

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141436.5 + phase: 0 /pseudo
(364 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
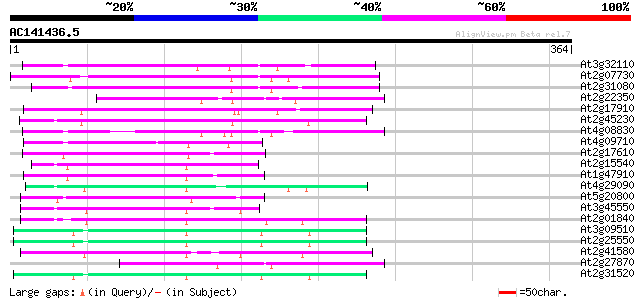
Score E
Sequences producing significant alignments: (bits) Value
At3g32110 non-LTR reverse transcriptase, putative 114 1e-25
At2g07730 putative non-LTR retroelement reverse transcriptase 109 2e-24
At2g31080 putative non-LTR retroelement reverse transcriptase 108 5e-24
At2g22350 putative non-LTR retroelement reverse transcriptase 96 3e-20
At2g17910 putative non-LTR retroelement reverse transcriptase 92 3e-19
At2g45230 putative non-LTR retroelement reverse transcriptase 91 1e-18
At4g08830 putative protein 86 4e-17
At4g09710 RNA-directed DNA polymerase -like protein 81 8e-16
At2g17610 putative non-LTR retroelement reverse transcriptase 81 1e-15
At2g15540 putative non-LTR retroelement reverse transcriptase 78 9e-15
At1g47910 reverse transcriptase, putative 76 3e-14
At4g29090 putative protein 76 3e-14
At5g20800 putative protein 75 6e-14
At3g45550 putative protein 71 1e-12
At2g01840 putative non-LTR retroelement reverse transcriptase 71 1e-12
At3g09510 putative non-LTR reverse transcriptase 69 3e-12
At2g25550 putative non-LTR retroelement reverse transcriptase 69 3e-12
At2g41580 putative non-LTR retroelement reverse transcriptase 69 4e-12
At2g27870 putative non-LTR retroelement reverse transcriptase 69 4e-12
At2g31520 putative non-LTR retroelement reverse transcriptase 67 2e-11
>At3g32110 non-LTR reverse transcriptase, putative
Length = 1911
Score = 114 bits (284), Expect = 1e-25
Identities = 69/248 (27%), Positives = 112/248 (44%), Gaps = 25/248 (10%)
Query: 9 SQDIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQETWQWIWKLHLPENLKHFIWLAM 68
+ D W + G ++VKS + LT++ Q + +WK+ PE ++ F+WL +
Sbjct: 1542 AHDRLAWGMTSDGRFTVKSAFAMLTND--DSPRQDMSSLYGRVWKVQAPERVRVFLWLVV 1599
Query: 69 HGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWDDLLFNK--DTNFNS 126
+ + TN+ R +HL C+ C IES+LH RDCP IWD ++ + + F
Sbjct: 1600 NQAIMTNSERKRRHLCDSDVCQVCRGGIESILHVLRDCPAMSGIWDRIVPRRLQQSFFTM 1659
Query: 127 QDFNWWFKHYALALN------GNIFVITCWVLWKARNHKVFNGNDWVCLSHI-------- 172
W + + L +F + W WK R +F G + C +
Sbjct: 1660 SLLEWLYSNLRQGLMTEGSDWSTMFAMAVWWGWKWRCSNIF-GENKTCRDRVRFIKDLAV 1718
Query: 173 ---YSLHSDVVAHISNDVIQKPIK*VKWIPPLDNIIKVNVDGISLSNHGRSVFGGLIRNN 229
+ +V +S + KPI +W PP++ K+N DG S N G + GG++RN+
Sbjct: 1719 EVSIAYSREVELRLSGLRVNKPI---RWTPPMEGWYKINTDGASRGNPGLASAGGVLRNS 1775
Query: 230 NGDWLLGF 237
G W GF
Sbjct: 1776 AGAWCGGF 1783
>At2g07730 putative non-LTR retroelement reverse transcriptase
