

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141111.4 - phase: 0 /pseudo
(1069 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
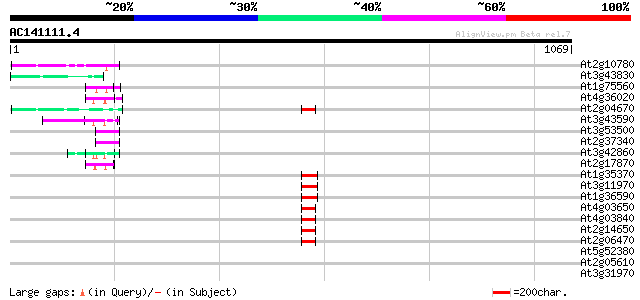
Score E
Sequences producing significant alignments: (bits) Value
At2g10780 pseudogene 62 2e-09
At3g43830 putative protein 57 4e-08
At1g75560 DNA-binding protein 50 5e-06
At4g36020 glycine-rich protein 49 2e-05
At2g04670 putative retroelement pol polyprotein 49 2e-05
At3g43590 putative protein 48 3e-05
At3g53500 splicing factor - like protein 46 1e-04
At2g37340 unknown protein 45 2e-04
At3g42860 unknown protein (At3g42860) 45 2e-04
At2g17870 putative glycine-rich, zinc-finger DNA-binding protein 45 3e-04
At1g35370 hypothetical protein 44 4e-04
At3g11970 hypothetical protein 44 5e-04
At1g36590 hypothetical protein 44 5e-04
At4g03650 putative reverse transcriptase 44 6e-04
At4g03840 putative transposon protein 43 8e-04
At2g14650 putative retroelement pol polyprotein 43 8e-04
At2g06470 putative retroelement pol polyprotein 43 8e-04
At5g52380 unknown protein 43 0.001
At2g05610 putative retroelement pol polyprotein 42 0.001
At3g31970 hypothetical protein 41 0.003
>At2g10780 pseudogene
Length = 1611
Score = 61.6 bits (148), Expect = 2e-09
Identities = 56/218 (25%), Positives = 90/218 (40%), Gaps = 22/218 (10%)
Query: 3 EFLNRYFPEDVRGKKEIEFLELKQGDMSVTEYAAKFVELAKFYPHYTAETAEFSKCIKIE 62
EF +YFP + + E +L+L QG+ +V EY +F L ++ E E ++ +
Sbjct: 248 EFNKKYFPPEAWDRLECAYLDLVQGNRTVREYDEEFNRLRRYVGRELEE--EQAQLRRFI 305
Query: 63 NGLRADIKRAIGYQKIRNFSELVSSCRIYEEDTKAHYKVMSERRGKGQLSRPKPYSAPPD 122
GLR +I+ + + SELV + EE + + E+ R + P D
Sbjct: 306 RGLRIEIRNHCLVRTFNSVSELVERAAMIEEGIEEERYLNREKAP----IRNNQSTKPAD 361
Query: 123 KGK*RLKDERRPKMRDAPTDIVCFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADCKRGD 182
K R D+ DA T C CG K H S C + C RCG K H + C + +
Sbjct: 362 KK--RKFDKVDNTKSDAKTG-ECVTCG-KNH-SGTCWKAIGACGRCGSKDHAIQSCPKME 416
Query: 183 -----------IVCYNFNEEGHISLQCTQPKKVRTGGK 209
C+ + GH+ +C + + G+
Sbjct: 417 PGQSKVLGEETRTCFYCGKTGHLKRECPKLTAEKQAGQ 454
Score = 41.2 bits (95), Expect = 0.003
Identities = 16/27 (59%), Positives = 24/27 (88%)
Query: 557 LDKFVVVFIDDILIYSETEEEHAEHLK 583
LD+FV++FI+DIL+YS++ E H EHL+
Sbjct: 804 LDEFVIIFINDILVYSKSWEAHQEHLR 830
>At3g43830 putative protein
Length = 329
Score = 57.4 bits (137), Expect = 4e-08
