

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141111.6 - phase: 0 /pseudo
(379 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
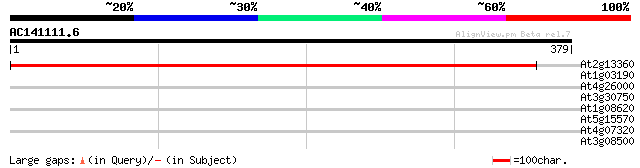
Score E
Sequences producing significant alignments: (bits) Value
At2g13360 alanine-glyoxylate aminotransferase 638 0.0
At1g03190 putative DNA repair protein 31 0.98
At4g26000 nucleic acid binding protein like 29 3.7
At3g30750 hypothetical protein 29 4.9
At1g08620 hypothetical protein 29 4.9
At5g15570 putative protein 28 8.3
At4g07320 28 8.3
At3g08500 putative protein 28 8.3
>At2g13360 alanine-glyoxylate aminotransferase
Length = 401
Score = 638 bits (1646), Expect = 0.0
Identities = 306/356 (85%), Positives = 339/356 (94%)
Query: 1 MDYVNGPGRNHLFVPGPVNIPDQVIRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTTTG 60
MDY+ GPGR+HLFVPGPVNIP+ VIRAM+RNNEDYRSPAIPALTKTLLEDVKKIFKTT+G
Sbjct: 1 MDYMYGPGRHHLFVPGPVNIPEPVIRAMNRNNEDYRSPAIPALTKTLLEDVKKIFKTTSG 60
Query: 61 TPFLIPTTGTGAWESALTNTLSPGDRIVSFVIGQFSLLWVDQQQRLKFNVDVVESEWGRG 120
TPFL PTTGTGAWESALTNTLSPGDRIVSF+IGQFSLLW+DQQ+RL FNVDVVES+WG+G
Sbjct: 61 TPFLFPTTGTGAWESALTNTLSPGDRIVSFLIGQFSLLWIDQQKRLNFNVDVVESDWGQG 120
Query: 121 ADLDILESKLASDSAHTIKAICIVHNETATGVTNNLAKVRQLLDAYQHPALLLVDGVSSI 180
A+L +L SKL+ D HTIKAICIVHNETATGVTN+++ VR LLD Y+HPALLLVDGVSSI
Sbjct: 121 ANLQVLASKLSQDENHTIKAICIVHNETATGVTNDISAVRTLLDHYKHPALLLVDGVSSI 180
Query: 181 CALDFRMDEWGVDVAITGSQKALSLPTGIGFVVASPKAIEASKSAKSLRVFFDWSDYLKF 240
CALDFRMDEWGVDVA+TGSQKALSLPTG+G V ASPKA+EA+K++KSL+VFFDW+DYLKF
Sbjct: 181 CALDFRMDEWGVDVALTGSQKALSLPTGLGIVCASPKALEATKTSKSLKVFFDWNDYLKF 240
Query: 241 YKMGTYWPYTPSIQLLYGLRAALDLIFEEGLENIIARHNRLGTATRLAVEAWGLKNCTQE 300
YK+GTYWPYTPSIQLLYGLRAALDLIFEEGLENIIARH RLG ATRLAVEAWGLKNCTQ+
Sbjct: 241 YKLGTYWPYTPSIQLLYGLRAALDLIFEEGLENIIARHARLGKATRLAVEAWGLKNCTQK 300
Query: 301 EEWFSDTVTAVVVPPYIDGAEIVRRAWKRYNLSLGLGLNKVAGKVFRIGHLGNLNE 356
EEW S+TVTAV+VPP+IDG+EIVRRAW+RYNLSLGLGLNKVAGKVFRIGHLGN+NE
Sbjct: 301 EEWISNTVTAVMVPPHIDGSEIVRRAWQRYNLSLGLGLNKVAGKVFRIGHLGNVNE 356
>At1g03190 putative DNA repair protein
Length = 758
Score = 31.2 bits (69), Expect = 0.98
Identities = 17/47 (36%), Positives = 23/47 (48%), Gaps = 6/47 (12%)
Query: 127 ESKLASDSAHTIKAICI------VHNETATGVTNNLAKVRQLLDAYQ 167
ES + D AH I +CI V T G NL K+RQ +D ++
Sbjct: 228 ESVVVFDEAHNIDNVCIEALSVSVRRVTLEGANRNLNKIRQEIDRFK 274
>At4g26000 nucleic acid binding protein like
Length = 495
