

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141111.3 + phase: 0 /pseudo
(64 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
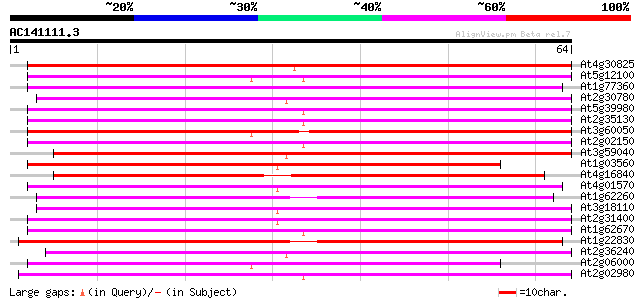
Score E
Sequences producing significant alignments: (bits) Value
At4g30825 puative protein 87 2e-18
At5g12100 unknown protein (At5g12100) 48 1e-06
At1g77360 47 2e-06
At2g30780 hypothetical protein 45 5e-06
At5g39980 putative protein 45 7e-06
At2g35130 hypothetical protein 44 2e-05
At3g60050 putative protein 43 3e-05
At2g02150 hypothetical protein 43 3e-05
At3g59040 unknown protein (At3g59040) 43 3e-05
At1g03560 hypothetical protein 43 3e-05
At4g16840 hypothetical protein 42 4e-05
At4g01570 unknown protein 42 4e-05
At1g62260 hypothetical protein 42 6e-05
At3g18110 hypothetical protein 42 7e-05
At2g31400 unknown protein 41 1e-04
At1g62670 PPR-repeat protein 41 1e-04
At1g22830 hypothetical protein, 5' partial 41 1e-04
At2g36240 putative salt-inducible protein 40 2e-04
At2g06000 unknown protein 40 2e-04
At2g02980 unknown protein 40 2e-04
>At4g30825 puative protein
Length = 904
Score = 87.0 bits (214), Expect = 2e-18
Identities = 41/63 (65%), Positives = 50/63 (79%), Gaps = 1/63 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGG-RIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
I+ YNT+ITGYGK M+ A+G+F L +EPDETSYRSMIEGWGRA NYE+A+ YY+
Sbjct: 349 IIAYNTLITGYGKIFKMEAAQGLFHRLCNIGLEPDETSYRSMIEGWGRADNYEEAKHYYQ 408
Query: 62 ELK 64
ELK
Sbjct: 409 ELK 411
Score = 30.8 bits (68), Expect = 0.13
Identities = 17/55 (30%), Positives = 29/55 (51%), Gaps = 2/55 (3%)
Query: 6 YNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNY-EKARW 58
YNT+I YG ++ A G+ + GR I PD+ +Y +++ R + E +W
Sbjct: 841 YNTLIKAYGIGGMVEEAVGLVKEMRGRNIIPDKVTYTNLVTALRRNDEFLEAIKW 895
Score = 30.4 bits (67), Expect = 0.17
Identities = 19/60 (31%), Positives = 30/60 (49%), Gaps = 1/60 (1%)
Query: 6 YNTMITGYGKASNMDGAEGVFLTLG-GRIEPDETSYRSMIEGWGRAGNYEKARWYYEELK 64
YN MI YG+ +D V L + PD SY ++I+ +G G E+A +E++
Sbjct: 806 YNIMINIYGEQGWIDEVADVLKELKESGLGPDLCSYNTLIKAYGIGGMVEEAVGLVKEMR 865
Score = 29.6 bits (65), Expect = 0.29
Identities = 17/60 (28%), Positives = 27/60 (44%), Gaps = 1/60 (1%)
Query: 6 YNTMITGYGKASNMDGAEGVFLTLGGRIE-PDETSYRSMIEGWGRAGNYEKARWYYEELK 64
YNT++ YGK M+ + + PD +Y MI +G G ++ +ELK
Sbjct: 771 YNTLLDAYGKDKQMEKFRSILKRMKKSTSGPDHYTYNIMINIYGEQGWIDEVADVLKELK 830
Score = 26.6 bits (57), Expect = 2.5
Identities = 14/50 (28%), Positives = 23/50 (46%)
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNY 53
V +N ++ YGKA +FL D SY ++I +G+ +Y
Sbjct: 700 VTFNVLLDVYGKAKLFKKVNELFLLAKRHGVVDVISYNTIIAAYGKNKDY 749
Score = 26.2 bits (56), Expect = 3.2
Identities = 17/65 (26%), Positives = 32/65 (49%), Gaps = 5/65 (7%)
Query: 3 IVVYNTMITGYGK---ASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKARWY 59
++ YNT+I YGK +NM A G + + +Y ++++ +G+ EK R
Sbjct: 733 VISYNTIIAAYGKNKDYTNMSSAIKNMQFDGFSVSLE--AYNTLLDAYGKDKQMEKFRSI 790
Query: 60 YEELK 64
+ +K
Sbjct: 791 LKRMK 795
>At5g12100 unknown protein (At5g12100)
Length = 816
Score = 47.8 bits (112), Expect = 1e-06
Identities = 23/63 (36%), Positives = 37/63 (58%), Gaps = 1/63 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+V YNT+I G + AE + L + + ++PD +Y S+I G+G AGN ++ YE
Sbjct: 564 LVTYNTLIDGLSMTGKLSEAEDLLLEISRKGLKPDVFTYNSLISGYGFAGNVQRCIALYE 623
Query: 62 ELK 64
E+K
Sbjct: 624 EMK 626
Score = 40.4 bits (93), Expect = 2e-04
Identities = 22/63 (34%), Positives = 36/63 (56%), Gaps = 1/63 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVF-LTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+ +YN +I G K M+ AE +F L R+ P +Y ++I+G+ +AGN EK+ E
