

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141115.10 + phase: 0 /pseudo
(173 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
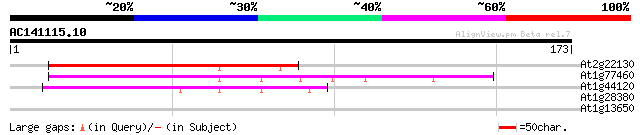
Score E
Sequences producing significant alignments: (bits) Value
At2g22130 unknown protein 92 1e-19
At1g77460 unknown protein 51 4e-07
At1g44120 hypothetical protein 44 4e-05
At1g28380 unknown protein 27 5.5
At1g13650 hypothetical protein 27 5.5
>At2g22130 unknown protein
Length = 2048
Score = 92.0 bits (227), Expect = 1e-19
Identities = 55/87 (63%), Positives = 60/87 (68%), Gaps = 10/87 (11%)
Query: 13 MDALFLLIQGWSACPVEVSRDQSNAAAYAIPLLQNLIQFGPVLFFEKAEFI-------LV 65
+DALFLL Q WSACP EVSR QS AAA AIPLLQ LIQ GP F EKAEF+ LV
Sbjct: 1922 LDALFLLRQAWSACPAEVSRAQSVAAADAIPLLQYLIQSGPPRFQEKAEFLLQCLPGTLV 1981
Query: 66 MIVKRGNNMRQCVGNQG---KITLEAN 89
+ +KRGNNM+Q VGN KITL N
Sbjct: 1982 VTIKRGNNMKQSVGNPSVFCKITLGNN 2008
>At1g77460 unknown protein
Length = 2110
Score = 50.8 bits (120), Expect = 4e-07
Identities = 46/179 (25%), Positives = 77/179 (42%), Gaps = 42/179 (23%)
Query: 13 MDALFLLIQGWSACPVEVSRDQSNAAAYAIPLLQNLIQFGPVLFFEKAEFI-------LV 65
+D L+LL W+ ++V++ Q+ AA AIP+LQ L++ P F +KA+ + L
Sbjct: 1926 LDILYLLRHSWTNMSIDVAKSQAMIAAEAIPVLQMLMKTCPPRFHDKADSLLHCLPGCLT 1985
Query: 66 MIVKRGNNMRQ---------------CVGNQGKITLEA-NQEWDERFTY---------MV 100
+ V R NN++Q C Q K+ + EW E FT+ +
Sbjct: 1986 VNVMRANNLKQSMATTNAFCQLTIGNCPPRQTKVVSNSTTPEWKEGFTWAFDVPPKGQKL 2045
Query: 101 L*ECSSRT--------EASYLLQKQA*SGKADEHTLL--PTSKSGQPRNLEVELKWSNK 149
C S++ + + K G+ L SK R+L++E+ WSN+
Sbjct: 2046 HIICKSKSTFGKTTLGRVTIQIDKVVTEGEYSGSLSLNHENSKDASSRSLDIEIAWSNR 2104
>At1g44120 hypothetical protein
Length = 2114
Score = 43.9 bits (102), Expect = 4e-05
Identities = 31/112 (27%), Positives = 53/112 (46%), Gaps = 24/112 (21%)
Query: 11 SWMDALFLLIQGWSACPVEVSRDQSNAAAYAIPLLQNLIQF-----GPVLFFEKAEFI-- 63
S MD ++ L Q W+ P E +R Q+ AA AIP+LQ +++ P F E+ +
Sbjct: 1930 SAMDTIYTLRQSWTTMPTETARSQAVLAADAIPVLQLMMKSKLKSPAPSSFHERGNSLLN 1989
Query: 64 -----LVMIVKRGNNMRQ-----------CVGNQGKITLEANQE-WDERFTY 98
L + +KRG+N+++ C + K+ ++ W E FT+
Sbjct: 1990 CLPGSLTVAIKRGDNLKRSNAFCRLIIDNCPTKKTKVVKRSSSPVWKESFTW 2041
>At1g28380 unknown protein
Length = 612
Score = 26.9 bits (58), Expect = 5.5
Identities = 18/66 (27%), Positives = 31/66 (46%), Gaps = 1/66 (1%)
Query: 28 VEVSRDQSNAAAYA-IPLLQNLIQFGPVLFFEKAEFILVMIVKRGNNMRQCVGNQGKITL 86
V+ +S A+ Y+ P +NL+Q+G F +VM K MR C+ G
Sbjct: 221 VDPVESKSPASVYSGKPKEENLLQWGLQPFGTSVSSAVVMHTKNEEIMRVCIRRGGVDLG 280
Query: 87 EANQEW 92
++++ W
Sbjct: 281 QSHERW 286
>At1g13650 hypothetical protein
Length = 281
Score = 26.9 bits (58), Expect = 5.5
Identities = 14/64 (21%), Positives = 33/64 (50%)
Query: 28 VEVSRDQSNAAAYAIPLLQNLIQFGPVLFFEKAEFILVMIVKRGNNMRQCVGNQGKITLE 87
+ + D ++A+++A+P+LQ+ PV + E L M++ +R+ + ++ E
Sbjct: 147 LSIRSDGTSASSFALPILQSEWNSSPVRMGKAEETQLRMVIAEERKVRKDKAEKTQLRKE 206
Query: 88 ANQE 91
+E
Sbjct: 207 KAEE 210
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.328 0.139 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,656,764
Number of Sequences: 26719
Number of extensions: 132660
Number of successful extensions: 292
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 283
Number of HSP's gapped (non-prelim): 9
length of query: 173
length of database: 11,318,596
effective HSP length: 92
effective length of query: 81
effective length of database: 8,860,448
effective search space: 717696288
effective search space used: 717696288
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC141115.10