Length = 970
Score = 109 bits (273), Expect = 2e-24
Identities = 77/252 (30%), Positives = 115/252 (45%), Gaps = 17/252 (6%)
Query: 1 QSIFIDESSQDIKIWTHSPTGMYSVKSGYQWLTSNCLS--FAGQTQQETWQWIWKLHLPE 58
+SI D D W + G ++V+S Y+ L G ++ IWKL PE
Sbjct: 599 ESISADALLSDELSWKGTQNGDFTVRSAYELLKPEAEERPLIGSFLKQ----IWKLVAPE 654
Query: 59 NLKHFIWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWDDLLF 118
++ FIWL H + TN R +HL+ ++C C + ES+LH RDCP IW LL
Sbjct: 655 RVRVFIWLVSHMVIMTNVERVRRHLSDIATCSVCNGADESILHVLRDCPAMTPIWQRLLP 714
Query: 119 NKDTNFNSQDFNWWFKHYALALNG--NIFVITCWVLWKARNHKVFNGNDWVC---LSHIY 173
+ N F W F + A +F + W WK R VF G +C L I
Sbjct: 715 QRRQNEFFSQFEWLFTNLDPAKGDWPTLFSMGIWWAWKWRCGDVF-GERKLCRDRLKFIK 773
Query: 174 SLHSDV----VAHISNDVIQKPI-K*VKWIPPLDNIIKVNVDGISLSNHGRSVFGGLIRN 228
+ +V V ++N V + + + ++W P D +K+ DG S + G + G I N
Sbjct: 774 DIAEEVRKAHVGTLNNHVKRARVERMIRWKAPSDRWVKLTTDGASRGHQGLAAASGAILN 833
Query: 229 NNGDWLLGFFMD 240
G+WL GF ++
Sbjct: 834 LQGEWLGGFALN 845
>At2g31080 putative non-LTR retroelement reverse transcriptase
Length = 1231
Score = 108 bits (270), Expect = 5e-24
Identities = 72/240 (30%), Positives = 106/240 (44%), Gaps = 19/240 (7%)
Query: 15 WTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQETWQWIWKLHLPENLKHFIWLAMHGNLPT 74
W + G ++V+S Y L + + IWKL PE ++ FIWL + T
Sbjct: 872 WKGTQDGAFTVRSAYSLLQGDVGD--RPNMGSFFNRIWKLITPERVRVFIWLVSQNVIMT 929
Query: 75 NAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWDDLL-FNKDTNFNSQDFNWWF 133
N R +HL+ ++ C C + E++LH RDCP IW LL + F SQ W
Sbjct: 930 NVERVRRHLSENAICSVCNGAEETILHVLRDCPAMEPIWRRLLPLRRHHEFFSQSLLEWL 989
Query: 134 KHYALALNG---NIFVITCWVLWKARNHKVFNGNDWVC----------LSHIYSLHSDVV 180
+ G +F + W WK R VF G +C + +H V
Sbjct: 990 FTNMDPVKGIWPTLFGMGIWWAWKWRCCDVF-GERKICRDRLKFIKDMAEEVRRVHVGAV 1048
Query: 181 AHISNDVIQKPIK*VKWIPPLDNIIKVNVDGISLSNHGRSVFGGLIRNNNGDWLLGFFMD 240
+ N V + + ++W P D +K+ DG S NHG + GG IRN G+WL GF ++
Sbjct: 1049 GNRPNGV--RVERMIRWQVPSDGWVKITTDGASRGNHGLAAAGGAIRNGQGEWLGGFALN 1106
>At2g22350 putative non-LTR retroelement reverse transcriptase
Length = 321
Score = 95.9 bits (237), Expect = 3e-20
Identities = 60/201 (29%), Positives = 98/201 (47%), Gaps = 18/201 (8%)
Query: 57 PENLKHFIWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWDDL 116
PE ++ F+WL + + TN R +HL+ C+ C E++LH RDCP IW L
Sbjct: 3 PERVRVFLWLVVQQVIITNVERYRRHLSDTRVCQICQGGEETILHVLRDCPAMAGIWSRL 62
Query: 117 LFNKDTN--FNSQDFNWWFKHYALALNGN---IFVITCWVLWKARNHKVFNGNDWVCLSH 171
+ F + W +K+ L G+ +FV+ W WK R +F GN C
Sbjct: 63 VPRDQIRQFFTASLLEWIYKN--LRERGSWPTVFVMAVWWGWKWRCGNIFGGNG-KCRDR 119
Query: 172 IYSLHSDVVAHIS--------NDV-IQKPIK*VKWIPPLDNIIKVNVDGISLSNHGRSVF 222