Sbjct: 214 VFIYNVLIDGLCKGKRMNDAEQLFDEMLARRLLPSLITYNTLIDGYCKAGNPEKSFKVRE 273
Query: 62 ELK 64
+K
Sbjct: 274 RMK 276
Score = 35.4 bits (80), Expect = 0.005
Identities = 20/63 (31%), Positives = 35/63 (54%), Gaps = 1/63 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
++ YNT+I GY KA N + + V + IEP ++ ++++G +AG E A +
Sbjct: 249 LITYNTLIDGYCKAGNPEKSFKVRERMKADHIEPSLITFNTLLKGLFKAGMVEDAENVLK 308
Query: 62 ELK 64
E+K
Sbjct: 309 EMK 311
Score = 32.0 bits (71), Expect = 0.059
Identities = 18/62 (29%), Positives = 31/62 (49%), Gaps = 1/62 (1%)
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
V+YNTMI GY + ++ GA + + ++PD +Y +I + G E A +
Sbjct: 390 VIYNTMIDGYCRKGDLVGARMKIEAMEKQGMKPDHLAYNCLIRRFCELGEMENAEKEVNK 449
Query: 63 LK 64
+K
Sbjct: 450 MK 451
Score = 26.9 bits (58), Expect = 1.9
Identities = 14/60 (23%), Positives = 29/60 (48%), Gaps = 1/60 (1%)
Query: 6 YNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYEELK 64
YN++I G K + + + R +EP+ +Y +++G +Y A +Y E++
Sbjct: 703 YNSLILGQLKVGKLCEVRSLIDEMNAREMEPEADTYNIIVKGHCEVKDYMSAYVWYREMQ 762
Score = 25.0 bits (53), Expect = 7.2
Identities = 11/30 (36%), Positives = 18/30 (59%)
Query: 35 PDETSYRSMIEGWGRAGNYEKARWYYEELK 64
P+E Y +MI+G+ R G+ AR E ++
Sbjct: 387 PNEVIYNTMIDGYCRKGDLVGARMKIEAME 416
>At1g77360
Length = 481
Score = 46.6 bits (109), Expect = 2e-06
Identities = 20/61 (32%), Positives = 35/61 (56%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
+V +N +++ K+ N+ A+ VF + R PD +Y ++EGWG+ N KAR + E
Sbjct: 167 LVAFNGLLSALCKSKNVRKAQEVFENMRDRFTPDSKTYSILLEGWGKEPNLPKAREVFRE 226
Query: 63 L 63
+
Sbjct: 227 M 227
>At2g30780 hypothetical protein
Length = 452
Score = 45.4 bits (106), Expect = 5e-06
Identities = 23/62 (37%), Positives = 33/62 (53%), Gaps = 1/62 (1%)
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTLG-GRIEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
V YN +I GY A N D E F + G +EPD +Y+ M+ G+ +GN + YE
Sbjct: 210 VTYNFLIAGYMTAWNWDKMEATFQEMKRGPVEPDTDTYQLMLRGYANSGNLNRMEEMYEV 269
Query: 63 LK 64
+K
Sbjct: 270 IK 271
Score = 35.4 bits (80), Expect = 0.005
Identities = 17/63 (26%), Positives = 34/63 (52%), Gaps = 1/63 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGG-RIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+V YN +++ YG+ + E F L ++ P+ +Y +I G+ A N++K ++
Sbjct: 174 VVTYNILVSVYGRLLMVKNMEAAFEELQKVKLPPNSVTYNFLIAGYMTAWNWDKMEATFQ 233
Query: 62 ELK 64
E+K
Sbjct: 234 EMK 236
>At5g39980 putative protein
Length = 678
Score = 45.1 bits (105), Expect = 7e-06
Identities = 22/63 (34%), Positives = 37/63 (57%), Gaps = 1/63 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+V YNTMI YGK + A + + R IEP+ +Y ++I WG+AG ++A ++
Sbjct: 400 VVTYNTMIKIYGKTMEHEKATNLVQEMQSRGIEPNAITYSTIISIWGKAGKLDRAATLFQ 459
Query: 62 ELK 64
+L+
Sbjct: 460 KLR 462
Score = 36.2 bits (82), Expect = 0.003
Identities = 21/62 (33%), Positives = 34/62 (53%), Gaps = 1/62 (1%)
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
+ Y+T+I+ +GKA +D A +F L +E D+ Y++MI + R G A+ E
Sbjct: 436 ITYSTIISIWGKAGKLDRAATLFQKLRSSGVEIDQVLYQTMIVAYERVGLMGHAKRLLHE 495
Query: 63 LK 64
LK
Sbjct: 496 LK 497
Score = 28.5 bits (62), Expect = 0.65
Identities = 19/60 (31%), Positives = 28/60 (46%), Gaps = 1/60 (1%)
Query: 6 YNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKARWYYEELK 64
Y+T+IT +GK D A + R+ D Y ++IE R +Y KA + LK
Sbjct: 193 YSTLITSFGKEGMFDSALSWLQKMEQDRVSGDLVLYSNLIELSRRLCDYSKAISIFSRLK 252
