

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140849.2 + phase: 0
(553 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
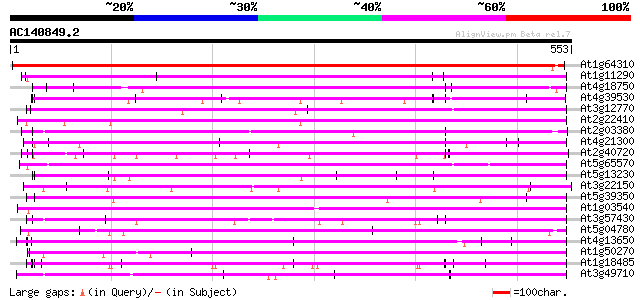
Score E
Sequences producing significant alignments: (bits) Value
At1g64310 hypothetical protein 541 e-154
At1g11290 hypothetical protein 313 2e-85
At4g18750 putative protein 309 2e-84
At4g39530 putative protein 307 9e-84
At3g12770 unknown protein 306 2e-83
At2g22410 putative protein 306 3e-83
At2g03380 unknown protein 303 2e-82
At4g21300 putative protein 298 4e-81
At2g40720 unknown protein 296 3e-80
At5g65570 putative protein 294 8e-80
At5g13230 putative protein 293 1e-79
At3g22150 hypothetical protein 286 2e-77
At5g39350 putative protein 286 2e-77
At1g03540 hypothetical protein 285 4e-77
At3g57430 putative protein 285 5e-77
At5g04780 selenium-binding protein-like 284 8e-77
At4g13650 unknown protein 283 1e-76
At1g50270 hypothetical protein 283 1e-76
At1g18485 hypothetical protein 283 1e-76
At3g49710 putative protein 283 2e-76
>At1g64310 hypothetical protein
Length = 552
Score = 541 bits (1393), Expect = e-154
Identities = 276/549 (50%), Positives = 372/549 (67%), Gaps = 6/549 (1%)
Query: 3 TQFHWLHSELTNVCKSLLRVKQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFD 62
TQ + E T ++ L ++LH+ + K+ L++DP++ATQ+ R YA N+ + A +FD
Sbjct: 5 TQLRLIIYEFTRKIQTRLNTQKLHSFVTKSKLARDPYFATQLARFYALNDDLISARKLFD 64
Query: 63 KTSTRSVFLWNSMIRAFAKARRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGM 122
RSVFLWNS+IRA+AKA +F+ +SLF +L D RPDN+TYAC R ++SFD
Sbjct: 65 VFPERSVFLWNSIIRAYAKAHQFTTVLSLFSQILRSDTRPDNFTYACLARGFSESFDTKG 124
Query: 123 LRVVHGSAVSVGLGLDPICCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGG 182
LR +HG A+ GLG D IC SA+V AYSK G++ EA ++F I +PDL LWN +I YG
Sbjct: 125 LRCIHGIAIVSGLGFDQICGSAIVKAYSKAGLIVEASKLFCSIPDPDLALWNVMILGYGC 184
Query: 183 SGMWEIGIQMFSSMRLAGKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHV 242
G W+ GI +F+ M+ G +P+ +T+ L G+ D SLL + +H K LDS +V
Sbjct: 185 CGFWDKGINLFNLMQHRGHQPNCYTMVALTSGLIDPSLLLVAWSVHAFCLKINLDSHSYV 244
Query: 243 GSLLVSMYSRCKCIDSAYRVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSK 302
G LV+MYSRC CI SA VF I PDLV S+LI+GYS+CG +++AL F +L M K
Sbjct: 245 GCALVNMYSRCMCIASACSVFNSISEPDLVACSSLITGYSRCGNHKEALHLFAELRMSGK 304
Query: 303 KLDSVLIATVLASITQMANVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTC 362
K D VL+A VL S ++++ + G E+H YV+R GLE D+KV SALIDMYSKCG L
Sbjct: 305 KPDCVLVAIVLGSCAELSDSVSGKEVHSYVIRLGLELDIKVCSALIDMYSKCGLLKCAMS 364
Query: 363 VFRIMLERNIISYNSMILAYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGL 422
+F + E+NI+S+NS+IL GLHG AS AF F E+L+ GL+PDE TFSALL CCH+GL
Sbjct: 365 LFAGIPEKNIVSFNSLILGLGLHGFASTAFEKFTEILEMGLIPDEITFSALLCTCCHSGL 424
Query: 423 VKDGRELFWRMKDEFNIKARPEHYVYMVKLLGGVGELEEAYNLTQSLPKPVDKAILGALL 482
+ G+E+F RMK EF I+ + EHYVYMVKL+G G+LEEA+ SL KP+D ILGALL
Sbjct: 425 LNKGQEIFERMKSEFGIEPQTEHYVYMVKLMGMAGKLEEAFEFVMSLQKPIDSGILGALL 484
Query: 483 SCCDSYGNSELAETVAQQIFKS-NPADNVYRVMLSNIYAGDGRWDDVKKLRD---KMVGG 538
SCC+ + N+ LAE VA+ I K+ +VY+VMLSN+YA GRWD+V++LRD + GG
Sbjct: 485 SCCEVHENTHLAEVVAENIHKNGEERRSVYKVMLSNVYARYGRWDEVERLRDGISESYGG 544
Query: 539 QKKMRGVSW 547
K+ G+SW
Sbjct: 545 --KLPGISW 551
>At1g11290 hypothetical protein
Length = 809
Score = 313 bits (801), Expect = 2e-85
Identities = 179/542 (33%), Positives = 306/542 (56%), Gaps = 4/542 (0%)
Query: 12 LTNVC--KSLLRV-KQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRS 68
L VC ++ LRV K++H L+K+ S D F T + +YA +N A VFD+ R
Sbjct: 141 LLKVCGDEAELRVGKEIHGLLVKSGFSLDLFAMTGLENMYAKCRQVNEARKVFDRMPERD 200
Query: 69 VFLWNSMIRAFAKARRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHG 128
+ WN+++ +++ A+ + ++M ++++P T + A + + + +HG
Sbjct: 201 LVSWNTIVAGYSQNGMARMALEMVKSMCEENLKPSFITIVSVLPAVSALRLISVGKEIHG 260
Query: 129 SAVSVGLGLDPICCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEI 188
A+ G +ALV Y+K G + AR++FDG++E ++V WNS+I AY + +
Sbjct: 261 YAMRSGFDSLVNISTALVDMYAKCGSLETARQLFDGMLERNVVSWNSMIDAYVQNENPKE 320
Query: 189 GIQMFSSMRLAGKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVS 248
+ +F M G KP ++ G L AD L G+ +H LS + GLD + V + L+S
Sbjct: 321 AMLIFQKMLDEGVKPTDVSVMGALHACADLGDLERGRFIHKLSVELGLDRNVSVVNSLIS 380
Query: 249 MYSRCKCIDSAYRVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVL 308
MY +CK +D+A +F + + LV+W+A+I G++Q G AL +F ++ ++ K D+
Sbjct: 381 MYCKCKEVDTAASMFGKLQSRTLVSWNAMILGFAQNGRPIDALNYFSQMRSRTVKPDTFT 440
Query: 309 IATVLASITQMANVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIML 368
+V+ +I +++ IHG V+R L+ +V V++AL+DMY+KCG + + +F +M
Sbjct: 441 YVSVITAIAELSITHHAKWIHGVVMRSCLDKNVFVTTALVDMYAKCGAIMIARLIFDMMS 500
Query: 369 ERNIISYNSMILAYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGRE 428
ER++ ++N+MI YG HG A +F+EM + + P+ TF +++SAC H+GLV+ G +
Sbjct: 501 ERHVTTWNAMIDGYGTHGFGKAALELFEEMQKGTIKPNGVTFLSVISACSHSGLVEAGLK 560