+ + D+ ++ N+V + + + V W+ P D +K+N DG S N G +
Sbjct: 120 VKFI-KDLAEEVAIANAFVKGNEVRVSRVERLVSWVSPEDGWVKLNTDGASRGNPGFATA 178
Query: 223 GGLIRNNNGDWLLGFFMDFVV 243
GG++R++NG W+ GF ++ V
Sbjct: 179 GGVLRDHNGAWIGGFAVNIGV 199
>At2g17910 putative non-LTR retroelement reverse transcriptase
Length = 1344
Score = 92.4 bits (228), Expect = 3e-19
Identities = 65/246 (26%), Positives = 106/246 (42%), Gaps = 22/246 (8%)
Query: 10 QDIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQ------ETWQWIWKLHLPENLKHF 63
+D W HS G+YSVK+GY++L+ Q + + IW LH ++ F
Sbjct: 977 EDSFCWLHSHNGLYSVKTGYEFLSKQVHHRLYQEAKVKPSVNSLFDKIWNLHTAPKIRIF 1036
Query: 64 IWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWD-DLLFNKDT 122
+W A+HG +P + + SD C C E++ H +CPL+ +W L + +
Sbjct: 1037 LWKALHGAIPVEDRLRTRGIRSDDGCLMCDTENETINHILFECPLARQVWAITHLSSAGS 1096
Query: 123 NFNSQDFNWWFKHYALALNGNI-----FVI--TCWVLWKARNHKVFNGNDWVCLSHI--- 172
F++ + + L ++ FV W LWK RN +F G + + +
Sbjct: 1097 EFSNSVYTNMSRLIDLTQQNDLPHHLRFVSPWILWFLWKNRNALLFEGKGSITTTLVDKA 1156
Query: 173 ---YSLHSDVVAHISNDVIQKPIK*VKWIPPLDNIIKVNVDGISLSNHGRSVFGGLIRNN 229
Y H+ ND +K +K KW PPL +K N+ H S ++R++
Sbjct: 1157 YEAYHEWFSAQTHMQND--EKHLKITKWCPPLPGELKCNIGFAWSKQHHFSGASWVVRDS 1214
Query: 230 NGDWLL 235
G LL
Sbjct: 1215 QGKVLL 1220
>At2g45230 putative non-LTR retroelement reverse transcriptase
Length = 1374
Score = 90.9 bits (224), Expect = 1e-18
Identities = 64/247 (25%), Positives = 106/247 (42%), Gaps = 23/247 (9%)
Query: 7 ESSQDIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQ-------ETWQWIWKLHLPEN 59
+ ++D W +S +G YSVKSGY W+ + ++ Q+ +Q IWKL +P
Sbjct: 1004 KETRDRFTWEYSRSGHYSVKSGY-WVMTEIINQRNNPQEVLQPSLDPIFQQIWKLDVPPK 1062
Query: 60 LKHFIWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWDDLLFN 119
+ HF+W ++ L + A +HL + SC RC + E++ H CP + W
Sbjct: 1063 IHHFLWRCVNNCLSVASNLAYRHLAREKSCVRCPSHGETVNHLLFKCPFARLTWAISPLP 1122
Query: 120 KDTNFNSQDFNWWFKHYALALNGN---------IFVITCWVLWKARNHKVFNGNDWVCLS 170
+ + H+ L+++ + + W LWK RN VF G ++
Sbjct: 1123 APPGGEWAESLFRNMHHVLSVHKSQPEESDHHALIPWILWRLWKNRNDLVFKGREFTAPQ 1182
Query: 171 HIYSLHSDVVAHISNDVIQKPI------K*VKWIPPLDNIIKVNVDGISLSNHGRSVFGG 224
I D+ A + Q + + VKW PP +K N DG + G G
Sbjct: 1183 VILKATEDMDAWNNRKEPQPQVTSSTRDRCVKWQPPSHGWVKCNTDGAWSKDLGNCGVGW 1242
Query: 225 LIRNNNG 231
++RN+ G
Sbjct: 1243 VLRNHTG 1249
>At4g08830 putative protein
Length = 947
Score = 85.5 bits (210), Expect = 4e-17
Identities = 68/256 (26%), Positives = 109/256 (42%), Gaps = 45/256 (17%)
Query: 9 SQDIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQETWQWIWKLHLPENLKHFIWLAM 68
