

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141111.5 - phase: 0 /pseudo
(1478 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
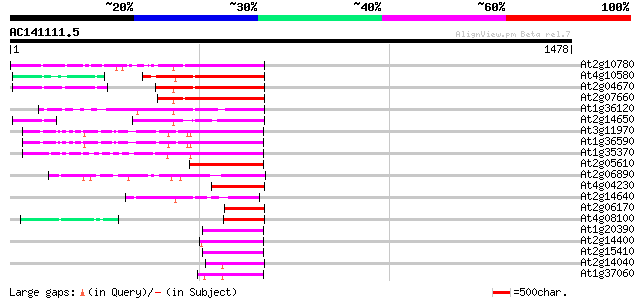
Score E
Sequences producing significant alignments: (bits) Value
At2g10780 pseudogene 355 8e-98
At4g10580 putative reverse-transcriptase -like protein 304 2e-82
At2g04670 putative retroelement pol polyprotein 292 9e-79
At2g07660 putative retroelement pol polyprotein 288 2e-77
At1g36120 putative reverse transcriptase gb|AAD22339.1 261 2e-69
At2g14650 putative retroelement pol polyprotein 256 7e-68
At3g11970 hypothetical protein 235 2e-61
At1g36590 hypothetical protein 235 2e-61
At1g35370 hypothetical protein 224 3e-58
At2g05610 putative retroelement pol polyprotein 205 1e-52
At2g06890 putative retroelement integrase 198 2e-50
At4g04230 putative transposon protein 171 3e-42
At2g14640 putative retroelement pol polyprotein 140 5e-33
At2g06170 putative Ty3-gypsy-like retroelement pol polyprotein 135 2e-31
At4g08100 putative polyprotein 127 5e-29
At1g20390 hypothetical protein 113 7e-25
At2g14400 putative retroelement pol polyprotein 112 1e-24
At2g15410 putative retroelement pol polyprotein 110 6e-24
At2g14040 putative retroelement pol polyprotein 105 1e-22
At1g37060 Athila retroelment ORF 1, putative 104 3e-22
>At2g10780 pseudogene
Length = 1611
Score = 355 bits (912), Expect = 8e-98
Identities = 254/718 (35%), Positives = 362/718 (50%), Gaps = 96/718 (13%)
Query: 3 AQAVGQQPAAGNGEVR-----MLETFLRNHPPAFKGRYDPDGAQTWLKEVERIFRVMQCS 57
++ V P +G R +LE R F G DP A W ++R F+ +C
Sbjct: 149 SEVVEVNPQPNHGRKRGDYLSLLEHVSRLGTRHFMGSTDPIVADEWRSRLKRNFKSTRCP 208
Query: 58 EVQKVRFGTHQLAEEADDWWVSLLPNLDQDGVDV-TWAVF*REFMRRYFPEDVRRKKEIE 116
E + H L +A +WW+++ + G +V ++A F EF ++YFP + + E
Sbjct: 209 EDYQRDIAVHFLEGDAHNWWLTVEK---RRGDEVRSFADFEDEFNKKYFPPEAWDRLECA 265
Query: 117 FLELKQGNMYVTEYAAKFVELAEFYPHYAAETVEFSKCIKFENGLRPDIKRAIEYQQIRV 176
+L+L QGN V EY +F L + E E ++ +F GLR +I+ +
Sbjct: 266 YLDLVQGNRTVREYDEEFNRLRRYVGRELEE--EQAQLRRFIRGLRIEIRNHCLVRTFNS 323
Query: 177 FPDLVNSCRIYEEDTKAHYKVVNERKTKGQQSRPKPYSAPADKGKQRMVDDRRPKKKDAP 236
+LV + EE + + +N K + ++ + PADK +R D K DA
Sbjct: 324 VSELVERAAMIEEGIEEE-RYLNREKAPIRNNQS---TKPADK--KRKFDKVDNTKSDAK 377
Query: 237 AEITCFNCGEKGHKSNVCPEEIKKCVRCGKKGHIVADCKRNDI-----------VCFNFN 285
C CG K H S C + I C RCG K H + C + + CF
Sbjct: 378 TG-ECVTCG-KNH-SGTCWKAIGACGRCGSKDHAIQSCPKMEPGQSKVLGEETRTCFYCG 434
Query: 286 EEGHIGSQCKQ---------------------PKKSPTTGRVFALDGTQTENEDRLIRCT 324
+ GH+ +C + PK+ RV+ L ++ N+ R
Sbjct: 435 KTGHLKRECPKLTAEKQAGQRDNRGGNGLPPPPKRQAVASRVYEL--SEEANDAGNFR-- 490
Query: 325 CYINNTPLVAIIDTGATHCFIAFDCVSALGLDLFDMNGEMVVETPAKGLVTTSLVCLRCL 384
AI + V A G G+ + T GLV V +
Sbjct: 491 ---------AITGGFRKEPNTDYGMVRAAG-------GQAMYPT---GLVRGISVVVN-- 529
Query: 385 LSMFGRDFEMDLVCLPLSGMDVILGMNWLEYNHVHINYFTKSVYFS---------SVEEE 435
G + DL+ +PL DVILGM+WL HI+ + F +
Sbjct: 530 ----GVNMPADLIIVPLKKHDVILGMDWLGKYKAHIDCHRGRIQFERDEGMLKFQGIRTT 585
Query: 436 SGAEFLSTKQLKQMERDGILMYPLMASLSFEN---QAVIDKLQVVCDFPEVFPDEIPDVP 492
SG+ +S Q ++M G Y +A+++ + A + + +V +F +VF + VP
Sbjct: 586 SGSLVISAIQAERMLGKGCEAY--LATITTKEVGASAELKDIPIVNEFSDVFA-AVSGVP 642
Query: 493 LEREVEFSIDLILGTKPVSMAPYRMSASELAKLKKQLEDLLEKKFVRPSVSPWGAPVLLV 552
+R F+I+L GT P+S APYRM+ +E+AKLKKQLE+LL+K F+RPS SPWGAPVL V
Sbjct: 643 PDRSDPFTIELEPGTTPISKAPYRMAPAEMAKLKKQLEELLDKGFIRPSSSPWGAPVLFV 702
Query: 553 KKKDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGAKVFSKIDLRSGYHQIKVKDE 612
KKKDGS RLCIDYR LNKVT+KN+YPLPRID+LMDQL GA+ FSKIDL SGYHQI ++
Sbjct: 703 KKKDGSFRLCIDYRGLNKVTVKNKYPLPRIDELMDQLGGAQWFSKIDLASGYHQIPIEPT 762
Query: 613 DMQKTAFSTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAFLDRFVVVFIDDILIYSK 670
D++KTAF TRY H+E+ VMPFG+TNAP FM+ MN +F FLD FV++FI+DIL+YSK
Sbjct: 763 DVRKTAFRTRYDHFEFVVMPFGLTNAPAAFMKMMNGVFRDFLDEFVIIFINDILVYSK 820
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 304 bits (779), Expect = 2e-82
Identities = 164/333 (49%), Positives = 216/333 (64%), Gaps = 22/333 (6%)
Query: 349 CVSALGLDLFDMNGE-MVVETPAKGLVTTSLVCLRCLLSMFGRDFEMDLVCLPLSGMDVI 407