Score = 28.5 bits (62), Expect = 0.65
Identities = 18/62 (29%), Positives = 32/62 (51%), Gaps = 1/62 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLG-GRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+V+Y+ +I + + A +F L I PD +Y SMI +G+A + +AR +
Sbjct: 225 LVLYSNLIELSRRLCDYSKAISIFSRLKRSGITPDLVAYNSMINVYGKAKLFREARLLIK 284
Query: 62 EL 63
E+
Sbjct: 285 EM 286
>At2g35130 hypothetical protein
Length = 591
Score = 43.9 bits (102), Expect = 2e-05
Identities = 22/63 (34%), Positives = 34/63 (53%), Gaps = 1/63 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+ VYN ++ Y +A GA +F + EPD SY M++ +GRAG + A +E
Sbjct: 334 VYVYNALMESYSRAGYPYGAAEIFSLMQHMGCEPDRASYNIMVDAYGRAGLHSDAEAVFE 393
Query: 62 ELK 64
E+K
Sbjct: 394 EMK 396
Score = 33.1 bits (74), Expect = 0.026
Identities = 20/62 (32%), Positives = 31/62 (49%), Gaps = 1/62 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYE 61
I YN +I YGKA ++ E +F+ L + PD ++ S I + R Y K +E
Sbjct: 474 ISTYNILINIYGKAGFLERIEELFVELKEKNFRPDVVTWTSRIGAYSRKKLYVKCLEVFE 533
Query: 62 EL 63
E+
Sbjct: 534 EM 535
Score = 32.0 bits (71), Expect = 0.059
Identities = 18/60 (30%), Positives = 33/60 (55%), Gaps = 1/60 (1%)
Query: 6 YNTMITGYGKASNMDGAEGVFLTLGG-RIEPDETSYRSMIEGWGRAGNYEKARWYYEELK 64
YN MI YGKAS + ++ + + +P+ +Y +++ + R G EKA +E+L+
Sbjct: 267 YNLMINLYGKASKSYMSWKLYCEMRSHQCKPNICTYTALVNAFAREGLCEKAEEIFEQLQ 326
Score = 30.8 bits (68), Expect = 0.13
Identities = 17/63 (26%), Positives = 32/63 (49%), Gaps = 1/63 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
++ +N +I YG+ AE +++ L R P E +Y +I+ + AG E+A
Sbjct: 155 VICFNLLIDAYGQKFQYKEAESLYVQLLESRYVPTEDTYALLIKAYCMAGLIERAEVVLV 214
Query: 62 ELK 64
E++
Sbjct: 215 EMQ 217
Score = 28.9 bits (63), Expect = 0.50
Identities = 17/61 (27%), Positives = 29/61 (46%), Gaps = 1/61 (1%)
Query: 5 VYNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKARWYYEEL 63
V N+M+ YG+ E + + G D ++Y +I +G+AG E+ + EL
Sbjct: 441 VLNSMLNLYGRLGQFTKMEKILAEMENGPCTADISTYNILINIYGKAGFLERIEELFVEL 500
Query: 64 K 64
K
Sbjct: 501 K 501
Score = 28.1 bits (61), Expect = 0.85
Identities = 14/50 (28%), Positives = 25/50 (50%), Gaps = 1/50 (2%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLG-GRIEPDETSYRSMIEGWGRAG 51
I Y ++ + + + AE +F L +EPD Y +++E + RAG
Sbjct: 299 ICTYTALVNAFAREGLCEKAEEIFEQLQEDGLEPDVYVYNALMESYSRAG 348
Score = 27.3 bits (59), Expect = 1.5
Identities = 14/57 (24%), Positives = 27/57 (46%), Gaps = 1/57 (1%)
Query: 9 MITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYEELK 64
+++ Y KA ++ E + + +EPD SM+ +GR G + K E++
Sbjct: 410 LLSAYSKARDVTKCEAIVKEMSENGVEPDTFVLNSMLNLYGRLGQFTKMEKILAEME 466
>At3g60050 putative protein
Length = 473
Score = 43.1 bits (100), Expect = 3e-05
Identities = 23/64 (35%), Positives = 39/64 (60%), Gaps = 3/64 (4%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVF--LTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYY 60
+V Y MITGY + +D A+ +F +T+ G++ P+ +Y SMI G AG + +A W
Sbjct: 359 VVCYTVMITGYVVSGELDKAKEMFREMTVKGQL-PNVFTYNSMIRGLCMAGEFREACWLL 417
Query: 61 EELK 64
+E++
Sbjct: 418 KEME 421
Score = 30.8 bits (68), Expect = 0.13
Identities = 18/63 (28%), Positives = 32/63 (50%), Gaps = 3/63 (4%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLT--LGGRIEPDETSYRSMIEGWGRAGNYEKARWYY 60
++ Y T+I G +A N++ + FL + PD Y MI G+ +G +KA+ +
Sbjct: 324 VLHYTTLIDGLSRAGNLEACK-YFLDEMVKAGCRPDVVCYTVMITGYVVSGELDKAKEMF 382
Query: 61 EEL 63
E+
Sbjct: 383 REM 385
Score = 30.4 bits (67), Expect = 0.17
Identities = 11/31 (35%), Positives = 20/31 (64%)