++D W +S G+++VKS Y+ LT + + +W++ E +K F+W
Sbjct: 656 ARDRLSWGYSADGVFTVKSAYRLLTED--HDPRPNMAAFFDRLWRVVALERVKTFLW--- 710
Query: 69 HGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWDDLLFNKDTN--FNS 126
H+ S C+ C E++LH +DCP IW L+ + + FN
Sbjct: 711 -------------HIGDTSVCQVCKGGDETILHVLKDCPSIAGIWRRLVQVQRSYDFFNG 757
Query: 127 QDFNWWFKHYAL--ALNG----NIFVITCWVLWKARNHKVFNGNDWVC------------ 168
F W + + + A G +F I W WK R VF G C
Sbjct: 758 SLFGWLYVNLGMKNAETGYAWATLFAIVVWWSWKWRCGYVF-GEVGKCRDRVKFFRDLAA 816
Query: 169 -LSHIYSLHSDVVAHISNDVIQKPIK*VKWIPPLDNIIKVNVDGISLSNHGRSVFGGLIR 227
+SH +++HS + + + + V W PP +K+N DG S N G + GG++R
Sbjct: 817 EVSHAHAIHSQ-----NGGLRTRVERLVAWKPPDGEWVKLNTDGASRGNLGLATTGGVLR 871
Query: 228 NNNGDWLLGFFMDFVV 243
+ G W GF +D V
Sbjct: 872 DGIGHWCGGFALDIGV 887
>At4g09710 RNA-directed DNA polymerase -like protein
Length = 1274
Score = 81.3 bits (199), Expect = 8e-16
Identities = 52/165 (31%), Positives = 80/165 (47%), Gaps = 13/165 (7%)
Query: 10 QDIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQETWQW-IWKLHLPENLKHFIWLAM 68
QD +W +G Y+ K+GY N SF WQ IWK+H +KHF+W AM
Sbjct: 934 QDSLVWLPVKSGEYTTKTGYALAKLN--SFPASQLDFNWQKNIWKIHTSPKVKHFLWKAM 991
Query: 69 HGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWD--DLLFN-KDTNFN 125
G LP + +++ ++ +CKRCG + ES LH CP + +W+ +LFN + +
Sbjct: 992 KGALPVGEALSRRNIEAEVTCKRCGQT-ESSLHLMLLCPYAKKVWELAPVLFNPSEATHS 1050
Query: 126 SQDFNWWFKHYALAL------NGNIFVITCWVLWKARNHKVFNGN 164
S +AL + ++ W LWKARN +F+ +
Sbjct: 1051 SVALLLVDAKRMVALPPTGLGSAPLYPWLLWHLWKARNRLIFDNH 1095
>At2g17610 putative non-LTR retroelement reverse transcriptase
Length = 773
Score = 80.9 bits (198), Expect = 1e-15
Identities = 50/174 (28%), Positives = 77/174 (43%), Gaps = 18/174 (10%)
Query: 9 SQDIKIWTHSPTGMYSVKSGYQWLT-------SNCLSFAGQTQQETWQWIWKLHLPENLK 61
+ D W ++ G YSVKSGY L ++ S + Q + IWK + P +K
Sbjct: 452 ANDAITWIYTHDGNYSVKSGYHLLRKLSQQQHASLPSPNEVSAQTVFTNIWKQNAPPKIK 511
Query: 62 HFIWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWD------- 114
HF W + H LPT + L +D +C+RCG + E + H C +S IW+
Sbjct: 512 HFWWRSAHNALPTAGNLKRRRLITDDTCQRCGEASEDVNHLLFQCRVSKEIWEQAHIKLC 571
Query: 115 --DLLFNKDTNFNSQDFNWWFKHYALALNGNIFVITCWVLWKARNHKVFNGNDW 166
D L + N N + + + + ++F W +WK RN +FN W
Sbjct: 572 PGDSLMSNSFNQNLESIQ--KLNQSARKDVSLFPFIGWRIWKMRNDLIFNNKRW 623
>At2g15540 putative non-LTR retroelement reverse transcriptase
Length = 1225
Score = 77.8 bits (190), Expect = 9e-15
Identities = 50/162 (30%), Positives = 76/162 (46%), Gaps = 16/162 (9%)
Query: 15 WTHSPTGMYSVKSGYQWLTSNC------LSFAGQTQQETWQWIWKLHLPENLKHFIWLAM 68
W+ + +G Y+VKSGY W + L F G + +WK+ KHF W +