C S LD D G+ + V AKG+ + + G DL+ P+ DVI
Sbjct: 331 CFSCGSLDHKDAGGQFLAVLGRAKGVD----------IQIAGESMPADLIISPVELYDVI 380
Query: 408 LGMNWLEYNHVHINYFTKSVYFSSVEEE---------SGAEFLSTKQLKQMERDGILMYP 458
LGM+WL+Y VH+++ V+F E SG+ +S Q ++M G Y
Sbjct: 381 LGMDWLDYYRVHLDWHRGRVFFERPEGRLVYQGVRPISGSLVISAVQAEKMIEKGCEAYL 440
Query: 459 LMASLSFE-NQAVIDKLQVVCDFPEVFPDEIPDVPLEREVEFSIDLILGTKPVSMAPYRM 517
+ S+ Q + ++VV +F +VF + +P + F+I+L GT P+S APYRM
Sbjct: 441 VTISMPESVGQVAVSDIRVVQEFQDVF-QSLQGLPPSQSDPFTIELEPGTAPLSKAPYRM 499
Query: 518 SASELAKLKKQLEDLLEKKFVRPSVSPWGAPVLLVKKKDGSMRLCIDYRQLNKVTIKNRY 577
+ +E+A+LKKQL+DLL K F+RPS SPWGAPVL VKKKDGS RLCIDYR+LN+VT+KNRY
Sbjct: 500 APAEMAELKKQLKDLLGKGFIRPSTSPWGAPVLFVKKKDGSFRLCIDYRELNRVTVKNRY 559
Query: 578 PLPRIDDLMDQLVGAKVFSKIDLRSGYHQIKVKDEDMQKTAFSTRYGHYEYKVMPFGVTN 637
PLPRID+L+DQL GA FSKIDL SGYHQI + + D++KTAF TRYGH+E+ VMPFG+TN
Sbjct: 560 PLPRIDELLDQLRGATCFSKIDLTSGYHQIPIAEADVRKTAFRTRYGHFEFVVMPFGLTN 619
Query: 638 APGVFMEYMNRIFHAFLDRFVVVFIDDILIYSK 670
AP VFM MN +F FLD FV++FIDDIL+YSK
Sbjct: 620 APAVFMRLMNSVFQEFLDEFVIIFIDDILVYSK 652
Score = 72.8 bits (177), Expect = 1e-12
Identities = 70/251 (27%), Positives = 100/251 (38%), Gaps = 43/251 (17%)
Query: 8 QQPAAGNGEV----RMLETFLRNHPPAFKGRYDPDGAQTWLKEVERIFRVMQCSEVQKVR 63
QQ AA EV RM+E R F G P+ A +W V R F +C +V
Sbjct: 125 QQRAAVAEEVLSYLRMMEQLQRIDTGYFSGGTSPEEADSWRSRVGRNFGSSRCPAEYRVD 184
Query: 64 FGTHQLAEEADDWWVSLLPNLDQDGVDVTWAVF*REFMRRYFPEDVRRKKEIEFLELKQG 123
H L +A WW S+ Q D++WA F EF +YFP++
Sbjct: 185 LAVHFLEGDAHLWWRSVTARRRQ--TDMSWADFVAEFKAKYFPQEA-------------- 228
Query: 124 NMYVTEYAAKFVELAEFYPHYAAETVEFSKCIKFENGLRPDIKRAIEYQQIRVFPDLVNS 183
+ YA + +E + ++ +F GLRPD++ Q LV +
Sbjct: 229 ---LDPYAGQGME------------DDQAQMRRFLRGLRPDLRVRCRVSQYATKAALVET 273
Query: 184 CRIYEEDTKAHY----KVVNERKTKGQQSRPKPYSAPADKGKQRMVDDRRPKKKDAPAEI 239
EED + VV +KT+ QQ P PA +G++R D P +
Sbjct: 274 AAEVEEDFQRQVVGVSPVVQPKKTQ-QQVTPSKGGKPA-QGQKRKWD--HPSRAGQGGRA 329
Query: 240 TCFNCGEKGHK 250
CF+CG HK
Sbjct: 330 RCFSCGSLDHK 340
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 292 bits (748), Expect = 9e-79
Identities = 151/296 (51%), Positives = 196/296 (66%), Gaps = 11/296 (3%)
Query: 385 LSMFGRDFEMDLVCLPLSGMDVILGMNWLEYNHVHINYFTKSVYFS---------SVEEE 435
+ + G DL+ P+ DVILGM+WL++ VH++ V F V
Sbjct: 384 IQIAGESMPADLIISPVELYDVILGMDWLDHYRVHLDCHRGRVSFERPEGRLVYQGVRPT 443
Query: 436 SGAEFLSTKQLKQMERDGILMYPLMASLSFE-NQAVIDKLQVVCDFPEVFPDEIPDVPLE 494
SG+ +S Q K+M G Y + S+ Q + ++V+ +F +VF + +P
Sbjct: 444 SGSLVISAVQAKKMIEKGCEAYLVTISMPESLGQVAVSDIRVIQEFEDVF-QSLQGLPPS 502
Query: 495 REVEFSIDLILGTKPVSMAPYRMSASELAKLKKQLEDLLEKKFVRPSVSPWGAPVLLVKK 554
R F+I+L GT P+S APYRM+ +E+ +LKKQLEDLL K F+RPS SPWGAPVL VKK
Sbjct: 503 RSDPFTIELEPGTAPLSKAPYRMAPAEMTELKKQLEDLLGKGFIRPSTSPWGAPVLFVKK 562
Query: 555 KDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGAKVFSKIDLRSGYHQIKVKDEDM 614
KDGS RLCIDYR LN VT+KN+YPLPRID+L+DQL GA FSKIDL SGYHQI + + D+
Sbjct: 563 KDGSFRLCIDYRGLNWVTVKNKYPLPRIDELLDQLRGATCFSKIDLTSGYHQIPIAEADV 622
Query: 615 QKTAFSTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAFLDRFVVVFIDDILIYSK 670
+KTAF TRYGH+E+ VMPF +TNAP FM MN +F FLD FV++FIDDIL+YSK
Sbjct: 623 RKTAFRTRYGHFEFVVMPFALTNAPAAFMRLMNSVFQEFLDEFVIIFIDDILVYSK 678
Score = 92.0 bits (227), Expect = 2e-18
Identities = 79/258 (30%), Positives = 109/258 (41%), Gaps = 18/258 (6%)
Query: 8 QQPAAGNGEV----RMLETFLRNHPPAFKGRYDPDGAQTWLKEVERIFRVMQCSEVQKVR 63
QQ AA EV RM+E R F P+ A +W V+R F +C +V
Sbjct: 111 QQRAAVAEEVPSYLRMMEQLQRIGTWYFSSGTSPEEADSWRSRVQRNFGSSRCPAEYRVD 170
Query: 64 FGTHQLAEEADDWWVSLLPNLDQDGVDVTWAVF*REFMRRYFPEDVRRKKEIEFLELKQG 123
H L +A WW S+ Q D++WA F EF +YFP++ + E FLEL QG
Sbjct: 171 LAVHFLEGDAHLWWRSVTARRRQ--ADMSWADFVAEFNAKYFPQEALDRMEARFLELTQG 228
Query: 124 NMYVTEYAAKFVELAEFYPHYAAETVE--FSKCIKFENGLRPDIKRAIEYQQIRVFPDLV 181
V EY +F L YA +E ++ +F GLRPD++ Q LV
Sbjct: 229 EWSVREYDREFNRLLA----YAGRGMEDDQAQMRRFLRGLRPDLRVQCRVSQYATKAALV 284
Query: 182 NSCRIYEEDTKAHYKVVN---ERKTKGQQSRPKPYSAPADKGKQRMVDDRRPKKKDAPAE 238
+ EED + V+ + K QQ P PA +G++R D P +
Sbjct: 285 ETAAEVEEDLQRQVVGVSPAVQTKNTQQQVTPSKGGKPA-QGQKRKWD--HPSRAGQGGR 341
Query: 239 ITCFNCGEKGHKSNVCPE 256
F+CG H C E
