

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141323.9 - phase: 0
(523 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
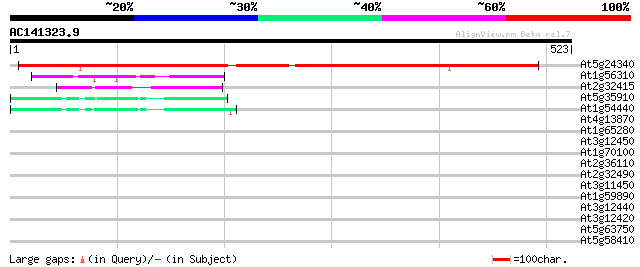
Score E
Sequences producing significant alignments: (bits) Value
At5g24340 unknown protein 582 e-166
At1g56310 hypothetical protein 80 3e-15
At2g32415 hypothetical protein 56 6e-08
At5g35910 nucleolar protein-like 52 1e-06
At1g54440 hypothetical protein 50 2e-06
At4g13870 Werner Syndrome-like exonuclease (WEX) 40 0.002
At1g65280 34 0.17
At3g12450 hypothetical protein 34 0.23
At1g70100 unknown protein (At1g70100) 33 0.39
At2g36110 hypothetical protein 32 0.86
At2g32490 hypothetical protein 32 0.86
At3g11450 putative cell division related protein 32 1.1
At1g59890 hypothetical protein 31 1.5
At3g12440 hypothetical protein 30 2.5
At3g12420 hypothetical protein 30 4.3
At5g63750 ARI-like RING zinc finger protein-like 28 9.5
At5g58410 unknown protein 28 9.5
>At5g24340 unknown protein
Length = 505
Score = 582 bits (1500), Expect = e-166
Identities = 298/492 (60%), Positives = 369/492 (74%), Gaps = 17/492 (3%)
Query: 9 LEIHFVTTTDSPEFTHLTRSLTQTSLVGLDAEWKPVRTHQNSFPTVSLLQIACQLG---D 65
L+I+ V++TDS EFTHL S T+++++ LDAEWKP ++ +SFPTV+LLQ+AC+L D
Sbjct: 5 LKIYLVSSTDSSEFTHLKWSFTRSTIIALDAEWKPQHSNTSSFPTVTLLQVACRLSHATD 64
Query: 66 DEVVFLLDLISLPLSSIWEPLREMLVSADILKLGFRFKQDLVYLSSTFCEQGCNPGFDKV 125
VFL+DL S+ L S+WE L +M VS D+LKLGFRFKQDLVYLSSTF + GC GF +V
Sbjct: 65 VSDVFLIDLSSIHLPSVWELLNDMFVSPDVLKLGFRFKQDLVYLSSTFTQHGCEGGFQEV 124
Query: 126 EPYLDITSVYNHLQFKKNGRIASKQNKSLSTICGELLGITLSKELQCSDWSQRPLTEEQM 185
+ YLDITS+YN+LQ K+ GR A K KSL+ IC E+L I+LSKELQCSDWS RPLTEEQ
Sbjct: 125 KQYLDITSIYNYLQHKRFGRKAPKDIKSLAAICKEMLDISLSKELQCSDWSYRPLTEEQK 184
Query: 186 TYAAMDAHCLLGIFKVFQATVAKEGELVNKTNILSIRSANLGLKELFRKHDTSDKVHSTQ 245
YAA DAHCLL IF VF+A LV + +R N+GL+E+ + D S K+ + +
Sbjct: 185 LYAATDAHCLLQIFDVFEA------HLVEGITVQDLRVINVGLQEILTESDYSSKIVTVK 238
Query: 246 FCEALAIVQATSCSDVVRVISSAGAVIQKSSCRDTKPMDEFLLKIVRKHGDRILLKESDR 305
C+A ++++ S + + A V+ + + +T PMDE LLKIVRK G+RILLKESD
Sbjct: 239 LCKATDVIRSMSENGQ----NIANGVVPRKTTLNTMPMDENLLKIVRKFGERILLKESDL 294
Query: 306 APKTSKKKRKKQLPINGIPKEKHLENFDEWQGTAPWDPLVGGDGFPKFLCDVMVEGLAKH 365
PK KKK ++++ + + K L +WQG PWD +GGDG PKFL DVMVEGLAKH
Sbjct: 295 LPKKLKKKTRRRVASSTMNTNKQLVCSADWQGPPPWDSSLGGDGCPKFLLDVMVEGLAKH 354
