

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141114.8 - phase: 0
(239 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
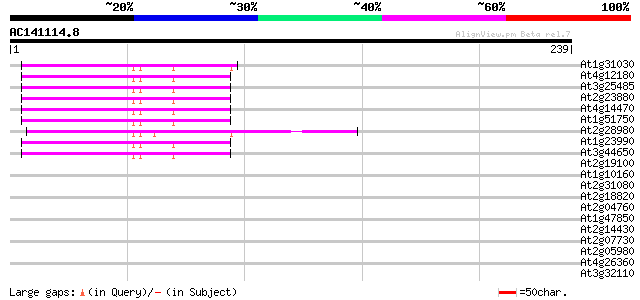
Sequences producing significant alignments: (bits) Value
At1g31030 F17F8.5 48 5e-06
At4g12180 putative reverse transcriptase 45 3e-05
At3g25485 unknown protein 45 3e-05
At2g23880 putative non-LTR retroelement reverse transcriptase 44 6e-05
At4g14470 reverse transcriptase like protein 44 8e-05
At1g51750 hypothetical protein 42 4e-04
At2g28980 putative non-LTR retroelement reverse transcriptase 41 5e-04
At1g23990 reverse transcriptase, putative 41 6e-04
At3g44650 putative protein 40 8e-04
At2g19100 putative non-LTR retroelement reverse transcriptase 40 0.001
At1g10160 putative reverse transcriptase 39 0.002
At2g31080 putative non-LTR retroelement reverse transcriptase 38 0.004
At2g18820 putative non-LTR retroelement reverse transcriptase 38 0.004
At2g04760 putative non-LTR retroelement reverse transcriptase 38 0.004
At1g47850 hypothetical protein 38 0.004
At2g14430 putative non-LTR retroelement reverse transcriptase 38 0.005
At2g07730 putative non-LTR retroelement reverse transcriptase 37 0.007
At2g05980 putative non-LTR retroelement reverse transcriptase 37 0.007
At4g26360 putative protein 35 0.027
At3g32110 non-LTR reverse transcriptase, putative 35 0.035
>At1g31030 F17F8.5
Length = 872
Score = 47.8 bits (112), Expect = 5e-06
Identities = 34/109 (31%), Positives = 55/109 (50%), Gaps = 17/109 (15%)
Query: 6 STSVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQELA--------PRC--- 54
S SV+ F GL QG +LSPYLF + +DVL++ + + A P+C
Sbjct: 284 SFSVQVNGDLVGYFQSKRGLRQGCSLSPYLFVICMDVLSKMLDKAAGVRKFGFHPKCQRL 343
Query: 55 ----MLFADDVVLVGESR-EEVNGRLETWRQALEAYGFRLSRSK-TEYM 97
+ FADD++++ + + + G LE + + + G R+S K T YM
Sbjct: 344 GLTHLSFADDLMVLSDGKTRSIEGILEVFDEFCKRSGLRISLEKSTLYM 392
>At4g12180 putative reverse transcriptase
Length = 662
Score = 45.4 bits (106), Expect = 3e-05
Identities = 30/105 (28%), Positives = 53/105 (49%), Gaps = 16/105 (15%)
Query: 6 STSVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQELA--------PRC--- 54
S SV+ F GL QG +L+PYLF +V+DVL++ + A P+C
Sbjct: 155 SFSVQVNRELAGYFNSLRGLRQGCSLTPYLFVIVMDVLSKKLDRAAGLRKFGYHPKCKNL 214
Query: 55 ----MLFADDVVLVGESR-EEVNGRLETWRQALEAYGFRLSRSKT 94
+ FADD++++ + + + G +E + + G ++S +KT
Sbjct: 215 GLTHLSFADDIMVLTDGKLRSLEGIVEVFDSFAKQSGLKISMAKT 259
>At3g25485 unknown protein
Length = 979
Score = 45.4 bits (106), Expect = 3e-05
Identities = 30/105 (28%), Positives = 52/105 (48%), Gaps = 16/105 (15%)
Query: 6 STSVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQELA--------PRC--- 54