Sbjct: 342 AGYFSCGSLDHTVADCTE 359
>At2g07660 putative retroelement pol polyprotein
Length = 949
Score = 288 bits (736), Expect = 2e-77
Identities = 152/294 (51%), Positives = 202/294 (68%), Gaps = 15/294 (5%)
Query: 389 GRDFEMDLVCLPLSGMDVILGMNWLEYNHVHINYFTKSVYFS---------SVEEESGAE 439
G + DL+ +PL DVILGM+WL HI+ + + F + SG+
Sbjct: 25 GVNMPADLIIVPLKKHDVILGMDWLGKYKGHIDCHRERIQFERDEGMLKFQGIRTTSGSL 84
Query: 440 FLSTKQLKQMERDGILMYPLMASLSFEN---QAVIDKLQVVCDFPEVFPDEIPDVPLERE 496
+S Q ++M G Y +A+++ + A + + +V +F +VF + VP +R
Sbjct: 85 VISAIQAERMLGKGCEAY--LATITTKEVGASAELKDILIVNEFSDVFA-AVSGVPPDRS 141
Query: 497 VEFSIDLILGTKPVSMAPYRMSASELAKLKKQLEDLLEKKFVRPSVSPWGAPVLLVKKKD 556
F+I+L GT P+S APYRM+ +E+A+LKKQLE+LL K F+RPS SPWGAPVL VKKKD
Sbjct: 142 DPFTIELEPGTTPISKAPYRMAPAEMAELKKQLEELLAKGFIRPSSSPWGAPVLFVKKKD 201
Query: 557 GSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGAKVFSKIDLRSGYHQIKVKDEDMQK 616
GS RLCIDYR LNKVT+KN+YPLPRID+LMDQL GA+ FSKIDL SGYHQI ++ D++K
Sbjct: 202 GSFRLCIDYRGLNKVTVKNKYPLPRIDELMDQLGGAQWFSKIDLASGYHQIPIEPTDVRK 261
Query: 617 TAFSTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAFLDRFVVVFIDDILIYSK 670
TAF TRYGH+E+ VMPFG+TNAP FM+ MN +F FLD FV++FIDDIL++SK
Sbjct: 262 TAFRTRYGHFEFVVMPFGLTNAPAAFMKMMNGVFRDFLDEFVIIFIDDILVHSK 315
>At1g36120 putative reverse transcriptase gb|AAD22339.1
Length = 1235
Score = 261 bits (667), Expect = 2e-69
Identities = 200/619 (32%), Positives = 290/619 (46%), Gaps = 108/619 (17%)
Query: 77 WVSLLPNLDQDGVDVTWAVF*REFMRRYFPEDVRRKKEIEFLELKQGNMYVTEYAAKFVE 136
++ ++ L + G D F EF +YFP++ + E FLEL +G + V EY +F
Sbjct: 143 YLRMMEQLQRIGTD-----FVAEFNAKYFPQEALDRMEARFLELTKGELSVREYGREFKR 197
Query: 137 LAEFYPHYAAETVEFSKCI--KFENGLRPDIKRAIEYQQIRVFPDLVNSCRIYEEDTKAH 194
L YA +E + +F GLRPD++ +R CR+ + TKA
Sbjct: 198 LLV----YAGRGMEDDQAQMRRFLRGLRPDLR-------VR--------CRVSQYATKAA 238
Query: 195 YKVVNERKTKGQQSRPKPYSAPADKGKQRMVDDRRPKKKDAPAEITCFNCGEKGHKSNVC 254
+A ++ QR V P +V
Sbjct: 239 LT-----------------AAEVEEDLQRQVVAVSP---------------------SVQ 260
Query: 255 PEEIKKCVRCGKKGHIVADCKRNDIVCFNFNEEGHIGSQCKQPKKSPTTGRVFALDGTQT 314
P++I++ V K G KR F + G GS S G + ++
Sbjct: 261 PKKIQQQVAPSKGGKPAQVQKRKWDHPFRAGQGGRAGSAADDSSTSTAGGAAQSAGAARS 320
Query: 315 ENEDRLIRCTC----YINNTPLVA-------IIDTGATHCFIAFDCVSALGLDLFDMNGE 363
E + C ++ L++ + D+GA+HCFI + SA ++ GE
Sbjct: 321 AAECAAVSTYCSGTTWLYYADLISGRCRGHVLFDSGASHCFITPE--SASRDNIRGEPGE 378
Query: 364 M--VVETPAKGLVTTSLVCLRCLLSMFGRDFEMDLVCLPLSGMDVILGMNWLEYNHVHIN 421
V+ +T + + DL+ P+ D ILGM+WL++ VH++
Sbjct: 379 QFGAVKIAGGQFLTVLGRAKGVNIQIEEESMPADLIINPVVLYDAILGMDWLDHYRVHLD 438
Query: 422 YFTKSVYFS---------SVEEESGAEFLSTKQLKQMERDGILMYPLMASLSFE-NQAVI 471
V F V S + +S Q+++M G Y + S+S Q +
Sbjct: 439 CHRGRVSFERPEGRLVYQGVRPTSRSLVISEVQVEKMIEKGCEAYLVTISMSESVGQVAV 498
Query: 472 DKLQVVCDFPEVFPDEIPDVPLEREVEFSIDLILGTKPVSMAPYRMSASELAKLKKQLED 531
+ VV +F +VF + +P R F+I+L LGT P+S PYRM +E+A+LKKQLED
Sbjct: 499 SDIWVVQEFEDVF-QSLQGLPPSRSDPFTIELELGTAPLSKTPYRMVPAEIAELKKQLED 557
Query: 532 LLEKKFVRPSVSPWGAPVLLVKKKDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVG 591
LL K F+RPS S WGAP LN+VT+KN+YPLPRID+L+DQL G
Sbjct: 558 LLGKGFIRPSTSRWGAP------------------GLNRVTLKNKYPLPRIDELLDQLRG 599
Query: 592 AKVFSKIDLRSGYHQIKVKDEDMQKTAFSTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFH 651
A FSKIDL GYHQ + + D++KTAF TRYGH+E+ VMPFG+TNAP M MN +F
Sbjct: 600 ATCFSKIDLTPGYHQFPIAEADVRKTAFRTRYGHFEFVVMPFGLTNAPTALMRLMNSVFQ 659
Query: 652 AFLDRFVVVFIDDILIYSK 670
FLD FV++FIDDIL+Y K
Sbjct: 660 EFLDEFVIIFIDDILVYFK 678
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 256 bits (654), Expect = 7e-68
Identities = 153/357 (42%), Positives = 201/357 (55%), Gaps = 44/357 (12%)
Query: 324 TCYINNTPLVAIIDTGATHCFIAFDCVSALGLDLFDMNGEMVVETPAKGLVTTSLVCLRC 383
T + + D+GA+HCFI + S G D ++ A G L +
Sbjct: 305 TLLVGGVEAHVLFDSGASHCFITPESASR-GNIRGDPGEQLGAVKVAGGQFVAVLGRTKG 363
Query: 384 L-LSMFGRDFEMDLVCLPLSGMDVILGMNWLEYNHVHINYFTKSVYFS---------SVE 433
+ + + G DL+ P+ DVILGM+WL++ VH++ V F V
Sbjct: 364 VDIQIAGESMPADLIISPVELYDVILGMDWLDHYRVHLDCHRGRVSFERPEGSLVYQGVR 423
Query: 434 EESGAEFLSTKQLKQMERDGILMYPLMASLSFENQAVIDKLQVVCDFPEVFPDEIPDVPL 493