Score = 29.3 bits (64), Expect = 3.7
Identities = 16/48 (33%), Positives = 25/48 (51%), Gaps = 9/48 (18%)
Query: 16 GPVNIPDQVIRAMSRNN-EDYRSPAIPALTKTLLEDVKKIFKTTTGTP 62
GPVN PD+++ + E Y SPA+ A V ++F+ +G P
Sbjct: 112 GPVNTPDRIVLISGKEEPEAYMSPAMDA--------VLRVFRRVSGLP 151
>At3g30750 hypothetical protein
Length = 111
Score = 28.9 bits (63), Expect = 4.9
Identities = 11/41 (26%), Positives = 19/41 (45%)
Query: 40 IPALTKTLLEDVKKIFKTTTGTPFLIPTTGTGAWESALTNT 80
+P + + L+DV+ +F +P G G W T+T
Sbjct: 4 VPHIDRQFLKDVEAVFTARARVNLPVPNPGLGTWVDRTTDT 44
>At1g08620 hypothetical protein
Length = 1183
Score = 28.9 bits (63), Expect = 4.9
Identities = 19/58 (32%), Positives = 26/58 (44%), Gaps = 7/58 (12%)
Query: 244 GTYWPY----TPSIQLLYGLRAALDL-IFEEGLENIIARHNRLGTATRLAVEAWGLKN 296
G YW T I++LYG A L+ +F G I + HN + + A W L N
Sbjct: 290 GEYWRIVDKATEEIEVLYG--ADLETGVFGSGFPKISSSHNASSSEDKYAKSGWNLNN 345
>At5g15570 putative protein
Length = 381
Score = 28.1 bits (61), Expect = 8.3
Identities = 10/27 (37%), Positives = 14/27 (51%)
Query: 280 RLGTATRLAVEAWGLKNCTQEEEWFSD 306
R TR A E W + C +E+ WF +
Sbjct: 332 RFKIGTRKASECWKINQCLEEKGWFKE 358
>At4g07320
Length = 437
Score = 28.1 bits (61), Expect = 8.3
Identities = 13/28 (46%), Positives = 17/28 (60%)
Query: 14 VPGPVNIPDQVIRAMSRNNEDYRSPAIP 41
VPGPVN P QVI + S + +D + P
Sbjct: 376 VPGPVNAPVQVILSDSEDGDDEEDDSNP 403
>At3g08500 putative protein
Length = 343
Score = 28.1 bits (61), Expect = 8.3
Identities = 17/79 (21%), Positives = 31/79 (38%)
Query: 4 VNGPGRNHLFVPGPVNIPDQVIRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTTTGTPF 63
VN G N L+ P +++PD M + + T+L+ + K F
Sbjct: 203 VNSVGLNQLYDPLMISVPDNGYHQMGNTVNVFSVNGLGDYGNTILDPISKRVSVEGDDWF 262
Query: 64 LIPTTGTGAWESALTNTLS 82
+ P+ T + +N L+
Sbjct: 263 IPPSENTNVIACSTSNNLN 281
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.138 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,305,031
Number of Sequences: 26719
Number of extensions: 336447
Number of successful extensions: 653
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 647
Number of HSP's gapped (non-prelim): 9
length of query: 379
length of database: 11,318,596
effective HSP length: 101
effective length of query: 278
effective length of database: 8,619,977
effective search space: 2396353606
effective search space used: 2396353606
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC141111.6