Identities = 46/182 (25%), Positives = 68/182 (37%), Gaps = 45/182 (24%)
Query: 1 RREFLNRYFPEDVRGKKEIEFLELKQGDMSVTEYAAKFVELAKFYPHYTAETAEFSKCIK 60
+ EF RY PE+ E++FL L+QG +V +Y +F L +F E K I
Sbjct: 181 KEEFTRRYLPEETIDDLEMKFLRLQQGTKTVRKYEKEFHSLERFERRKRGEHELIHKFI- 239
Query: 61 IENGLRADIKRAIGYQKIRNFSELV---SSCRIYEEDTKAHYKVMSERRGKGQLSRPKPY 117
+GLR DI+ + N +LV +S I E+ + ++ E+ G
Sbjct: 240 --SGLRVDIRLCCHVRDFDNMIDLVEKAASLEIGLEEEARYKRIAQEKEAMGSY------ 291
Query: 118 SAPPDKGK*RLKDERRPKMRDAPTDIVCFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLAD 177
G+ GH C E C+RCG GH D
Sbjct: 292 -------------------------------GQTGHSKRRC--QEVTCYRCGVAGHIARD 318
Query: 178 CK 179
C+
Sbjct: 319 CR 320
>At1g75560 DNA-binding protein
Length = 257
Score = 50.4 bits (119), Expect = 5e-06
Identities = 26/75 (34%), Positives = 36/75 (47%), Gaps = 10/75 (13%)
Query: 145 CFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADCKRGDI------VCYNFNEEGHISLQC 198
C+ C E GH ++ C +E C CGK GH DC D +C N ++GH++ C
Sbjct: 95 CWNCREPGHVASNCS-NEGICHSCGKSGHRARDCSNSDSRAGDLRLCNNCFKQGHLAADC 153
Query: 199 TQP---KKVRTGGKV 210
T K RT G +
Sbjct: 154 TNDKACKNCRTSGHI 168
Score = 47.8 bits (112), Expect = 3e-05
Identities = 21/61 (34%), Positives = 29/61 (47%), Gaps = 7/61 (11%)
Query: 144 VCFKCGEKGHKSNVCDRDEKK------CFRCGKKGHTLADCKRGDIVCYNFNEEGHISLQ 197
+C CG+ GH++ C + + C C K+GH ADC D C N GHI+
Sbjct: 113 ICHSCGKSGHRARDCSNSDSRAGDLRLCNNCFKQGHLAADC-TNDKACKNCRTSGHIARD 171
Query: 198 C 198
C
Sbjct: 172 C 172
Score = 40.4 bits (93), Expect = 0.005
Identities = 24/73 (32%), Positives = 33/73 (44%), Gaps = 10/73 (13%)
Query: 144 VCFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADCKRGDIVCYNFNEEGHISLQC----- 198
+C C ++GH + C D K C C GH DC R D VC + GH++ C
Sbjct: 139 LCNNCFKQGHLAADCTND-KACKNCRTSGHIARDC-RNDPVCNICSISGHVARHCPKGDS 196
Query: 199 ---TQPKKVRTGG 208
+ +VR GG
Sbjct: 197 NYSDRGSRVRDGG 209
Score = 31.2 bits (69), Expect = 3.3
Identities = 14/35 (40%), Positives = 17/35 (48%), Gaps = 1/35 (2%)
Query: 165 CFRCGKKGHTLADCKRGDIVCYNFNEEGHISLQCT 199
C C + GH DC VC N GHI+ +CT
Sbjct: 57 CNNCKRPGHFARDCSNVS-VCNNCGLPGHIAAECT 90
>At4g36020 glycine-rich protein
Length = 299
Score = 48.5 bits (114), Expect = 2e-05
Identities = 30/95 (31%), Positives = 39/95 (40%), Gaps = 25/95 (26%)
Query: 145 CFKCGEKGHKSNVC------------DRDEKKCFRCGKKGHTLADC------------KR 180