SG+ +S Q ++M G Y +F +VF + +P
Sbjct: 424 PTSGSLVISAVQAEKMIEKGCEAY--------------------LEFEDVF-QSLQGLPP 462
Query: 494 EREVEFSIDLILGTKPVSMAPYRMSASELAKLKKQLEDLLEKKFVRPSVSPWGAPVLLVK 553
R F+I+L GT P+S APYRM+ +E+A+LKKQLEDLL GAPVL VK
Sbjct: 463 SRSDPFTIELEPGTAPLSKAPYRMAPAEMAELKKQLEDLL------------GAPVLFVK 510
Query: 554 KKDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGAKVFSKIDLRSGYHQIKVKDED 613
KKDGS RLCIDYR LN VT+KN+YPLPRID+L+DQL GA FSKIDL SGYH I + + D
Sbjct: 511 KKDGSFRLCIDYRGLNWVTVKNKYPLPRIDELLDQLRGATCFSKIDLTSGYHLIPIAEAD 570
Query: 614 MQKTAFSTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAFLDRFVVVFIDDILIYSK 670
++KTAF TRYGH+E+ VMPFG+TNAP FM MN +F LD FV++FIDDIL+YSK
Sbjct: 571 VRKTAFRTRYGHFEFVVMPFGLTNAPAAFMRLMNSVFQEVLDEFVIIFIDDILVYSK 627
Score = 67.8 bits (164), Expect = 4e-11
Identities = 43/120 (35%), Positives = 57/120 (46%), Gaps = 6/120 (5%)
Query: 8 QQPAAGNGEV----RMLETFLRNHPPAFKGRYDPDGAQTWLKEVERIFRVMQCSEVQKVR 63
QQ AA EV RM+E R F G P+ A +W VER F +C ++
Sbjct: 129 QQRAAVAEEVPSYLRMMEQLQRIGTGYFSGGTSPEVADSWRSRVERNFGSSRCPAEYRID 188
Query: 64 FGTHQLAEEADDWWVSLLPNLDQDGVDVTWAVF*REFMRRYFPEDVRRKKEIEFLELKQG 123
H L +A WW S+ Q D++WA F EF +YFP++ + E FLEL QG
Sbjct: 189 LAVHFLEGDAHLWWRSVTARRRQ--ADISWADFVAEFNAKYFPQEALDRMEAHFLELTQG 246
>At3g11970 hypothetical protein
Length = 1499
Score = 235 bits (599), Expect = 2e-61
Identities = 204/700 (29%), Positives = 321/700 (45%), Gaps = 118/700 (16%)
Query: 34 RYDPDGAQTWLKEVERIFRVMQCSEVQKVRFGTHQLAEEADDWWVSLLPNLDQDGVDVT- 92
R+D + WL +VE F V E KV+ A W S + Q GV +
Sbjct: 116 RFDGTRLKEWLFKVEEFFGVDSTPEDMKVKMAAIHFDSHASTWHQSFI----QSGVGLEV 171
Query: 93 ---WAVF*REFMRRYFPEDVRRKKEIEFLELKQGNMYVTEYAAKFVELAEFYPHYAAETV 149
W + + R+ + E++ L+ G + +Y KF EL + + + E +
Sbjct: 172 LYDWKGYVKLLKERFEDDCDDPMAELKHLQETDG---IIDYHQKF-ELIKTRVNLSEEYL 227
Query: 150 EFSKCIKFENGLRPDIKRAIEYQQIRVF-PDLVNSC----RIYEEDTKAH---------- 194
+ GLR D + +R+F P V C + YE KAH
Sbjct: 228 ----VSVYLAGLRTDTQ-----MHVRMFQPQTVRHCLFLGKTYE---KAHPKKPANTTWS 275
Query: 195 ------------YKVVNERKTK--GQQSRPKPYSAPADKGKQRMVDDRRPKKKDAPAEIT 240
Y+ E KT G + KP S K Q+ + DRR K
Sbjct: 276 TNRSAPTGGYNKYQKEGESKTDHYGNKGNFKPVSQQPKKMSQQEMSDRRSKG-------L 328
Query: 241 CFNCGEKGHKSNVCPEEIKKCVRCGKKGHIVADCKRNDIVCFNFNEEGHIGSQCKQPKKS 300
C+ C EK + + + R + R ++V N++ Q S
Sbjct: 329 CYFCDEKYTPEHYLVHKKTQLFRMDVDEEF--EDAREELV----NDDDEHMPQISVNAVS 382
Query: 301 PTTG-RVFALDGTQTENEDRLIRCTCYINNTPLVAIIDTGATHCFIAFDCVSALG--LDL 357
G + + GT D+ I + +ID+G+TH F+ + + LG +D
Sbjct: 383 GIAGYKTMRVKGTY----DKKI----------IFILIDSGSTHNFLDPNTAAKLGCKVDT 428
Query: 358 FDMNGEMVVE---TPAKGLVTTSLVCLRCLLSMFGRDFEMDLVCLPLSGMDVILGMNWLE 414
+ V + +G VT L+ F+ D++ +PL G+D++LG+ WLE
Sbjct: 429 AGLTRVSVADGRKLRVEGKVTDFSWKLQTTT------FQSDILLIPLQGIDMVLGVQWLE 482
Query: 415 YN----------HVHINYFTKSVYFSSVEEESGAEFLSTKQLKQMERDGILMYPLMASLS 464
+ + + V + S E + ++L++++ D + + L
Sbjct: 483 TLGRISWEFKKLEMRFKFNNQKVLLHGLTSGSVRE-VKAQKLQKLQEDQVQLAMLCVQEV 541
Query: 465 FEN--------QAVIDKL-------QVVCDFPEVFPDEIPDVPLEREVEFSIDLILGTKP 509
E+ A+ +L +V+ ++P++F + P + I L+ G+ P
Sbjct: 542 SESTEGELCTINALTSELGEESVVEEVLNEYPDIFIEPTALPPFREKHNHKIKLLEGSNP 601
Query: 510 VSMAPYRMSASELAKLKKQLEDLLEKKFVRPSVSPWGAPVLLVKKKDGSMRLCIDYRQLN 569
V+ PYR S + ++ K +EDLL V+ S SP+ +PV+LVKKKDG+ RLC+DYR+LN
Sbjct: 602 VNQRPYRYSIHQKNEIDKLVEDLLTNGTVQASSSPYASPVVLVKKKDGTWRLCVDYRELN 661
Query: 570 KVTIKNRYPLPRIDDLMDQLVGAKVFSKIDLRSGYHQIKVKDEDMQKTAFSTRYGHYEYK 629
+T+K+ +P+P I+DLMD+L GA +FSKIDLR+GYHQ+++ +D+QKTAF T GH+EY
Sbjct: 662 GMTVKDSFPIPLIEDLMDELGGAVIFSKIDLRAGYHQVRMDPDDIQKTAFKTHSGHFEYL 721
Query: 630 VMPFGVTNAPGVFMEYMNRIFHAFLDRFVVVFIDDILIYS 669
VMPFG+TNAP F MN IF FL +FV+VF DDIL+YS
Sbjct: 722 VMPFGLTNAPATFQGLMNFIFKPFLRKFVLVFFDDILVYS 761
>At1g36590 hypothetical protein
Length = 1499
Score = 235 bits (599), Expect = 2e-61
Identities = 204/700 (29%), Positives = 321/700 (45%), Gaps = 118/700 (16%)
Query: 34 RYDPDGAQTWLKEVERIFRVMQCSEVQKVRFGTHQLAEEADDWWVSLLPNLDQDGVDVT- 92
R+D + WL +VE F V E KV+ A W S + Q GV +
Sbjct: 116 RFDGTRLKEWLFKVEEFFGVDSTPEDMKVKMAAIHFDSHASTWHQSFI----QSGVGLEV 171
Query: 93 ---WAVF*REFMRRYFPEDVRRKKEIEFLELKQGNMYVTEYAAKFVELAEFYPHYAAETV 149