Query: 33 IEPDETSYRSMIEGWGRAGNYEKARWYYEEL 63
I+P Y ++I+G RAGN E +++ +E+
Sbjct: 320 IDPSVLHYTTLIDGLSRAGNLEACKYFLDEM 350
>At2g02150 hypothetical protein
Length = 1107
Score = 43.1 bits (100), Expect = 3e-05
Identities = 21/63 (33%), Positives = 36/63 (56%), Gaps = 1/63 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+ YN MI K +++ A G+F + R + PD +Y SMI+G+G+ G + ++E
Sbjct: 130 VFTYNIMIDCMCKEGDVEAARGLFEEMKFRGLVPDTVTYNSMIDGFGKVGRLDDTVCFFE 189
Query: 62 ELK 64
E+K
Sbjct: 190 EMK 192
Score = 34.3 bits (77), Expect = 0.012
Identities = 19/62 (30%), Positives = 30/62 (47%), Gaps = 1/62 (1%)
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTLGGRI-EPDETSYRSMIEGWGRAGNYEKARWYYEE 62
V YN+MI G+GK +D F + EPD +Y ++I + + G +Y E
Sbjct: 166 VTYNSMIDGFGKVGRLDDTVCFFEEMKDMCCEPDVITYNALINCFCKFGKLPIGLEFYRE 225
Query: 63 LK 64
+K
Sbjct: 226 MK 227
Score = 33.9 bits (76), Expect = 0.016
Identities = 23/60 (38%), Positives = 30/60 (49%), Gaps = 1/60 (1%)
Query: 6 YNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYEELK 64
YN +I G+ KA NMD A + L GR I+PD Y + I G E A+ E+K
Sbjct: 343 YNALIHGFVKAKNMDRALELLNELKGRGIKPDLLLYGTFIWGLCSLEKIEAAKVVMNEMK 402
Score = 32.7 bits (73), Expect = 0.035
Identities = 20/63 (31%), Positives = 32/63 (50%), Gaps = 1/63 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLG-GRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+V Y +I G A M AE +F + + P+ SY ++I G+ +A N ++A
Sbjct: 305 VVTYTALIDGLCDAERMKEAEELFGKMDTAGVIPNLASYNALIHGFVKAKNMDRALELLN 364
Query: 62 ELK 64
ELK
Sbjct: 365 ELK 367
Score = 28.1 bits (61), Expect = 0.85
Identities = 15/53 (28%), Positives = 30/53 (56%), Gaps = 1/53 (1%)
Query: 5 VYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKA 56
++ MI G K + ++ A +F + + + PD T+Y S+++G + GN +A
Sbjct: 483 IFTAMIDGLCKDNQVEAATTLFEQMVQKGLVPDRTAYTSLMDGNFKQGNVLEA 535
Score = 26.9 bits (58), Expect = 1.9
Identities = 16/59 (27%), Positives = 27/59 (45%), Gaps = 1/59 (1%)
Query: 7 NTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKARWYYEELK 64
N ++ + K D + F + G P +Y MI+ + G+ E AR +EE+K
Sbjct: 99 NGLLHRFAKLGKTDDVKRFFKDMIGAGARPTVFTYNIMIDCMCKEGDVEAARGLFEEMK 157
>At3g59040 unknown protein (At3g59040)
Length = 526
Score = 42.7 bits (99), Expect = 3e-05
Identities = 21/60 (35%), Positives = 37/60 (61%), Gaps = 1/60 (1%)
Query: 6 YNTMITGYGKASNMDGAEGVFLTLG-GRIEPDETSYRSMIEGWGRAGNYEKARWYYEELK 64
Y TM++ Y AS+M+GAE F + EP+ +Y ++I+G+ +A + EK YE+++
Sbjct: 330 YTTMLSAYVNASDMEGAEKFFKRIKVDGFEPNIVTYGTLIKGYAKANDVEKMMEVYEKMR 389
Score = 36.6 bits (83), Expect = 0.002
Identities = 19/57 (33%), Positives = 29/57 (50%), Gaps = 1/57 (1%)
Query: 9 MITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYEELK 64
+IT YGK N +GAE V L P+ SY +++E +GR G A + ++
Sbjct: 88 LITAYGKLGNFNGAERVLSVLSKMGSTPNVISYTALMESYGRGGKCNNAEAIFRRMQ 144
Score = 36.2 bits (82), Expect = 0.003
Identities = 20/64 (31%), Positives = 39/64 (60%), Gaps = 3/64 (4%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVF--LTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYY 60
IV Y T+I GY KA++++ V+ + L G I+ ++T ++++ GR N+ A +Y
Sbjct: 362 IVTYGTLIKGYAKANDVEKMMEVYEKMRLSG-IKANQTILTTIMDASGRCKNFGSALGWY 420
Query: 61 EELK 64
+E++
Sbjct: 421 KEME 424
Score = 36.2 bits (82), Expect = 0.003
Identities = 20/64 (31%), Positives = 32/64 (49%), Gaps = 4/64 (6%)
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTL----GGRIEPDETSYRSMIEGWGRAGNYEKARWY 59
+ Y ++ + + AE VF TL ++PD+ Y MI + +AGNYEKAR
Sbjct: 153 ITYQIILKTFVEGDKFKEAEEVFETLLDEKKSPLKPDQKMYHMMIYMYKKAGNYEKARKV 212
Query: 60 YEEL 63