C+ CGE GH S C R + C+ CG GH DC K
Sbjct: 102 CYNCGELGHISKDCGIGGGGGGGERRSRGGEGCYNCGDTGHFARDCTSAGNGDQRGATKG 161
Query: 181 GDIVCYNFNEEGHISLQCTQPKKVRTGGKVFALTG 215
G+ CY + GH++ CTQ K V G + A+ G
Sbjct: 162 GNDGCYTCGDVGHVARDCTQ-KSVGNGDQRGAVKG 195
Score = 46.2 bits (108), Expect = 1e-04
Identities = 22/65 (33%), Positives = 33/65 (49%), Gaps = 10/65 (15%)
Query: 145 CFKCGEKGHKSNVCD---RDEKKCFRCGKKGHTLADC-KRG------DIVCYNFNEEGHI 194
C+ CG GH + C + + C++CG GH DC +RG D CY +EGH
Sbjct: 232 CYSCGGVGHIARDCATKRQPSRGCYQCGGSGHLARDCDQRGSGGGGNDNACYKCGKEGHF 291
Query: 195 SLQCT 199
+ +C+
Sbjct: 292 ARECS 296
Score = 42.7 bits (99), Expect = 0.001
Identities = 28/93 (30%), Positives = 36/93 (38%), Gaps = 29/93 (31%)
Query: 145 CFKCGEKGHKSNVC-------DRDEKK-----CFRCGKKGHTLADC-------------- 178
C+ CG+ GH + C R K C+ CG GH DC
Sbjct: 134 CYNCGDTGHFARDCTSAGNGDQRGATKGGNDGCYTCGDVGHVARDCTQKSVGNGDQRGAV 193
Query: 179 KRGDIVCYNFNEEGHISLQCTQ---PKKVRTGG 208
K G+ CY + GH + CTQ VR+GG
Sbjct: 194 KGGNDGCYTCGDVGHFARDCTQKVAAGNVRSGG 226
Score = 37.7 bits (86), Expect = 0.035
Identities = 19/71 (26%), Positives = 28/71 (38%), Gaps = 15/71 (21%)
Query: 145 CFKCGEKGHKSNVCDRD------------EKKCFRCGKKGHTLADC---KRGDIVCYNFN 189
C+ CG+ GH + C + C+ CG GH DC ++ CY
Sbjct: 200 CYTCGDVGHFARDCTQKVAAGNVRSGGGGSGTCYSCGGVGHIARDCATKRQPSRGCYQCG 259
Query: 190 EEGHISLQCTQ 200
GH++ C Q
Sbjct: 260 GSGHLARDCDQ 270
Score = 37.0 bits (84), Expect = 0.060
Identities = 22/89 (24%), Positives = 34/89 (37%), Gaps = 26/89 (29%)
Query: 145 CFKCGEKGHKSNVCDR------DEKK--------CFRCGKKGHTLADCKR---------- 180
C+ CG+ GH + C + D++ C+ CG GH DC +
Sbjct: 166 CYTCGDVGHVARDCTQKSVGNGDQRGAVKGGNDGCYTCGDVGHFARDCTQKVAAGNVRSG 225
Query: 181 --GDIVCYNFNEEGHISLQCTQPKKVRTG 207
G CY+ GHI+ C ++ G
Sbjct: 226 GGGSGTCYSCGGVGHIARDCATKRQPSRG 254
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 48.5 bits (114), Expect = 2e-05
Identities = 49/213 (23%), Positives = 80/213 (37%), Gaps = 39/213 (18%)
Query: 3 EFLNRYFPEDVRGKKEIEFLELKQGDMSVTEYAAKFVELAKFYPHYTAETAEFSKCIKIE 62
EF +YFP++ + E FLEL QG+ SV EY +F L + + + ++ +
Sbjct: 204 EFNAKYFPQEALDRMEARFLELTQGEWSVREYDREFNRLLAYAGRGMED--DQAQMRRFL 261
Query: 63 NGLRADIKRAIGYQKIRNFSELVSSCRIYEEDTKAHYKVMSERRGKGQLSRPKPYSAPPD 122
GLR D++ + + LV + EED + +S ++ P
Sbjct: 262 RGLRPDLRVQCRVSQYATKAALVETAAEVEEDLQRQVVGVS----PAVQTKNTQQQVTPS 317
