

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140913.9 - phase: 2 /pseudo
(273 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
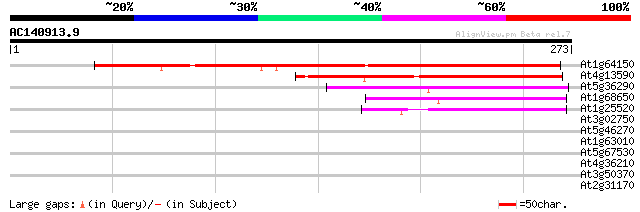
Sequences producing significant alignments: (bits) Value
At1g64150 unknown protein 246 1e-65
At4g13590 unknown protein 94 1e-19
At5g36290 transmembrane protein FT27/PFT27-like 72 3e-13
At1g68650 transmembrane like protein 63 2e-10
At1g25520 unknown protein 59 4e-09
At3g02750 protein phosphatase-2C (PP2C) like protein 35 0.033
At5g46270 disease resistance protein-like 29 3.1
At1g63010 tetracycline resistance efflux protein like protein 29 3.1
At5g67530 cyclophilin (ROC23) 27 9.0
At4g36210 putative protein 27 9.0
At3g50370 putative protein 27 9.0
At2g31170 putative cysteinyl-tRNA synthetase 27 9.0
>At1g64150 unknown protein
Length = 370
Score = 246 bits (627), Expect = 1e-65
Identities = 148/237 (62%), Positives = 168/237 (70%), Gaps = 13/237 (5%)
Query: 42 VLKFMLFSAFFALQDAFPAVAASDFATGLNS-IPIFGDVGDLSTGFASYHRHFC*YFSLN 100
VL F+ S AL PA AAS S + FGD+GD+S+GFAS +FS
Sbjct: 114 VLMFLAVSGSVALLGTDPAFAASSIPNVTQSLVTSFGDLGDISSGFAS--AFLLIFFSEL 171
Query: 101 WETRLFSLQHC*QLEIQPVLF-----SLGHLAH----LRRTFHYVDELLPFRFGETDLPI 151
+ F +F +LG + L RTFHYVDE+LPFRFG TDLPI
Sbjct: 172 GDKTFFIAALLAARNSAATVFVGTFGALGIMTIISVVLGRTFHYVDEVLPFRFGGTDLPI 231
Query: 152 DDIAAVCLLVYFGVSTLLDASSSDSQKSDDEQKEAELAVSDFSGDGAGILAAASTIVSTF 211
DDIAAVCLLVYFGVSTLLDA S D K+D+EQKEAELAVS+ SG+GAGI+AAA+TI+STF
Sbjct: 232 DDIAAVCLLVYFGVSTLLDAVS-DEGKADEEQKEAELAVSELSGNGAGIVAAANTIISTF 290
Query: 212 LLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEK 268
LVFVAEWGDKSFFSTIALAAASSPLGVIAG+LAGHG ATL+AVLGGSLLG FLSEK
Sbjct: 291 ALVFVAEWGDKSFFSTIALAAASSPLGVIAGALAGHGAATLLAVLGGSLLGNFLSEK 347
>At4g13590 unknown protein
Length = 359
Score = 93.6 bits (231), Expect = 1e-19
Identities = 53/139 (38%), Positives = 85/139 (61%), Gaps = 12/139 (8%)
Query: 140 LPFRFGETDLPIDDIAAVCLLVYFGVSTLLDA---------SSSDSQKSDDEQKEAELAV 190
+P +F +T LPI + AA+ LL++FG+ ++ DA + ++ E EAE V
Sbjct: 203 VPAQF-QTTLPIGEYAAIALLMFFGLKSIKDAWDLPPVEAKNGEETGIELGEYSEAEELV 261
Query: 191 SDFSGDGAGILAAASTIVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVA 250
+ + + + +F LVF AEWGD+S +T+AL AA SPLGV +G++AGH VA
Sbjct: 262 KEKASKK--LTNPLEILWKSFSLVFFAEWGDRSMLATVALGAAQSPLGVASGAIAGHLVA 319
Query: 251 TLIAVLGGSLLGTFLSEKV 269
T++A++GG+ L ++SEK+
Sbjct: 320 TVLAIMGGAFLANYISEKL 338
Score = 39.3 bits (90), Expect = 0.002
Identities = 23/68 (33%), Positives = 37/68 (53%), Gaps = 2/68 (2%)
Query: 195 GDGAGILAAA--STIVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATL 252
G + +LAA S + F L+FV+E GDK+FF LA V+ GS+ + T+
Sbjct: 132 GGPSSVLAAVAKSGFTAAFSLIFVSEIGDKTFFIAALLAMQYEKTLVLLGSMGALSLMTI 191
Query: 253 IAVLGGSL 260
++V+ G +
Sbjct: 192 LSVVIGKI 199
>At5g36290 transmembrane protein FT27/PFT27-like
Length = 293
Score = 72.0 bits (175), Expect = 3e-13
