAQTNAPWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYY-SDEPYLFKRG*DGIFRR 173
++ AD + L+ L E + K N+ Y S+ + G R
Sbjct: 547 KEVESLGLGAADQADLLSRIRLAQERDEEIKGWALNNKTEYQTSNNGTIVVNG-----RV 601
Query: 174 CIPENEV--SSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQ 230
C+P + IL H S + H + K ++ L + W + KDV +++KC CQ
Sbjct: 602 CVPNDRALKEEILREAHQSKFSIHPGSNKM-YRDLKRYYHWVGMKKDVARWVAKCPTCQ 659
>orf119 -mitochondrial genome-
Length = 119
Score = 85.1 bits (209), Expect = 9e-17
Identities = 36/56 (64%), Positives = 44/56 (78%)
Query: 205 ILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKRNEMPLNNILEVEIFDVWVIDFI 260
+L +GF+WP+ FKD H F+S CD CQR GN TKRNEMP + ILEVE+FDVW I F+
Sbjct: 35 VLQAGFYWPTTFKDAHGFVSSCDACQRKGNFTKRNEMPQHFILEVEVFDVWGIYFM 90
>At2g13230 pseudogene
Length = 353
Score = 76.6 bits (187), Expect = 3e-14
Identities = 58/237 (24%), Positives = 94/237 (39%), Gaps = 24/237 (10%)
Query: 120 NAPWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIPENE 179
++ W NF++ + L + +K + + E LFK G C+ +
Sbjct: 89 DSEWIEPIRNFISKRKV--RLDKWEARKLKAEAARFVLVEEKLFKWRLSGPLMTCVEKEA 146
Query: 180 VSSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKRN 239
++ H S G ++ + + KI G++WP++ KD C+KCQR
Sbjct: 147 ARKVMKEVHGGSCGNNSGGRALAIKIKRLGYFWPTMIKD-------CEKCQRNAPTIHLP 199
Query: 240 EMPLNNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQVVIKL 299
L++I W +D IGP +S + +LV DY SK E + + V
Sbjct: 200 AELLSSIASPYPLMRWSMDIIGPMHASKQKKLVLVLTDYFSKLREAESYASIKEAQVESF 259
Query: 300 FKKNHFSRFGVPRVVISDGGSQNILKNFCKNLE*GTKLQHPITHKLVDKWRSQIDKS 356
K+ R GVP +++D GSQ I F DKW ++ KS
Sbjct: 260 MWKHILCRHGVPYEIVTDNGSQFISTRF---------------QGFCDKWGIRLSKS 301
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.336 0.148 0.464
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,204,287
Number of Sequences: 26719
Number of extensions: 466329
Number of successful extensions: 1824
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 47
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 1712
Number of HSP's gapped (non-prelim): 81
length of query: 547
length of database: 11,318,596
effective HSP length: 104
effective length of query: 443
effective length of database: 8,539,820
effective search space: 3783140260
effective search space used: 3783140260
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC141865.9