W + + R+ + E++ L+ G + +Y KF EL + + + E +
Sbjct: 172 LYDWKGYVKLLKERFEDDCDDPMAELKHLQETDG---IIDYHQKF-ELIKTRVNLSEEYL 227
Query: 150 EFSKCIKFENGLRPDIKRAIEYQQIRVF-PDLVNSC----RIYEEDTKAH---------- 194
+ GLR D + +R+F P V C + YE KAH
Sbjct: 228 ----VSVYLAGLRTDTQ-----MHVRMFQPQTVRHCLFLGKTYE---KAHLKKPANTTWS 275
Query: 195 ------------YKVVNERKTK--GQQSRPKPYSAPADKGKQRMVDDRRPKKKDAPAEIT 240
Y+ E KT G + KP S K Q+ + DRR K
Sbjct: 276 TNRSAPTGGYNKYQKEGESKTDHYGNKGNFKPVSQQPKKMSQQEMSDRRSKG-------L 328
Query: 241 CFNCGEKGHKSNVCPEEIKKCVRCGKKGHIVADCKRNDIVCFNFNEEGHIGSQCKQPKKS 300
C+ C EK + + + R + R ++V N++ Q S
Sbjct: 329 CYFCDEKYTPEHYLVHKKTQLFRMDVDEEF--EDAREELV----NDDDEHMPQISVNAVS 382
Query: 301 PTTG-RVFALDGTQTENEDRLIRCTCYINNTPLVAIIDTGATHCFIAFDCVSALG--LDL 357
G + + GT D+ I + +ID+G+TH F+ + + LG +D
Sbjct: 383 GIAGYKTMRVKGTY----DKKI----------IFILIDSGSTHNFLDPNTAAKLGCKVDT 428
Query: 358 FDMNGEMVVE---TPAKGLVTTSLVCLRCLLSMFGRDFEMDLVCLPLSGMDVILGMNWLE 414
+ V + +G VT L+ F+ D++ +PL G+D++LG+ WLE
Sbjct: 429 AGLTRVSVADGRKLRVEGKVTDFSWKLQTTT------FQSDILLIPLQGIDMVLGVQWLE 482
Query: 415 YN----------HVHINYFTKSVYFSSVEEESGAEFLSTKQLKQMERDGILMYPLMASLS 464
+ + + V + S E + ++L++++ D + + L
Sbjct: 483 TLGRISWEFKKLEMRFKFNNQKVLLHGLTSGSVRE-VKAQKLQKLQEDQVQLAMLCVQEV 541
Query: 465 FEN--------QAVIDKL-------QVVCDFPEVFPDEIPDVPLEREVEFSIDLILGTKP 509
E+ A+ +L +V+ ++P++F + P + I L+ G+ P
Sbjct: 542 SESTEGELCTINALTSELGEESVVEEVLNEYPDIFIEPTALPPFREKHNHKIKLLEGSNP 601
Query: 510 VSMAPYRMSASELAKLKKQLEDLLEKKFVRPSVSPWGAPVLLVKKKDGSMRLCIDYRQLN 569
V+ PYR S + ++ K +EDLL V+ S SP+ +PV+LVKKKDG+ RLC+DYR+LN
Sbjct: 602 VNQRPYRYSIHQKNEIDKLVEDLLTNGTVQASSSPYASPVVLVKKKDGTWRLCVDYRELN 661
Query: 570 KVTIKNRYPLPRIDDLMDQLVGAKVFSKIDLRSGYHQIKVKDEDMQKTAFSTRYGHYEYK 629
+T+K+ +P+P I+DLMD+L GA +FSKIDLR+GYHQ+++ +D+QKTAF T GH+EY
Sbjct: 662 GMTVKDSFPIPLIEDLMDELGGAVIFSKIDLRAGYHQVRMDPDDIQKTAFKTHSGHFEYL 721
Query: 630 VMPFGVTNAPGVFMEYMNRIFHAFLDRFVVVFIDDILIYS 669
VMPFG+TNAP F MN IF FL +FV+VF DDIL+YS
Sbjct: 722 VMPFGLTNAPATFQGLMNFIFKPFLRKFVLVFFDDILVYS 761
>At1g35370 hypothetical protein
Length = 1447
Score = 224 bits (571), Expect = 3e-58
Identities = 189/662 (28%), Positives = 308/662 (45%), Gaps = 68/662 (10%)
Query: 34 RYDPDGAQTWLKEVERIFRVMQCSEVQKVRFGTHQLAEEADDWWVSLLPNLDQDGVDVTW 93
R+D WL +VE F V E KV+ A W S + + V W
Sbjct: 121 RFDGSRINEWLFKVEEFFGVDFTPEEMKVKMVAIHFDSHAATWHHSFIQSGIGLDVFFNW 180
Query: 94 AVF*REFMRRYFPEDVRRKKEIEFLELKQGNMYVTEYAAKFVELAEFYPHYAAETVEFSK 153
+ + R+ ED E +L++ + V EY +F EL + + + E +
Sbjct: 181 PEYVKLLKDRF--EDACDDPMAELKKLQETDGIV-EYHQQF-ELIKVRLNLSEEYL---- 232
Query: 154 CIKFENGLRPDIKRAIEYQQIRVFPDLVNSCRIYEEDTKAHYK--VVNERKTKGQQSRPK 211
+ GLR D + + + + D + + YE +AH K V + KG +S
Sbjct: 233 VSVYLAGLRTDTQMHVRMFEPKTVRDCLRLGKYYE---RAHPKKTVSSTWSQKGTRSG-- 287
Query: 212 PYSAPADKGKQRMVDDRRPKKKDAPAEITCFNCGEKGHKSNVCPEEIKKCVRCGKKGHIV 271
G R V + K C+ C EK + + + R
Sbjct: 288 --------GSYRPVKEVEQKSDHLGL---CYFCDEKFTPEHYLVHKKTQLFRMD------ 330
Query: 272 ADCKRNDIVCFNFNEEGHIGSQCKQPKKSPTTGRVFALDGTQTENEDRLIRCTCYINNTP 331
D + D V +++ ++P + V + G +T ++ T ++
Sbjct: 331 VDEEFEDAVEVLSDDDHE-----QKPMPQISVNAVSGISGYKTMG----VKGT--VDKRD 379
Query: 332 LVAIIDTGATHCFIAFDCVSALGLDLFDMNGEMVVETPAKGLVTTSLVCLRCLLSMFGRD 391
L +ID+G+TH FI + LG + V + L + +
Sbjct: 380 LFILIDSGSTHNFIDSTVAAKLGCHVESAGLTKVAVADGRKLNVDGQI-KGFTWKLQSTT 438
Query: 392 FEMDLVCLPLSGMDVILGMNWLE--------YNHVHINYFTKS--VYFSSVEEESGAEFL 441
F+ D++ +PL G+D++LG+ WLE + + + +F K+ V+ + S +
Sbjct: 439 FQSDILLIPLQGVDMVLGVQWLETLGRISWEFKKLEMQFFYKNQRVWLHGIITGSVRDIK 498
Query: 442 STK-QLKQMERDGILMYPLMASLSFENQAV--IDKL-----------QVVCDFPEVFPDE 487
+ K Q Q ++ + M + +S E Q + I L +V +FP+VF +
Sbjct: 499 AHKLQKTQADQIQLAMVCVREVVSDEEQEIGSISALTSDVVEESVVQNIVEEFPDVFAEP 558
Query: 488 IPDVPLEREVEFSIDLILGTKPVSMAPYRMSASELAKLKKQLEDLLEKKFVRPSVSPWGA 547
P + + I L+ G PV+ PYR + ++ K ++D+++ ++ S SP+ +
Sbjct: 559 TDLPPFREKHDHKIKLLEGANPVNQRPYRYVVHQKDEIDKIVQDMIKSGTIQVSSSPFAS 618
Query: 548 PVLLVKKKDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGAKVFSKIDLRSGYHQI 607
PV+LVKKKDG+ RLC+DY +LN +T+K+R+ +P I+DLMD+L G+ VFSKIDLR+GYHQ+
Sbjct: 619 PVVLVKKKDGTWRLCVDYTELNGMTVKDRFLIPLIEDLMDELGGSVVFSKIDLRAGYHQV 678
Query: 608 KVKDEDMQKTAFSTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAFLDRFVVVFIDDILI 667
++ +D+QKTAF T GH+EY VM FG+TNAP F MN +F FL +FV+VF DDILI
Sbjct: 679 RMDPDDIQKTAFKTHNGHFEYLVMLFGLTNAPATFQSLMNSVFRDFLRKFVLVFFDDILI 738
Query: 668 YS 669
YS
Sbjct: 739 YS 740
>At2g05610 putative retroelement pol polyprotein
Length = 780
Score = 205 bits (522), Expect = 1e-52
Identities = 96/195 (49%), Positives = 137/195 (70%)
Query: 475 QVVCDFPEVFPDEIPDVPLEREVEFSIDLILGTKPVSMAPYRMSASELAKLKKQLEDLLE 534
+VV +FP++F + P + I+L+ + PV+ PYR + ++ K ++D+L
Sbjct: 9 KVVTEFPDIFVEPTDLPPFRAPHDHKIELLEDSNPVNQRPYRYVVHQKNEIDKIVDDMLA 68
Query: 535 KKFVRPSVSPWGAPVLLVKKKDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGAKV 594
++ S SP+ +PV+LVKKKDG+ RLC+DYR+LN +T+K+R+P+P I+DLMD+L G+ V
Sbjct: 69 SGTIQASSSPYASPVVLVKKKDGTWRLCVDYRELNGMTVKDRFPIPLIEDLMDELGGSNV 128
Query: 595 FSKIDLRSGYHQIKVKDEDMQKTAFSTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAFL 654
+SKIDLR+GYHQ+++ D+ KTAF T GHYEY VMPFG+TNAP F MN F FL
Sbjct: 129 YSKIDLRAGYHQVRMDPLDIHKTAFKTHNGHYEYLVMPFGLTNAPASFQSLMNSFFKPFL 188
Query: 655 DRFVVVFIDDILIYS 669
+FV+VF DDILIYS
Sbjct: 189 RKFVLVFFDDILIYS 203
>At2g06890 putative retroelement integrase
Length = 1215
Score = 198 bits (504), Expect = 2e-50
Identities = 168/639 (26%), Positives = 273/639 (42%), Gaps = 129/639 (20%)
Query: 102 RRYFPEDVRRKKEIEFLELKQGNMYVTEYAAKFVELAEFYPHYAAETVEFSKCIKFENGL 161
+R+ P R+ ++ L QGN V EY + E+ + +F L
Sbjct: 7 KRFVPSHYHRELHLKLRNLTQGNRSVEEY---YKEMETLMLRADISEDREATLSRFLGDL 63
Query: 162 RPDIKRAIEYQQIRVFPDLVNSCRIYEEDTK--------------AHYKVVNERKTKGQQ 207
DI+ +E Q ++++ ++E+ K A E +T
Sbjct: 64 NRDIQDRLETQYYVQIEEMLHKAILFEQQVKRKSSSRSSYGSGTIAKPTYQREERTSSYH 123
Query: 208 SRP--------KPYSAPAD-KGKQRMVDDRRPKKKDAPAEITCFNCGEKGHKSNVCPEEI 258
++P KPY+A D KGK + R ++ C+ C KGH +N CP
Sbjct: 124 NKPIVSPRAESKPYAAVQDHKGKAEISTSR-------VRDVRCYKCQGKGHYANECPN-- 174
Query: 259 KKCVRCGKKGHIVADCKRNDIVCFNFNEEGHIGSQCKQPKKSPTTGRVFALDGT------ 312
K+ + G I + E S K+ ++ P G + T
Sbjct: 175 KRVMILLDNGEIEPE-----------EEIPDSPSSLKENEELPAQGELLVARRTLSVQTK 223
Query: 313 --QTENEDRLIRCTCYINNTPLVAIIDTGATHCFIAFDCVSALGLDLFDMNGEMVVETPA 370
+ E L C+++ IID G+ + V LGL + +G+M V
Sbjct: 224 TDEQEQRKNLFHTRCHVHGKVCSLIIDGGSCTNVASETMVKKLGLKWLNDSGKMRV---- 279
Query: 371 KGLVTTSLVCLRCLLSMFGRDFEMDLVC--LPLSGMDVILGMNWLEYNHVHINYFT---- 424
K V +V + +E +++C LP+ ++LG W V + FT
Sbjct: 280 KNQVVVPIVIGK---------YEDEILCDVLPMEAGHILLGRPWQSDRKVMHDGFTNRHS 330
Query: 425 ------KSVYFSSVEEESGAEFLSTKQLKQM-------------------ERDGILMYPL 459
K++ S E + + KQ K+ + +L++
Sbjct: 331 FEFKGGKTILVSMTPHEVYQDQIHLKQKKEQVVKQPNFFAKSGEVKSAYSSKQPMLLFVF 390
Query: 460 MASL-SFENQAVI---DKLQVVCDFPEVFPDEIPD-VPLEREVEFSIDLILGTKPVSMAP 514
+L S N A + + ++ D+ +VFP++ P +P R +E ID + G +
Sbjct: 391 KEALTSLTNFAPVLPSEMTSLLQDYKDVFPEDNPKGLPPIRGIEHQIDFVPGASLPNRPA 450
Query: 515 YRMSASELAKLKKQLEDLLEKKFVRPSVSPWGAPVLLVKKKDGSMRLCIDYRQLNKVTIK 574
YR + E +L++Q KDGS R+C D R +N VT+K
Sbjct: 451 YRTNPVETKELQRQ--------------------------KDGSWRMCFDCRAINNVTVK 484
Query: 575 NRYPLPRIDDLMDQLVGAKVFSKIDLRSGYHQIKVKDEDMQKTAFSTRYGHYEYKVMPFG 634
+P+PR+DD++D+L G+ +FSKIDL+SGYHQI++ + D KTAF T++G YE+ VMPFG
Sbjct: 485 YCHPIPRLDDMLDELHGSSIFSKIDLKSGYHQIRMNEGDEWKTAFKTKHGLYEWLVMPFG 544
Query: 635 VTNAPGVFMEYMNRIFHAFLDRFVVVFIDDILIYSKD*R 673
+T+AP FM MN + AF+ FV+V+ DDIL+YS+ R
Sbjct: 545 LTHAPSTFMRLMNHVLRAFIGIFVIVYFDDILVYSESLR 583
>At4g04230 putative transposon protein
Length = 315
Score = 171 bits (433), Expect = 3e-42
Identities = 76/138 (55%), Positives = 107/138 (77%)
Query: 533 LEKKFVRPSVSPWGAPVLLVKKKDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGA 592
+ K ++R S+SP PVLLV KKDG+ R+C+D R +N +TIK R+P+PR+ D++D+L GA
Sbjct: 125 IHKGYIRESLSPCAVPVLLVPKKDGTWRMCLDCRAINNITIKYRHPIPRLYDMLDELSGA 184
Query: 593 KVFSKIDLRSGYHQIKVKDEDMQKTAFSTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHA 652
+FSK+DLRSGYHQ+++++ D KTAF T+ G YE VMPFG+TNAP FM MN++ +
Sbjct: 185 IIFSKVDLRSGYHQVRMREGDEWKTAFKTKQGLYECLVMPFGLTNAPSTFMRLMNQVLRS 244
Query: 653 FLDRFVVVFIDDILIYSK 670
F+ +FVVV+ DDILIY+K
Sbjct: 245 FIGKFVVVYFDDILIYNK 262
