

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141322.10 + phase: 0 /pseudo
(409 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
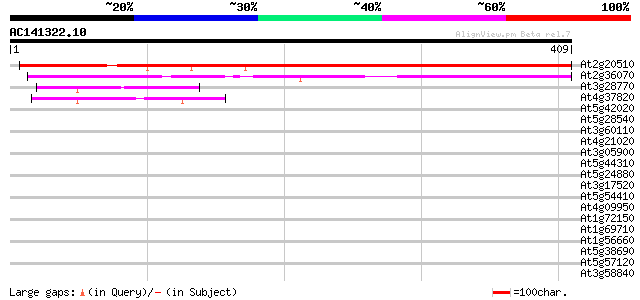
Score E
Sequences producing significant alignments: (bits) Value
At2g20510 putative mitochondrial inner membrane translocating pr... 320 1e-87
At2g36070 hypothetical protein 291 6e-79
At3g28770 hypothetical protein 42 6e-04
At4g37820 unknown protein 42 8e-04
At5g42020 luminal binding protein (dbj|BAA13948.1) 39 0.007
At5g28540 luminal binding protein 39 0.007
At3g60110 unknown protein 39 0.007
At4g21020 unknown protein (At4g21020) 38 0.012
At3g05900 unknown protein 37 0.015
At5g44310 late embryogenesis abundant protein-like 37 0.026
At5g24880 glutamic acid-rich protein 37 0.026
At3g17520 unknown protein 36 0.044
At5g54410 unknown protein 35 0.057
At4g09950 AIG1-like protein 35 0.075
At1g72150 putative cytosolic factor protein 35 0.075
At1g69710 putative regulator of chromosome condensation 35 0.075
At1g56660 hypothetical protein 35 0.075
At5g38690 unknown protein 35 0.098
At5g57120 unknown protein 34 0.13
At3g58840 unknown protein 34 0.13
>At2g20510 putative mitochondrial inner membrane translocating
protein
Length = 533
Score = 320 bits (819), Expect = 1e-87
Identities = 176/418 (42%), Positives = 252/418 (60%), Gaps = 23/418 (5%)
Query: 8 VNSVDRRGYSVFNEFSKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQ 67
+ R +SVF EFSK ++ E NPEF+++VKELKE+ EE KG+ E LK +TKQTTE+
Sbjct: 105 IGYTSHRRFSVFTEFSKNIRGEAHSNPEFERTVKELKERTEEFKGVTEDLKVRTKQTTEK 164
Query: 68 LYRQFDSVWKEAEAAAKKVSHNVKEKISAATD-----FSTKQNADAKQGSQKSPEEEKNE 122
LY+Q A+A +VS +VK+K+SAA++ F + +A+ S + E
Sbjct: 165 LYKQ-------ADALTVQVSSSVKDKLSAASEEVKESFKLGKEENAESASSSGTRASQGE 217
Query: 123 ESPSGNASE--SLFGKFKSTFSSPMVSTSFQKLKDAKLWT*PRKVMTY*R--------RS 172
+ SG+ E + F KFKS+ SSP VS F +LK+AK + ++ + + R
Sbjct: 218 KQQSGSTEELHTFFAKFKSSLSSPKVSEVFYRLKEAKPFDIVKQALDIVKDELRGNPSRK 277
Query: 173 *VVTHLRDSVFVSPAK*VQKLILLLCPLTNLGGVRRLMSSGIRRHPVSKNFVKYIDPVKT 232
+ H F ++++ + L ++ PV K +PV
Sbjct: 278 KFLEHTPPPPFTGERSMRTEMVVTQTKQSKLQQKWESFREKMQGSPVFKRLSGMSEPVVN 337
Query: 233 TSRDIVDDVRDIIDRIDNPIIDKIQSNMSRTFQETDAALTYREIRKRDPKFSLPDFVGEV 292
S++I +DVR+I + DNPI+ KIQ + +ETD+A TY+EIR RDP FSLPDF E+
Sbjct: 338 KSQEIAEDVREIWETSDNPIVHKIQDMNEKFLKETDSASTYKEIRSRDPSFSLPDFAAEI 397
Query: 293 QEAIKPVLNAYIKGDFETLKKYCAPQLIERCKAEHGAYKDRGIFYDNKILHISDADVREV 352
+E IKPVLNAY +GD ETLKKYC+ ++IERC AE AY+ G+ +DNK+LHIS+ V
Sbjct: 398 EEVIKPVLNAYSEGDVETLKKYCSKEVIERCTAERTAYQTHGVLFDNKLLHISEVSVSVT 457
Query: 353 KILESSPFIIVVFQTQQIHCVRDRNGEITEGGKDTIHSVYYLWAL-QMDSEDHAEDGI 409
K++ SP II FQTQ+I+CVRD NGEI EGG+DTIH+VY+ WA+ Q+++ + ED I
Sbjct: 458 KMMGDSPIIIAKFQTQEIYCVRDENGEIQEGGQDTIHTVYHEWAMQQVETTELGEDAI 515
>At2g36070 hypothetical protein
Length = 499
Score = 291 bits (744), Expect = 6e-79
Identities = 169/414 (40%), Positives = 242/414 (57%), Gaps = 61/414 (14%)
Query: 14 RGYSVFNEFSKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFD 73
R +SVF+EFSKK++ E NPEFQK+VKE KE+AEEL+G+KE LK +TKQTTE+LY+Q
Sbjct: 111 RRFSVFSEFSKKIRGEADSNPEFQKTVKEFKERAEELQGVKEDLKVRTKQTTEKLYKQGQ 170
Query: 74 SVWKEAEAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSGNASESL 133
VW EAE+ AKKV + K + + ++ +G E++ ++S S ++
Sbjct: 171 GVWTEAESVAKKVKESFKLGKEESAESASSSGTGTTEG------EKQQQQSGSTEEQDTF 224
Query: 134 FGKFKSTFSSPMVSTSFQKLKDAKLWT*PRKVMTY*RRS*VVTHLRDSVFVSPAK*VQKL 193
FGKFKS+ SSP +S +F K D +K + ++D + +P+K
Sbjct: 225 FGKFKSSISSPKLSEAFHKPLDFA-----KKGLDI---------VKDELRGNPSKRKHLE 270
Query: 194 ILLLCPLTNLGGVRRLM-----------------SSGIRRHPVSKNFVKYIDPVKTTSRD 236
P T R M ++ +PV K +PV S++
Sbjct: 271 YTPPPPFTGERSTRTEMVIMPTKQSKWQKKWESLREKMQGYPVFKRLSGMSEPVVNKSQE 330
Query: 237 IVDDVRDIIDRIDNPIIDKIQSNMSRTFQETDAALTYREIRKRDPKFSLPDFVGEVQEAI 296
I +DVR+ + DNPI+ KIQ + FSLPDFV E+QEAI
Sbjct: 331 IAEDVREKWETSDNPIVHKIQES-----------------------FSLPDFVSEIQEAI 367
