

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140916.4 - phase: 0
(250 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
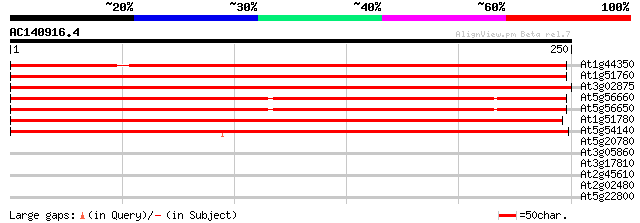
Score E
Sequences producing significant alignments: (bits) Value
At1g44350 IAA-amino acid hydrolase (At1g44350) 345 1e-95
At1g51760 unknown protein 266 6e-72
At3g02875 IAA-amino acid hydrolase (ILR1) 243 5e-65
At5g56660 IAA-amino acid hydrolase (gb|AAC04866.1) 237 5e-63
At5g56650 IAA-amino acid hydrolase homolog 1 precursor (sp|P54969) 236 9e-63
At1g51780 hypothetical protein 230 5e-61
At5g54140 IAA-amino acid hydrolase homolog ILL3 (gb|AAC31939.1) 207 3e-54
At5g20780 putative protein 30 1.2
At3g05860 putative DNA-binding protein 30 1.2
At3g17810 dihydroorotate dehydrogenase like protein 28 4.6
At2g45610 unknown protein 28 6.0
At2g02480 STICHEL (STI) 28 6.0
At5g22800 alanyl-tRNA synthetase 27 7.9
>At1g44350 IAA-amino acid hydrolase (At1g44350)
Length = 464
Score = 345 bits (886), Expect = 1e-95
Identities = 166/248 (66%), Positives = 209/248 (83%), Gaps = 5/248 (2%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
MI+DGAL+DVEAIFAVHVSH HPTG+IGSR GPLLAGCG FRAVI+ + + R +A+
Sbjct: 219 MIEDGALDDVEAIFAVHVSHIHPTGVIGSRSGPLLAGCGIFRAVITSE-----DSRGAAN 273
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
+LAAS+AVIS+QGIVSRE++PLDSQVVSVTSF+GG+S D+ PD+VV+GGTFRAFSN+SF
Sbjct: 274 LLLAASSAVISLQGIVSREASPLDSQVVSVTSFDGGHSLDVAPDTVVLGGTFRAFSNSSF 333
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
Y L +RI++V++ Q V+ C A V+FFEK+ IYPPT N+D Y H+KKV+IDLLG +F
Sbjct: 334 YYLKKRIQEVLMDQVGVFGCQATVNFFEKQNAIYPPTTNNDATYNHLKKVTIDLLGDSHF 393
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHAT 240
+ P MMGAED++FYS++IP+AF++IGIRNE LGS H HSPHF IDED+LP+GAAVHA
Sbjct: 394 TLAPQMMGAEDFAFYSEIIPAAFYFIGIRNEELGSVHIAHSPHFMIDEDSLPVGAAVHAA 453
Query: 241 IAERYLNE 248
+AERYLN+
Sbjct: 454 VAERYLND 461
>At1g51760 unknown protein
Length = 440
Score = 266 bits (681), Expect = 6e-72
Identities = 129/248 (52%), Positives = 182/248 (73%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
+++ G LE+V AIF +HV+++ G + SR GP+LAG GFF+A ISGK AA P+++ D
Sbjct: 178 IVEAGVLENVSAIFGLHVTNQLALGQVSSREGPMLAGSGFFKAKISGKGGHAALPQHTID 237
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
P+LAAS ++S+Q +VSRE++PLDSQVV+V F GG + ++IPDSV IGGTFRAFS SF
Sbjct: 238 PILAASNVIVSLQHLVSREADPLDSQVVTVAKFEGGGAFNVIPDSVTIGGTFRAFSTKSF 297
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
QL +RIEQVI +QASV C A VDF E+E +PPTVND +++ K VS D+LG +N+
Sbjct: 298 MQLKKRIEQVITRQASVNMCNATVDFIEEEKPFFPPTVNDKALHQFFKNVSGDMLGIENY 357
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHAT 240