Query: 123 KGK*RLKDERRPKMRDAPTDIVCFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADCKRGD 182
KG + ++R + H S F CG HT+ADC
Sbjct: 318 KGGKPAQGQKR----------------KWDHPSRAGQGGRAGYFSCGSLDHTVADC---- 357
Query: 183 IVCYNFNEEGHISLQCTQPKKVRTGGKVFALTG 215
E GH+ ++ GG+ A+ G
Sbjct: 358 ------TERGHLHVKV-------AGGQFLAVLG 377
Score = 43.5 bits (101), Expect = 6e-04
Identities = 17/27 (62%), Positives = 24/27 (87%)
Query: 557 LDKFVVVFIDDILIYSETEEEHAEHLK 583
LD+FV++FIDDIL+YS++ EEH HL+
Sbjct: 662 LDEFVIIFIDDILVYSKSPEEHEVHLR 688
>At3g43590 putative protein
Length = 551
Score = 47.8 bits (112), Expect = 3e-05
Identities = 43/157 (27%), Positives = 64/157 (40%), Gaps = 13/157 (8%)
Query: 63 NGLRADIKRAIGYQKI-RNFSELVSSCRIYEEDTKAHYKVMSERRGKGQLSRPKPYSAPP 121
NG+++ +K++ +K+ RN E I D V +G+G+ + P
Sbjct: 79 NGVKSKVKKSESSKKMKRNKLEADHEIPIVWNDQDEEKVVEEIVKGEGEDDEVERSDEPK 138
Query: 122 DK---GK*RLKDERRPKMRDAPTD---IVCFKCGEKGHKSNVC---DRDEKKCFRCGKKG 172
+ LK R P D + C+ CGE+GH S C + K CF CG
Sbjct: 139 TEETASNLVLKKLLRGARYFDPPDAGWVSCYSCGEQGHTSFNCPTPTKRRKPCFICGSLE 198
Query: 173 HTLADCKRGDIVCYNFNEEGHISLQCTQPKKVRTGGK 209
H C +G CY + GH + C P K + G K
Sbjct: 199 HGAKQCSKGH-DCYICKKTGHRAKDC--PDKYKNGSK 232
Score = 45.4 bits (106), Expect = 2e-04
Identities = 21/93 (22%), Positives = 36/93 (38%), Gaps = 31/93 (33%)
Query: 143 IVCFKCGEKGHKSNVC--------------------DRDEKKCFRCGKKGHTLADC---- 178
+ C++CG+ GH C R+ +C+RCG++GH +C
Sbjct: 285 VSCYRCGQLGHSGLACGRHYEESNENDSATPERLFNSREASECYRCGEEGHFARECPNSS 344
Query: 179 -------KRGDIVCYNFNEEGHISLQCTQPKKV 204
+ +CY N GH + +C +V
Sbjct: 345 SISTSHGRESQTLCYRCNGSGHFARECPNSSQV 377
Score = 35.4 bits (80), Expect = 0.17
Identities = 21/76 (27%), Positives = 33/76 (42%), Gaps = 13/76 (17%)
Query: 135 KMRDAPTDIVCFKCGEKGHKSNVC-------DRDEKKCFRCGKKGHTLADCKRGD----- 182
K ++ VC +CG+ GH +C D + +C+ C GH L + G+
Sbjct: 226 KYKNGSKGAVCLRCGDFGHDMILCKYEYSKEDLKDVQCYICKSFGH-LCCVEPGNSLSWA 284
Query: 183 IVCYNFNEEGHISLQC 198
+ CY + GH L C
Sbjct: 285 VSCYRCGQLGHSGLAC 300
>At3g53500 splicing factor - like protein
Length = 243
Score = 46.2 bits (108), Expect = 1e-04
Identities = 23/50 (46%), Positives = 24/50 (48%), Gaps = 5/50 (10%)
Query: 164 KCFRCGKKGHTLADCKRGD--IVCYNFNEEGHISLQC---TQPKKVRTGG 208
+CF CG GH DC GD CY E GHI C PKK R GG
Sbjct: 59 RCFNCGVDGHWARDCTAGDWKNKCYRCGERGHIERNCKNSPSPKKARQGG 108
>At2g37340 unknown protein