Sbjct: 885 WSFTKSGNYTVKSGY-WAARDLSRPTCDLPFQGPSVSALQAQVWKIKTTRKFKHFEWQCL 943
Query: 69 HGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIW-------DDLLFNKD 121
G L TN +H+ ++ C RCGA ES+ H CP S IW + +F ++
Sbjct: 944 SGCLATNQRLFSRHIGTEKVCPRCGAEEESINHLLFLCPPSRQIWALSPIPSSEYIFPRN 1003
Query: 122 TNFNSQDFNW-WFKHYALALN-GNIFVITCWVLWKARNHKVF 161
+ F + DF K + +A + IF W +WK+RN +F
Sbjct: 1004 SLFYNFDFLLSRGKEFDIAEDIMEIFPWILWYIWKSRNRFIF 1045
>At1g47910 reverse transcriptase, putative
Length = 1142
Score = 76.3 bits (186), Expect = 3e-14
Identities = 47/166 (28%), Positives = 72/166 (43%), Gaps = 12/166 (7%)
Query: 10 QDIKIWTHSPTGMYSVKSGYQWLTSNC---LSFAGQTQQETWQWIWKLHLPENLKHFIWL 66
+D W + G Y+VKSGY + + G +IWK+ P L+HF+W
Sbjct: 784 EDTLGWHFTKAGNYTVKSGYHTARLDLNEGTTLIGPDLTTLKAYIWKVQCPPKLRHFLWQ 843
Query: 67 AMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIW-------DDLLFN 119
+ G +P + + + D C CGAS ES+ HT C + IW +F
Sbjct: 844 ILSGCVPVSENLRKRGILCDKGCVSCGASEESINHTLFQCHPARQIWALSQIPTAPGIFP 903
Query: 120 KDTNFNSQDFNWWFKHYALALNGNIFVITCWVLWKARNHKVFNGND 165
++ F + D +W ++ + W +WKARN KVF D
Sbjct: 904 SNSIFTNLDHLFW--RIPSGVDSAPYPWIIWYIWKARNEKVFENVD 947
>At4g29090 putative protein
Length = 575
Score = 75.9 bits (185), Expect = 3e-14
Identities = 61/252 (24%), Positives = 101/252 (39%), Gaps = 37/252 (14%)
Query: 11 DIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQET-------WQWIWKLHLPENLKHF 63
D W ++ +G Y+VKSGY W+ + ++ Q+ + +Q IWK ++HF
Sbjct: 211 DSYTWDYTSSGDYTVKSGY-WVLTQIINKRSSPQEVSEPSLNPIYQKIWKSQTSPKIQHF 269
Query: 64 IWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIW---------- 113
+W + +LP A +HL+ +S+C RC + E++ H C + W
Sbjct: 270 LWKCLSNSLPVAGALAYRHLSKESACIRCPSCKETVNHLLFKCTFARLTWAISSIPIPLG 329
Query: 114 ----DDLLFNKDTNFNSQDFN-WWFKHYALALNGNIFVITCWVLWKARNHKVFNGNDWVC 168
D + N FN + N W K + W LWK RN VF G ++
Sbjct: 330 GEWADSIYVNLYWVFNLGNGNPQWEK------ASQLVPWLLWRLWKNRNELVFRGREFNA 383
Query: 169 LSHIYSLHSDV----VAHISNDVIQKP----IK*VKWIPPLDNIIKVNVDGISLSNHGRS 220
+ D+ + + KP +W PP +K N D ++ R
Sbjct: 384 QEVLRRAEDDLEEWRIRTEAESCGTKPQVNRSSCGRWRPPPHQWVKCNTDATWNRDNERC 443
Query: 221 VFGGLIRNNNGD 232
G ++RN G+
Sbjct: 444 GIGWVLRNEKGE 455
>At5g20800 putative protein
Length = 359
Score = 75.1 bits (183), Expect = 6e-14
Identities = 47/170 (27%), Positives = 80/170 (46%), Gaps = 15/170 (8%)
Query: 8 SSQDIKIWTHSPTGMYSVKSGY-----QWLTSNCLSFAGQTQQETWQWIWKLHLPENLKH 62
+ +D W + +G Y+ KSGY +++ +N LSF G + WK+ P L+H
Sbjct: 80 NKEDTLGWHFTKSGKYTFKSGYHTSRIEYIDTN-LSFIGPEITLLKAYSWKVQCPPKLRH 138
Query: 63 FIWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWDDL------ 116
F+W + +P + + D C RCGA+ +++ HT C + IW