+ P+MG+ED+SFY Q IP F ++G++N+ + HSP+F ++E+ LP GA++HA+
Sbjct: 358 VEMQPLMGSEDFSFYQQAIPGHFSFVGMQNKARSPMASPHSPYFEVNEELLPYGASLHAS 417
Query: 241 IAERYLNE 248
+A RYL E
Sbjct: 418 MATRYLLE 425
>At3g02875 IAA-amino acid hydrolase (ILR1)
Length = 442
Score = 243 bits (621), Expect = 5e-65
Identities = 120/250 (48%), Positives = 174/250 (69%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
M++D L+D++ I +VHV P+G IGSRPG +LAG G F + G+ + AA P S D
Sbjct: 182 MLKDEILDDLDGILSVHVFPSIPSGGIGSRPGTVLAGAGLFTVTVHGQGSHAATPHFSKD 241
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
PVLAAS+AV+++Q IVSRE +PL++ VV+V GG++ ++IP S GGTFR+ SN
Sbjct: 242 PVLAASSAVVALQQIVSRELDPLEAGVVTVGYIEGGHAQNVIPQSAKFGGTFRSLSNDGL 301
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
+ RI+++ QASVY C AEV+F EK+ +++P ND+ +YEH KKV+ ++G+ NF
Sbjct: 302 LFIQRRIKEISEAQASVYRCKAEVNFEEKKPSLHPVMNNDEGLYEHGKKVAEAMIGKNNF 361
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHAT 240
P MG ED+SF++Q +A F +G++NETLG+ HSP+F +DE+ALP+GAA+HA
Sbjct: 362 HDFPVTMGGEDFSFFTQKTKAAIFVLGVKNETLGAGKPLHSPYFFVDEEALPVGAALHAA 421
Query: 241 IAERYLNEHG 250
+A YL+EHG
Sbjct: 422 MAVSYLDEHG 431
>At5g56660 IAA-amino acid hydrolase (gb|AAC04866.1)
Length = 439
Score = 237 bits (604), Expect = 5e-63
Identities = 122/248 (49%), Positives = 176/248 (70%), Gaps = 3/248 (1%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
M ++GAL++VEAIF +H+S P G SR G LAG G F AVI+GK AA P+++ D
Sbjct: 181 MREEGALKNVEAIFGIHLSARIPFGKAASRAGSFLAGAGVFEAVITGKGGHAAIPQHTID 240
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
PV+AAS+ V+S+Q +VSRE++PLDS+VV+V+ NGGN+ ++IPDS+ IGGT RAF T F
Sbjct: 241 PVVAASSIVLSLQQLVSRETDPLDSKVVTVSKVNGGNAFNVIPDSITIGGTLRAF--TGF 298
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
QL +R+++VI +QA+V+ C A V+ PPTVN+ +Y+ KKV DLLGQ+ F
Sbjct: 299 TQLQQRVKEVITKQAAVHRCNASVNLTPNGREPMPPTVNNKDLYKQFKKVVRDLLGQEAF 358
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHAT 240
P+MG+ED+S++++ IP F +G+++ET G + HSP + I+ED LP GAA+HA+
Sbjct: 359 VEAAPVMGSEDFSYFAETIPGHFSLLGMQDETNGYA-SSHSPLYRINEDVLPYGAAIHAS 417
Query: 241 IAERYLNE 248
+A +YL E
Sbjct: 418 MAVQYLKE 425
>At5g56650 IAA-amino acid hydrolase homolog 1 precursor (sp|P54969)
Length = 438
Score = 236 bits (602), Expect = 9e-63
Identities = 122/248 (49%), Positives = 174/248 (69%), Gaps = 3/248 (1%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
M ++GAL++VEAIF +H+S P G S G +AG G F AVI+GK AA P+++ D
Sbjct: 180 MREEGALKNVEAIFGIHLSPRTPFGKAASLAGSFMAGAGAFEAVITGKGGHAAIPQHTID 239
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
PV+AAS+ V+S+Q +VSRE++P DS+VV+VT NGGN+ ++IPDS+ IGGT RAF T F
Sbjct: 240 PVVAASSIVLSLQHLVSRETDPSDSKVVTVTKVNGGNAFNVIPDSITIGGTLRAF--TGF 297