Query: 366 LRCVGIDAAVPSSKKPEPRELITKAQKEKRVLLTRDAKLLRHD----YQIYKVKSLLKNE 421
LRCVGIDAA+P SKKP+ REL+ +A KE RVLLTRD KLLRH +QIY+VKSLLKNE
Sbjct: 355 LRCVGIDAAIPHSKKPDSRELLDQAFKENRVLLTRDTKLLRHQDLAKHQIYRVKSLLKNE 414
Query: 422 QLLEIIETFQLNINEDQLMSRCTKCNGRFIQKPLSTEEAIEAAKGFQKIPNCLFNKNLEF 481
QLLE+IETFQL I+ +QLMSRCTKCNG+FIQKPLS EEAIEAAKGFQ+IPNCLFNKNLEF
Sbjct: 415 QLLEVIETFQLKISGNQLMSRCTKCNGKFIQKPLSIEEAIEAAKGFQRIPNCLFNKNLEF 474
Query: 482 WQCMDCHQLYWE 493
WQCM+CHQLYWE
Sbjct: 475 WQCMNCHQLYWE 486
>At1g56310 hypothetical protein
Length = 582
Score = 80.1 bits (196), Expect = 3e-15
Identities = 62/197 (31%), Positives = 98/197 (49%), Gaps = 36/197 (18%)
Query: 21 EFTHLTRSLTQTSLVGLDAEWKPVRTHQNSFPTVSLLQIACQLGDDEVVFLLDLISL--P 78
E T L +VG+D EWKP + VS++QI G D +F+LDLI L
Sbjct: 367 ELRKATSFLEGCRVVGIDCEWKPNYIKGSKQNKVSIMQI----GSDTKIFILDLIKLYND 422
Query: 79 LSSIWEP-LREMLVSADILKL--------------GFRFKQDLVYLSSTFCEQGCNPGFD 123
S I + L +L S LKL G+ F+ D+ L+ ++ + C F+
Sbjct: 423 ASEILDNCLSHILQSKSTLKLVSLTEDYPDHKLSSGYNFQCDIKQLALSYGDLKC---FE 479
Query: 124 KVEPYLDITSVYNHLQFKKNGRIASKQNKSLSTICGELLGITLSKELQCSDWSQRPLTEE 183
+ + LDI +V+N + L+ + ++LG++L+K + SDW QRPL++
Sbjct: 480 RYDMLLDIQNVFN------------EPFGGLAGLTKKILGVSLNKTRRNSDWEQRPLSQN 527
Query: 184 QMTYAAMDAHCLLGIFK 200
Q+ YAA+DA L+ IF+
Sbjct: 528 QLEYAALDAAVLIHIFR 544
>At2g32415 hypothetical protein
Length = 237
Score = 55.8 bits (133), Expect = 6e-08
Identities = 45/155 (29%), Positives = 66/155 (42%), Gaps = 19/155 (12%)
Query: 44 VRTHQNSFPTVSLLQIACQLGDDEVVFLLDLISLPLSSIWEPLREMLVSADILKLGFRFK 103
V T Q+S + Q+ E FL+D I+L + LR + +I K+
Sbjct: 79 VDTEQHSLRSFLGFTALIQISTHEEDFLVDTIAL--HDVMSILRPVFSDPNICKVFHGAD 136
Query: 104 QDLVYLSSTFCEQGCNPGFDKVEPYLDITSVYNHLQFKKNGRIASKQNKSLSTICGELLG 163
D+++L F V N K + SK +SL+ + + G
Sbjct: 137 NDVIWLQRDFH-----------------IYVVNMFDTAKACEVLSKPQRSLAYLLETVCG 179
Query: 164 ITLSKELQCSDWSQRPLTEEQMTYAAMDAHCLLGI 198
+ +K LQ DW QRPL+EE + YA DAH LL I
Sbjct: 180 VATNKLLQREDWRQRPLSEEMVRYARTDAHYLLYI 214
>At5g35910 nucleolar protein-like
Length = 834
Score = 51.6 bits (122), Expect = 1e-06
Identities = 52/204 (25%), Positives = 81/204 (39%), Gaps = 26/204 (12%)
Query: 1 MDPPNQKPLE-IHFVTTTDSPEFTHLTRSLTQTSLVGLDAEWKPVRTHQNSFPTVSLLQI 59
M+P PLE F + + L L +D E R+ Q L+QI
Sbjct: 210 MEPVKPLPLEQTPFKFVQEVKDLKELVAKLRSVEEFAVDLEHNQYRSFQG---LTCLMQI 266
Query: 60 ACQLGDDEVVFLLDLISLPLSSIWEPLREMLVSADILKLGFRFKQDLVYLSSTFCEQGCN 119