S SV+ F + GL QG +LSPYLF + +DVL++ + A PRC
Sbjct: 553 SFSVQVNGELVGFFNSSRGLRQGCSLSPYLFVIAMDVLSKMLDRAAGFKKFGYHPRCQTI 612
Query: 55 ----MLFADDVVLVGESR-EEVNGRLETWRQALEAYGFRLSRSKT 94
+ FADD++++ + + V G + + + + G ++S K+
Sbjct: 613 GLTHLTFADDLMVLSDGKVRSVEGIVSVFDEFAKKSGLKISMEKS 657
>At2g23880 putative non-LTR retroelement reverse transcriptase
Length = 1216
Score = 44.3 bits (103), Expect = 6e-05
Identities = 30/105 (28%), Positives = 50/105 (47%), Gaps = 16/105 (15%)
Query: 6 STSVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQELA--------PRC--- 54
S S++ F GL QG +LSPYLF + +DVL+ + + A PRC
Sbjct: 358 SFSIQVNGELAGYFRSARGLRQGCSLSPYLFVISMDVLSRMLDKAAGAREFGYHPRCKTL 417
Query: 55 ----MLFADDVVLVGESR-EEVNGRLETWRQALEAYGFRLSRSKT 94
+ FADD++++ + + V+G ++ Q G ++ KT
Sbjct: 418 GLTHLCFADDLMILTDGKIRSVDGIVKVLNQFAAKLGLKICMEKT 462
>At4g14470 reverse transcriptase like protein
Length = 318
Score = 43.9 bits (102), Expect = 8e-05
Identities = 30/105 (28%), Positives = 52/105 (48%), Gaps = 16/105 (15%)
Query: 6 STSVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQELA--------PRC--- 54
S SV+ F + GL QG +LSPYLF +V+DVL++ + A P+C
Sbjct: 73 SFSVQVNGELAGYFNSSRGLRQGCSLSPYLFVIVMDVLSKKLDRAAGLRKFGYHPKCKNL 132
Query: 55 ----MLFADDVVLVGESR-EEVNGRLETWRQALEAYGFRLSRSKT 94
+ FADD++++ + + + G +E + + ++S KT
Sbjct: 133 GLTHLSFADDIMVLTDGKLRSLEGIVEVFDSFAKQSCLKISMEKT 177
>At1g51750 hypothetical protein
Length = 629
Score = 41.6 bits (96), Expect = 4e-04
Identities = 28/105 (26%), Positives = 51/105 (47%), Gaps = 16/105 (15%)
Query: 6 STSVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQELA--------PRC--- 54
S SV+ F G+ QG LSPYLF + ++VL++ + + A P+C
Sbjct: 42 SFSVQVNGELAGYFRSARGIRQGCALSPYLFVISMEVLSKMLDQAAGGKRFGFHPKCKNL 101
Query: 55 ----MLFADDVVLVGESR-EEVNGRLETWRQALEAYGFRLSRSKT 94
+ FADD++++ + + V+G +E + G +++ KT
Sbjct: 102 GLTHLCFADDLMILTDGKVRSVDGIVEVMNLFAKRSGLQINMEKT 146
>At2g28980 putative non-LTR retroelement reverse transcriptase
Length = 1529
Score = 41.2 bits (95), Expect = 5e-04
Identities = 42/158 (26%), Positives = 68/158 (42%), Gaps = 21/158 (13%)
Query: 8 SVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQELA--------PRC----- 54
SV+ F + GL QG LSPYLF + ++VL+ I E A P+C
Sbjct: 936 SVQVNGELAGFFGSSRGLRQGCALSPYLFVICMNVLSHMIDEAAVHRNIGYHPKCEKIGL 995
Query: 55 --MLFADD-VVLVGESREEVNGRLETWRQALEAYGFRLSRSK-TEYMEWNFSGRRSRSTL 110
+ FADD +V V + + G + +++ G ++S K T Y+ + R ++
Sbjct: 996 THLCFADDLMVFVDGHQWSIEGVINVFKEFAGRSGLQISLEKSTIYLAGVSASDRVQTLS 1055
Query: 111 EVKVGDHIIPQVTRFKYLGSFVQNDGEIEADVSHRIQA 148
+ +P +YLG + AD S I+A
Sbjct: 1056 SFPFANGQLP----VRYLGLPLLTKQMTTADYSPLIEA 1089