Query: 429 LFWRMKDEFNIKARPEHYVYMVKLLGGVGELEEAYNLTQSLPKPVDKAILGALLSCCDSY 488
F+ MK+ ++I+ +HY MV LLG G L EA++ +P + GA+L C +
Sbjct: 561 CFYMMKENYSIELSMDHYGAMVDLLGRAGRLNEAWDFIMQMPVKPAVNVYGAMLGACQIH 620
Query: 489 GNSELAETVAQQIFKSNPADNVYRVMLSNIYAGDGRWDDVKKLRDKMV-GGQKKMRGVSW 547
N AE A+++F+ NP D Y V+L+NIY W+ V ++R M+ G +K G S
Sbjct: 621 KNVNFAEKAAERLFELNPDDGGYHVLLANIYRAASMWEKVGQVRVSMLRQGLRKTPGCSM 680
Query: 548 IE 549
+E
Sbjct: 681 VE 682
Score = 225 bits (573), Expect = 6e-59
Identities = 116/401 (28%), Positives = 220/401 (53%)
Query: 16 CKSLLRVKQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSM 75
C SL ++Q+ + K L ++ F+ T+++ L+ ++ A VF+ ++ L+++M
Sbjct: 47 CSSLKELRQILPLVFKNGLYQEHFFQTKLVSLFCRYGSVDEAARVFEPIDSKLNVLYHTM 106
Query: 76 IRAFAKARRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGL 135
++ FAK A+ F M DD+ P Y + ++ C D + + + +HG V G
Sbjct: 107 LKGFAKVSDLDKALQFFVRMRYDDVEPVVYNFTYLLKVCGDEAELRVGKEIHGLLVKSGF 166
Query: 136 GLDPICCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSS 195
LD + L + Y+K V+EAR+VFD + E DLV WN++++ Y +GM + ++M S
Sbjct: 167 SLDLFAMTGLENMYAKCRQVNEARKVFDRMPERDLVSWNTIVAGYSQNGMARMALEMVKS 226
Query: 196 MRLAGKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKC 255
M KP T+ +L ++ L+S+G+E+HG + +SG DS ++ + LV MY++C
Sbjct: 227 MCEENLKPSFITIVSVLPAVSALRLISVGKEIHGYAMRSGFDSLVNISTALVDMYAKCGS 286
Query: 256 IDSAYRVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLAS 315
+++A ++F G+ ++V+W+++I Y Q ++A+L F+K+ + K V + L +
Sbjct: 287 LETARQLFDGMLERNVVSWNSMIDAYVQNENPKEAMLIFQKMLDEGVKPTDVSVMGALHA 346
Query: 316 ITQMANVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISY 375
+ ++ G IH + GL+ +V V ++LI MY KC + +F + R ++S+
Sbjct: 347 CADLGDLERGRFIHKLSVELGLDRNVSVVNSLISMYCKCKEVDTAASMFGKLQSRTLVSW 406
Query: 376 NSMILAYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSA 416
N+MIL + +G A F +M + + PD T+ ++++A
Sbjct: 407 NAMILGFAQNGRPIDALNYFSQMRSRTVKPDTFTYVSVITA 447
Score = 172 bits (437), Expect = 3e-43
Identities = 98/283 (34%), Positives = 161/283 (56%)
Query: 145 LVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSMRLAGKKPD 204
LVS + + G V EA RVF+ I VL+++++ + + +Q F MR +P
Sbjct: 75 LVSLFCRYGSVDEAARVFEPIDSKLNVLYHTMLKGFAKVSDLDKALQFFVRMRYDDVEPV 134
Query: 205 GFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDSAYRVFC 264
+ LL D + L +G+E+HGL KSG D + L +MY++C+ ++ A +VF
Sbjct: 135 VYNFTYLLKVCGDEAELRVGKEIHGLLVKSGFSLDLFAMTGLENMYAKCRQVNEARKVFD 194
Query: 265 GIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLASITQMANVLP 324
+ DLV+W+ +++GYSQ G + AL + + ++ K + I +VL +++ + +
Sbjct: 195 RMPERDLVSWNTIVAGYSQNGMARMALEMVKSMCEENLKPSFITIVSVLPAVSALRLISV 254
Query: 325 GCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSMILAYGL 384
G EIHGY +R G +S V +S+AL+DMY+KCG L +F MLERN++S+NSMI AY
Sbjct: 255 GKEIHGYAMRSGFDSLVNISTALVDMYAKCGSLETARQLFDGMLERNVVSWNSMIDAYVQ 314
Query: 385 HGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGR 427
+ +A +F +ML +G+ P + + L AC G ++ GR
Sbjct: 315 NENPKEAMLIFQKMLDEGVKPTDVSVMGALHACADLGDLERGR 357
>At4g18750 putative protein
Length = 871
Score = 309 bits (792), Expect = 2e-84
Identities = 178/530 (33%), Positives = 283/530 (52%), Gaps = 4/530 (0%)
Query: 23 KQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMIRAFAKA 82
+QLH +LK+ + ++ Y N ++ A VFD+ + R V WNS+I +
Sbjct: 215 EQLHGFILKSGFGERNSVGNSLVAFYLKNQRVDSARKVFDEMTERDVISWNSIINGYVSN 274
Query: 83 RRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGLGLDPICC 142
+S+F MLV I D T CADS + R VH V + C
Sbjct: 275 GLAEKGLSVFVQMLVSGIEIDLATIVSVFAGCADSRLISLGRAVHSIGVKACFSREDRFC 334
Query: 143 SALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSMRLAGKK 202
+ L+ YSK G + A+ VF + + +V + S+I+ Y G+ +++F M G
Sbjct: 335 NTLLDMYSKCGDLDSAKAVFREMSDRSVVSYTSMIAGYAREGLAGEAVKLFEEMEEEGIS 394
Query: 203 PDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDSAYRV 262
PD +T+ +L A LL G+ +H +++ L D V + L+ MY++C + A V
Sbjct: 395 PDVYTVTAVLNCCARYRLLDEGKRVHEWIKENDLGFDIFVSNALMDMYAKCGSMQEAELV 454
Query: 263 FCGIFNPDLVTWSALISGYSQ-CGEYQKALLFFRKLNMKSKKLDSVLIATVLASITQMAN 321
F + D+++W+ +I GYS+ C + LF L K D +A VL + ++
Sbjct: 455 FSEMRVKDIISWNTIIGGYSKNCYANEALSLFNLLLEEKRFSPDERTVACVLPACASLSA 514
Query: 322 VLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSMILA 381
G EIHGY++R+G SD V+++L+DMY+KCG L L +F + ++++S+ MI
Sbjct: 515 FDKGREIHGYIMRNGYFSDRHVANSLVDMYAKCGALLLAHMLFDDIASKDLVSWTVMIAG 574
Query: 382 YGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGRELFWRMKDEFNIKA 441
YG+HG +A +F++M Q G+ DE +F +LL AC H+GLV +G F M+ E I+
Sbjct: 575 YGMHGFGKEAIALFNQMRQAGIEADEISFVSLLYACSHSGLVDEGWRFFNIMRHECKIEP 634
Query: 442 RPEHYVYMVKLLGGVGELEEAYNLTQSLPKPVDKAILGALLSCCDSYGNSELAETVAQQI 501
EHY +V +L G+L +AY +++P P D I GALL C + + +LAE VA+++
Sbjct: 635 TVEHYACIVDMLARTGDLIKAYRFIENMPIPPDATIWGALLCGCRIHHDVKLAEKVAEKV 694