+ + D +++D L + I LRE+ K+ +D+++L F CN
Sbjct: 267 STRTED----YIVDTFKLRIH-IGPYLREIFKDPKKKKVMHGADRDIIWLQRDFGIYVCN 321
Query: 120 PGFDKVEPYLDITSVYNHLQFKKNGRIASKQNKSLSTICGELLGITLSKELQCSDWSQRP 179
FD + R+ + + SL + G+T +KE Q +DW RP
Sbjct: 322 L-FDTGQA----------------SRVLNLERNSLEFLLQHFCGVTANKEYQNADWRIRP 364
Query: 180 LTEEQMTYAAMDAHCLLGIFKVFQ 203
L EE YA D H LL I+ + +
Sbjct: 365 LPEEMTRYAREDTHYLLYIYDLIK 388
>At1g54440 hypothetical protein
Length = 738
Score = 50.4 bits (119), Expect = 2e-06
Identities = 56/215 (26%), Positives = 84/215 (39%), Gaps = 29/215 (13%)
Query: 1 MDPPNQKPLE-IHFVTTTDSPEFTHLTRSLTQTSLVGLDAEWKPVRTHQNSFPTVSLLQI 59
M P PLE F + + L +L +D E RT Q L+QI
Sbjct: 215 MRPVKPLPLEETPFKLVEEVKDLEDLAAALQSVEEFAVDLEHNQYRTFQG---LTCLMQI 271
Query: 60 ACQLGDDEVVFLLDLISLPLSSIWEPLREMLVSADILKLGFRFKQDLVYLSSTFCEQGCN 119
+ + D +++D+ L I LRE+ K+ +D+++L F CN
Sbjct: 272 STRTED----YIVDIFKL-WDHIGPYLRELFKDPKKKKVIHGADRDIIWLQRDFGIYVCN 326
Query: 120 PGFDKVEPYLDITSVYNHLQFKKNGRIASKQNKSLSTICGELLGITLSKELQCSDWSQRP 179
FD + R+ + SL + G+ +KE Q +DW RP
Sbjct: 327 L-FDTGQA----------------SRVLKLERNSLEFLLKHYCGVAANKEYQKADWRIRP 369
Query: 180 LTEEQMTYAAMDAHCLLGIFKVFQA---TVAKEGE 211
L + YA D H LL I+ V + T+AKE E
Sbjct: 370 LPDVMKRYAREDTHYLLYIYDVMRMELHTMAKEDE 404
>At4g13870 Werner Syndrome-like exonuclease (WEX)
Length = 288
Score = 40.4 bits (93), Expect = 0.002
Identities = 41/172 (23%), Positives = 78/172 (44%), Gaps = 21/172 (12%)
Query: 33 SLVGLDAEWKPVRTHQNSFPTVSLLQIACQLGDDEVVFLLDLISLPLSSIWEPLREMLVS 92
+ VGLD EW+P V+ +QI +V+ + S I + L+ ++
Sbjct: 128 AFVGLDIEWRPSFRKGVLPGKVATVQICVDSNYCDVMHIFH------SGIPQSLQHLIED 181
Query: 93 ADILKLGFRFKQDLVYLSSTFCEQGCNPGFDKVEPYLDITSVYNH-LQFKKNGRIASKQN 151
+ ++K+G D V L F + G + ++ D++ + N + K +AS
Sbjct: 182 STLVKVGIGIDGDSVKL---FHDYGVS-----IKDVEDLSDLANQKIGGDKKWGLASLTE 233
Query: 152 KSLSTICGELLGITLSKELQCSDWSQRPLTEEQMTYAAMDAHCLLGIFKVFQ 203
+ +C ELL ++ +W PL+++Q+ YAA DA+ ++KV +
Sbjct: 234 ---TLVCKELLK---PNRIRLGNWEFYPLSKQQLQYAATDAYASWHLYKVLK 279
>At1g65280
Length = 605
Score = 34.3 bits (77), Expect = 0.17
Identities = 27/80 (33%), Positives = 35/80 (43%), Gaps = 18/80 (22%)
Query: 290 IVRKHGDRILLKESDRAPKTSKKKRKKQLPINGIPKEKHLENFDEWQGTAPWDP------ 343
+V KH + D + +S+ K+KK+L + EK DEW G PW P
Sbjct: 525 LVEKHRE-------DSSSSSSRLKKKKKLSSSKEKTEK-----DEWVGKHPWKPWDREND 572
Query: 344 LVGGDGFPKFLCDVMVEGLA 363