>At1g23990 reverse transcriptase, putative
Length = 657
Score = 40.8 bits (94), Expect = 6e-04
Identities = 29/105 (27%), Positives = 52/105 (48%), Gaps = 16/105 (15%)
Query: 6 STSVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQELA--------PRC--- 54
S SV+ F GL QG +LSPYLF + +DVL++ + + A RC
Sbjct: 86 SFSVQVNGELVGFFQSKRGLRQGCSLSPYLFVMSMDVLSKLLDQAASAKKFGYHSRCKEL 145
Query: 55 ----MLFADDVVLVGESR-EEVNGRLETWRQALEAYGFRLSRSKT 94
+ FADD++++ + + ++G +E + + G ++S K+
Sbjct: 146 SLTHLSFADDLMVLSDGKVRSIDGIVEVFDIFAKFSGLKISMEKS 190
>At3g44650 putative protein
Length = 762
Score = 40.4 bits (93), Expect = 8e-04
Identities = 28/105 (26%), Positives = 51/105 (47%), Gaps = 16/105 (15%)
Query: 6 STSVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQELA--------PRC--- 54
S SV+ F + GL QG LSPYLF + +DVL++ + ++ P C
Sbjct: 190 SFSVQVNGELAGFFQSSRGLRQGCALSPYLFVICMDVLSKLLDKVVGIGRIGYHPHCKRM 249
Query: 55 ----MLFADDVVLVGESR-EEVNGRLETWRQALEAYGFRLSRSKT 94
+ FADD++++ + + + G +E + + G ++S K+
Sbjct: 250 GLTHLSFADDLMILTDGQCRSIEGIIEVFDLFSKWSGLKISMEKS 294
>At2g19100 putative non-LTR retroelement reverse transcriptase
Length = 1447
Score = 39.7 bits (91), Expect = 0.001
Identities = 27/98 (27%), Positives = 45/98 (45%), Gaps = 17/98 (17%)
Query: 6 STSVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQE-------------LAP 52
S SV+ F GL QG +LSPYLF + +D+L++ + + L
Sbjct: 991 SFSVQVNGELAGFFQSARGLRQGCSLSPYLFVISMDILSKMLDKAAGQLPVRYLGLPLVT 1050
Query: 53 RCMLFADDVVLVGESREEVNGRLETWRQALEAYGFRLS 90
+ + AD + L+ E++ R+ TW +Y RL+
Sbjct: 1051 KRLSSADSLPLI----EQIRHRIGTWTSRFLSYAGRLN 1084
>At1g10160 putative reverse transcriptase
Length = 1118
Score = 39.3 bits (90), Expect = 0.002
Identities = 30/101 (29%), Positives = 48/101 (46%), Gaps = 18/101 (17%)
Query: 24 GLHQGSTLSPYLFTLVLDVLTEHIQ--------ELAPRCML-------FADDVVLVGE-S 67
G+ QG +S +LF LV+DVL++ + L P C+ FADDV++ + +
Sbjct: 649 GIRQGDPMSSHLFVLVMDVLSKSLDLGALNGLFNLHPNCLAPIITHLSFADDVLVFSDGA 708
Query: 68 REEVNGRLETWRQALEAYGFRLSRSKTEYM--EWNFSGRRS 106
+ G L + G ++R KTE + NF+ RS
Sbjct: 709 ASSIAGILTILDDFRQGSGLGINREKTELLLDGGNFARNRS 749
>At2g31080 putative non-LTR retroelement reverse transcriptase
Length = 1231
Score = 38.1 bits (87), Expect = 0.004
Identities = 47/169 (27%), Positives = 70/169 (40%), Gaps = 27/169 (15%)
Query: 6 STSVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQ---------ELAPRC-- 54
S SV T+ F GL QG LSPYLF L L+ L I+ +A C
Sbjct: 477 SMSVLWNGERTDSFVPARGLRQGDPLSPYLFVLCLERLCHLIEASVGKREWKPIAVSCGG 536
Query: 55 -----MLFADDVVLVGE-SREEVNGRLETWRQALEAYGFRLSRSKTEYMEWNFSGRRSRS 108
+ FADD++L E S ++ + EA G ++S K++ + R
Sbjct: 537 SKLSHVCFADDLILFAEASVAQIRIIRRVLERFCEASGQKVSLEKSKIFFSHNVSREMEQ 596