Query: 297 KPVLNAYIKGDFETLKKYCAPQLIERCKAEHGAYKDRGIFYDNKILHISDADVREVKILE 356
+PVLNAY KGD +TLKKYC+ +LIERC AEH A+ +G F+D+K+LH+S+ D++E K++
Sbjct: 368 RPVLNAYSKGDAKTLKKYCSKELIERCTAEHRAFTSQGYFFDHKLLHVSEVDIQETKMMG 427
Query: 357 SSPFIIVVFQTQQIHCVRDRNGEITEGGKDTIHSVYYLWAL-QMDSEDHAEDGI 409
++P IIV FQTQ+I CVRD++G+I EGG+DTIH+VYY WA+ Q+D+ + ED I
Sbjct: 428 TTPVIIVRFQTQEIFCVRDQDGKIKEGGQDTIHTVYYDWAMQQVDAAELGEDAI 481
>At3g28770 hypothetical protein
Length = 2081
Score = 42.0 bits (97), Expect = 6e-04
Identities = 31/124 (25%), Positives = 58/124 (46%), Gaps = 7/124 (5%)
Query: 20 NEFSKKVKDETVKNPEFQKSVKELKEKAE-----ELKGIKEGLKEKTKQTTEQLYRQFDS 74
NE K + VK +K KE +EK+E K K + +K K++++ ++ +
Sbjct: 1136 NEKKKSQHVKLVKKESDKKEKKENEEKSETKEIESSKSQKNEVDKKEKKSSKDQQKKKEK 1195
Query: 75 VWKEAEAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSGNASESLF 134
KE+E KK+ N +++ + K+ + K+ K +++KN SG ES+
Sbjct: 1196 EMKESEE--KKLKKNEEDRKKQTSVEENKKQKETKKEKNKPKDDKKNTTKQSGGKKESME 1253
Query: 135 GKFK 138
+ K
Sbjct: 1254 SESK 1257
Score = 36.6 bits (83), Expect = 0.026
Identities = 39/148 (26%), Positives = 65/148 (43%), Gaps = 11/148 (7%)
Query: 12 DRRGYSVFNEFSKKV--KDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLY 69
D++ Y V NE K+ K ET K+ +LKE+ ++ K KE +K ++ Y
Sbjct: 962 DKKEY-VNNELKKQEDNKKETTKSEN-----SKLKEENKDNKEKKESEDSASKNREKKEY 1015
Query: 70 RQFDSVWKEAEAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSGNA 129
+ S KE KK S + K + D +++ K+ S+ ++K EE+
Sbjct: 1016 EEKKSKTKEEAKKEKKKSQDKKRE---EKDSEERKSKKEKEESRDLKAKKKEEETKEKKE 1072
Query: 130 SESLFGKFKSTFSSPMVSTSFQKLKDAK 157
SE+ K K + S +K +D K
Sbjct: 1073 SENHKSKKKEDKKEHEDNKSMKKEEDKK 1100
Score = 34.7 bits (78), Expect = 0.098
Identities = 31/113 (27%), Positives = 49/113 (42%), Gaps = 12/113 (10%)
Query: 19 FNEFSKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKE 78
+ E K K+E K + + K ++ +EE K KE KE++ R + KE
Sbjct: 1015 YEEKKSKTKEEAKKEKKKSQDKKREEKDSEERKSKKE--KEES--------RDLKAKKKE 1064
Query: 79 AEAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSGNASE 131
E KK S N K K D ++ + + + E++K+EES S E
Sbjct: 1065 EETKEKKESENHKSK--KKEDKKEHEDNKSMKKEEDKKEKKKHEESKSRKKEE 1115
Score = 33.5 bits (75), Expect = 0.22
Identities = 39/157 (24%), Positives = 60/157 (37%), Gaps = 23/157 (14%)
Query: 20 NEFSKKVKDETVKNPEFQKSVKELKEKAEELKG--IKEGLKEKTKQTTEQLYRQFDS--- 74
+E K K E K E KS+K+ ++K E+ K K KE+ K+ E+L Q +
Sbjct: 1073 SENHKSKKKEDKKEHEDNKSMKKEEDKKEKKKHEESKSRKKEEDKKDMEKLEDQNSNKKK 1132
Query: 75 --------------VWKEAEAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEK 120
V KE++ KK + E + S K D K+ +++K
Sbjct: 1133 EDKNEKKKSQHVKLVKKESDKKEKKENEEKSETKEIESSKSQKNEVDKKEKKSSKDQQKK 1192
Query: 121 NEESPSGNASESLFGKFKSTFSSPMVSTSFQKLKDAK 157
E+ ES K K TS ++ K K
Sbjct: 1193 KEK----EMKESEEKKLKKNEEDRKKQTSVEENKKQK 1225
Score = 33.1 bits (74), Expect = 0.28
Identities = 36/138 (26%), Positives = 63/138 (45%), Gaps = 15/138 (10%)
Query: 21 EFSKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKE-KTKQTTEQLYRQFDSVWKEA 79
E +K+ K+E+ + K + G KE K+ K ++ E + +S+ K+
Sbjct: 822 EDNKEDKEESKDYQSVEAKEKNENGGVDTNVGNKEDSKDLKDDRSVEVKANKEESMKKKR 881
Query: 80 EAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSGNASESLFGKFKS 139
E +V N K DF+ + D ++GS +S + +K+E+ GN E+ K
Sbjct: 882 E----EVQRNDKSSTKEVRDFANNMDIDVQKGSGESVKYKKDEKK-EGNKEEN-----KD 931
Query: 140 TFSSPMVSTSFQKLKDAK 157
T + ++S QK KD K
Sbjct: 932 TIN----TSSKQKGKDKK 945
Score = 33.1 bits (74), Expect = 0.28
Identities = 27/108 (25%), Positives = 47/108 (43%), Gaps = 10/108 (9%)
Query: 27 KDETVKNPEFQKSVKE--------LKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKE 78
K K+ E +KS KE K+K EE K KE K+K+ ++ + + K+
Sbjct: 1036 KKREEKDSEERKSKKEKEESRDLKAKKKEEETKEKKESENHKSKKKEDKKEHEDNKSMKK 1095
Query: 79 AEAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEESPS 126
E +K H + D + + + ++K +E+KNE+ S
Sbjct: 1096 EEDKKEKKKHEESKSRKKEEDKKDMEKLEDQNSNKK--KEDKNEKKKS 1141