Query: 502 FKSNPADNVYRVMLSNIYAGDGRWDDVKKLRDKMVG--GQKKMRGVSWIE 549
F+ P + Y V+++NIYA +W+ VK+LR K +G G +K G SWIE
Sbjct: 695 FELEPENTGYYVLMANIYAEAEKWEQVKRLR-KRIGQRGLRKNPGCSWIE 743
Score = 201 bits (510), Expect = 1e-51
Identities = 121/400 (30%), Positives = 211/400 (52%), Gaps = 13/400 (3%)
Query: 37 DPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMIRAFAKARRFSNAISLFRTML 96
D +++ +Y + A VFD+ WN ++ AK+ FS +I LF+ M+
Sbjct: 128 DSNLGSKLSLMYTNCGDLKEASRVFDEVKIEKALFWNILMNELAKSGDFSGSIGLFKKMM 187
Query: 97 VDDIRPDNYTYACAIRACADSFDFGMLRVVHGS------AVSVGLGLDPICCSALVSAYS 150
+ D+YT++C S F LR VHG + G G ++LV+ Y
Sbjct: 188 SSGVEMDSYTFSCV------SKSFSSLRSVHGGEQLHGFILKSGFGERNSVGNSLVAFYL 241
Query: 151 KLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSMRLAGKKPDGFTLAG 210
K V AR+VFD + E D++ WNS+I+ Y +G+ E G+ +F M ++G + D T+
Sbjct: 242 KNQRVDSARKVFDEMTERDVISWNSIINGYVSNGLAEKGLSVFVQMLVSGIEIDLATIVS 301
Query: 211 LLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDSAYRVFCGIFNPD 270
+ G ADS L+S+G+ +H + K+ + + L+ MYS+C +DSA VF + +
Sbjct: 302 VFAGCADSRLISLGRAVHSIGVKACFSREDRFCNTLLDMYSKCGDLDSAKAVFREMSDRS 361
Query: 271 LVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLASITQMANVLPGCEIHG 330
+V+++++I+GY++ G +A+ F ++ + D + VL + + G +H
Sbjct: 362 VVSYTSMIAGYAREGLAGEAVKLFEEMEEEGISPDVYTVTAVLNCCARYRLLDEGKRVHE 421
Query: 331 YVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSMILAYGLHGCASQ 390
++ + L D+ VS+AL+DMY+KCG + VF M ++IIS+N++I Y + A++
Sbjct: 422 WIKENDLGFDIFVSNALMDMYAKCGSMQEAELVFSEMRVKDIISWNTIIGGYSKNCYANE 481
Query: 391 AFTMFDEML-QKGLVPDEGTFSALLSACCHAGLVKDGREL 429
A ++F+ +L +K PDE T + +L AC GRE+
Sbjct: 482 ALSLFNLLLEEKRFSPDERTVACVLPACASLSAFDKGREI 521
Score = 172 bits (436), Expect = 4e-43
Identities = 110/372 (29%), Positives = 184/372 (48%), Gaps = 2/372 (0%)
Query: 64 TSTRSVFLWNSMIRAFAKARRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGML 123
T RSV N+ +R F ++ NA+ L DI P T ++ CADS
Sbjct: 56 TFDRSVTDANTQLRRFCESGNLENAVKLLCVSGKWDIDPR--TLCSVLQLCADSKSLKDG 113
Query: 124 RVVHGSAVSVGLGLDPICCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGS 183
+ V G +D S L Y+ G + EA RVFD + + WN L++ S
Sbjct: 114 KEVDNFIRGNGFVIDSNLGSKLSLMYTNCGDLKEASRVFDEVKIEKALFWNILMNELAKS 173
Query: 184 GMWEIGIQMFSSMRLAGKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVG 243
G + I +F M +G + D +T + + + + G++LHG KSG VG
Sbjct: 174 GDFSGSIGLFKKMMSSGVEMDSYTFSCVSKSFSSLRSVHGGEQLHGFILKSGFGERNSVG 233
Query: 244 SLLVSMYSRCKCIDSAYRVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKK 303
+ LV+ Y + + +DSA +VF + D+++W+++I+GY G +K L F ++ + +
Sbjct: 234 NSLVAFYLKNQRVDSARKVFDEMTERDVISWNSIINGYVSNGLAEKGLSVFVQMLVSGIE 293
Query: 304 LDSVLIATVLASITQMANVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCV 363
+D I +V A + G +H ++ + + + L+DMYSKCG L V
Sbjct: 294 IDLATIVSVFAGCADSRLISLGRAVHSIGVKACFSREDRFCNTLLDMYSKCGDLDSAKAV 353
Query: 364 FRIMLERNIISYNSMILAYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLV 423
FR M +R+++SY SMI Y G A +A +F+EM ++G+ PD T +A+L+ C L+
Sbjct: 354 FREMSDRSVVSYTSMIAGYAREGLAGEAVKLFEEMEEEGISPDVYTVTAVLNCCARYRLL 413
Query: 424 KDGRELFWRMKD 435
+G+ + +K+
Sbjct: 414 DEGKRVHEWIKE 425
>At4g39530 putative protein
Length = 834
Score = 307 bits (787), Expect = 9e-84
Identities = 179/530 (33%), Positives = 280/530 (52%), Gaps = 5/530 (0%)
Query: 23 KQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMIRAFAKA 82
KQ+HA +L+ L D +I Y + AH +F+ +++ W +++ + +
Sbjct: 269 KQIHAHILRYGLEMDASLMNVLIDSYVKCGRVIAAHKLFNGMPNKNIISWTTLLSGYKQN 328
Query: 83 RRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGLGLDPICC 142
A+ LF +M ++PD Y + + +CA G VH + LG D
Sbjct: 329 ALHKEAMELFTSMSKFGLKPDMYACSSILTSCASLHALGFGTQVHAYTIKANLGNDSYVT 388
Query: 143 SALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSG-MWEI--GIQMFSSMRLA 199
++L+ Y+K + +AR+VFD D+VL+N++I Y G WE+ + +F MR
Sbjct: 389 NSLIDMYAKCDCLTDARKVFDIFAAADVVLFNAMIEGYSRLGTQWELHEALNIFRDMRFR 448
Query: 200 GKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDSA 259
+P T LL A + L + +++HGL K GL+ D GS L+ +YS C C+ +
Sbjct: 449 LIRPSLLTFVSLLRASASLTSLGLSKQIHGLMFKYGLNLDIFAGSALIDVYSNCYCLKDS 508
Query: 260 YRVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLASITQM 319
VF + DLV W+++ +GY Q E ++AL F +L + ++ D A ++ + +
Sbjct: 509 RLVFDEMKVKDLVIWNSMFAGYVQQSENEEALNLFLELQLSRERPDEFTFANMVTAAGNL 568
Query: 320 ANVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSMI 379
A+V G E H +L+ GLE + +++AL+DMY+KCG F R+++ +NS+I
Sbjct: 569 ASVQLGQEFHCQLLKRGLECNPYITNALLDMYAKCGSPEDAHKAFDSAASRDVVCWNSVI 628
Query: 380 LAYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGRELFWRMKDEFNI 439
+Y HG +A M ++M+ +G+ P+ TF +LSAC HAGLV+DG + F M F I
Sbjct: 629 SSYANHGEGKKALQMLEKMMSEGIEPNYITFVGVLSACSHAGLVEDGLKQFELML-RFGI 687
Query: 440 KARPEHYVYMVKLLGGVGELEEAYNLTQSLPKPVDKAILGALLSCCDSYGNSELAETVAQ 499
+ EHYV MV LLG G L +A L + +P + +LLS C GN ELAE A+