+ +
Sbjct: 213 FSSM 216
Score = 31.2 bits (69), Expect = 0.10
Identities = 24/92 (26%), Positives = 37/92 (40%), Gaps = 33/92 (35%)
Query: 5 VYNTMITGYGKASNMDGAEGVFLTLGGR-------------------------------- 32
+Y+ MI Y KA N + A VF ++ G+
Sbjct: 192 MYHMMIYMYKKAGNYEKARKVFSSMVGKGVPQSTVTYNSLMSFETSYKEVSKIYDQMQRS 251
Query: 33 -IEPDETSYRSMIEGWGRAGNYEKARWYYEEL 63
I+PD SY +I+ +GRA E+A +EE+
Sbjct: 252 DIQPDVVSYALLIKAYGRARREEEALSVFEEM 283
>At1g03560 hypothetical protein
Length = 660
Score = 42.7 bits (99), Expect = 3e-05
Identities = 22/55 (40%), Positives = 34/55 (61%), Gaps = 1/55 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKA 56
+ YN ++ G A +D AE VF + GRI+PD +Y +MI+G+ +AG +KA
Sbjct: 222 LYTYNFLMNGLVSAMFVDSAERVFEVMESGRIKPDIVTYNTMIKGYCKAGQTQKA 276
Score = 30.4 bits (67), Expect = 0.17
Identities = 17/62 (27%), Positives = 32/62 (51%), Gaps = 1/62 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLG-GRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+V Y+ ++ G K ++ A F T + + Y S+I+G G+AG ++A +E
Sbjct: 397 VVTYSVVVNGLCKNGRVEEALDYFHTCRFDGLAINSMFYSSLIDGLGKAGRVDEAERLFE 456
Query: 62 EL 63
E+
Sbjct: 457 EM 458
>At4g16840 hypothetical protein
Length = 661
Score = 42.4 bits (98), Expect = 4e-05
Identities = 20/56 (35%), Positives = 35/56 (61%), Gaps = 3/56 (5%)
Query: 6 YNTMITGYGKASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+NTMITGY + M+ A +F ++ +E +E S+ +MI G+ G+ EKA +++
Sbjct: 158 WNTMITGYARRGEMEKARELFYSM---MEKNEVSWNAMISGYIECGDLEKASHFFK 210
Score = 30.8 bits (68), Expect = 0.13
Identities = 17/62 (27%), Positives = 31/62 (49%), Gaps = 1/62 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFL-TLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+V +N MI+GY + N D A +F + +I PD ++ +++ AG Y+E
Sbjct: 350 VVAWNAMISGYAQHGNADKALCLFREMIDNKIRPDWITFVAVLLACNHAGLVNIGMAYFE 409
Query: 62 EL 63
+
Sbjct: 410 SM 411
Score = 29.3 bits (64), Expect = 0.38
Identities = 18/61 (29%), Positives = 33/61 (53%), Gaps = 4/61 (6%)
Query: 4 VVYNTMITGYGK-ASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
+ +N+++ G K S M A +F + EPD SY M+ + R N+EKA+ +++
Sbjct: 93 ITWNSLLIGISKDPSRMMEAHQLFDEIP---EPDTFSYNIMLSCYVRNVNFEKAQSFFDR 149
Query: 63 L 63
+
Sbjct: 150 M 150
Score = 25.0 bits (53), Expect = 7.2
Identities = 10/25 (40%), Positives = 15/25 (60%)
Query: 32 RIEPDETSYRSMIEGWGRAGNYEKA 56
++EP Y M++ GRAG E+A
Sbjct: 416 KVEPQPDHYTCMVDLLGRAGKLEEA 440
>At4g01570 unknown protein
Length = 351
Score = 42.4 bits (98), Expect = 4e-05
Identities = 24/62 (38%), Positives = 36/62 (57%), Gaps = 1/62 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
IV+YNT+I GKA+ +D A +F + I PD SY +MIE +AG ++A Y +
Sbjct: 246 IVMYNTLINALGKATRLDEATQLFDHMKSNGINPDVVSYNTMIEVNSKAGKLKEAYKYLK 305
Query: 62 EL 63
+
Sbjct: 306 AM 307
Score = 30.4 bits (67), Expect = 0.17
Identities = 20/65 (30%), Positives = 30/65 (45%), Gaps = 5/65 (7%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTL---GGRIEPDETSYRSMIEGWGRAGNYEKARWY 59
I YN +I G GK D A V L GG + D Y ++I G+A ++A
Sbjct: 211 IATYNVIIQGLGKMGRADLASAVLDRLTKQGGYL--DIVMYNTLINALGKATRLDEATQL 268
Query: 60 YEELK 64
++ +K
Sbjct: 269 FDHMK 273
>At1g62260 hypothetical protein
Length = 656
Score = 42.0 bits (97), Expect = 6e-05
Identities = 22/59 (37%), Positives = 32/59 (53%), Gaps = 3/59 (5%)
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
+ +NTMI GY S M+ A +F + R D S+ M+ G+ GN E AR Y+E+
Sbjct: 313 ISWNTMIDGYVHVSRMEDAFALFSEMPNR---DAHSWNMMVSGYASVGNVELARHYFEK 368
Score = 33.1 bits (74), Expect = 0.026