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
QL ERI+++I +QA+V+ C A V+ PPTVN+ +Y+ KKV DLLGQ+ F
Sbjct: 298 TQLQERIKEIITKQAAVHRCNASVNLAPNGNQPMPPTVNNMDLYKKFKKVVRDLLGQEAF 357
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHAT 240
P MG+ED+S++++ IP F +G+++ET G + HSPH+ I+ED LP GAA+HAT
Sbjct: 358 VEAVPEMGSEDFSYFAETIPGHFSLLGMQDETQGYA-SSHSPHYRINEDVLPYGAAIHAT 416
Query: 241 IAERYLNE 248
+A +YL +
Sbjct: 417 MAVQYLKD 424
>At1g51780 hypothetical protein
Length = 424
Score = 230 bits (587), Expect = 5e-61
Identities = 120/246 (48%), Positives = 169/246 (67%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
+++ G LE+V AIF +HVS+ G + SR G L+AG G F+A ISGK AA P+ + D
Sbjct: 167 IVEAGVLENVGAIFGLHVSNLLGLGQLSSREGLLMAGSGRFKATISGKGGHAALPQFAID 226
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
PVLAAS ++S+Q +VSRE++PLDSQVV+V +F G ++ ++IPDSV IGGTFRA SF
Sbjct: 227 PVLAASNVILSLQHLVSREADPLDSQVVTVATFEGSDAFNVIPDSVTIGGTFRALLPKSF 286
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
QL +RI QVI QASV C A VDF E E +PPTVN+ ++ K VS+D+LG +N+
Sbjct: 287 EQLKQRIVQVITTQASVNMCNATVDFLEDETPPFPPTVNNKTLHLFYKNVSVDMLGIENY 346
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHAT 240
P+M +ED++FY Q IP F ++G++N++ HSP F ++E+ LP GA++ A+
Sbjct: 347 VETLPVMVSEDFAFYQQAIPGHFSFVGMQNKSHSPMANPHSPFFEVNEELLPYGASLLAS 406
Query: 241 IAERYL 246
+A RYL
Sbjct: 407 LATRYL 412
>At5g54140 IAA-amino acid hydrolase homolog ILL3 (gb|AAC31939.1)
Length = 428
Score = 207 bits (528), Expect = 3e-54
Identities = 108/250 (43%), Positives = 159/250 (63%), Gaps = 1/250 (0%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
MI++GAL D EAIF +HV PTG + + GP LA F +SGK +++ + D
Sbjct: 171 MIKEGALGDSEAIFGMHVHTGLPTGELATISGPALASTSIFSVRMSGKSPASSETYSCVD 230
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSF-NGGNSHDMIPDSVVIGGTFRAFSNTS 119
PVLAAS+ ++++Q I+SRE +PL S V+SVT +GG+ D+IP V GGT R+ +
Sbjct: 231 PVLAASSTILALQLIISREVDPLLSHVLSVTFMKSGGSEFDVIPAYVEFGGTLRSLTTNG 290
Query: 120 FYQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKN 179
L++R+++V+ QA V C A++D E ++ +YP TVND +++E +KV LLG +
Sbjct: 291 INWLIKRLKEVVEGQAEVQRCKADIDMHEDDHPMYPATVNDHKLHEFTEKVLKLLLGPEK 350
Query: 180 FRVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHA 239
+ +M ED++FY Q IP + IGIRNE +GS + HSP+F +DE+ LPIG+A A
Sbjct: 351 VKPANKVMAGEDFAFYQQKIPGYYIGIGIRNEEIGSVRSVHSPYFFLDENVLPIGSATFA 410
Query: 240 TIAERYLNEH 249
+AE YL EH
Sbjct: 411 ALAEMYLQEH 420
>At5g20780 putative protein
Length = 818
Score = 30.0 bits (66), Expect = 1.2
Identities = 12/34 (35%), Positives = 19/34 (55%)
Query: 158 VNDDQMYEHVKKVSIDLLGQKNFRVVPPMMGAED 191
+NDD + EHV+ V + LLG + V + +D