Sbjct: 688 EPETEHYVCMVSLLGRAGRLNKARELIEKMPTKPAAIVWRSLLSGCAKAGNVELAEHAAE 747
Query: 500 QIFKSNPADNVYRVMLSNIYAGDGRWDDVKKLRDKM-VGGQKKMRGVSWI 548
S+P D+ MLSNIYA G W + KK+R++M V G K G SWI
Sbjct: 748 MAILSDPKDSGSFTMLSNIYASKGMWTEAKKVRERMKVEGVVKEPGRSWI 797
Score = 200 bits (509), Expect = 1e-51
Identities = 127/491 (25%), Positives = 241/491 (48%), Gaps = 5/491 (1%)
Query: 22 VKQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMIRAFAK 81
V QL + L+K+ +D + T +I Y + +I+YA VFD +S W +MI K
Sbjct: 167 VFQLQSFLVKSGFDRDVYVGTLLIDFYLKDGNIDYARLVFDALPEKSTVTWTTMISGCVK 226
Query: 82 ARRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGLGLDPIC 141
R ++ LF ++ D++ PD Y + + AC+ + +H + GL +D
Sbjct: 227 MGRSYVSLQLFYQLMEDNVVPDGYILSTVLSACSILPFLEGGKQIHAHILRYGLEMDASL 286
Query: 142 CSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSMRLAGK 201
+ L+ +Y K G V A ++F+G+ +++ W +L+S Y + + + +++F+SM G
Sbjct: 287 MNVLIDSYVKCGRVIAAHKLFNGMPNKNIISWTTLLSGYKQNALHKEAMELFTSMSKFGL 346
Query: 202 KPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDSAYR 261
KPD + + +L A L G ++H + K+ L +D +V + L+ MY++C C+ A +
Sbjct: 347 KPDMYACSSILTSCASLHALGFGTQVHAYTIKANLGNDSYVTNSLIDMYAKCDCLTDARK 406
Query: 262 VFCGIFNPDLVTWSALISGYSQCG---EYQKALLFFRKLNMKSKKLDSVLIATVLASITQ 318
VF D+V ++A+I GYS+ G E +AL FR + + + + ++L +
Sbjct: 407 VFDIFAAADVVLFNAMIEGYSRLGTQWELHEALNIFRDMRFRLIRPSLLTFVSLLRASAS 466
Query: 319 MANVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSM 378
+ ++ +IHG + ++GL D+ SALID+YS C L VF M ++++ +NSM
Sbjct: 467 LTSLGLSKQIHGLMFKYGLNLDIFAGSALIDVYSNCYCLKDSRLVFDEMKVKDLVIWNSM 526
Query: 379 ILAYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGRELFWRMKDEFN 438
Y +A +F E+ PDE TF+ +++A + V+ G+E ++
Sbjct: 527 FAGYVQQSENEEALNLFLELQLSRERPDEFTFANMVTAAGNLASVQLGQEFHCQLLKR-G 585
Query: 439 IKARPEHYVYMVKLLGGVGELEEAYNLTQSLPKPVDKAILGALLSCCDSYGNSELAETVA 498
++ P ++ + G E+A+ S D +++S ++G + A +
Sbjct: 586 LECNPYITNALLDMYAKCGSPEDAHKAFDSAASR-DVVCWNSVISSYANHGEGKKALQML 644
Query: 499 QQIFKSNPADN 509
+++ N
Sbjct: 645 EKMMSEGIEPN 655
Score = 157 bits (398), Expect = 1e-38
Identities = 104/398 (26%), Positives = 195/398 (48%), Gaps = 6/398 (1%)
Query: 25 LHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMIRAFAKARR 84
+H ++ L D + + +I LY+ + YA VF+K R++ W++M+ A
Sbjct: 66 VHGQIIVWGLELDTYLSNILINLYSRAGGMVYARKVFEKMPERNLVSWSTMVSACNHHGI 125
Query: 85 FSNAISLFRTML-VDDIRPDNYTYACAIRACA--DSFDFGMLRVVHGSAVSVGLGLDPIC 141
+ ++ +F P+ Y + I+AC+ D M+ + V G D
Sbjct: 126 YEESLVVFLEFWRTRKDSPNEYILSSFIQACSGLDGRGRWMVFQLQSFLVKSGFDRDVYV 185
Query: 142 CSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSMRLAGK 201
+ L+ Y K G + AR VFD + E V W ++IS G + +Q+F +
Sbjct: 186 GTLLIDFYLKDGNIDYARLVFDALPEKSTVTWTTMISGCVKMGRSYVSLQLFYQLMEDNV 245
Query: 202 KPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDSAYR 261
PDG+ L+ +L + L G+++H + GL+ D + ++L+ Y +C + +A++
Sbjct: 246 VPDGYILSTVLSACSILPFLEGGKQIHAHILRYGLEMDASLMNVLIDSYVKCGRVIAAHK 305
Query: 262 VFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLASITQMAN 321
+F G+ N ++++W+ L+SGY Q +++A+ F ++ K D +++L S +
Sbjct: 306 LFNGMPNKNIISWTTLLSGYKQNALHKEAMELFTSMSKFGLKPDMYACSSILTSCASLHA 365
Query: 322 VLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSMILA 381
+ G ++H Y ++ L +D V+++LIDMY+KC L VF I +++ +N+MI
Sbjct: 366 LGFGTQVHAYTIKANLGNDSYVTNSLIDMYAKCDCLTDARKVFDIFAAADVVLFNAMIEG 425
Query: 382 YGLHGC---ASQAFTMFDEMLQKGLVPDEGTFSALLSA 416
Y G +A +F +M + + P TF +LL A
Sbjct: 426 YSRLGTQWELHEALNIFRDMRFRLIRPSLLTFVSLLRA 463
Score = 148 bits (373), Expect = 9e-36
Identities = 95/298 (31%), Positives = 159/298 (52%), Gaps = 7/298 (2%)
Query: 125 VVHGSAVSVGLGLDPICCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSG 184
VVHG + GL LD + L++ YS+ G + AR+VF+ + E +LV W++++SA G
Sbjct: 65 VVHGQIIVWGLELDTYLSNILINLYSRAGGMVYARKVFEKMPERNLVSWSTMVSACNHHG 124
Query: 185 MWEIGIQMFSSM-RLAGKKPDGFTLAGLLGGIADSSLLSIGQ----ELHGLSQKSGLDSD 239
++E + +F R P+ + L+ + A S L G+ +L KSG D D
Sbjct: 125 IYEESLVVFLEFWRTRKDSPNEYILSSFIQ--ACSGLDGRGRWMVFQLQSFLVKSGFDRD 182
Query: 240 CHVGSLLVSMYSRCKCIDSAYRVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNM 299
+VG+LL+ Y + ID A VF + VTW+ +ISG + G +L F +L
Sbjct: 183 VYVGTLLIDFYLKDGNIDYARLVFDALPEKSTVTWTTMISGCVKMGRSYVSLQLFYQLME 242
Query: 300 KSKKLDSVLIATVLASITQMANVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHL 359
+ D +++TVL++ + + + G +IH ++LR+GLE D + + LID Y KCG +
Sbjct: 243 DNVVPDGYILSTVLSACSILPFLEGGKQIHAHILRYGLEMDASLMNVLIDSYVKCGRVIA 302
Query: 360 GTCVFRIMLERNIISYNSMILAYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSAC 417
+F M +NIIS+ +++ Y + +A +F M + GL PD S++L++C
Sbjct: 303 AHKLFNGMPNKNIISWTTLLSGYKQNALHKEAMELFTSMSKFGLKPDMYACSSILTSC 360
Score = 85.9 bits (211), Expect = 5e-17
Identities = 60/227 (26%), Positives = 115/227 (50%), Gaps = 9/227 (3%)
Query: 209 AGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDSAYRVFCGIFN 268