Score = 30.4 bits (67), Expect = 1.8
Identities = 27/112 (24%), Positives = 54/112 (48%), Gaps = 7/112 (6%)
Query: 21 EFSKKVKDETVKNPEFQKSVKELKEKAEELKG--IKEGLKEKTKQTTEQLYRQFDS---V 75
E +K+ +T+ QK K+ K+K +E K +K+ ++K + +L +Q D+
Sbjct: 923 EGNKEENKDTINTSSKQKG-KDKKKKKKESKNSNMKKKEEDKKEYVNNELKKQEDNKKET 981
Query: 76 WKEAEAAAKKVSHNVKEKISAATDFS-TKQNADAKQGSQKSPEEEKNEESPS 126
K + K+ + + KEK + S ++ + ++ K+ EE K E+ S
Sbjct: 982 TKSENSKLKEENKDNKEKKESEDSASKNREKKEYEEKKSKTKEEAKKEKKKS 1033
Score = 30.0 bits (66), Expect = 2.4
Identities = 26/101 (25%), Positives = 47/101 (45%), Gaps = 4/101 (3%)
Query: 24 KKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAA 83
+ VK+E ++N E +K +K+ E + L+EK +QT + +S K +
Sbjct: 595 ESVKNENLENKEDKKELKD-DESVGAKTNNETSLEEKREQTQKGHDNSINS--KIVDNKG 651
Query: 84 KKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEES 124
N KEK D + N ++K+ ++ E +KN+ S
Sbjct: 652 GNADSN-KEKEVHVGDSTNDNNMESKEDTKSEVEVKKNDGS 691
Score = 29.3 bits (64), Expect = 4.1
Identities = 25/121 (20%), Positives = 51/121 (41%), Gaps = 8/121 (6%)
Query: 24 KKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDS------VWK 77
+ ++ E+ + QKS + ++E K E L + Q Q DS +
Sbjct: 1361 ESMESESKEAENQQKSQATTQADSDESKN--EILMQADSQADSHSDSQADSDESKNEILM 1418
Query: 78 EAEAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSGNASESLFGKF 137
+A++ A +N +++ + K+ + K+ K +++KN SG ES+ +
Sbjct: 1419 QADSQATTQRNNEEDRKKQTSVAENKKQKETKEEKNKPKDDKKNTTEQSGGKKESMESES 1478
Query: 138 K 138
K
Sbjct: 1479 K 1479
Score = 29.3 bits (64), Expect = 4.1
Identities = 25/121 (20%), Positives = 51/121 (41%), Gaps = 8/121 (6%)
Query: 24 KKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDS------VWK 77
+ ++ E+ + QKS + ++E K E L + Q Q DS +
Sbjct: 1250 ESMESESKEAENQQKSQATTQADSDESKN--EILMQADSQADSHSDSQADSDESKNEILM 1307
Query: 78 EAEAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSGNASESLFGKF 137
+A++ A +N +++ + K+ + K+ K +++KN SG ES+ +
Sbjct: 1308 QADSQATTQRNNEEDRKKQTSVAENKKQKETKEEKNKPKDDKKNTTKQSGGKKESMESES 1367
Query: 138 K 138
K
Sbjct: 1368 K 1368
Score = 28.9 bits (63), Expect = 5.4
Identities = 29/117 (24%), Positives = 51/117 (42%), Gaps = 4/117 (3%)
Query: 24 KKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAA 83
+K E+VK ++K K+ K E I K+K K ++ +S K+ E
Sbjct: 907 QKGSGESVK---YKKDEKKEGNKEENKDTINTSSKQKGKDKKKKKKESKNSNMKKKEEDK 963
Query: 84 KKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEK-NEESPSGNASESLFGKFKS 139
K+ +N +K +TK + K +E+K +E+S S N + + + KS
Sbjct: 964 KEYVNNELKKQEDNKKETTKSENSKLKEENKDNKEKKESEDSASKNREKKEYEEKKS 1020
Score = 28.1 bits (61), Expect = 9.1
Identities = 43/169 (25%), Positives = 68/169 (39%), Gaps = 37/169 (21%)
Query: 20 NEFSKKVKDETVKNPEFQKSVKELKEKAEELK--------GIKEGLKEKTK----QTTEQ 67
NE +K + +N + QK KE K K ++ K G KE ++ ++K Q Q
Sbjct: 1319 NEEDRKKQTSVAENKK-QKETKEEKNKPKDDKKNTTKQSGGKKESMESESKEAENQQKSQ 1377
Query: 68 LYRQFDS------VWKEAEAAAKKVSHNV------KEKISAATDFST-----------KQ 104
Q DS + +A++ A S + K +I D KQ
Sbjct: 1378 ATTQADSDESKNEILMQADSQADSHSDSQADSDESKNEILMQADSQATTQRNNEEDRKKQ 1437
Query: 105 NADAKQGSQKSPEEEKNE-ESPSGNASESLFGKFKSTFSSPMVSTSFQK 152
+ A+ QK +EEKN+ + N +E GK +S S + + QK
Sbjct: 1438 TSVAENKKQKETKEEKNKPKDDKKNTTEQSGGKKESMESESKEAENQQK 1486
>At4g37820 unknown protein
Length = 532
Score = 41.6 bits (96), Expect = 8e-04
Identities = 36/148 (24%), Positives = 68/148 (45%), Gaps = 12/148 (8%)
Query: 17 SVFNEFSKKVKDETVKNPEFQKSVKELKEKAE-----ELKGIKEGLKEKTKQTTEQLYRQ 71
S E K KD + E ++ E K+K E E K + +EK ++++ ++
Sbjct: 310 SKVKESGKNEKDASSSQDESKEEKPERKKKEESSSQGEGKEEEPEKREKEDSSSQEESKE 369
Query: 72 FDSVWKEAEAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEES--PSGNA 129
+ KE EA++ + + +KE T+ K+ + +++G++ E+K+ ES
Sbjct: 370 EEPENKEKEASSSQEENEIKE-----TEIKEKEESSSQEGNENKETEKKSSESQRKENTN 424
Query: 130 SESLFGKFKSTFSSPMVSTSFQKLKDAK 157
SE + +ST SS QK ++K
Sbjct: 425 SEKKIEQVESTDSSNTQKGDEQKTDESK 452
Score = 34.7 bits (78), Expect = 0.098