L G K D M EGLA
Sbjct: 573 LTAGRQNVKLDADGMAEGLA 592
>At3g12450 hypothetical protein
Length = 171
Score = 33.9 bits (76), Expect = 0.23
Identities = 13/31 (41%), Positives = 21/31 (66%)
Query: 163 GITLSKELQCSDWSQRPLTEEQMTYAAMDAH 193
G+ L +E+ SDWS L ++Q+ A++DAH
Sbjct: 127 GVRLDREISMSDWSAYELCDDQILQASIDAH 157
>At1g70100 unknown protein (At1g70100)
Length = 467
Score = 33.1 bits (74), Expect = 0.39
Identities = 28/99 (28%), Positives = 43/99 (43%), Gaps = 1/99 (1%)
Query: 213 VNKTNILSIRSANLGLKELFRKHDTSDKVHSTQFCEALA-IVQATSCSDVVRVISSAGAV 271
VN+T + KE DT D + + + E L IVQ V VI V
Sbjct: 148 VNETCNHKPLEETMDFKECRSSVDTGDDLSTLKLEEKLEEIVQVEDKEKVEEVICMKEEV 207
Query: 272 IQKSSCRDTKPMDEFLLKIVRKHGDRILLKESDRAPKTS 310
+ +DT M E L+K +K D +K++D+ +T+
Sbjct: 208 KEDVPSKDTGEMSETLMKETKKEKDHNPIKKTDKNVRTN 246
>At2g36110 hypothetical protein
Length = 239
Score = 32.0 bits (71), Expect = 0.86
Identities = 41/169 (24%), Positives = 68/169 (39%), Gaps = 30/169 (17%)
Query: 34 LVGLDAEWKPVRTHQNSFPTVSLLQIACQLGDDEVVFLLDLISLPLSSIWEPLREMLVSA 93
+VGLD +W P S P +LQ+ +G+ ++ L I E LR L
Sbjct: 57 VVGLDVQWTP----GGSDPPPDILQLC--VGNRCLIIQLS----HCKRIPEVLRSFLEDE 106
Query: 94 DILKLGFRFKQDLVYLSSTFCEQGCNPGFDKVEPYLDITSVYNHLQFKKN--GRIASKQN 151
I +G QD QG K+E + ++ L + R+ +
Sbjct: 107 TITFVGVWNSQD----------QG------KLERFRHQLEIWRLLDIRHYLPTRLLNSSF 150
Query: 152 KSLSTICGELLGITLSKELQCSDWSQRPLTEEQMTYAAMDAH--CLLGI 198
+ + C G+ KE+ S+W R L+ +Q+ A+ D + C LG+
Sbjct: 151 EKIVEECLGYKGVRKDKEICMSNWGARSLSHDQIVQASDDVYVCCKLGV 199
>At2g32490 hypothetical protein
Length = 217
Score = 32.0 bits (71), Expect = 0.86
Identities = 15/38 (39%), Positives = 23/38 (60%), Gaps = 2/38 (5%)
Query: 163 GITLSKELQCSDWSQRPLTEEQMTYAAMDAH--CLLGI 198
G+T KEL S+W R L+ +Q+ A+ D + C LG+
Sbjct: 171 GVTKDKELCMSNWGARSLSHDQIVQASHDVYVCCKLGV 208
>At3g11450 putative cell division related protein
Length = 663
Score = 31.6 bits (70), Expect = 1.1
Identities = 27/115 (23%), Positives = 51/115 (43%), Gaps = 14/115 (12%)
Query: 356 DVMVEGLAKHLRCVGIDAAVPSSKKPEPRELITKAQKEKRVLLTRDAKLLRHDYQIYKVK 415
D ++ K I A +K E + ++ ++++ R+ KLLR + +
Sbjct: 330 DAKIQAKKKQEEDAAIAAEEEKRRKEEEEKRAAESAQQQKKTKEREKKLLRKERNRLRTL 389
Query: 416 SL-LKNEQLLEI----IETFQLNINEDQLMSRCTKCNGRFIQKPLSTEEAIEAAK 465
S L ++LL+I IE +++N +QL + C K + +E +E AK
Sbjct: 390 SAPLVAQRLLDISEEDIENLCMSLNTEQLQNLCDK---------MGNKEGLELAK 435