Identities = 20/74 (27%), Positives = 37/74 (49%), Gaps = 12/74 (16%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTL--------GG----RIEPDETSYRSMIEGWGRA 50
+ YNT+I GYG+ ++ A +F + GG R + S+ SMI+ + +
Sbjct: 235 VYAYNTLIVGYGQRGQVEAARCLFDQIPDLCGDDHGGEFRERFCKNVVSWNSMIKAYLKV 294
Query: 51 GNYEKARWYYEELK 64
G+ AR ++++K
Sbjct: 295 GDVVSARLLFDQMK 308
Score = 32.7 bits (73), Expect = 0.035
Identities = 21/63 (33%), Positives = 34/63 (53%), Gaps = 6/63 (9%)
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGN---YEKARWYY 60
V +NTMI+GY K M+ A +F + R D ++ +MI G+ G E+AR +
Sbjct: 72 VTWNTMISGYVKRREMNQARKLFDVMPKR---DVVTWNTMISGYVSCGGIRFLEEARKLF 128
Query: 61 EEL 63
+E+
Sbjct: 129 DEM 131
Score = 27.3 bits (59), Expect = 1.5
Identities = 18/64 (28%), Positives = 33/64 (51%), Gaps = 6/64 (9%)
Query: 3 IVVYNTMITGY---GKASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKARWY 59
+V +NTMI+GY G ++ A +F + R D S+ +MI G+ + +A
Sbjct: 102 VVTWNTMISGYVSCGGIRFLEEARKLFDEMPSR---DSFSWNTMISGYAKNRRIGEALLL 158
Query: 60 YEEL 63
+E++
Sbjct: 159 FEKM 162
>At3g18110 hypothetical protein
Length = 1429
Score = 41.6 bits (96), Expect = 7e-05
Identities = 22/62 (35%), Positives = 36/62 (57%), Gaps = 1/62 (1%)
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
+ YNT+++ + SN+DGA VF + R +PD +Y +MI +GR G +A + E
Sbjct: 298 ITYNTLLSACSRDSNLDGAVKVFEDMEAHRCQPDLWTYNAMISVYGRCGLAAEAERLFME 357
Query: 63 LK 64
L+
Sbjct: 358 LE 359
Score = 36.2 bits (82), Expect = 0.003
Identities = 19/60 (31%), Positives = 33/60 (54%), Gaps = 1/60 (1%)
Query: 6 YNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYEELK 64
YN MI+ YG+ AE +F+ L + PD +Y S++ + R N EK + Y++++
Sbjct: 335 YNAMISVYGRCGLAAEAERLFMELELKGFFPDAVTYNSLLYAFARERNTEKVKEVYQQMQ 394
Score = 32.7 bits (73), Expect = 0.035
Identities = 20/58 (34%), Positives = 29/58 (49%), Gaps = 3/58 (5%)
Query: 2 CIVVYNTMITGYGKASNMDGAEGVF--LTLGGRIEPDETSYRSMIEGWGRAGNYEKAR 57
C +Y +I YGK AE V L GR PD ++ S++ + + G YE+AR
Sbjct: 751 CSPMYTDIIEAYGKQKLWQKAESVVGNLRQSGRT-PDLKTWNSLMSAYAQCGCYERAR 807
Score = 29.6 bits (65), Expect = 0.29
Identities = 20/63 (31%), Positives = 31/63 (48%), Gaps = 4/63 (6%)
Query: 4 VVYNTMITGYGKASNMDGAEGVFLT---LGGRIEPDETSYRSMIEGWGRAGNYEKARWYY 60
+ YNT+I YGK +D A ++ L GR PD +Y +I+ G+A +A
Sbjct: 403 MTYNTIIHMYGKQGQLDLALQLYKDMKGLSGR-NPDAITYTVLIDSLGKANRTVEAAALM 461
Query: 61 EEL 63
E+
Sbjct: 462 SEM 464
Score = 27.3 bits (59), Expect = 1.5
Identities = 19/61 (31%), Positives = 31/61 (50%), Gaps = 5/61 (8%)
Query: 6 YNTMITGYGKASNMDGAEGVFLTLGGR---IEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
YNT+I Y + + EG L R ++P +Y+S+I +G+ E+A +EE
Sbjct: 965 YNTLIIMYCRDRRPE--EGYLLMQQMRNLGLDPKLDTYKSLISAFGKQKCLEQAEQLFEE 1022
Query: 63 L 63
L
Sbjct: 1023 L 1023
Score = 26.9 bits (58), Expect = 1.9
Identities = 16/54 (29%), Positives = 29/54 (53%), Gaps = 1/54 (1%)
Query: 4 VVYNTMITGYGKASN-MDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKA 56
+ Y +I GKA+ ++ A + L I+P +Y ++I G+ +AG E+A
Sbjct: 439 ITYTVLIDSLGKANRTVEAAALMSEMLDVGIKPTLQTYSALICGYAKAGKREEA 492
>At2g31400 unknown protein
Length = 918
Score = 41.2 bits (95), Expect = 1e-04
Identities = 19/63 (30%), Positives = 35/63 (55%), Gaps = 1/63 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+V YN ++ GYGK D + VF + + P+ +Y ++I+G+ + G Y++A +
Sbjct: 479 VVTYNALLGGYGKQGKYDEVKKVFTEMKREHVLPNLLTYSTLIDGYSKGGLYKEAMEIFR 538
Query: 62 ELK 64
E K
Sbjct: 539 EFK 541
Score = 35.4 bits (80), Expect = 0.005
Identities = 16/62 (25%), Positives = 34/62 (54%), Gaps = 1/62 (1%)