Identities = 38/150 (25%), Positives = 69/150 (45%), Gaps = 16/150 (10%)
Query: 21 EFSKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAE 80
E SK+ + E K E S +E + K E+K +E ++ + E + +S KE
Sbjct: 365 EESKEEEPEN-KEKEASSSQEENEIKETEIKEKEESSSQEGNENKETEKKSSESQRKENT 423
Query: 81 AAAKKV-------SHNVKEKISAATDFSTKQ------NADAKQGSQKSPEEEKNEESPSG 127
+ KK+ S N ++ TD S ++ N + + S K+ E+K E + +G
Sbjct: 424 NSEKKIEQVESTDSSNTQKGDEQKTDESKRESGNDTSNKETEDDSSKTESEKKEENNRNG 483
Query: 128 NASESLFGKFKSTFSSPMVSTSFQKLKDAK 157
E+ + + T S+ +S + Q +KDA+
Sbjct: 484 ETEETQ-NEQEQTKSALEISHT-QDVKDAR 511
Score = 28.1 bits (61), Expect = 9.1
Identities = 28/125 (22%), Positives = 48/125 (38%), Gaps = 11/125 (8%)
Query: 17 SVFNEFSKKVKDETVKNPEFQKSVKELKEKAEELKGIKEG---LKEKTKQTTEQLYRQFD 73
SV E S + + E Q++ EL K E KG + L E T+
Sbjct: 217 SVIKEVSLNTTENGSDDGEQQETKSELDSKTGE-KGFSDSNGELPETNLSTSNATETTES 275
Query: 74 SVWKEAEAAAKKVSH-------NVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEESPS 126
S E+ ++ K + + KEK+ ++ + S + + + S ++E EE P
Sbjct: 276 SGSDESGSSGKSTGYQQTKNEEDEKEKVQSSEEESKVKESGKNEKDASSSQDESKEEKPE 335
Query: 127 GNASE 131
E
Sbjct: 336 RKKKE 340
>At5g42020 luminal binding protein (dbj|BAA13948.1)
Length = 668
Score = 38.5 bits (88), Expect = 0.007
Identities = 30/89 (33%), Positives = 45/89 (49%), Gaps = 3/89 (3%)
Query: 35 EFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAAKKVSHNVKEKI 94
E + VKE +E AEE K +KE + + T +Y + V + + A K+ + KEKI
Sbjct: 545 EIDRMVKEAEEFAEEDKKVKEKIDARNALET-YVYNMKNQV-SDKDKLADKLEGDEKEKI 602
Query: 95 SAATDFSTKQNADAKQGSQKSPEEEKNEE 123
AAT + D Q S+K +EK +E
Sbjct: 603 EAATK-EALEWLDENQNSEKEEYDEKLKE 630
>At5g28540 luminal binding protein
Length = 669
Score = 38.5 bits (88), Expect = 0.007
Identities = 30/89 (33%), Positives = 45/89 (49%), Gaps = 3/89 (3%)
Query: 35 EFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAAKKVSHNVKEKI 94
E + VKE +E AEE K +KE + + T +Y + V + + A K+ + KEKI
Sbjct: 545 EIDRMVKEAEEFAEEDKKVKEKIDARNALET-YVYNMKNQV-NDKDKLADKLEGDEKEKI 602
Query: 95 SAATDFSTKQNADAKQGSQKSPEEEKNEE 123
AAT + D Q S+K +EK +E
Sbjct: 603 EAATK-EALEWLDENQNSEKEEYDEKLKE 630
>At3g60110 unknown protein
Length = 644
Score = 38.5 bits (88), Expect = 0.007
Identities = 41/146 (28%), Positives = 63/146 (43%), Gaps = 15/146 (10%)
Query: 21 EFSKKVKDETVKNPEFQKSVKELKEK--AEELKGIKEGLKEKTKQTTEQLYRQFDSVWKE 78
E S DE +N + +K K+K A+ LK IK+ +K K+TT + ++ D K+
Sbjct: 508 ESSDDDDDEKEENSKTEKKTVADKKKSVADFLKRIKKNSPQKGKETTSKNQKKNDGNVKK 567
Query: 79 AEAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSGNASESLFGKFK 138
KK NVK K+N+ K +S +K E + N+ S K K
Sbjct: 568 ENDHQKKSDGNVK-----------KENSKVKPRELRSSTGKKKVEVENNNSKSS--SKRK 614
Query: 139 STFSSPMVSTSFQKLKDAKLWT*PRK 164
T + V+T + + K PRK
Sbjct: 615 QTKETAEVATGKRGRESGKDDKQPRK 640
Score = 28.1 bits (61), Expect = 9.1
Identities = 32/147 (21%), Positives = 58/147 (38%), Gaps = 11/147 (7%)
Query: 9 NSVDRRGYSVFNEFSKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQL 68
+SV R+ SV + K K +K S ++ EK + E +EK TT
Sbjct: 417 SSVSRQKSSVLSLVPCKKKSSALKKTSPSSSSRQKDEKKSQ-----EVSEEKIVTTTATT 471
Query: 69 YRQFDSVWKEAEAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSGN 128
+ + +K+++ K+ + + K+ D K S ++EK E S +
Sbjct: 472 SA------RSSRRTSKEIAVVAKDTKTGRAKNNIKKQTDTKTESSDDDDDEKEENSKTEK 525
Query: 129 ASESLFGKFKSTFSSPMVSTSFQKLKD 155
+ + K + F + S QK K+
Sbjct: 526 KTVADKKKSVADFLKRIKKNSPQKGKE 552
>At4g21020 unknown protein (At4g21020)
Length = 266
Score = 37.7 bits (86), Expect = 0.012
Identities = 30/109 (27%), Positives = 50/109 (45%), Gaps = 6/109 (5%)
Query: 21 EFSKKVKDETVKNP-EFQKSVKE-LKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKE 78
E +K ++T+ N E + KE +KE E+ K EG KE K E+L + K
Sbjct: 152 EKAKDYAEDTMDNAKEKARHAKEKVKEYGEDTKEKAEGFKETVKGKAEELGEKTKETVKG 211
Query: 79 AEAAAKKVSHNVKEKISAATDFSTKQNADAKQG----SQKSPEEEKNEE 123
A + K + V E + + + K AD +G +K E+++ E+
Sbjct: 212 AWESTKNAAQTVTEAVVGPEEDAEKARADMNKGVEDHRKKKAEKDQKED 260