Length = 249
Score = 45.4 bits (106), Expect = 2e-04
Identities = 22/48 (45%), Positives = 24/48 (49%), Gaps = 3/48 (6%)
Query: 164 KCFRCGKKGHTLADCKRGD--IVCYNFNEEGHISLQC-TQPKKVRTGG 208
+CF CG GH DC GD CY E GHI C PKK+R G
Sbjct: 59 RCFNCGVDGHWARDCTAGDWKNKCYRCGERGHIERNCKNSPKKLRRSG 106
Score = 41.2 bits (95), Expect = 0.003
Identities = 16/37 (43%), Positives = 22/37 (59%), Gaps = 2/37 (5%)
Query: 145 CFKCGEKGHKSNVCDRDE--KKCFRCGKKGHTLADCK 179
CF CG GH + C + KC+RCG++GH +CK
Sbjct: 60 CFNCGVDGHWARDCTAGDWKNKCYRCGERGHIERNCK 96
Score = 30.4 bits (67), Expect = 5.6
Identities = 11/25 (44%), Positives = 14/25 (56%)
Query: 145 CFKCGEKGHKSNVCDRDEKKCFRCG 169
C++CGE+GH C KK R G
Sbjct: 82 CYRCGERGHIERNCKNSPKKLRRSG 106
>At3g42860 unknown protein (At3g42860)
Length = 372
Score = 45.1 bits (105), Expect = 2e-04
Identities = 30/100 (30%), Positives = 37/100 (37%), Gaps = 37/100 (37%)
Query: 145 CFKCGEKGHKSNVC----------------DRDEKKCFRCGKKGHTLADC---------- 178
CFKCG+ GH S C +C++CGK+GH DC
Sbjct: 270 CFKCGKPGHWSRDCTAQSGNPKYEPGQMKSSSSSGECYKCGKQGHWSRDCTGQSSNQQFQ 329
Query: 179 --------KRGDIVCYNFNEEGHISLQCTQP-KKVRTGGK 209
GD CY + GH S CT P + T GK
Sbjct: 330 SGQAKSTSSTGD--CYKCGKAGHWSRDCTSPAQTTNTPGK 367
Score = 43.9 bits (102), Expect = 5e-04
Identities = 32/116 (27%), Positives = 43/116 (36%), Gaps = 30/116 (25%)
Query: 111 LSRPKPYSAPPDKGK*RLKDERRPKMRDAPTDIVCFKCGEKGHKSNVC----DRDEKK-- 164
+S P YS + D ++A T C+KCG++GH + C D K
Sbjct: 208 VSSPTSYSVTKNSN---FGDSDTRGYQNAKTGTPCYKCGKEGHWARDCTVQSDTGPVKST 264
Query: 165 -----CFRCGKKGHTLADC----------------KRGDIVCYNFNEEGHISLQCT 199
CF+CGK GH DC CY ++GH S CT
Sbjct: 265 SAAGDCFKCGKPGHWSRDCTAQSGNPKYEPGQMKSSSSSGECYKCGKQGHWSRDCT 320
>At2g17870 putative glycine-rich, zinc-finger DNA-binding protein
Length = 301
Score = 44.7 bits (104), Expect = 3e-04
Identities = 24/81 (29%), Positives = 32/81 (38%), Gaps = 27/81 (33%)
Query: 145 CFKCGEKGHKSNVCD----------------RDEKKCFRCGKKGHTLADCKR-------- 180
CF CGE GH + CD E +C+ CG GH DC++
Sbjct: 96 CFNCGEVGHMAKDCDGGSGGKSFGGGGGRRSGGEGECYMCGDVGHFARDCRQSGGGNSGG 155
Query: 181 ---GDIVCYNFNEEGHISLQC 198
G CY+ E GH++ C
Sbjct: 156 GGGGGRPCYSCGEVGHLAKDC 176
Score = 43.1 bits (100), Expect = 8e-04
Identities = 22/73 (30%), Positives = 30/73 (40%), Gaps = 18/73 (24%)
Query: 145 CFKCGEKGHKSNVCDRD--------EKKCFRCGKKGHTLADCKR----------GDIVCY 186