A LL A LL +HG GL+ D ++ ++L+++YSR + A +VF +
Sbjct: 48 ARLLQLRASDDLLHYQNVVHGQIIVWGLELDTYLSNILINLYSRAGGMVYARKVFEKMPE 107
Query: 269 PDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLASITQMANVLPG--- 325
+LV+WS ++S + G Y+++L+ F + K + I L+S Q + L G
Sbjct: 108 RNLVSWSTMVSACNHHGIYEESLVVFLEFWRTRKDSPNEYI---LSSFIQACSGLDGRGR 164
Query: 326 ---CEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSMILAY 382
++ ++++ G + DV V + LID Y K G + VF + E++ +++ +MI
Sbjct: 165 WMVFQLQSFLVKSGFDRDVYVGTLLIDFYLKDGNIDYARLVFDALPEKSTVTWTTMISGC 224
Query: 383 GLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGREL 429
G + + +F ++++ +VPD S +LSAC ++ G+++
Sbjct: 225 VKMGRSYVSLQLFYQLMEDNVVPDGYILSTVLSACSILPFLEGGKQI 271
Score = 41.2 bits (95), Expect = 0.002
Identities = 31/123 (25%), Positives = 51/123 (41%), Gaps = 43/123 (34%)
Query: 328 IHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSMILAYGLHGC 387
+HG ++ GLE D +S+ LI++YS+ G + VF M ERN++S+
Sbjct: 66 VHGQIIVWGLELDTYLSNILINLYSRAGGMVYARKVFEKMPERNLVSW------------ 113
Query: 388 ASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGREL---FWRMKDEFNIKARPE 444
S ++SAC H G+ ++ + FWR + K P
Sbjct: 114 -----------------------STMVSACNHHGIYEESLVVFLEFWRTR-----KDSPN 145
Query: 445 HYV 447
Y+
Sbjct: 146 EYI 148
>At3g12770 unknown protein
Length = 719
Score = 306 bits (784), Expect = 2e-83
Identities = 172/532 (32%), Positives = 297/532 (55%), Gaps = 4/532 (0%)
Query: 21 RVKQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMIRAFA 80
++KQ+HA LL L F T++I + I +A VFD +F WN++IR ++
Sbjct: 36 QLKQIHARLLVLGLQFSGFLITKLIHASSSFGDITFARQVFDDLPRPQIFPWNAIIRGYS 95
Query: 81 KARRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGLGLDPI 140
+ F +A+ ++ M + + PD++T+ ++AC+ M R VH +G D
Sbjct: 96 RNNHFQDALLMYSNMQLARVSPDSFTFPHLLKACSGLSHLQMGRFVHAQVFRLGFDADVF 155
Query: 141 CCSALVSAYSKLGVVHEARRVFDGIVEPD--LVLWNSLISAYGGSGMWEIGIQMFSSMRL 198
+ L++ Y+K + AR VF+G+ P+ +V W +++SAY +G +++FS MR
Sbjct: 156 VQNGLIALYAKCRRLGSARTVFEGLPLPERTIVSWTAIVSAYAQNGEPMEALEIFSQMRK 215
Query: 199 AGKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDS 258
KPD L +L L G+ +H K GL+ + + L +MY++C + +
Sbjct: 216 MDVKPDWVALVSVLNAFTCLQDLKQGRSIHASVVKMGLEIEPDLLISLNTMYAKCGQVAT 275
Query: 259 AYRVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLASITQ 318
A +F + +P+L+ W+A+ISGY++ G ++A+ F ++ K + D++ I + +++ Q
Sbjct: 276 AKILFDKMKSPNLILWNAMISGYAKNGYAREAIDMFHEMINKDVRPDTISITSAISACAQ 335
Query: 319 MANVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSM 378
+ ++ ++ YV R DV +SSALIDM++KCG + VF L+R+++ +++M
Sbjct: 336 VGSLEQARSMYEYVGRSDYRDDVFISSALIDMFAKCGSVEGARLVFDRTLDRDVVVWSAM 395
Query: 379 ILAYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGRELFWRMKDEFN 438
I+ YGLHG A +A +++ M + G+ P++ TF LL AC H+G+V++G F RM D
Sbjct: 396 IVGYGLHGRAREAISLYRAMERGGVHPNDVTFLGLLMACNHSGMVREGWWFFNRMADH-K 454
Query: 439 IKARPEHYVYMVKLLGGVGELEEAYNLTQSLPKPVDKAILGALLSCCDSYGNSELAETVA 498
I + +HY ++ LLG G L++AY + + +P + GALLS C + + EL E A
Sbjct: 455 INPQQQHYACVIDLLGRAGHLDQAYEVIKCMPVQPGVTVWGALLSACKKHRHVELGEYAA 514
Query: 499 QQIFKSNPADNVYRVMLSNIYAGDGRWDDVKKLRDKM-VGGQKKMRGVSWIE 549
QQ+F +P++ + V LSN+YA WD V ++R +M G K G SW+E
Sbjct: 515 QQLFSIDPSNTGHYVQLSNLYAAARLWDRVAEVRVRMKEKGLNKDVGCSWVE 566
Score = 116 bits (291), Expect = 3e-26
Identities = 78/283 (27%), Positives = 127/283 (44%), Gaps = 6/283 (2%)
Query: 17 KSLLRVKQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMI 76
+ L + + +HA ++K L +P + +YA + A +FDK + ++ LWN+MI
Sbjct: 236 QDLKQGRSIHASVVKMGLEIEPDLLISLNTMYAKCGQVATAKILFDKMKSPNLILWNAMI 295
Query: 77 RAFAKARRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGLG 136
+AK AI +F M+ D+RPD + AI ACA R ++
Sbjct: 296 SGYAKNGYAREAIDMFHEMINKDVRPDTISITSAISACAQVGSLEQARSMYEYVGRSDYR 355
Query: 137 LDPICCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSM 196
D SAL+ ++K G V AR VFD ++ D+V+W+++I YG G I ++ +M
Sbjct: 356 DDVFISSALIDMFAKCGSVEGARLVFDRTLDRDVVVWSAMIVGYGLHGRAREAISLYRAM 415
Query: 197 RLAGKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCI 256
G P+ T GLL S ++ G ++ + ++ + R +
Sbjct: 416 ERGGVHPNDVTFLGLLMACNHSGMVREGWWFFNRMADHKINPQQQHYACVIDLLGRAGHL 475
Query: 257 DSAYRVF-CGIFNPDLVTWSALISG-----YSQCGEYQKALLF 293
D AY V C P + W AL+S + + GEY LF
Sbjct: 476 DQAYEVIKCMPVQPGVTVWGALLSACKKHRHVELGEYAAQQLF 518
>At2g22410 putative protein
Length = 681
Score = 306 bits (783), Expect = 3e-83
Identities = 185/582 (31%), Positives = 305/582 (51%), Gaps = 40/582 (6%)
Query: 8 LHSELTNV---CKSLLRVKQLHACLLKTHLSKDPFYATQIIRLYAFNN--HINYAHHVFD 62
LH+ L ++ CK LL +KQ+ A ++ L DPF ++++I A + +++Y+ +
Sbjct: 52 LHNPLLSLLEKCKLLLHLKQIQAQMIINGLILDPFASSRLIAFCALSESRYLDYSVKILK 111
Query: 63 KTSTRSVFLWNSMIRAFAKARRFSNAISLFRTMLVD---DIRPDNYTYACAIRACADSFD 119
++F WN IR F+++ + L++ ML + RPD++TY + CAD
Sbjct: 112 GIENPNIFSWNVTIRGFSESENPKESFLLYKQMLRHGCCESRPDHFTYPVLFKVCADLRL 171