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTLGG-RIEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
V YNT+++ Y K + A + + I+ D +Y +++ G+G+ G Y++ + + E
Sbjct: 445 VSYNTLLSIYTKVGRSEEALDILREMASVGIKKDVVTYNALLGGYGKQGKYDEVKKVFTE 504
Query: 63 LK 64
+K
Sbjct: 505 MK 506
Score = 34.7 bits (78), Expect = 0.009
Identities = 18/60 (30%), Positives = 34/60 (56%), Gaps = 1/60 (1%)
Query: 6 YNTMITGYGKASNMDGAEGVFLTLG-GRIEPDETSYRSMIEGWGRAGNYEKARWYYEELK 64
YNT++ K MD A + + RI P+ SY ++I+G+ +AG +++A + E++
Sbjct: 377 YNTLLDAICKGGQMDLAFEILAQMPVKRIMPNVVSYSTVIDGFAKAGRFDEALNLFGEMR 436
Score = 27.7 bits (60), Expect = 1.1
Identities = 17/63 (26%), Positives = 29/63 (45%), Gaps = 2/63 (3%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGR--IEPDETSYRSMIEGWGRAGNYEKARWYY 60
+V YN +I GK F R ++PD ++ S++ R G +E AR +
Sbjct: 303 LVTYNAVIDACGKGGMEFKQVAKFFDEMQRNGVQPDRITFNSLLAVCSRGGLWEAARNLF 362
Query: 61 EEL 63
+E+
Sbjct: 363 DEM 365
Score = 24.6 bits (52), Expect = 9.4
Identities = 15/59 (25%), Positives = 31/59 (52%), Gaps = 3/59 (5%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVF--LTLGGRIEPDETSYRSMIEGWGRAGNYEKARWY 59
+V+Y+ +I K + A + +T G I P+ +Y S+I+ +GR+ +++ Y
Sbjct: 549 VVLYSALIDALCKNGLVGSAVSLIDEMTKEG-ISPNVVTYNSIIDAFGRSATMDRSADY 606
>At1g62670 PPR-repeat protein
Length = 630
Score = 40.8 bits (94), Expect = 1e-04
Identities = 22/63 (34%), Positives = 36/63 (56%), Gaps = 1/63 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYE 61
I YN MI G KA ++ +F L + ++PD +Y +MI G+ R G+ E+A ++
Sbjct: 501 IYTYNIMIEGMCKAGKVEDGWDLFCNLSLKGVKPDVVAYNTMISGFCRKGSKEEADALFK 560
Query: 62 ELK 64
E+K
Sbjct: 561 EMK 563
Score = 33.1 bits (74), Expect = 0.026
Identities = 20/53 (37%), Positives = 29/53 (53%), Gaps = 1/53 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLG-GRIEPDETSYRSMIEGWGRAGNYE 54
I+ YNT++ G K ++ A VF L ++EP +Y MIEG +AG E
Sbjct: 466 IMTYNTLLDGLCKNGKLEKAMVVFEYLQRSKMEPTIYTYNIMIEGMCKAGKVE 518
Score = 32.3 bits (72), Expect = 0.045
Identities = 17/62 (27%), Positives = 35/62 (56%), Gaps = 1/62 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+V YNT+I G+ K ++ VF + R + + +Y +I+G +AG+ + A+ ++
Sbjct: 396 VVTYNTLIKGFCKYKRVEEGMEVFREMSQRGLVGNTVTYNILIQGLFQAGDCDMAQEIFK 455
Query: 62 EL 63
E+
Sbjct: 456 EM 457
Score = 30.0 bits (66), Expect = 0.22
Identities = 17/55 (30%), Positives = 29/55 (51%), Gaps = 1/55 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKA 56
+++YNT+I G K +MD A +F + + I P+ +Y S+I G + A
Sbjct: 256 VLIYNTIIDGLCKYKHMDDALNLFKEMETKGIRPNVVTYSSLISCLCNYGRWSDA 310
Score = 25.0 bits (53), Expect = 7.2
Identities = 15/64 (23%), Positives = 35/64 (54%), Gaps = 5/64 (7%)
Query: 3 IVVYNTMIT---GYGKASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKARWY 59
+V Y+++I+ YG+ S D + + + +I PD ++ ++I+ + + G +A
Sbjct: 291 VVTYSSLISCLCNYGRWS--DASRLLSDMIERKINPDVFTFSALIDAFVKEGKLVEAEKL 348
Query: 60 YEEL 63
Y+E+
Sbjct: 349 YDEM 352
>At1g22830 hypothetical protein, 5' partial
Length = 312
Score = 40.8 bits (94), Expect = 1e-04
Identities = 18/62 (29%), Positives = 40/62 (64%), Gaps = 3/62 (4%)
Query: 2 CIVVYNTMITGYGKASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
C++++N+++ Y K+ + A+ VF ++ R D+ +Y S+I+G+GR G E A +++
Sbjct: 67 CLILWNSLVDMYAKSGEIIAAKRVFDSMRKR---DKVTYTSLIDGYGRLGKGEVALAWFK 123
Query: 62 EL 63
++
Sbjct: 124 DM 125
>At2g36240 putative salt-inducible protein
Length = 497
Score = 40.4 bits (93), Expect = 2e-04
Identities = 20/61 (32%), Positives = 36/61 (58%), Gaps = 1/61 (1%)
Query: 5 VYNTMITGYGKASNMDGAEGVFLTLG-GRIEPDETSYRSMIEGWGRAGNYEKARWYYEEL 63