C+ CG GH + VC + C+ CG GH DC R G C+
Sbjct: 226 CYTCGGVGHIAKVCTSKIPSGGGGGGRACYECGGTGHLARDCDRRGSGSSGGGGGSNKCF 285
Query: 187 NFNEEGHISLQCT 199
+EGH + +CT
Sbjct: 286 ICGKEGHFARECT 298
Score = 40.8 bits (94), Expect = 0.004
Identities = 25/87 (28%), Positives = 33/87 (37%), Gaps = 25/87 (28%)
Query: 145 CFKCGEKGHKSNVCDRDE---------------KKCFRCGKKGHTLADCKR--------G 181
C+ CGE GH + C C+ CG GH DC++ G
Sbjct: 163 CYSCGEVGHLAKDCRGGSGGNRYGGGGGRGSGGDGCYMCGGVGHFARDCRQNGGGNVGGG 222
Query: 182 DIVCYNFNEEGHISLQCTQPKKVRTGG 208
CY GHI+ CT K+ +GG
Sbjct: 223 GSTCYTCGGVGHIAKVCT--SKIPSGG 247
Score = 36.2 bits (82), Expect = 0.10
Identities = 22/90 (24%), Positives = 31/90 (34%), Gaps = 26/90 (28%)
Query: 145 CFKCGEKGHKSNVCDRDE-----------KKCFRCGKKGHTLADCK-------------- 179
C+ CG+ GH + C + + C+ CG+ GH DC+
Sbjct: 132 CYMCGDVGHFARDCRQSGGGNSGGGGGGGRPCYSCGEVGHLAKDCRGGSGGNRYGGGGGR 191
Query: 180 -RGDIVCYNFNEEGHISLQCTQPKKVRTGG 208
G CY GH + C Q GG
Sbjct: 192 GSGGDGCYMCGGVGHFARDCRQNGGGNVGG 221
Score = 34.7 bits (78), Expect = 0.30
Identities = 20/80 (25%), Positives = 29/80 (36%), Gaps = 16/80 (20%)
Query: 145 CFKCGEKGHKSNVCDRDE--------KKCFRCGKKGHTLADCKR--------GDIVCYNF 188
C+ CG GH + C ++ C+ CG GH C G CY
Sbjct: 198 CYMCGGVGHFARDCRQNGGGNVGGGGSTCYTCGGVGHIAKVCTSKIPSGGGGGGRACYEC 257
Query: 189 NEEGHISLQCTQPKKVRTGG 208
GH++ C + +GG
Sbjct: 258 GGTGHLARDCDRRGSGSSGG 277
Score = 33.1 bits (74), Expect = 0.87
Identities = 20/76 (26%), Positives = 25/76 (32%), Gaps = 16/76 (21%)
Query: 149 GEKGHKSNVCDRDEKKCFRCGKKGHTLADC----------------KRGDIVCYNFNEEG 192
G K N CF CG+ GH DC G+ CY + G
Sbjct: 80 GSLNKKENSSRGSGGNCFNCGEVGHMAKDCDGGSGGKSFGGGGGRRSGGEGECYMCGDVG 139
Query: 193 HISLQCTQPKKVRTGG 208
H + C Q +GG
Sbjct: 140 HFARDCRQSGGGNSGG 155
>At1g35370 hypothetical protein
Length = 1447
Score = 44.3 bits (103), Expect = 4e-04
Identities = 21/30 (70%), Positives = 24/30 (80%)
Query: 557 LDKFVVVFIDDILIYSETEEEHAEHLK*VF 586
L KFV+VF DDILIYS + EEH EHL+ VF
Sbjct: 725 LRKFVLVFFDDILIYSSSIEEHKEHLRLVF 754
>At3g11970 hypothetical protein
Length = 1499
Score = 43.9 bits (102), Expect = 5e-04
Identities = 20/30 (66%), Positives = 24/30 (79%)
Query: 557 LDKFVVVFIDDILIYSETEEEHAEHLK*VF 586
L KFV+VF DDIL+YS + EEH +HLK VF
Sbjct: 746 LRKFVLVFFDDILVYSSSLEEHRQHLKQVF 775
>At1g36590 hypothetical protein
Length = 1499
Score = 43.9 bits (102), Expect = 5e-04