Score = 33.9 bits (76), Expect = 0.17
Identities = 33/139 (23%), Positives = 58/139 (40%), Gaps = 10/139 (7%)
Query: 20 NEFSKKVKDETVKNPEFQKS-VKELKEKA--------EELKGIKEGLKEKTKQTTEQLYR 70
NE + K D+ + E K E KEKA E+ K E K+K + +
Sbjct: 86 NEGASKAADKAYETKEQAKDKAYETKEKAKDTAYNAKEKAKDYAERTKDKVNEGAYKAAD 145
Query: 71 QFDSVWKEAEAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSG-NA 129
+ + ++A+ A+ N KEK A + + D K+ ++ E K + G
Sbjct: 146 KAEDTKEKAKDYAEDTMDNAKEKARHAKEKVKEYGEDTKEKAEGFKETVKGKAEELGEKT 205
Query: 130 SESLFGKFKSTFSSPMVST 148
E++ G ++ST ++ T
Sbjct: 206 KETVKGAWESTKNAAQTVT 224
>At3g05900 unknown protein
Length = 660
Score = 37.4 bits (85), Expect = 0.015
Identities = 31/121 (25%), Positives = 61/121 (49%), Gaps = 13/121 (10%)
Query: 24 KKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKT-----KQTTEQLYRQFDSVWKE 78
K V +E +K+ Q++V E K+ +L E +K+ T + +E+ ++ D+ E
Sbjct: 514 KHVVEEPLKDE--QENVSEAKDVVTKLAAEDENIKKDTDTPVAEGKSEETLKETDTESVE 571
Query: 79 AEAAAKKVSHNVKEKISAATDFS--TKQNADAKQ----GSQKSPEEEKNEESPSGNASES 132
EAAA K + EK++ + + K+ +AKQ ++++P ++K+ S +S
Sbjct: 572 KEAAANKQEEPITEKVAEVVETAPVAKEIDEAKQQPEVTTKEAPAKQKHSNSIISKVKQS 631
Query: 133 L 133
L
Sbjct: 632 L 632
>At5g44310 late embryogenesis abundant protein-like
Length = 331
Score = 36.6 bits (83), Expect = 0.026
Identities = 28/108 (25%), Positives = 52/108 (47%), Gaps = 11/108 (10%)
Query: 20 NEFSKKVKDETVKNPEFQKSVKE-LKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKE 78
NE + + D+ + + K E KEKAE++ G KEK + E+ VW+
Sbjct: 225 NEGASRAADKAEETKDKAKDYAEDSKEKAEDMA---HGFKEKAQDIGEKTMDTVKDVWET 281
Query: 79 AEAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQ---KSPEEEKNEE 123
A++ A+KV+ E + + + + K D +G + K +E +N++
Sbjct: 282 AKSTAQKVT----EAVVGSGEEADKARDDVDKGLEDLSKKAKENRNKD 325
Score = 30.4 bits (67), Expect = 1.8
Identities = 34/118 (28%), Positives = 54/118 (44%), Gaps = 12/118 (10%)
Query: 23 SKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQ--------LYRQFDS 74
SKK ++E+ + E K K K K EE K +KE+TK EQ R D
Sbjct: 68 SKKWREESGEYAEAGKG-KAHKTK-EEAKDKAYDMKERTKDYAEQTKNKVNEGASRAADK 125
Query: 75 VWKEAEAAAKKVSHNVKEKI-SAATDFSTKQNADAKQGSQKSPEEEKNEESPSGNASE 131
+ E + AK +++VKEK A + K N A + + K+ E ++ + + + E
Sbjct: 126 AY-ETKEKAKDKAYDVKEKTKDYAEEAKDKVNEGASRAADKAYETKEKAKDKAYDVKE 182
Score = 29.6 bits (65), Expect = 3.1
Identities = 25/112 (22%), Positives = 52/112 (46%), Gaps = 7/112 (6%)
Query: 21 EFSKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAE 80
+++++ KD+ N ++ + E E+ K +KEKTK E+ + + E
Sbjct: 146 DYAEEAKDKV--NEGASRAADKAYETKEKAKDKAYDVKEKTKDFAEETKEKVN----EGA 199
Query: 81 AAAKKVSHNVKEKI-SAATDFSTKQNADAKQGSQKSPEEEKNEESPSGNASE 131
+ A +++VKEK + A K N A + + K+ E + + + ++ E
Sbjct: 200 SRAADKAYDVKEKTKNYAEQTKDKVNEGASRAADKAEETKDKAKDYAEDSKE 251
>At5g24880 glutamic acid-rich protein
Length = 443
Score = 36.6 bits (83), Expect = 0.026
Identities = 36/162 (22%), Positives = 73/162 (44%), Gaps = 21/162 (12%)
Query: 20 NEFSKKVKDETVKNPE-FQKSVKELKE-------KAEELKGIKEGLKE-KTKQTTEQLYR 70
N +++++ +KN + ++ +E+KE K+EE + +K+ + E +T + + +
Sbjct: 264 NAVVAELEEKLIKNEDDIEEKTEEMKEQDNNQANKSEEEEDVKKKIDENETPEKVDTESK 323
Query: 71 QFDSVWKEAEAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSGN-- 128
+ +SV + + ++V KE++ K+ K+ QK EE+ +E G+
Sbjct: 324 EVESVEETTQEKEEEVKEEGKERVEE----EEKEKEKVKEDDQKEKVEEEEKEKVKGDEE 379
Query: 129 -----ASESLFGKFKSTFSSPMVSTS-FQKLKDAKLWT*PRK 164
ES GK K S S + + +K+ PRK
Sbjct: 380 KEKVKEEESAEGKKKEVVKGKKESPSAYNDVIASKMQENPRK 421
Score = 31.2 bits (69), Expect = 1.1
Identities = 30/142 (21%), Positives = 54/142 (37%), Gaps = 43/142 (30%)
Query: 25 KVKDETVKNPEFQKSVKELKEKA-----------------------------------EE 49
+V D ++ P+++K KE++EK E+
Sbjct: 214 QVSDHDIEEPKYEKEEKEVQEKVVQANESVEEKAESSGPTPVASPVGKDCNAVVAELEEK 273
Query: 50 LKGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAAK--------KVSHNVKEKISAATDFS 101
L ++ ++EKT++ EQ Q + +E + K KV KE S