>At1g59890 hypothetical protein
Length = 1125
Score = 31.2 bits (69), Expect = 1.5
Identities = 17/72 (23%), Positives = 31/72 (42%)
Query: 344 LVGGDGFPKFLCDVMVEGLAKHLRCVGIDAAVPSSKKPEPRELITKAQKEKRVLLTRDAK 403
++G + F D +V+ KHL V D + E K + ++ +A+
Sbjct: 903 IIGAQSYVLFTLDKLVQKFVKHLHAVAADETDTKLLQLYAYENYRKPGRFFDIVYHENAR 962
Query: 404 LLRHDYQIYKVK 415
L HD IY+++
Sbjct: 963 ALLHDQNIYRIE 974
>At3g12440 hypothetical protein
Length = 353
Score = 30.4 bits (67), Expect = 2.5
Identities = 15/47 (31%), Positives = 26/47 (54%), Gaps = 3/47 (6%)
Query: 150 QNKSLSTICGELLG---ITLSKELQCSDWSQRPLTEEQMTYAAMDAH 193
+ +S + GE LG + L K + SDWS L+ +Q+ A++D +
Sbjct: 293 KGRSFEVVVGECLGYRGVRLEKAIGMSDWSVYDLSYDQILQASIDVY 339
>At3g12420 hypothetical protein
Length = 308
Score = 29.6 bits (65), Expect = 4.3
Identities = 19/62 (30%), Positives = 29/62 (46%), Gaps = 7/62 (11%)
Query: 144 GRIASKQNKSLSTICGELLG-----ITLSKELQCSDWSQRPLTEEQMTYAAMDAHC--LL 196
G++ + S I E LG + L + SDWS L +Q+ A++D H LL
Sbjct: 243 GKLLDIRGCSFEEIVDEFLGSPGSGLRLDPAISMSDWSVHVLDRDQVLQASIDVHVCHLL 302
Query: 197 GI 198
G+
Sbjct: 303 GV 304
>At5g63750 ARI-like RING zinc finger protein-like
Length = 536
Score = 28.5 bits (62), Expect = 9.5
Identities = 13/27 (48%), Positives = 15/27 (55%)
Query: 446 CNGRFIQKPLSTEEAIEAAKGFQKIPN 472
CNGRF K + EEA + GF K N
Sbjct: 299 CNGRFCWKCMQPEEAHKTESGFYKFCN 325
>At5g58410 unknown protein
Length = 1873
Score = 28.5 bits (62), Expect = 9.5
Identities = 17/70 (24%), Positives = 31/70 (44%), Gaps = 12/70 (17%)
Query: 320 INGIPKEKHLENFDEWQGTAP--WDPLVGGDGFPKFLCDVMVEGLAKHLRCVGIDAAVPS 377
+N + K +E D + A WDP+ D ++EG+ + C G+ A+ +
Sbjct: 1376 LNALVNHKSVEGEDNLKSLADETWDPVK----------DRILEGVLRLFFCTGLTEAIAA 1425
Query: 378 SKKPEPRELI 387
S PE ++
Sbjct: 1426 SYSPEAASIV 1435
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.136 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,932,497
Number of Sequences: 26719
Number of extensions: 512320
Number of successful extensions: 1450
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 1433
Number of HSP's gapped (non-prelim): 17
length of query: 523
length of database: 11,318,596
effective HSP length: 104
effective length of query: 419
effective length of database: 8,539,820
effective search space: 3578184580
effective search space used: 3578184580
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC141323.9