Identities = 20/30 (66%), Positives = 24/30 (79%)
Query: 557 LDKFVVVFIDDILIYSETEEEHAEHLK*VF 586
L KFV+VF DDIL+YS + EEH +HLK VF
Sbjct: 746 LRKFVLVFFDDILVYSSSLEEHRQHLKQVF 775
>At4g03650 putative reverse transcriptase
Length = 839
Score = 43.5 bits (101), Expect = 6e-04
Identities = 17/27 (62%), Positives = 24/27 (87%)
Query: 557 LDKFVVVFIDDILIYSETEEEHAEHLK 583
LD+FV++FIDDIL+YS++ EEH HL+
Sbjct: 561 LDEFVIIFIDDILVYSKSPEEHEVHLR 587
Score = 38.5 bits (88), Expect = 0.021
Identities = 27/92 (29%), Positives = 42/92 (45%), Gaps = 2/92 (2%)
Query: 3 EFLNRYFPEDVRGKKEIEFLELKQGDMSVTEYAAKFVELAKFYPHYTAETAEFSKCIKIE 62
EF +YFP + + E FLEL QG SV EY KF L + + + ++ +
Sbjct: 222 EFNAKYFPREALDRMEARFLELTQGVRSVREYDRKFNRLLVYAGRGMED--DQAQMRRFL 279
Query: 63 NGLRADIKRAIGYQKIRNFSELVSSCRIYEED 94
GLR +++ + + LV + EED
Sbjct: 280 RGLRPNLRVRCRVSQYATKAALVETAAEVEED 311
>At4g03840 putative transposon protein
Length = 973
Score = 43.1 bits (100), Expect = 8e-04
Identities = 17/27 (62%), Positives = 24/27 (87%)
Query: 557 LDKFVVVFIDDILIYSETEEEHAEHLK 583
LD+FV++FIDDIL+YS++ E H EHL+
Sbjct: 328 LDEFVIIFIDDILVYSKSWEAHQEHLR 354
Score = 34.3 bits (77), Expect = 0.39
Identities = 15/42 (35%), Positives = 25/42 (58%)
Query: 3 EFLNRYFPEDVRGKKEIEFLELKQGDMSVTEYAAKFVELAKF 44
EF +YFP + + E +L+L QG+ +V EY +F L ++
Sbjct: 213 EFNKKYFPPEAWDRLECAYLDLVQGNRTVREYDEEFNRLRRY 254
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 43.1 bits (100), Expect = 8e-04
Identities = 17/28 (60%), Positives = 25/28 (88%)
Query: 556 MLDKFVVVFIDDILIYSETEEEHAEHLK 583
+LD+FV++FIDDIL+YS++ EEH HL+
Sbjct: 610 VLDEFVIIFIDDILVYSKSLEEHEVHLR 637
>At2g06470 putative retroelement pol polyprotein
Length = 899
Score = 43.1 bits (100), Expect = 8e-04
Identities = 17/27 (62%), Positives = 24/27 (87%)
Query: 557 LDKFVVVFIDDILIYSETEEEHAEHLK 583
LD+FV++FIDDIL+YS++ E H EHL+
Sbjct: 328 LDEFVIIFIDDILVYSKSWEAHQEHLR 354
Score = 34.3 bits (77), Expect = 0.39
Identities = 15/42 (35%), Positives = 25/42 (58%)
Query: 3 EFLNRYFPEDVRGKKEIEFLELKQGDMSVTEYAAKFVELAKF 44
EF +YFP + + E +L+L QG+ +V EY +F L ++
Sbjct: 213 EFNKKYFPPEAWDRLECAYLDLVQGNRTVREYDEEFNRLRRY 254
>At5g52380 unknown protein
Length = 268
Score = 42.7 bits (99), Expect = 0.001
Identities = 24/79 (30%), Positives = 35/79 (43%), Gaps = 13/79 (16%)
Query: 145 CFKCGEKGHKSNVCDRD-----EKKCFRCGKKGHTLADCKRGD------IVCYNFNEEGH 193