Query: 120 FGMLRVVHGSAVSVGLGLDPICCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISA 179
+ ++ G + + L L +A + ++ G + AR+VFD DLV WN LI+
Sbjct: 172 SSLGHMILGHVLKLRLELVSHVHNASIHMFASCGDMENARKVFDESPVRDLVSWNCLING 231
Query: 180 YGGSGMWEIGIQMFSSMRLAGKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSD 239
Y G E I ++ M G KPD T+ GL+ + L+ G+E + +++GL
Sbjct: 232 YKKIGEAEKAIYVYKLMESEGVKPDDVTMIGLVSSCSMLGDLNRGKEFYEYVKENGLRMT 291
Query: 240 CHVGSLLVSMYSRCKCIDSAYRVFCGIFNPDLVTWSALISGYSQCG-------------- 285
+ + L+ M+S+C I A R+F + +V+W+ +ISGY++CG
Sbjct: 292 IPLVNALMDMFSKCGDIHEARRIFDNLEKRTIVSWTTMISGYARCGLLDVSRKLFDDMEE 351
Query: 286 -----------------EYQKALLFFRKLNMKSKKLDSVLIATVLASITQMANVLPGCEI 328
Q AL F+++ + K D + + L++ +Q+ + G I
Sbjct: 352 KDVVLWNAMIGGSVQAKRGQDALALFQEMQTSNTKPDEITMIHCLSACSQLGALDVGIWI 411
Query: 329 HGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSMILAYGLHGCA 388
H Y+ ++ L +V + ++L+DMY+KCG + VF + RN ++Y ++I LHG A
Sbjct: 412 HRYIEKYSLSLNVALGTSLVDMYAKCGNISEALSVFHGIQTRNSLTYTAIIGGLALHGDA 471
Query: 389 SQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGRELFWRMKDEFNIKARPEHYVY 448
S A + F+EM+ G+ PDE TF LLSACCH G+++ GR+ F +MK FN+ + +HY
Sbjct: 472 STAISYFNEMIDAGIAPDEITFIGLLSACCHGGMIQTGRDYFSQMKSRFNLNPQLKHYSI 531
Query: 449 MVKLLGGVGELEEAYNLTQSLPKPVDKAILGALLSCCDSYGNSELAETVAQQIFKSNPAD 508
MV LLG G LEEA L +S+P D A+ GALL C +GN EL E A+++ + +P+D
Sbjct: 532 MVDLLGRAGLLEEADRLMESMPMEADAAVWGALLFGCRMHGNVELGEKAAKKLLELDPSD 591
Query: 509 NVYRVMLSNIYAGDGRWDDVKKLRDKM-VGGQKKMRGVSWIE 549
+ V+L +Y W+D K+ R M G +K+ G S IE
Sbjct: 592 SGIYVLLDGMYGEANMWEDAKRARRMMNERGVEKIPGCSSIE 633
>At2g03380 unknown protein
Length = 689
Score = 303 bits (775), Expect = 2e-82
Identities = 176/530 (33%), Positives = 282/530 (53%), Gaps = 9/530 (1%)
Query: 23 KQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMIRAFAKA 82
K++H L+K S D T ++ +YA I AH VF+ + R+V W SMI + K
Sbjct: 162 KKIHCQLVKVP-SFDNVVLTGLLDMYAKCGEIKSAHKVFNDITLRNVVCWTSMIAGYVKN 220
Query: 83 RRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGLGLDPICC 142
+ LF M +++ + YTY I AC + HG V G+ L
Sbjct: 221 DLCEEGLVLFNRMRENNVLGNEYTYGTLIMACTKLSALHQGKWFHGCLVKSGIELSSCLV 280
Query: 143 SALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSMRLAGKK 202
++L+ Y K G + ARRVF+ DLV+W ++I Y +G + +F M+ K
Sbjct: 281 TSLLDMYVKCGDISNARRVFNEHSHVDLVMWTAMIVGYTHNGSVNEALSLFQKMKGVEIK 340
Query: 203 PDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDSAYRV 262
P+ T+A +L G L +G+ +HGLS K G+ D +V + LV MY++C A V
Sbjct: 341 PNCVTIASVLSGCGLIENLELGRSVHGLSIKVGI-WDTNVANALVHMYAKCYQNRDAKYV 399
Query: 263 FCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLASITQMANV 322
F D+V W+++ISG+SQ G +AL F ++N +S + V +A++ ++ + ++
Sbjct: 400 FEMESEKDIVAWNSIISGFSQNGSIHEALFLFHRMNSESVTPNGVTVASLFSACASLGSL 459
Query: 323 LPGCEIHGYVLRHGL--ESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSMIL 380
G +H Y ++ G S V V +AL+D Y+KCG +F + E+N I++++MI
Sbjct: 460 AVGSSLHAYSVKLGFLASSSVHVGTALLDFYAKCGDPQSARLIFDTIEEKNTITWSAMIG 519
Query: 381 AYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGRELFWRMKDEFNIK 440
YG G + +F+EML+K P+E TF+++LSAC H G+V +G++ F M ++N
Sbjct: 520 GYGKQGDTIGSLELFEEMLKKQQKPNESTFTSILSACGHTGMVNEGKKYFSSMYKDYNFT 579
Query: 441 ARPEHYVYMVKLLGGVGELEEAYNLTQSLPKPVDKAILGALLSCCDSYGNSELAETVAQQ 500
+HY MV +L GELE+A ++ + +P D GA L C + +L E V ++
Sbjct: 580 PSTKHYTCMVDMLARAGELEQALDIIEKMPIQPDVRCFGAFLHGCGMHSRFDLGEIVIKK 639
Query: 501 IFKSNPADNVYRVMLSNIYAGDGRWDDVKKLRDKMVGGQKKMRGVSWIEG 550
+ +P D Y V++SN+YA DGRW+ K++R+ M K RG+S I G
Sbjct: 640 MLDLHPDDASYYVLVSNLYASDGRWNQAKEVRNLM-----KQRGLSKIAG 684
Score = 193 bits (491), Expect = 2e-49
Identities = 125/419 (29%), Positives = 212/419 (49%), Gaps = 4/419 (0%)
Query: 12 LTNVCKSLLRVKQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFL 71
L + C ++ ++Q H L L D AT+++ LY F + A VFD+ +L
Sbjct: 50 LLSKCTNIDSLRQSHGVLTGNGLMGDISIATKLVSLYGFFGYTKDARLVFDQIPEPDFYL 109
Query: 72 WNSMIRAFAKARRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAV 131
W M+R + + + L+ ++ R D+ ++ A++AC + D + +H V
Sbjct: 110 WKVMLRCYCLNKESVEVVKLYDLLMKHGFRYDDIVFSKALKACTELQDLDNGKKIHCQLV 169
Query: 132 SVGLGLDPICCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQ 191
V D + + L+ Y+K G + A +VF+ I ++V W S+I+ Y + + E G+
Sbjct: 170 KVP-SFDNVVLTGLLDMYAKCGEIKSAHKVFNDITLRNVVCWTSMIAGYVKNDLCEEGLV 228
Query: 192 MFSSMRLAGKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLD-SDCHVGSLLVSMY 250
+F+ MR + +T L+ S L G+ HG KSG++ S C V SLL MY
Sbjct: 229 LFNRMRENNVLGNEYTYGTLIMACTKLSALHQGKWFHGCLVKSGIELSSCLVTSLL-DMY 287
Query: 251 SRCKCIDSAYRVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIA 310
+C I +A RVF + DLV W+A+I GY+ G +AL F+K+ K + V IA
Sbjct: 288 VKCGDISNARRVFNEHSHVDLVMWTAMIVGYTHNGSVNEALSLFQKMKGVEIKPNCVTIA 347
Query: 311 TVLASITQMANVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLER 370
+VL+ + N+ G +HG ++ G+ D V++AL+ MY+KC VF + E+