Sbjct: 139 FLWQILASCVPVRENLRKRGINCDIGCARCGATEKTINHTLFQCHPARQIWALSQIPTVI 198
Query: 117 -LFNKDTNFNSQDFNWWFKHYALALNGNIFVITCWVLWKARNHKVFNGND 165
+F D+ + + D +W A+ ++I W +WKARN K+F+ D
Sbjct: 199 GIFPSDSIYENLDHLFWRVQSAVDSFSYPWII--WYIWKARNAKLFDNVD 246
>At3g45550 putative protein
Length = 851
Score = 70.9 bits (172), Expect = 1e-12
Identities = 51/172 (29%), Positives = 74/172 (42%), Gaps = 20/172 (11%)
Query: 8 SSQDIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQETW------QWIWKLHLPENLK 61
S D IW+++ TG Y+V+SGY WL+++ S T + IW L + LK
Sbjct: 619 SKTDRLIWSYNSTGDYTVRSGY-WLSTHDPSNTIPTMAKPHGSVDLKTKIWNLPIMPKLK 677
Query: 62 HFIWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIW---DDLLF 118
HF+W + LPT + + D C RC ES+ H CP + W D L+
Sbjct: 678 HFLWRILSKALPTTDRLTTRGMRIDPGCPRCRRENESINHALFTCPFATMAWRLSDTPLY 737
Query: 119 NKDTNFNSQDFNWWFKHYALALNGNIFVIT--------CWVLWKARNHKVFN 162
N+ + N + L L + W +WKARN+ VFN
Sbjct: 738 RSSILSNNIEDN--ISNILLLLQNTTITDSQKLIPFWLLWRIWKARNNVVFN 787
>At2g01840 putative non-LTR retroelement reverse transcriptase
Length = 1715
Score = 70.9 bits (172), Expect = 1e-12
Identities = 61/256 (23%), Positives = 107/256 (40%), Gaps = 37/256 (14%)
Query: 8 SSQDIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQETW----------QWIWKLHLP 57
+++D W ++ Y+V+SGY W+ ++ T++E Q IW+L +
Sbjct: 1340 AARDSYKWAYTRNTQYTVRSGY-WVATH----VNLTEEEIINPLEGDVPLKQEIWRLKIT 1394
Query: 58 ENLKHFIWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIW---- 113
+KHFIW + G L T +++ +D +C+RC + E++ H C + +W
Sbjct: 1395 PKIKHFIWRCLSGALSTTTQLRNRNIPADPTCQRCCNADETINHIIFTCSYAQVVWRSAN 1454
Query: 114 ----DDLLFNKDTNFNSQDFNWWFKHYAL-ALNGNIFVITCWVLWKARNHKVFNGND--- 165
+ L F + N + K+ L LNG + W LWK+RN +F D
Sbjct: 1455 FSGSNRLCFTDNLEENIRLILQGKKNQNLPILNGLMPFWIMWRLWKSRNEYLFQQLDRFP 1514
Query: 166 WVCLSHIYSLHSDVVAHISNDVI---------QKPIK*VK-WIPPLDNIIKVNVDGISLS 215
W ++ V + ND +P+ K W P + +K N D +
Sbjct: 1515 WKVAQKAEQEATEWVETMVNDTAISHNTAQSNDRPLSRSKQWSSPPEGFLKCNFDSGYVQ 1574
Query: 216 NHGRSVFGGLIRNNNG 231
+ G ++R+ NG
Sbjct: 1575 GRDYTSTGWILRDCNG 1590
>At3g09510 putative non-LTR reverse transcriptase
Length = 484
Score = 69.3 bits (168), Expect = 3e-12
Identities = 68/255 (26%), Positives = 102/255 (39%), Gaps = 28/255 (10%)
Query: 3 IFIDESSQDIKI-WTHSPTGMYSVKSGYQWLTSNCLSFA-------GQTQQETWQWIWKL 54
I++ +S + KI W ++ TG Y+V+SGY LT + + G +T IW L
Sbjct: 108 IYLAKSKKPDKIIWNYNTTGEYTVRSGYWLLTHDPSTNIPAINPPHGSIDLKTR--IWNL 165
Query: 55 HLPENLKHFIWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIW- 113
+ LKHF+W A+ L T + + D SC RC ES+ H CP + W
Sbjct: 166 PIMPKLKHFLWRALSQALATTERLTTRGMRIDPSCPRCHRENESINHALFTCPFATMAWR 225