CF C K H + +C K C +C ++GH+L +C + +CYN + GH
Sbjct: 76 CFICHSKTHIAKLCPEKSEWERNKICLQCRRRGHSLKNCPEKNNESSEKKLCYNCGDTGH 135
Query: 194 ISLQCTQPKKVRTGGKVFA 212
C P + GG FA
Sbjct: 136 SLSHCPYP--MEDGGTKFA 152
Score = 40.8 bits (94), Expect = 0.004
Identities = 21/82 (25%), Positives = 35/82 (42%), Gaps = 13/82 (15%)
Query: 134 PKMRDAPTDIVCFKCGEKGHKSNVCDR------DEKKCFRCGKKGHTLADCK-------R 180
P+ + + +C +C +GH C ++K C+ CG GH+L+ C
Sbjct: 90 PEKSEWERNKICLQCRRRGHSLKNCPEKNNESSEKKLCYNCGDTGHSLSHCPYPMEDGGT 149
Query: 181 GDIVCYNFNEEGHISLQCTQPK 202
C+ +GHIS C + K
Sbjct: 150 KFASCFICKGQGHISKNCPENK 171
>At2g05610 putative retroelement pol polyprotein
Length = 780
Score = 42.4 bits (98), Expect = 0.001
Identities = 20/30 (66%), Positives = 24/30 (79%)
Query: 557 LDKFVVVFIDDILIYSETEEEHAEHLK*VF 586
L KFV+VF DDILIYS + EEH +HL+ VF
Sbjct: 188 LRKFVLVFFDDILIYSTSMEEHKKHLEAVF 217
>At3g31970 hypothetical protein
Length = 1329
Score = 41.2 bits (95), Expect = 0.003
Identities = 27/92 (29%), Positives = 44/92 (47%), Gaps = 2/92 (2%)
Query: 3 EFLNRYFPEDVRGKKEIEFLELKQGDMSVTEYAAKFVELAKFYPHYTAETAEFSKCIKIE 62
EF +YFP++ + E FLEL QG+ SV EY +F L + + + ++ +
Sbjct: 224 EFNAKYFPQEALDRMEARFLELTQGERSVREYEREFNRLLVYAGRGMQD--DQAQMRRFL 281
Query: 63 NGLRADIKRAIGYQKIRNFSELVSSCRIYEED 94
GLR D++ + + LV + EED
Sbjct: 282 RGLRPDLRVRCRVLQYATKAALVETAAEVEED 313
Score = 39.3 bits (90), Expect = 0.012
Identities = 15/22 (68%), Positives = 21/22 (95%)
Query: 557 LDKFVVVFIDDILIYSETEEEH 578
LD+FV++FIDDIL+YS++ EEH
Sbjct: 620 LDEFVIIFIDDILVYSKSPEEH 641
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.369 0.167 0.628
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,731,386
Number of Sequences: 26719
Number of extensions: 665320
Number of successful extensions: 3858
Number of sequences better than 10.0: 62
Number of HSP's better than 10.0 without gapping: 41
Number of HSP's successfully gapped in prelim test: 22
Number of HSP's that attempted gapping in prelim test: 3573
Number of HSP's gapped (non-prelim): 198
length of query: 1069
length of database: 11,318,596
effective HSP length: 110
effective length of query: 959
effective length of database: 8,379,506
effective search space: 8035946254
effective search space used: 8035946254
T: 11
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.7 bits)
S2: 65 (29.6 bits)
Medicago: description of AC141111.4