Sbjct: 274 LIKNEDDIEEKTEEMKEQDNNQANKSEEEEDVKKKIDENETPEKVDTESKEVESVEETTQ 333
Query: 102 TKQNADAKQGSQKSPEEEKNEE 123
K+ ++G ++ EEEK +E
Sbjct: 334 EKEEEVKEEGKERVEEEEKEKE 355
>At3g17520 unknown protein
Length = 298
Score = 35.8 bits (81), Expect = 0.044
Identities = 29/116 (25%), Positives = 57/116 (49%), Gaps = 6/116 (5%)
Query: 45 EKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAAKKVSHNVKEKISAATDFSTKQ 104
+KAE+ K E K K ++ + S ++ A++ ++ +VK+K S + D + +
Sbjct: 74 QKAEDAK---EAAKRKAEEAVGAAKEKAGSAYETAKSKVEEGLASVKDKASQSYDSAGQV 130
Query: 105 NADAKQGSQKSPEE---EKNEESPSGNASESLFGKFKSTFSSPMVSTSFQKLKDAK 157
D S++ + ++N+ES +G A E + K + S + T +K K+AK
Sbjct: 131 KDDVSHKSKQVKDSLSGDENDESWTGWAKEKIGIKNEDINSPNLGETVSEKAKEAK 186
Score = 35.0 bits (79), Expect = 0.075
Identities = 29/113 (25%), Positives = 49/113 (42%), Gaps = 14/113 (12%)
Query: 26 VKDETVKNPEFQKSV----KELKEKA--------EELKGIKEGLKEKTKQTTEQLYRQFD 73
+K+E + +P ++V KE KE A E+L E KEK T + +
Sbjct: 164 IKNEDINSPNLGETVSEKAKEAKEAAKRKAGDAKEKLAETVETAKEKASDMTSAAKEKAE 223
Query: 74 SVWKEAEAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEE--EKNEES 124
+ +EAE +K +KE A + + AK + +S + K+EE+
Sbjct: 224 KLKEEAERESKSAKEKIKESYETAKSKADETLESAKDKASQSYDSAARKSEEA 276
Score = 33.1 bits (74), Expect = 0.28
Identities = 27/84 (32%), Positives = 38/84 (45%), Gaps = 7/84 (8%)
Query: 17 SVFNEFSKKVKDETVKNPEFQKSVKE-LKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSV 75
S E ++K+K+E + KS KE +KE E K + E K Q Y DS
Sbjct: 216 SAAKEKAEKLKEEAERE---SKSAKEKIKESYETAKSKADETLESAKDKASQSY---DSA 269
Query: 76 WKEAEAAAKKVSHNVKEKISAATD 99
+++E A VSH K + TD
Sbjct: 270 ARKSEEAKDTVSHKSKRVKESLTD 293
Score = 31.6 bits (70), Expect = 0.83
Identities = 18/67 (26%), Positives = 38/67 (55%), Gaps = 1/67 (1%)
Query: 77 KEAEAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNE-ESPSGNASESLFG 135
KEA+ AAK+ + + KEK++ + + ++ +D +++ E+ K E E S +A E +
Sbjct: 183 KEAKEAAKRKAGDAKEKLAETVETAKEKASDMTSAAKEKAEKLKEEAERESKSAKEKIKE 242
Query: 136 KFKSTFS 142
+++ S
Sbjct: 243 SYETAKS 249
Score = 31.6 bits (70), Expect = 0.83
Identities = 18/76 (23%), Positives = 42/76 (54%), Gaps = 4/76 (5%)
Query: 38 KSVKELKEKAEEL----KGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAAKKVSHNVKEK 93
++V+ KEKA ++ K E LKE+ ++ ++ + ++ A++ A + + K+K
Sbjct: 202 ETVETAKEKASDMTSAAKEKAEKLKEEAERESKSAKEKIKESYETAKSKADETLESAKDK 261
Query: 94 ISAATDFSTKQNADAK 109
S + D + +++ +AK
Sbjct: 262 ASQSYDSAARKSEEAK 277
>At5g54410 unknown protein
Length = 219
Score = 35.4 bits (80), Expect = 0.057
Identities = 26/103 (25%), Positives = 47/103 (45%), Gaps = 5/103 (4%)
Query: 23 SKKVKDETVK-NPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWK-EAE 80
+KK+++ET++ N E +++ + K E K ++ EK KQ + D +K + E
Sbjct: 39 NKKLREETLQSNEEANDAMETFRRKTNEQKRLEN---EKRKQALKDAKDLKDLTYKTKVE 95
Query: 81 AAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEE 123
KK D + ++ D + +K P EEK +E
Sbjct: 96 NKLKKTQPEKDRAEEEEKDLTEEKKKDPTEEEEKDPTEEKKKE 138
>At4g09950 AIG1-like protein
Length = 336
Score = 35.0 bits (79), Expect = 0.075
Identities = 34/145 (23%), Positives = 65/145 (44%), Gaps = 16/145 (11%)
Query: 8 VNSVDRRGYSVFN---EFSKK------VKDETVKNPEFQKSVKELKEKAEELKGIKEGLK 58
V+ DR+ + + N + SKK + D + + E + ++KE +++ EE+KG
Sbjct: 186 VSKKDRQVHDLLNLVEQISKKNNGKSYMADLSHELRENEATIKEKQKQIEEMKGWS---- 241
Query: 59 EKTKQTTEQLYRQFDSVWKEA-EAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPE 117
+KQ Q+ ++ + E E +K+S+ +KE + + K A+ ++ +K E
Sbjct: 242 --SKQEISQMKKELEKSHNEMLEGIKEKISNQLKESLEDVKEQLAKAQAEREETEKKMNE 299
Query: 118 EEKNEESPSGNASESLFGKFKSTFS 142
+K E L K T S
Sbjct: 300 IQKLSSDEIRRLREQLNKAEKETAS 324
>At1g72150 putative cytosolic factor protein
Length = 573
Score = 35.0 bits (79), Expect = 0.075
Identities = 34/148 (22%), Positives = 61/148 (40%), Gaps = 12/148 (8%)
Query: 2 VKTRLFVNSVDRRGYSVFNEFSKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKT 61