Query: 109 TLEVKVGDHIIPQVTRFKYLGSFV-------QNDGEIEADVSHRIQAGW 150
+ + G ++ KYLG + + GE+ VS R+ AGW
Sbjct: 597 LISEESGIGCTKELG--KYLGMPILQKRMNKETFGEVLERVSARL-AGW 642
>At2g18820 putative non-LTR retroelement reverse transcriptase
Length = 1412
Score = 38.1 bits (87), Expect = 0.004
Identities = 32/113 (28%), Positives = 50/113 (43%), Gaps = 17/113 (15%)
Query: 15 TTEVFPITI-GLHQGSTLSPYLFTLVLDVLTEHIQELA--------PRC-------MLFA 58
+T F + + GL QG +LSPYLF + ++VL+ + + A PRC + FA
Sbjct: 900 STASFSVQVNGLRQGCSLSPYLFVICMNVLSAMLDKGAVEKRFGYHPRCRNMGLTHLCFA 959
Query: 59 DDV-VLVGESREEVNGRLETWRQALEAYGFRLSRSKTEYMEWNFSGRRSRSTL 110
DD+ V S + G L ++ G +S K+ + S S L
Sbjct: 960 DDIMVFSAGSAHSLEGVLAIFKDFAAFSGLNISLEKSTLFMASISSETCASIL 1012
>At2g04760 putative non-LTR retroelement reverse transcriptase
Length = 631
Score = 38.1 bits (87), Expect = 0.004
Identities = 25/87 (28%), Positives = 40/87 (45%), Gaps = 16/87 (18%)
Query: 24 GLHQGSTLSPYLFTLVLDVLTEHIQELA--------PRC-------MLFADD-VVLVGES 67
GL QG LSPYLF + ++VL+ I E A P+C + F DD +V +
Sbjct: 225 GLRQGCALSPYLFVICMNVLSHMIDEAAVRRNIGYHPKCKKLSLTHLCFVDDLMVFIDGQ 284
Query: 68 REEVNGRLETWRQALEAYGFRLSRSKT 94
+ + G + + + G +S K+
Sbjct: 285 QRSIEGVINIFHEFAGKSGLHISLEKS 311
>At1g47850 hypothetical protein
Length = 799
Score = 38.1 bits (87), Expect = 0.004
Identities = 26/87 (29%), Positives = 41/87 (46%), Gaps = 16/87 (18%)
Query: 24 GLHQGSTLSPYLFTLVLDVLTEHIQELA--------PRC-------MLFADD-VVLVGES 67
GL QG LSPYLF + ++VL+ I A P+C + FADD +V +
Sbjct: 228 GLRQGCALSPYLFVICMNVLSHMIDVAAVHRNIGYHPKCKKLSLTHLCFADDLMVFIDGQ 287
Query: 68 REEVNGRLETWRQALEAYGFRLSRSKT 94
+ V G + +++ G +S K+
Sbjct: 288 QRSVEGVINIFKEFAGKSGLHISLEKS 314
>At2g14430 putative non-LTR retroelement reverse transcriptase
Length = 1277
Score = 37.7 bits (86), Expect = 0.005
Identities = 29/103 (28%), Positives = 44/103 (42%), Gaps = 16/103 (15%)
Query: 8 SVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQELA--------PRC----- 54
SV+ F GL QG LSPYLF + ++VL+ I A P+C
Sbjct: 781 SVQVNSEQAGFFGSKRGLRQGCALSPYLFVICMNVLSHMIDVAAVHRNIGYHPKCKKLSL 840
Query: 55 --MLFADD-VVLVGESREEVNGRLETWRQALEAYGFRLSRSKT 94
+ FADD +V + + V G + ++ G +S K+
Sbjct: 841 THLCFADDLMVFIDGQQRSVEGVINIFKDFAGKSGLHISLEKS 883
>At2g07730 putative non-LTR retroelement reverse transcriptase
Length = 970
Score = 37.4 bits (85), Expect = 0.007
Identities = 49/172 (28%), Positives = 67/172 (38%), Gaps = 37/172 (21%)
Query: 6 STSVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQ---------ELAPRCML 56
S SV G T+ F GL QG LSPYLF L L+ L I+ +A C
Sbjct: 327 SMSVLWNGGKTDSFVPARGLRQGDPLSPYLFVLCLERLCHLIEASVGTWDWKPMALSCEA 386