VYNT++ GY K+ +MD A + +G R +PD ++ +I G+ R+ ++ A + E+
Sbjct: 195 VYNTVVNGYVKSGDMDKALRFYQRMGKERAKPDVCTFNILINGYCRSSKFDLALDLFREM 254
Query: 64 K 64
K
Sbjct: 255 K 255
Score = 30.4 bits (67), Expect = 0.17
Identities = 16/61 (26%), Positives = 34/61 (55%), Gaps = 2/61 (3%)
Query: 5 VYNTMITGYGKASNMDGAEGVFLTLGGRIE--PDETSYRSMIEGWGRAGNYEKARWYYEE 62
++ + I Y +A MD A F T+ I+ P+ Y +++ G+ ++G+ +KA +Y+
Sbjct: 159 IFRSAIDAYCRARKMDYALLAFDTMKRLIDGKPNVGVYNTVVNGYVKSGDMDKALRFYQR 218
Query: 63 L 63
+
Sbjct: 219 M 219
Score = 25.0 bits (53), Expect = 7.2
Identities = 9/30 (30%), Positives = 17/30 (56%)
Query: 34 EPDETSYRSMIEGWGRAGNYEKARWYYEEL 63
EPDET+Y ++ G+ + G ++ E+
Sbjct: 435 EPDETTYHVLVSGFTKEGRRKEGEVLVNEM 464
>At2g06000 unknown protein
Length = 536
Score = 40.4 bits (93), Expect = 2e-04
Identities = 21/56 (37%), Positives = 33/56 (58%), Gaps = 2/56 (3%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVF--LTLGGRIEPDETSYRSMIEGWGRAGNYEKA 56
IV YNT+I G+ K++ ++ A +F + G PD +Y SMI G+ +AG +A
Sbjct: 241 IVTYNTLIQGFCKSNELNKASEMFKDVKSGSVCSPDVVTYTSMISGYCKAGKMREA 296
Score = 34.7 bits (78), Expect = 0.009
Identities = 20/63 (31%), Positives = 33/63 (51%), Gaps = 5/63 (7%)
Query: 4 VVYNTMITGYGKASNMDGAE---GVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYY 60
V +N ++ GY KA M AE G ++ G PD ++ S+I+G+ R G + +
Sbjct: 313 VTFNVLVDGYAKAGEMLTAEEIRGKMISFG--CFPDVVTFTSLIDGYCRVGQVSQGFRLW 370
Query: 61 EEL 63
EE+
Sbjct: 371 EEM 373
Score = 32.7 bits (73), Expect = 0.035
Identities = 18/52 (34%), Positives = 29/52 (55%), Gaps = 5/52 (9%)
Query: 3 IVVYNTMITGYGKASNMDGAEGV---FLTLGGRIEPDETSYRSMIEGWGRAG 51
+V Y +MI+GY KA M A + L LG I P ++ +++G+ +AG
Sbjct: 277 VVTYTSMISGYCKAGKMREASSLLDDMLRLG--IYPTNVTFNVLVDGYAKAG 326
Score = 28.5 bits (62), Expect = 0.65
Identities = 16/55 (29%), Positives = 28/55 (50%), Gaps = 2/55 (3%)
Query: 10 ITGYGKASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYEELK 64
+ G GKA GV G EPD +Y ++I+G+ ++ KA ++++K
Sbjct: 216 LCGVGKAEKALELLGVMSGFG--CEPDIVTYNTLIQGFCKSNELNKASEMFKDVK 268
>At2g02980 unknown protein
Length = 603
Score = 40.4 bits (93), Expect = 2e-04
Identities = 18/64 (28%), Positives = 36/64 (56%), Gaps = 1/64 (1%)
Query: 2 CIVVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYY 60
C+V YN MITGY + + + A +F + G+ ++P+E + S++ G+ + +W +
Sbjct: 194 CVVCYNAMITGYARRNRPNEALSLFREMQGKYLKPNEITLLSVLSSCALLGSLDLGKWIH 253
Query: 61 EELK 64
+ K
Sbjct: 254 KYAK 257
Score = 25.8 bits (55), Expect = 4.2
Identities = 16/56 (28%), Positives = 29/56 (51%), Gaps = 1/56 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFL-TLGGRIEPDETSYRSMIEGWGRAGNYEKAR 57
IV++N+M GY + +N +F+ L I PD ++ S+++ A E+ R
Sbjct: 94 IVIFNSMARGYSRFTNPLEVFSLFVEILEDGILPDNYTFPSLLKACAVAKALEEGR 149
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.137 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,469,130
Number of Sequences: 26719
Number of extensions: 46780
Number of successful extensions: 1254
Number of sequences better than 10.0: 323
Number of HSP's better than 10.0 without gapping: 189
Number of HSP's successfully gapped in prelim test: 134
Number of HSP's that attempted gapping in prelim test: 324
Number of HSP's gapped (non-prelim): 960
length of query: 64
length of database: 11,318,596
effective HSP length: 40
effective length of query: 24
effective length of database: 10,249,836
effective search space: 245996064
effective search space used: 245996064
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 52 (24.6 bits)
Medicago: description of AC141111.3