V ++ V + + EF + V+ E + EF V +KE+ E K +E KE+
Sbjct: 80 VAEKVVVLTAEEVQKKALEEFKELVR-EALNKREFTAPVTPVKEEKTEEKKTEEETKEEE 138
Query: 62 KQTTEQLYRQFDSVWKEAEAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKN 121
K TE+ K+ E + K + AA + + + A + S++ PEE+
Sbjct: 139 K--TEE---------KKEETTTEVKVEEEKPAVPAAEEEKSSEAAPVETKSEEKPEEKAE 187
Query: 122 EESPSGNASESLFGKFKSTFSSPMVSTS 149
+ +++E K +VS S
Sbjct: 188 VTTEKASSAEEDGTKTVEAIEESIVSVS 215
>At1g69710 putative regulator of chromosome condensation
Length = 1028
Score = 35.0 bits (79), Expect = 0.075
Identities = 30/112 (26%), Positives = 55/112 (48%), Gaps = 10/112 (8%)
Query: 13 RRGYSVFNEFSKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQF 72
RR Y V SK++KD F + + LKE+ E+L L+E+ ++T QL
Sbjct: 825 RRSYEVAAAESKQLKDS------FNQDMAGLKEQVEQLASKAHQLEEELEKTKRQLKVVT 878
Query: 73 DSVWKEAE--AAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNE 122
EAE +AK+V ++ ++ + +++ D+ + K ++EK+E
Sbjct: 879 AMAADEAEENRSAKEVIRSLTTQLKEMAEKQSQK--DSISTNSKHTDKEKSE 928
>At1g56660 hypothetical protein
Length = 522
Score = 35.0 bits (79), Expect = 0.075
Identities = 29/114 (25%), Positives = 52/114 (45%), Gaps = 5/114 (4%)
Query: 24 KKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLK-EKTKQTTEQLYRQFDSVWKEAEAA 82
KK KDE+ + +K KE K+K E + +K +K K L ++ + KE +
Sbjct: 179 KKEKDESGTEEKKKKPKKEKKQKEESKSNEDKKVKGKKEKGEKGDLEKEDEEKKKEHD-- 236
Query: 83 AKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSGNASESLFGK 136
+ +KEK S K + A++ +K +E+K ++ + + L GK
Sbjct: 237 --ETDQEMKEKDSKKNKKKEKDESCAEEKKKKPDKEKKEKDESTEKEDKKLKGK 288
Score = 34.7 bits (78), Expect = 0.098
Identities = 33/126 (26%), Positives = 57/126 (45%), Gaps = 24/126 (19%)
Query: 25 KVKDETVKNPEFQKSVKELKEKA-EELKGIKEGLK-------------EKTKQTTEQLYR 70
+VK+ VK E +K K+ KEK EEL+ KEG K EK K+ ++ +
Sbjct: 100 EVKESDVKVEEHEKEHKKGKEKKHEELEEEKEGKKKKNKKEKDESGPEEKNKKADKE--K 157
Query: 71 QFDSVWKEAEAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSGNAS 130
+ + V +E E ++ K+K K + ++ +K +E+K +E N
Sbjct: 158 KHEDVSQEKEELEEEDGKKNKKK--------EKDESGTEEKKKKPKKEKKQKEESKSNED 209
Query: 131 ESLFGK 136
+ + GK
Sbjct: 210 KKVKGK 215
Score = 33.9 bits (76), Expect = 0.17
Identities = 30/118 (25%), Positives = 52/118 (43%), Gaps = 9/118 (7%)
Query: 23 SKKVKDETVKNPEFQKSVKELK--------EKAEELKGIKEGLKEKTKQTTEQLYRQFDS 74
+KK KDE+ + +K+ KE K E+ EE G K KEK + TE+ ++
Sbjct: 137 NKKEKDESGPEEKNKKADKEKKHEDVSQEKEELEEEDGKKNKKKEKDESGTEEKKKKPKK 196
Query: 75 VWKEAEAAAKKVSHNVK-EKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSGNASE 131
K+ E + VK +K K++ + K+ ++ +E K ++S E
Sbjct: 197 EKKQKEESKSNEDKKVKGKKEKGEKGDLEKEDEEKKKEHDETDQEMKEKDSKKNKKKE 254
Score = 33.9 bits (76), Expect = 0.17
Identities = 27/110 (24%), Positives = 46/110 (41%), Gaps = 5/110 (4%)
Query: 24 KKVKDETVKNPEFQKSVKELKEKAEEL--KGIKEGLKEKTKQTTEQLYRQFDSVWKEAEA 81
K+ KD+ E ++ + KEK E K +KE K++ TE + R EAE
Sbjct: 346 KETKDKDDDEGETKQKKNKKKEKKSEKGEKDVKEDKKKENPLETEVMSRDIKLEEPEAE- 404
Query: 82 AAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSGNASE 131
KK + +EK + + + K+ K ++K+ + P E
Sbjct: 405 --KKEEDDTEEKKKSKVEGGESEEGKKKKKKDKKKNKKKDTKEPKMTEDE 452
Score = 32.0 bits (71), Expect = 0.63
Identities = 32/117 (27%), Positives = 47/117 (39%), Gaps = 17/117 (14%)
Query: 24 KKVKDETVKNPEFQKSVKELKEKAE-------ELKGIK--------EGLKEKTKQTTEQL 68
KK KDE+ + +K KE KEK E +LKG K E +KTK+
Sbjct: 252 KKEKDESCAEEKKKKPDKEKKEKDESTEKEDKKLKGKKGKGEKPEKEDEGKKTKEHDATE 311
Query: 69 YRQFDSVWKEAEAAAKKVSHNVKEKISAATDFSTKQ--NADAKQGSQKSPEEEKNEE 123
D E KK K+K + + K+ + D +G K + +K E+
Sbjct: 312 QEMDDEAADHKEGKKKKNKDKAKKKETVIDEVCEKETKDKDDDEGETKQKKNKKKEK 368
Score = 31.2 bits (69), Expect = 1.1
Identities = 31/134 (23%), Positives = 53/134 (39%), Gaps = 5/134 (3%)
Query: 24 KKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAA 83
K K ETV + +K K+ + E K K KEK + E+ ++ KE
Sbjct: 332 KAKKKETVIDEVCEKETKDKDDDEGETKQKKNKKKEKKSEKGEKDVKEDKK--KENPLET 389