Sbjct: 185 INDDILLEHVQAVELQLLGGRELEAVENLFDGQD 218
>At3g05860 putative DNA-binding protein
Length = 249
Score = 30.0 bits (66), Expect = 1.2
Identities = 24/92 (26%), Positives = 41/92 (44%), Gaps = 9/92 (9%)
Query: 94 NGGNSHDMIPDSVVIGGTFRAFSNTSFYQLLERIEQVIVQQASVYSCFAEVDFFEKEYTI 153
+ G ++ M P SVV F S+ YQ ++ I+ Q +Y+ +++ +E+
Sbjct: 167 SSGATNAMTPTSVVENPNFNHLSHNQ-YQYQQQFGYPILVQDGIYNP-SQIQNQHEEWL- 223
Query: 154 YPPTVNDDQMYEHVKKVSIDLLGQKNFRVVPP 185
DD M H K++S L+ NF P
Sbjct: 224 ------DDHMMNHSKEISHPLMDDNNFYYQQP 249
>At3g17810 dihydroorotate dehydrogenase like protein
Length = 426
Score = 28.1 bits (61), Expect = 4.6
Identities = 11/39 (28%), Positives = 23/39 (58%), Gaps = 1/39 (2%)
Query: 103 PDSVVIGGTFRAFSNTSFYQLLERIEQVIVQQASV-YSC 140
PD ++I ++ T++ +L++R+EQ V + +SC
Sbjct: 153 PDRILIASVMEEYNKTAWEELIDRVEQTGVDALEINFSC 191
>At2g45610 unknown protein
Length = 324
Score = 27.7 bits (60), Expect = 6.0
Identities = 11/31 (35%), Positives = 18/31 (57%)
Query: 37 GCGFFRAVISGKRASAANPRNSADPVLAASA 67
GC F++ + GK + + +N ADPV+ A
Sbjct: 196 GCVFYQPLFGGKTRTKSELKNFADPVMPVPA 226
>At2g02480 STICHEL (STI)
Length = 1218
Score = 27.7 bits (60), Expect = 6.0
Identities = 29/113 (25%), Positives = 50/113 (43%), Gaps = 11/113 (9%)
Query: 27 IGSRPGPLLAGCGFFRAVISGKRASAANPRNSADPVLAASAAVISIQGIVSRESNPLDSQ 86
+GS P P G R S RA+ +P + + V+A + + S+ ++P
Sbjct: 799 LGSMPSPGTTHTGSSRRQSS--RATDDDPASVSREVMAYKQRIGGLH--FSKSASP---- 850
Query: 87 VVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSFYQLLERIEQVIVQQASVYS 139
SV NG +SH+ P S VI + ++S Q++E + + S+ S
Sbjct: 851 -ASVIKRNGNHSHEAKPFSRVIDN--NCYKSSSSSQMIESEGSIASHENSIAS 900
>At5g22800 alanyl-tRNA synthetase
Length = 954
Score = 27.3 bits (59), Expect = 7.9
Identities = 18/77 (23%), Positives = 32/77 (41%), Gaps = 7/77 (9%)
Query: 39 GFFRAVISGKRASAANPRNSADPVLAASAAVISIQGIVSRESNPLDSQVVSVTS------ 92
GF +G S PR++ P + A ++ +S+Q + S+ VS S
Sbjct: 5 GFVTRTSAGVSPSILLPRSTQSPQIIAKSSSVSVQPVSEDAKEDYQSKDVSGDSIRRRFL 64
Query: 93 -FNGGNSHDMIPDSVVI 108
F H ++P S ++
Sbjct: 65 EFFASRGHKVLPSSSLV 81
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.135 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,570,996
Number of Sequences: 26719
Number of extensions: 223685
Number of successful extensions: 484
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 468
Number of HSP's gapped (non-prelim): 14
length of query: 250
length of database: 11,318,596
effective HSP length: 97
effective length of query: 153
effective length of database: 8,726,853
effective search space: 1335208509
effective search space used: 1335208509
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC140916.4