>At2g14640 putative retroelement pol polyprotein
Length = 945
Score = 140 bits (353), Expect = 5e-33
Identities = 112/365 (30%), Positives = 176/365 (47%), Gaps = 48/365 (13%)
Query: 306 VFALDGTQTENEDRLIRCTCYINNTPLVAIIDTGATHCFIAFDCVSALGLDLFDMNGEMV 365
++AL+G T + R +R T + N L+A+I++G+TH FI V + L V
Sbjct: 306 LYALEGVDTTSTIR-VRATIHRNR--LIALINSGSTHNFIGEKAVRGMNLKATTTKPFTV 362
Query: 366 VETPAKGLVTTSLVCLRCLLSMFGRDFEMDLVCLPLSGMDVILGMNWLE-YNHVHINYFT 424
LV S ++ M G F + L LPL G+D+ +G+ WL N+
Sbjct: 363 RVVNGMPLVCRSRYEAIPVV-MGGVVFPVTLYALPLMGLDLAMGVQWLSTLGPTLCNWKE 421
Query: 425 KSVYFSSVEEE--------SGAEFLSTKQLKQMERDGILMYPL-MASLSFENQAVIDKLQ 475
+++ F +E +G + K + + R G ++ + MA S + +++
Sbjct: 422 QTLQFHWAGDEVRLMGIKPTGLRGVEHKTITKKARMGHTIFAITMAHNSSDPNTPDGEIK 481
Query: 476 -VVCDFPEVFPDEIP-DVPLEREVEFSIDLILGTKPVSMAPYRMSASELAKLKKQLEDLL 533
++ +F +F E P +P ER + I L GT PV++ PYR + Q +++
Sbjct: 482 KLIEEFAGLF--ETPTQLPSERPIVHRIALKKGTDPVNVRPYRYAFY-------QKDEMS 532
Query: 534 EKKFVRPSVSPWGAPVLLVKKKDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGAK 593
+RPS SP+ +PVLL R+P+P +DD++D+L GA
Sbjct: 533 RAGIIRPSSSPFLSPVLL-----------------------ERFPIPTVDDMLDELNGAV 569
Query: 594 VFSKIDLRSGYHQIKVKDEDMQKTAFSTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAF 653
F+K+DL +GY Q+++ D+ K AF T GHYEY VMPFG+ NAP F MN IF
Sbjct: 570 YFTKLDLTAGYQQVRMHSPDIPKIAFQTHNGHYEYLVMPFGLCNAPSTFQALMNEIFWPL 629
Query: 654 LDRFV 658
L R V
Sbjct: 630 LSRSV 634
>At2g06170 putative Ty3-gypsy-like retroelement pol polyprotein
Length = 587
Score = 135 bits (340), Expect = 2e-31
Identities = 61/105 (58%), Positives = 83/105 (78%)
Query: 566 RQLNKVTIKNRYPLPRIDDLMDQLVGAKVFSKIDLRSGYHQIKVKDEDMQKTAFSTRYGH 625
+ +NK+TIK R+P+P++DD++D+L G+KVFSKIDLRSGYHQI+++ D KT F ++ G
Sbjct: 401 KAINKITIKYRFPIPQLDDMLDELSGSKVFSKIDLRSGYHQIRIRPGDEWKTDFKSKDGL 460
Query: 626 YEYKVMPFGVTNAPGVFMEYMNRIFHAFLDRFVVVFIDDILIYSK 670
YE++VMPFG++NAP FM MN+I F FVVV+ DDILIYSK
Sbjct: 461 YEWQVMPFGMSNAPSTFMRLMNQILRPFTGSFVVVYFDDILIYSK 505
>At4g08100 putative polyprotein
Length = 1054
Score = 127 bits (319), Expect = 5e-29
Identities = 56/108 (51%), Positives = 79/108 (72%)
Query: 564 DYRQLNKVTIKNRYPLPRIDDLMDQLVGAKVFSKIDLRSGYHQIKVKDEDMQKTAFSTRY 623
D R +N +T+K R+P+PR+DD +D+L G+ +FSKIDL+SGYHQ ++K+ D KTA T+
Sbjct: 527 DCRAINNITVKYRHPIPRLDDTLDKLHGSSIFSKIDLKSGYHQTRMKEGDEWKTAIKTKQ 586
Query: 624 GHYEYKVMPFGVTNAPGVFMEYMNRIFHAFLDRFVVVFIDDILIYSKD 671
YE+ VMPFG+TNAP FM MN + + FV+V+ DDIL+YSK+
Sbjct: 587 RLYEWLVMPFGLTNAPNTFMRLMNHVLRKHIGVFVIVYFDDILVYSKN 634
Score = 58.2 bits (139), Expect = 4e-08
Identities = 59/264 (22%), Positives = 100/264 (37%), Gaps = 19/264 (7%)
Query: 29 PAFKGRYDPDGAQTWLKEVERIFRVMQCSEVQKVRFGTHQLAEEADDWWVSLLPN--LDQ 86
PAF G +PD W +++E +F +C + KV+ + A WW L+ + +
Sbjct: 97 PAFHGTDNPDTYLEWEQKIELVFLCQECLQSNKVKIAATKFYNYALSWWDQLVTSRRRTR 156
Query: 87 DGVDVTWAVF*REFMRRYFPEDVRRKKEIEFLELKQGNMYVTEYAAKFVELAEFYPHYAA 146
D TW +R+ P R+ L QG+ V EY F+E+
Sbjct: 157 DYPIKTWNQLKFVMRKRFVPSYYHRELHQRLRNLVQGSKTVEEY---FLEMETLMLRADL 213
Query: 147 ETVEFSKCIKFENGLRPDIKRAIEYQQIRVFPDLVNSCRIYEEDTKAHYKVVNERKTKGQ 206
+ + +F GL +I+ +E Q ++++ ++E+ K + KT
Sbjct: 214 QEDGEAVMSRFMGGLNREIQDRLETQHYVELEEMLHKAVMFEQQIKRKNARSSHTKTNYS 273
Query: 207 QSRPKPYSAPADKGKQRMVDDRRPKKKDAPAEITCFNCGEKGHKSNVCPEEIKKCVRCGK 266
+P Y G Q +D +P K P ++ KG V + +RC K
Sbjct: 274 SGKPS-YQKEEKFGYQ---NDSKPFVKPKPVDL-----DPKGKGKEVITR--ARDIRCFK 322
Query: 267 K---GHIVADCKRNDIVCFNFNEE 287
H + C I+ N E
Sbjct: 323 SQGLRHYASKCSNKRIMVHRDNGE 346
>At1g20390 hypothetical protein
Length = 1791
Score = 113 bits (283), Expect = 7e-25
Identities = 60/163 (36%), Positives = 94/163 (56%), Gaps = 1/163 (0%)
Query: 508 KPVSMAPYRMSASELAKLKKQLEDLLEK-KFVRPSVSPWGAPVLLVKKKDGSMRLCIDYR 566
KPV ++ + +++E LL+ + + W A ++VKKK+G R+C+DY
Sbjct: 776 KPVKQKRRKLGPERARAVNEEVEKLLKAGQIIEVKYPEWLANPVVVKKKNGKWRVCVDYT 835
Query: 567 QLNKVTIKNRYPLPRIDDLMDQLVGAKVFSKIDLRSGYHQIKVKDEDMQKTAFSTRYGHY 626
LNK K+ YPLP ID L++ G + S +D SGY+QI + +D +KT+F T G Y
Sbjct: 836 DLNKACPKDSYPLPHIDRLVEATSGNGLLSFMDAFSGYNQILMHKDDQEKTSFVTDRGTY 895
Query: 627 EYKVMPFGVTNAPGVFMEYMNRIFHAFLDRFVVVFIDDILIYS 669
YKVM FG+ NA + ++N++ + R V V+IDD+L+ S