Sbjct: 348 SVLSGCGLIENLELGRSVHGLSIKVGI-WDTNVANALVHMYAKCYQNRDAKYVFEMESEK 406
Query: 371 NIISYNSMILAYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGREL 429
+I+++NS+I + +G +A +F M + + P+ T ++L SAC G + G L
Sbjct: 407 DIVAWNSIISGFSQNGSIHEALFLFHRMNSESVTPNGVTVASLFSACASLGSLAVGSSL 465
>At4g21300 putative protein
Length = 857
Score = 298 bits (764), Expect = 4e-81
Identities = 167/541 (30%), Positives = 288/541 (52%), Gaps = 5/541 (0%)
Query: 14 NVCKSLLRVK---QLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVF 70
+VC S L + QLH ++ + + + ++ +Y+ + A +F S
Sbjct: 247 SVCASKLLIDLGVQLHGLVVVSGVDFEGSIKNSLLSMYSKCGRFDDASKLFRMMSRADTV 306
Query: 71 LWNSMIRAFAKARRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSA 130
WN MI + ++ +++ F M+ + PD T++ + + + + + +H
Sbjct: 307 TWNCMISGYVQSGLMEESLTFFYEMISSGVLPDAITFSSLLPSVSKFENLEYCKQIHCYI 366
Query: 131 VSVGLGLDPICCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGI 190
+ + LD SAL+ AY K V A+ +F D+V++ ++IS Y +G++ +
Sbjct: 367 MRHSISLDIFLTSALIDAYFKCRGVSMAQNIFSQCNSVDVVVFTAMISGYLHNGLYIDSL 426
Query: 191 QMFSSMRLAGKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMY 250
+MF + P+ TL +L I L +G+ELHG K G D+ C++G ++ MY
Sbjct: 427 EMFRWLVKVKISPNEITLVSILPVIGILLALKLGRELHGFIIKKGFDNRCNIGCAVIDMY 486
Query: 251 SRCKCIDSAYRVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIA 310
++C ++ AY +F + D+V+W+++I+ +Q A+ FR++ + D V I+
Sbjct: 487 AKCGRMNLAYEIFERLSKRDIVSWNSMITRCAQSDNPSAAIDIFRQMGVSGICYDCVSIS 546
Query: 311 TVLASITQMANVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLER 370
L++ + + G IHG++++H L SDV S LIDMY+KCG L VF+ M E+
Sbjct: 547 AALSACANLPSESFGKAIHGFMIKHSLASDVYSESTLIDMYAKCGNLKAAMNVFKTMKEK 606
Query: 371 NIISYNSMILAYGLHGCASQAFTMFDEMLQK-GLVPDEGTFSALLSACCHAGLVKDGREL 429
NI+S+NS+I A G HG + +F EM++K G+ PD+ TF ++S+CCH G V +G
Sbjct: 607 NIVSWNSIIAACGNHGKLKDSLCLFHEMVEKSGIRPDQITFLEIISSCCHVGDVDEGVRF 666
Query: 430 FWRMKDEFNIKARPEHYVYMVKLLGGVGELEEAYNLTQSLPKPVDKAILGALLSCCDSYG 489
F M +++ I+ + EHY +V L G G L EAY +S+P P D + G LL C +
Sbjct: 667 FRSMTEDYGIQPQQEHYACVVDLFGRAGRLTEAYETVKSMPFPPDAGVWGTLLGACRLHK 726
Query: 490 NSELAETVAQQIFKSNPADNVYRVMLSNIYAGDGRWDDVKKLRDKMVGGQ-KKMRGVSWI 548
N ELAE + ++ +P+++ Y V++SN +A W+ V K+R M + +K+ G SWI
Sbjct: 727 NVELAEVASSKLMDLDPSNSGYYVLISNAHANAREWESVTKVRSLMKEREVQKIPGYSWI 786
Query: 549 E 549
E
Sbjct: 787 E 787
Score = 202 bits (515), Expect = 3e-52
Identities = 130/482 (26%), Positives = 246/482 (50%), Gaps = 6/482 (1%)
Query: 23 KQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTR--SVFLWNSMIRAFA 80
KQ+HA L+ +S D + +I+ +YA + +F + R S+ WNS+I +F
Sbjct: 55 KQVHAFLIVNSISGDSYTDERILGMYAMCGSFSDCGKMFYRLDLRRSSIRPWNSIISSFV 114
Query: 81 KARRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGLGLDPI 140
+ + A++ + ML + PD T+ C ++AC +F + + + S+G+ +
Sbjct: 115 RNGLLNQALAFYFKMLCFGVSPDVSTFPCLVKACVALKNFKGIDFLSDTVSSLGMDCNEF 174
Query: 141 CCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSMRLAG 200
S+L+ AY + G + ++FD +++ D V+WN +++ Y G + I+ FS MR+
Sbjct: 175 VASSLIKAYLEYGKIDVPSKLFDRVLQKDCVIWNVMLNGYAKCGALDSVIKGFSVMRMDQ 234
Query: 201 KKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDSAY 260
P+ T +L A L+ +G +LHGL SG+D + + + L+SMYS+C D A
Sbjct: 235 ISPNAVTFDCVLSVCASKLLIDLGVQLHGLVVVSGVDFEGSIKNSLLSMYSKCGRFDDAS 294
Query: 261 RVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLASITQMA 320
++F + D VTW+ +ISGY Q G +++L FF ++ D++ +++L S+++
Sbjct: 295 KLFRMMSRADTVTWNCMISGYVQSGLMEESLTFFYEMISSGVLPDAITFSSLLPSVSKFE 354
Query: 321 NVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSMIL 380
N+ +IH Y++RH + D+ ++SALID Y KC + + +F +++ + +MI
Sbjct: 355 NLEYCKQIHCYIMRHSISLDIFLTSALIDAYFKCRGVSMAQNIFSQCNSVDVVVFTAMIS 414
Query: 381 AYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGRELF-WRMKDEFNI 439
Y +G + MF +++ + P+E T ++L +K GREL + +K F+
Sbjct: 415 GYLHNGLYIDSLEMFRWLVKVKISPNEITLVSILPVIGILLALKLGRELHGFIIKKGFDN 474
Query: 440 KARPEHYVYMVKLLGGVGELEEAYNLTQSLPKPVDKAILGALLSCCDSYGNSELAETVAQ 499
+ V + + G + AY + + L K D ++++ C N A + +
Sbjct: 475 RCNIGCAV--IDMYAKCGRMNLAYEIFERLSKR-DIVSWNSMITRCAQSDNPSAAIDIFR 531
Query: 500 QI 501
Q+
Sbjct: 532 QM 533
Score = 194 bits (494), Expect = 8e-50
Identities = 124/455 (27%), Positives = 220/455 (48%), Gaps = 10/455 (2%)
Query: 39 FYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMIRAFAKARRFSNAISLFRTMLVD 98
F A+ +I+ Y I+ +FD+ + +WN M+ +AK + I F M +D
Sbjct: 174 FVASSLIKAYLEYGKIDVPSKLFDRVLQKDCVIWNVMLNGYAKCGALDSVIKGFSVMRMD 233
Query: 99 DIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGLGLDPICCSALVSAYSKLGVVHEA 158
I P+ T+ C + CA + +HG V G+ + ++L+S YSK G +A
Sbjct: 234 QISPNAVTFDCVLSVCASKLLIDLGVQLHGLVVVSGVDFEGSIKNSLLSMYSKCGRFDDA 293
Query: 159 RRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSMRLAGKKPDGFTLAGLLGGIADS 218
++F + D V WN +IS Y SG+ E + F M +G PD T + LL ++