Query: 57 FADDVVLVGESREEVNGRLETWRQALEAYGFRLSRSKTEYMEWNFSGRRSRSTLEVKVGD 116
++++ E Q EA G +S K++ + FS SR + G+
Sbjct: 387 SVAQILIIRRVLE----------QFCEASGQNVSLEKSKIV---FSNNVSRDMERLISGE 433
Query: 117 HIIPQVTRF-KYLGSFV-------QNDGEIEADVSHRIQAGW------LKWR 154
I KYLG + + GE+ VS + AGW L WR
Sbjct: 434 SGIGCTRELGKYLGMPILQKRMNKETSGEVLEHVSSSL-AGWKSRSLSLAWR 484
>At2g05980 putative non-LTR retroelement reverse transcriptase
Length = 1352
Score = 37.4 bits (85), Expect = 0.007
Identities = 30/122 (24%), Positives = 54/122 (43%), Gaps = 16/122 (13%)
Query: 6 STSVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQELA--------PRC--- 54
S SV+ + F GL QG +LSPYL+ + ++VL+ + + A PRC
Sbjct: 784 SFSVQVNGELSGFFRSERGLRQGCSLSPYLYVICMNVLSCMLDKAAVEKKISYHPRCRNM 843
Query: 55 ----MLFADDVVLVGE-SREEVNGRLETWRQALEAYGFRLSRSKTEYMEWNFSGRRSRST 109
+ FADD+++ + + + + G L + + ++S K+ S S
Sbjct: 844 NLTHLCFADDIMVFSDGTSKSIQGTLAIFEKFAAMSWLKISLEKSTIFMAGISPNAKTSI 903
Query: 110 LE 111
L+
Sbjct: 904 LQ 905
>At4g26360 putative protein
Length = 1141
Score = 35.4 bits (80), Expect = 0.027
Identities = 23/89 (25%), Positives = 42/89 (46%), Gaps = 16/89 (17%)
Query: 24 GLHQGSTLSPYLFTLVLDVLTEHI---------------QELAPRCMLFADDVVLVGE-S 67
GL QG LSPYLF L ++V ++ + +L+ ++FADDV++ +
Sbjct: 627 GLRQGDPLSPYLFVLAMEVFSKLLNSRFDSGYIRYHPKASDLSISHLMFADDVMIFFDGG 686
Query: 68 REEVNGRLETWRQALEAYGFRLSRSKTEY 96
++G ET G +++ K+ +
Sbjct: 687 SSSLHGICETLEDFASWSGLKVNNDKSHF 715
>At3g32110 non-LTR reverse transcriptase, putative
Length = 1911
Score = 35.0 bits (79), Expect = 0.035
Identities = 26/81 (32%), Positives = 35/81 (43%), Gaps = 16/81 (19%)
Query: 3 EGVSTSVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTE--------------HIQ 48
EG S + T+ F GL QG LSPYLF L ++ L I
Sbjct: 1150 EGPSMRLLWNGEKTDAFKPLRGLRQGDPLSPYLFVLCIERLCHLIESSIAAKKWKPIKIS 1209
Query: 49 ELAPRC--MLFADDVVLVGES 67
+ PR + FADD++L E+
Sbjct: 1210 QSGPRLSHICFADDLILFAEA 1230
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.136 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,226,039
Number of Sequences: 26719
Number of extensions: 209944
Number of successful extensions: 501
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 38
Number of HSP's successfully gapped in prelim test: 17
Number of HSP's that attempted gapping in prelim test: 458
Number of HSP's gapped (non-prelim): 58
length of query: 239
length of database: 11,318,596
effective HSP length: 96
effective length of query: 143
effective length of database: 8,753,572
effective search space: 1251760796
effective search space used: 1251760796
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC141114.8