Query: 114 --------DDLLFNKDTNFNSQDFNWWFKHYALALNGNIFVITCWVLWKARNHKVFN--- 162
+ L+ N S N+ + + V W +WKARN+ VFN
Sbjct: 226 LSDSSLIRNQLMSNDFEENISNILNFVQDTTMSDFHKLLPVWLIWRIWKARNNVVFNKFR 285
Query: 163 -GNDWVCLSHIYSLHSDVVAHISNDVIQKPIK-----*VKWIPPLDNIIKVNVDGISLSN 216
LS H + A S+ P + ++W P +K N D
Sbjct: 286 ESPSKTVLSAKAETHDWLNATQSHKKTPSPTRQIAENKIEWRNPPATYVKCNFDAGFDVQ 345
Query: 217 HGRSVFGGLIRNNNG 231
+ G +IRN+ G
Sbjct: 346 KLEATGGWIIRNHYG 360
>At2g25550 putative non-LTR retroelement reverse transcriptase
Length = 1750
Score = 69.3 bits (168), Expect = 3e-12
Identities = 68/255 (26%), Positives = 102/255 (39%), Gaps = 28/255 (10%)
Query: 3 IFIDESSQDIKI-WTHSPTGMYSVKSGYQWLTSNCLSFA-------GQTQQETWQWIWKL 54
I++ +S + KI W ++ TG Y+V+SGY LT + + G +T IW L
Sbjct: 1374 IYLAKSKKPDKIIWNYNTTGEYTVRSGYWLLTHDPSTNIPAINPPHGSIDLKTR--IWNL 1431
Query: 55 HLPENLKHFIWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIW- 113
+ LKHF+W A+ L T + + D SC RC ES+ H CP + W
Sbjct: 1432 PIMPKLKHFLWRALSQALATTERLTTRGMRIDPSCPRCHRENESINHALFTCPFATMAWR 1491
Query: 114 --------DDLLFNKDTNFNSQDFNWWFKHYALALNGNIFVITCWVLWKARNHKVFN--- 162
+ L+ N S N+ + + V W +WKARN+ VFN
Sbjct: 1492 LSDSSLIRNQLMSNDFEENISNILNFVQDTTMSDFHKLLPVWLIWRIWKARNNVVFNKFR 1551
Query: 163 -GNDWVCLSHIYSLHSDVVAHISNDVIQKPIK-----*VKWIPPLDNIIKVNVDGISLSN 216
LS H + A S+ P + ++W P +K N D
Sbjct: 1552 ESPSKTVLSAKAETHDWLNATQSHKKTPSPTRQIAENKIEWRNPPATYVKCNFDAGFDVQ 1611
Query: 217 HGRSVFGGLIRNNNG 231
+ G +IRN+ G
Sbjct: 1612 KLEATGGWIIRNHYG 1626
>At2g41580 putative non-LTR retroelement reverse transcriptase
Length = 1094
Score = 68.9 bits (167), Expect = 4e-12
Identities = 63/252 (25%), Positives = 105/252 (41%), Gaps = 31/252 (12%)
Query: 8 SSQDIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQET------WQWIWKLHLPENLK 61
S QD W + G+Y+VKS Y + + +E + IW L+ +K
Sbjct: 726 SRQDSFCWFGTNHGLYTVKSEYDLCSRQVHKQMFKEAEEQPSLNPLFGKIWNLNSAPKIK 785
Query: 62 HFIWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIW-------- 113
F+W + G + + + + C C E+L H CPL+ +W
Sbjct: 786 VFLWKVLKGAVAVEDRLRTRGVLIEDGCSMCPEKNETLNHILFQCPLARQVWALTPMQSP 845
Query: 114 ----DDLLFNKDTNFNSQDFNWWFKHYALALNGNIFVIT---CWVLWKARNHKVFNGNDW 166
D +F TN N N + L+ ++ ++ W+LWK RN ++F G
Sbjct: 846 NHGFGDSIF---TNVNHVIGNC----HNTELSPHLRYVSPWIIWILWKNRNKRLFEGIGS 898
Query: 167 VCLSHIYSLHSDVVAHI-SNDVI--QKPIK*VKWIPPLDNIIKVNVDGISLSNHGRSVFG 223
V LS + D + ++++I ++P K + WIPPL N +K N+ H +
Sbjct: 899 VSLSIVGKALEDCKEWLKAHELICSKEPTKDLTWIPPLMNELKCNIGIAWSKKHQMAGVS 958
Query: 224 GLIRNNNGDWLL 235
++RN G LL
Sbjct: 959 WVVRNWKGRVLL 970
>At2g27870 putative non-LTR retroelement reverse transcriptase