Identities = 42/124 (33%), Positives = 67/124 (53%), Gaps = 6/124 (4%)
Query: 155 AAVCLLVYFGVSTLLDASSSDSQKSDDEQKEAELAVSDFSGDGAGILA------AASTIV 208
AA L +FG+ L A S KS+ +++ E+ SG G +
Sbjct: 151 AATVLYAFFGLRLLYIAWRSTDSKSNQKKEMEEVEEKLESGQGKTPFRRLFSRFCTPIFL 210
Query: 209 STFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEK 268
+F+L F+AEWGD+S +TIALA + +GV G+ GH V T +AV+GGS+L + +S++
Sbjct: 211 ESFILTFLAEWGDRSQIATIALATHKNAIGVAIGASIGHTVCTSLAVVGGSMLASRISQR 270
Query: 269 VTSS 272
++
Sbjct: 271 TVAT 274
Score = 34.3 bits (77), Expect = 0.074
Identities = 18/66 (27%), Positives = 36/66 (54%)
Query: 207 IVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLS 266
+ S+F ++ V E GD++F +A V++G+L+ V T+++ G ++ +S
Sbjct: 85 LFSSFSMILVTEIGDETFIIAALMAMRHPKATVLSGALSALFVMTILSTGLGRIVPNLIS 144
Query: 267 EKVTSS 272
K T+S
Sbjct: 145 RKHTNS 150
>At1g68650 transmembrane like protein
Length = 228
Score = 62.8 bits (151), Expect = 2e-10
Identities = 35/99 (35%), Positives = 55/99 (55%), Gaps = 1/99 (1%)
Query: 174 SDSQKSDDEQKEAELAVSDFSGDGAGILAAASTI-VSTFLLVFVAEWGDKSFFSTIALAA 232
SD +K++D+ K +++ + A S I + F + F EWGDKS +TI LAA
Sbjct: 109 SDLKKTNDQSKNSKIEDEQKKQKRPFLTAFFSPIFLKAFSINFFGEWGDKSQLATIGLAA 168
Query: 233 ASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEKVTS 271
+PLGV+ G + + T AVLGG L + +SE++ +
Sbjct: 169 DENPLGVVLGGIVAQTLCTTAAVLGGKSLASQISERIVA 207
Score = 31.2 bits (69), Expect = 0.62
Identities = 19/58 (32%), Positives = 30/58 (50%)
Query: 213 LVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEKVT 270
+ F++E GDK+FF+ LA V+AG L+ V T+++ G +S K T
Sbjct: 14 MTFLSEIGDKTFFAAAILAMRYPRRLVLAGCLSALIVMTILSATLGWAAPNLISRKWT 71
>At1g25520 unknown protein
Length = 230
Score = 58.5 bits (140), Expect = 4e-09
Identities = 34/101 (33%), Positives = 52/101 (50%), Gaps = 10/101 (9%)
Query: 172 SSSDSQKSDDEQKEAELA-VSDFSGDGAGILAAASTIVSTFLLVFVAEWGDKSFFSTIAL 230
S DS K +DE K+ A ++ F + + F + F EWGDKS +TI L
Sbjct: 118 SPKDSSKREDENKKQNRAFLTQFF---------SPIFLKAFSINFFGEWGDKSQLATIGL 168
Query: 231 AAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEKVTS 271
AA +P GV+ G + + T AV+GG L + +SE++ +
Sbjct: 169 AADENPFGVVLGGVVAQFLCTTAAVIGGKSLASQISERIVA 209
Score = 32.3 bits (72), Expect = 0.28
Identities = 20/58 (34%), Positives = 30/58 (51%)
Query: 213 LVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEKVT 270
+ FV+E GDK+FF+ LA V+AG L+ V T+++ G +S K T
Sbjct: 14 MTFVSEIGDKTFFAAAILAMRYPRRLVLAGCLSALIVMTILSATLGWAAPNLISRKWT 71
>At3g02750 protein phosphatase-2C (PP2C) like protein
Length = 492
Score = 35.4 bits (80), Expect = 0.033
Identities = 19/59 (32%), Positives = 30/59 (50%)
Query: 151 IDDIAAVCLLVYFGVSTLLDASSSDSQKSDDEQKEAELAVSDFSGDGAGILAAASTIVS 209
+DD AAVCL + + + +SS S+ D E++E + D G L +ST+ S
Sbjct: 383 VDDCAAVCLYLDSSNTNAISTASSISKLEDGEEEELKATTEDDDASGPSGLGRSSTVRS 441
>At5g46270 disease resistance protein-like
Length = 1145
Score = 28.9 bits (63), Expect = 3.1