Query: 84 KKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSGNASESLFGKFKSTFSS 143
+ +S ++K + A K+ D ++ + E ++EE + K K T
Sbjct: 390 EVMSRDIKLEEPEA---EKKEEDDTEEKKKSKVEGGESEEGKKKKKKDKKKNKKKDTKEP 446
Query: 144 PMVSTSFQKLKDAK 157
M +K D+K
Sbjct: 447 KMTEDEEEKKDDSK 460
Score = 30.4 bits (67), Expect = 1.8
Identities = 25/111 (22%), Positives = 53/111 (47%), Gaps = 1/111 (0%)
Query: 21 EFSKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAE 80
E SK +D+ VK + + +L+++ EE K + ++ K+ + ++ + AE
Sbjct: 202 EESKSNEDKKVKGKKEKGEKGDLEKEDEEKKKEHDETDQEMKEKDSKKNKKKEKDESCAE 261
Query: 81 AAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSGNASE 131
KK KEK +T+ K+ K +K +E++ +++ +A+E
Sbjct: 262 EKKKKPDKEKKEK-DESTEKEDKKLKGKKGKGEKPEKEDEGKKTKEHDATE 311
Score = 28.9 bits (63), Expect = 5.4
Identities = 25/109 (22%), Positives = 53/109 (47%), Gaps = 5/109 (4%)
Query: 20 NEFSKKVKDET---VKNPEFQKSVKELKEK--AEELKGIKEGLKEKTKQTTEQLYRQFDS 74
+E KK DET +K + +K+ K+ K++ AEE K + K++ ++TE+ ++
Sbjct: 228 DEEKKKEHDETDQEMKEKDSKKNKKKEKDESCAEEKKKKPDKEKKEKDESTEKEDKKLKG 287
Query: 75 VWKEAEAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEE 123
+ E K+ ++ A + AD K+G +K +++ ++
Sbjct: 288 KKGKGEKPEKEDEGKKTKEHDATEQEMDDEAADHKEGKKKKNKDKAKKK 336
>At5g38690 unknown protein
Length = 572
Score = 34.7 bits (78), Expect = 0.098
Identities = 43/168 (25%), Positives = 69/168 (40%), Gaps = 42/168 (25%)
Query: 4 TRLFVNSVDRRGYSVFNE-----------FSKKVKDETV----------------KNPEF 36
T + V++ + G+S +E ++KKVK E V K+
Sbjct: 104 TGILVHTAKKTGFSSVSELLKTSGSDKYFYTKKVKPEGVVVVSPLKLDQENSIEQKHVSI 163
Query: 37 QKSVKELKEKAEELKGIKEGLKEK---TKQTTEQLYRQFDSVWKEAEAAAKKVSHNVKEK 93
+KS K K EELK I G + K++ + + DSV E K ++ K+K
Sbjct: 164 KKSRK---TKREELKDINNGCSNENVVVKKSNPKKIKLSDSVHATTEVIKKVLAEGKKKK 220
Query: 94 ISAATDFSTKQNADAKQGSQKSPEEEKNEES------PSGNASESLFG 135
+A K+N A++ + P +K EE P GN S ++ G
Sbjct: 221 TTAK---GIKENNVAEKIKRIKPALKKKEEDQVEIKIPQGNISITVSG 265
>At5g57120 unknown protein
Length = 330
Score = 34.3 bits (77), Expect = 0.13
Identities = 35/140 (25%), Positives = 60/140 (42%), Gaps = 20/140 (14%)
Query: 7 FVNSVDRRGYSVFNEFSKKVKD-ETVKNPEFQKSVKELKEKAEELKGIKE-------GLK 58
F+N D + N + V+ E VK + +K KE K + E + +KE G+K
Sbjct: 63 FLNKRDHEAAANGNTEANVVEAVENVKKDKKKKKNKETKVEVTEEEKVKETDAVIEDGVK 122
Query: 59 EKTKQ-------TTEQLYRQFDSVWKEAEAAAKKVSHNVKEKISAATDFSTKQNADAKQG 111
EK K+ T E+ ++ D+V ++ KK K+ S + + + +K+
Sbjct: 123 EKKKKKETKVKVTEEEKVKETDAVIEDGVKEKKK-----KKSKSKSVEADDDKEKVSKKR 177
Query: 112 SQKSPEEEKNEESPSGNASE 131
+ PEE K E S+
Sbjct: 178 KRSEPEETKEETEDDDEESK 197
>At3g58840 unknown protein
Length = 318
Score = 34.3 bits (77), Expect = 0.13
Identities = 30/130 (23%), Positives = 53/130 (40%), Gaps = 4/130 (3%)
Query: 8 VNSVDRRGYSVFNEFSKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQ 67
++ D+ G E +K++D KN E + +ELKE+ E L G E +K+ + Q
Sbjct: 12 ISDYDQGGVKT-TELERKIEDMENKNQELTRENRELKERLERLTGEIEEMKD-VEAEMNQ 69
Query: 68 LYRQFDSVWKEAEAAAKKVSHNVKEKISAATDFSTKQN--ADAKQGSQKSPEEEKNEESP 125
+ + + +E E K + + T+ S + + G K+ EE +
Sbjct: 70 RFGEMEKEIEEYEEEKKALEAISTRAVELETEVSNLHDDLITSLNGVDKTAEEVAELKKA 129
Query: 126 SGNASESLFG 135
E L G
Sbjct: 130 LAEIVEKLEG 139
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.136 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,818,172
Number of Sequences: 26719
Number of extensions: 379346
Number of successful extensions: 2335
Number of sequences better than 10.0: 187
Number of HSP's better than 10.0 without gapping: 42
Number of HSP's successfully gapped in prelim test: 147
Number of HSP's that attempted gapping in prelim test: 2059
Number of HSP's gapped (non-prelim): 354
length of query: 409
length of database: 11,318,596
effective HSP length: 102
effective length of query: 307
effective length of database: 8,593,258
effective search space: 2638130206
effective search space used: 2638130206
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC141322.10