Sbjct: 896 CYKVMSFGLKNAGATYQRFVNKMLADQIGRTVEVYIDDMLVKS 938
>At2g14400 putative retroelement pol polyprotein
Length = 1466
Score = 112 bits (281), Expect = 1e-24
Identities = 64/178 (35%), Positives = 100/178 (55%), Gaps = 8/178 (4%)
Query: 500 SIDLI-------LGTKPVSMAPYRMSASELAKLKKQLEDLLEKKFVRPSVSP-WGAPVLL 551
S+DLI +G + ++ + +++ LL+ +R P W A ++
Sbjct: 448 SVDLIFLSTLERMGISRADVNRRKLGVERAKAVNDEVDKLLKIGSIREVQYPDWLANTVV 507
Query: 552 VKKKDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGAKVFSKIDLRSGYHQIKVKD 611
VKKK+G R+CID+ LNK K+ +PLP ID L++ G ++ S +D SGY+QI +
Sbjct: 508 VKKKNGKDRVCIDFTDLNKACPKDSFPLPHIDRLVESTAGNELLSFMDAFSGYNQIMMNP 567
Query: 612 EDMQKTAFSTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAFLDRFVVVFIDDILIYS 669
ED +KT F T G Y YKVMPFG+ NA + +N++F + + + V+IDD+LI S
Sbjct: 568 EDQEKTLFITDRGIYCYKVMPFGLRNAGATYPRLVNKMFSEHVGKTMEVYIDDMLIKS 625
>At2g15410 putative retroelement pol polyprotein
Length = 1787
Score = 110 bits (275), Expect = 6e-24
Identities = 59/163 (36%), Positives = 92/163 (56%), Gaps = 1/163 (0%)
Query: 508 KPVSMAPYRMSASELAKLKKQLEDLLEK-KFVRPSVSPWGAPVLLVKKKDGSMRLCIDYR 566
KP+ ++ + ++++ LL+ V W ++VKKK+G R+CID+
Sbjct: 781 KPLKQKRRKLGPDRTKDVNEEVKKLLDAGSIVEVRYPDWLRNPVVVKKKNGKWRVCIDFT 840
Query: 567 QLNKVTIKNRYPLPRIDDLMDQLVGAKVFSKIDLRSGYHQIKVKDEDMQKTAFSTRYGHY 626
LNK K+ +PLP ID L++ G ++ S +D SGY+QI + D +KT F T G Y
Sbjct: 841 DLNKACPKDSFPLPHIDRLVEATAGNELLSFMDAFSGYNQILMHQNDREKTVFITDQGTY 900
Query: 627 EYKVMPFGVTNAPGVFMEYMNRIFHAFLDRFVVVFIDDILIYS 669
YKVMPFG+ NA + +N++F LD + V+IDD+L+ S
Sbjct: 901 CYKVMPFGLKNAGATYPRLVNQMFTDQLDHSMEVYIDDMLVKS 943
>At2g14040 putative retroelement pol polyprotein
Length = 841
Score = 105 bits (263), Expect = 1e-22
Identities = 59/174 (33%), Positives = 94/174 (53%), Gaps = 19/174 (10%)
Query: 516 RMSASELAKLKKQLEDLLEKKFVRP-SVSPWGAPVLLVKKKDGSM--------------- 559
R++++ +KK++ LL+ + P S S W +PV +V KK G +
Sbjct: 381 RLNSNLRDVVKKEIMKLLDAGIIYPISDSTWVSPVHVVPKKGGVIVIKNEKNELIPTRTV 440
Query: 560 ---RLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGAKVFSKIDLRSGYHQIKVKDEDMQK 616
R+CIDYR+LN T K+ +PL ID ++++L + +D G+ QI + +D +K
Sbjct: 441 TGHRMCIDYRKLNSATRKDNFPLSFIDQMLERLSNQPYYCFLDGYLGFFQILIHPDDQEK 500
Query: 617 TAFSTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAFLDRFVVVFIDDILIYSK 670
T F+ YG + Y+ MPFG+ NAP F M IF ++ F+ VFIDD L+ +
Sbjct: 501 TTFTCPYGTFAYRRMPFGLCNAPATFQHCMKYIFSDMIEDFMEVFIDDFLVLQR 554
>At1g37060 Athila retroelment ORF 1, putative
Length = 1734
Score = 104 bits (260), Expect = 3e-22
Identities = 67/208 (32%), Positives = 99/208 (47%), Gaps = 34/208 (16%)
Query: 495 REVEFSIDLILGTKPV--------------SMAPYRMSASELAKL-KKQLEDLLEKKFVR 539
+ + +S+D I G P S+ P R L ++ KK++ LL+ +
Sbjct: 870 KAIGYSLDDIKGISPTLCTHRIHLENESYSSIEPQRRLNPNLKEVVKKEILKLLDAGVIY 929
Query: 540 P-SVSPWGAPVLLVKKKDGSM------------------RLCIDYRQLNKVTIKNRYPLP 580
P S S W +PV V KK G R+CI+YR+LN + K +PLP
Sbjct: 930 PISDSTWVSPVHCVPKKGGMTVVKNSKDELIPTRTITGHRMCIEYRKLNVASRKEHFPLP 989
Query: 581 RIDDLMDQLVGAKVFSKIDLRSGYHQIKVKDEDMQKTAFSTRYGHYEYKVMPFGVTNAPG 640
ID ++++L + +D SG+ QI + D KT F+ YG + YK MPFG+ NAP
Sbjct: 990 FIDHMLERLANHPYYCFLDSYSGFFQIPIHPNDQGKTTFTCPYGTFAYKRMPFGLCNAPA 1049
Query: 641 VFMEYMNRIFHAFLDRFVVVFIDDILIY 668
F M IF ++ V VF+DD +Y
Sbjct: 1050 TFQRCMTSIFSDLIEEMVEVFMDDFSVY 1077
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.349 0.155 0.537
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,264,599
Number of Sequences: 26719
Number of extensions: 1194634
Number of successful extensions: 5416
Number of sequences better than 10.0: 84
Number of HSP's better than 10.0 without gapping: 63
Number of HSP's successfully gapped in prelim test: 21
Number of HSP's that attempted gapping in prelim test: 5054
Number of HSP's gapped (non-prelim): 226
length of query: 1478
length of database: 11,318,596
effective HSP length: 112
effective length of query: 1366
effective length of database: 8,326,068
effective search space: 11373408888
effective search space used: 11373408888
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.8 bits)
S2: 67 (30.4 bits)
Medicago: description of AC141111.5