Identities = 23/86 (26%), Positives = 41/86 (46%), Gaps = 3/86 (3%)
Query: 108 LQHC*QLEIQPVLFSLGHLAHLRRTFHYVDELLPFRFGETDLPIDDIAAVCLLVYFGVST 167
++ C LEI P F+L L HL F Y EL F T++ + + + + +
Sbjct: 681 MEFCHSLEILPTGFNLKSLDHL--NFRYCSELRTFPEFSTNISVLMLFGTNIEEFPNLEN 738
Query: 168 LLDASSSDSQKSDDEQKEAELAVSDF 193
L++ S S ++SD +Q + ++ F
Sbjct: 739 LVELSLS-KEESDGKQWDGVKPLTPF 763
>At1g63010 tetracycline resistance efflux protein like protein
Length = 699
Score = 28.9 bits (63), Expect = 3.1
Identities = 12/59 (20%), Positives = 30/59 (50%)
Query: 199 GILAAASTIVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLG 257
G++ + + F V+ + W +KS+F + ++ + +G + +LA + + +LG
Sbjct: 286 GVVIGSMAVAQVFSSVYFSAWSNKSYFKPLVFSSIALFIGNLMYALAYDANSIALLLLG 344
>At5g67530 cyclophilin (ROC23)
Length = 595
Score = 27.3 bits (59), Expect = 9.0
Identities = 17/62 (27%), Positives = 30/62 (47%), Gaps = 2/62 (3%)
Query: 146 ETDLPIDDIAAVCLLVYFGVSTLLD--ASSSDSQKSDDEQKEAELAVSDFSGDGAGILAA 203
E+D P+++I + V+ T LD ++K +E K+ E S +S G+G A
Sbjct: 481 ESDRPLEEIKIIEASVFVNPYTELDEEEEKEKAEKEKNEDKDIEKIGSWYSNPGSGTTEA 540
Query: 204 AS 205
+
Sbjct: 541 GA 542
>At4g36210 putative protein
Length = 516
Score = 27.3 bits (59), Expect = 9.0
Identities = 15/39 (38%), Positives = 22/39 (55%), Gaps = 3/39 (7%)
Query: 231 AAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEKV 269
AAA+S G +AGS+A VA G L GT ++ ++
Sbjct: 244 AAAASAAGTVAGSVA---VAASFGAAGAGLTGTKMARRI 279
>At3g50370 putative protein
Length = 2152
Score = 27.3 bits (59), Expect = 9.0
Identities = 23/88 (26%), Positives = 40/88 (45%), Gaps = 17/88 (19%)
Query: 123 LGHLAHLRRTFHYVDELLPFRFGETDLPIDDIAAVCLLVYFGVSTLLDASSSDSQKSDD- 181
+G + R FH+V+++ R GE D P + L ++S QK +
Sbjct: 120 VGAAEPMERAFHHVEKVATLR-GE-DFP-------------SLKASLPSASVSGQKQKEG 164
Query: 182 -EQKEAELAVSDFSGDGAGILAAASTIV 208
QK+ + A DFS + G+ +S++V
Sbjct: 165 LNQKQKQAAGEDFSKEPRGVSGMSSSLV 192
>At2g31170 putative cysteinyl-tRNA synthetase
Length = 563
Score = 27.3 bits (59), Expect = 9.0
Identities = 17/56 (30%), Positives = 26/56 (46%), Gaps = 1/56 (1%)
Query: 185 EAELAVSDFSGDGAGILAAASTIVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVI 240
E+ L D + + + + T ++TF FVA D + + LAA S PL I
Sbjct: 389 ESALGEKDSTFENGSVPSDTLTSINTFRTEFVASMSD-DLLTPVTLAAMSEPLKTI 443
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.325 0.138 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,550,206
Number of Sequences: 26719
Number of extensions: 216359
Number of successful extensions: 912
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 895
Number of HSP's gapped (non-prelim): 19
length of query: 273
length of database: 11,318,596
effective HSP length: 98
effective length of query: 175
effective length of database: 8,700,134
effective search space: 1522523450
effective search space used: 1522523450
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC140913.9