Length = 314
Score = 68.9 bits (167), Expect = 4e-12
Identities = 50/189 (26%), Positives = 82/189 (42%), Gaps = 18/189 (9%)
Query: 72 LPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWDDLL-FNKDTNFNSQDFN 130
L TNA R +HL+ C+ C + ++++H RDCP IW L+ K F +Q
Sbjct: 5 LMTNAERRRRHLSDSDICQICKGAEKTIIHILRDCPAMEGIWIRLVPAGKRREFFTQSLL 64
Query: 131 WWF-------KHYALALNGNIFVITCWVLWKARNHKVFNGNDWVC---------LSHIYS 174
W + + +F ++ W WK R +F D C L+ S
Sbjct: 65 EWLFANLGDRRKTCESTWSTLFALSIWWAWKWRCGNIFGVQD-KCRDRVRFLKDLARETS 123
Query: 175 LHSDVVAHISNDVIQKPIK*VKWIPPLDNIIKVNVDGISLSNHGRSVFGGLIRNNNGDWL 234
+ +V +S ++ + + W P + K+N DG S N G + GG++R+ G W
Sbjct: 124 MAHVIVRTLSGGHGERVERLIAWSKPEEGWWKLNTDGASRGNPGLASAGGVLRDEEGAWR 183
Query: 235 LGFFMDFVV 243
GF ++ V
Sbjct: 184 GGFALNIGV 192
>At2g31520 putative non-LTR retroelement reverse transcriptase
Length = 1524
Score = 67.0 bits (162), Expect = 2e-11
Identities = 67/255 (26%), Positives = 101/255 (39%), Gaps = 28/255 (10%)
Query: 3 IFIDESSQDIKI-WTHSPTGMYSVKSGYQWLTSNCLSFA-------GQTQQETWQWIWKL 54
I++ +S + KI W ++ TG Y+V+SGY LT + + G +T IW L
Sbjct: 1148 IYLAKSKKPDKIIWNYNTTGEYTVRSGYWLLTHDPSTNIPAINPPHGSIDLKTR--IWNL 1205
Query: 55 HLPENLKHFIWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIW- 113
+ LKHF+W A+ L T + + D C RC ES+ H CP + W
Sbjct: 1206 PIMPKLKHFLWRALSQALATTERLTTRGMRIDPICPRCHRENESINHALFTCPFATMAWW 1265
Query: 114 --------DDLLFNKDTNFNSQDFNWWFKHYALALNGNIFVITCWVLWKARNHKVFN--- 162
+ L+ N S N+ + + V W +WKARN+ VFN
Sbjct: 1266 LSDSSLIRNQLMSNDFEENISNILNFVQDTTMSDFHKLLPVWLIWRIWKARNNVVFNKFR 1325
Query: 163 -GNDWVCLSHIYSLHSDVVAHISNDVIQKPIK-----*VKWIPPLDNIIKVNVDGISLSN 216
LS H + A S+ P + ++W P +K N D
Sbjct: 1326 ESPSKTVLSAKAETHDWLNATQSHKKTPSPTRQIAENKIEWRNPPATYVKCNFDAGFDVQ 1385
Query: 217 HGRSVFGGLIRNNNG 231
+ G +IRN+ G
Sbjct: 1386 KLEATGGWIIRNHYG 1400
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.337 0.147 0.506
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,212,219
Number of Sequences: 26719
Number of extensions: 344013
Number of successful extensions: 1213
Number of sequences better than 10.0: 88
Number of HSP's better than 10.0 without gapping: 64
Number of HSP's successfully gapped in prelim test: 24
Number of HSP's that attempted gapping in prelim test: 1062
Number of HSP's gapped (non-prelim): 106
length of query: 364
length of database: 11,318,596
effective HSP length: 101
effective length of query: 263
effective length of database: 8,619,977
effective search space: 2267053951
effective search space used: 2267053951
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC141436.5