Sbjct: 294 SKLFRMMSRADTVTWNCMISGYVQSGLMEESLTFFYEMISSGVLPDAITFSSLLPSVSKF 353
Query: 219 SLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDSAYRVFCGIFNPDLVTWSALI 278
L +++H + + D + S L+ Y +C+ + A +F + D+V ++A+I
Sbjct: 354 ENLEYCKQIHCYIMRHSISLDIFLTSALIDAYFKCRGVSMAQNIFSQCNSVDVVVFTAMI 413
Query: 279 SGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLASITQMANVLPGCEIHGYVLRHGLE 338
SGY G Y +L FR L + + + ++L I + + G E+HG++++ G +
Sbjct: 414 SGYLHNGLYIDSLEMFRWLVKVKISPNEITLVSILPVIGILLALKLGRELHGFIIKKGFD 473
Query: 339 SDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSMILAYGLHGCASQAFTMFDEM 398
+ + A+IDMY+KCG ++L +F + +R+I+S+NSMI S A +F +M
Sbjct: 474 NRCNIGCAVIDMYAKCGRMNLAYEIFERLSKRDIVSWNSMITRCAQSDNPSAAIDIFRQM 533
Query: 399 LQKGLVPDEGTFSALLSACCHAGLVKDGRELFWRMKDEFNIKARPEHYVY----MVKLLG 454
G+ D + SA LSAC + E F + F IK VY ++ +
Sbjct: 534 GVSGICYDCVSISAALSACANL-----PSESFGKAIHGFMIKHSLASDVYSESTLIDMYA 588
Query: 455 GVGELEEAYNLTQSLPKPVDKAILGALLSCCDSYG 489
G L+ A N+ +++ K + ++++ C ++G
Sbjct: 589 KCGNLKAAMNVFKTM-KEKNIVSWNSIIAACGNHG 622
Score = 62.8 bits (151), Expect = 5e-10
Identities = 47/224 (20%), Positives = 99/224 (43%), Gaps = 2/224 (0%)
Query: 208 LAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDSAYRVF--CG 265
L+ LL ++ +LL G+++H + + D + ++ MY+ C ++F
Sbjct: 38 LSLLLQACSNPNLLRQGKQVHAFLIVNSISGDSYTDERILGMYAMCGSFSDCGKMFYRLD 97
Query: 266 IFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLASITQMANVLPG 325
+ + W+++IS + + G +AL F+ K+ D ++ + + N
Sbjct: 98 LRRSSIRPWNSIISSFVRNGLLNQALAFYFKMLCFGVSPDVSTFPCLVKACVALKNFKGI 157
Query: 326 CEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSMILAYGLH 385
+ V G++ + V+S+LI Y + G + + + +F +L+++ + +N M+ Y
Sbjct: 158 DFLSDTVSSLGMDCNEFVASSLIKAYLEYGKIDVPSKLFDRVLQKDCVIWNVMLNGYAKC 217
Query: 386 GCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGREL 429
G F M + P+ TF +LS C L+ G +L
Sbjct: 218 GALDSVIKGFSVMRMDQISPNAVTFDCVLSVCASKLLIDLGVQL 261
>At2g40720 unknown protein
Length = 860
Score = 296 bits (757), Expect = 3e-80
Identities = 177/532 (33%), Positives = 283/532 (52%), Gaps = 3/532 (0%)
Query: 23 KQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMIRAFAKA 82
+Q+H ++K L DP+ T ++ +Y+ + A VF + + +WN+M+ A+A+
Sbjct: 292 RQIHCDVVKMGLHNDPYVCTSLLSMYSKCGMVGEAETVFSCVVDKRLEIWNAMVAAYAEN 351
Query: 83 RRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGLGLDPICC 142
+A+ LF M + PD++T + I C+ + + VH +
Sbjct: 352 DYGYSALDLFGFMRQKSVLPDSFTLSNVISCCSVLGLYNYGKSVHAELFKRPIQSTSTIE 411
Query: 143 SALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSMRLAGK- 201
SAL++ YSK G +A VF + E D+V W SLIS +G ++ +++F M+
Sbjct: 412 SALLTLYSKCGCDPDAYLVFKSMEEKDMVAWGSLISGLCKNGKFKEALKVFGDMKDDDDS 471
Query: 202 -KPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDSAY 260
KPD + + A L G ++HG K+GL + VGS L+ +YS+C + A
Sbjct: 472 LKPDSDIMTSVTNACAGLEALRFGLQVHGSMIKTGLVLNVFVGSSLIDLYSKCGLPEMAL 531
Query: 261 RVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLASITQMA 320
+VF + ++V W+++IS YS+ + ++ F + + DSV I +VL +I+ A
Sbjct: 532 KVFTSMSTENMVAWNSMISCYSRNNLPELSIDLFNLMLSQGIFPDSVSITSVLVAISSTA 591
Query: 321 NVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSMIL 380
++L G +HGY LR G+ SD + +ALIDMY KCGF +F+ M +++I++N MI
Sbjct: 592 SLLKGKSLHGYTLRLGIPSDTHLKNALIDMYVKCGFSKYAENIFKKMQHKSLITWNLMIY 651
Query: 381 AYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGRELFWRMKDEFNIK 440
YG HG A ++FDEM + G PD+ TF +L+SAC H+G V++G+ +F MK ++ I+
Sbjct: 652 GYGSHGDCITALSLFDEMKKAGESPDDVTFLSLISACNHSGFVEEGKNIFEFMKQDYGIE 711
Query: 441 ARPEHYVYMVKLLGGVGELEEAYNLTQSLPKPVDKAILGALLSCCDSYGNSELAETVAQQ 500
EHY MV LLG G LEEAY+ +++P D +I LLS ++ N EL A++
Sbjct: 712 PNMEHYANMVDLLGRAGLLEEAYSFIKAMPIEADSSIWLCLLSASRTHHNVELGILSAEK 771
Query: 501 IFKSNPADNVYRVMLSNIYAGDGRWDDVKKLRDKM-VGGQKKMRGVSWIEGS 551
+ + P V L N+Y G ++ KL M G K G SWIE S
Sbjct: 772 LLRMEPERGSTYVQLINLYMEAGLKNEAAKLLGLMKEKGLHKQPGCSWIEVS 823
Score = 212 bits (539), Expect = 5e-55
Identities = 129/411 (31%), Positives = 220/411 (53%), Gaps = 5/411 (1%)
Query: 23 KQLHACLLKTHLSKDPFYATQIIRLY-AFNNHINYAHHVFDKTSTRS-VFLWNSMIRAFA 80
KQ+H +L+ L D F T +I +Y F I+ A VF + +S V LWN MI F
Sbjct: 190 KQIHGFMLRNSLDTDSFLKTALIDMYFKFGLSID-AWRVFVEIEDKSNVVLWNVMIVGFG 248
Query: 81 KARRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGLGLDPI 140
+ +++ L+ + ++ + ++ A+ AC+ S + G R +H V +GL DP
Sbjct: 249 GSGICESSLDLYMLAKNNSVKLVSTSFTGALGACSQSENSGFGRQIHCDVVKMGLHNDPY 308
Query: 141 CCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSMRLAG 200
C++L+S YSK G+V EA VF +V+ L +WN++++AY + + +F MR
Sbjct: 309 VCTSLLSMYSKCGMVGEAETVFSCVVDKRLEIWNAMVAAYAENDYGYSALDLFGFMRQKS 368
Query: 201 KKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDSAY 260
PD FTL+ ++ + L + G+ +H K + S + S L+++YS+C C AY
Sbjct: 369 VLPDSFTLSNVISCCSVLGLYNYGKSVHAELFKRPIQSTSTIESALLTLYSKCGCDPDAY 428
Query: 261 RVFCGIFNPDLVTWSALISGYSQCGEYQKALLFF--RKLNMKSKKLDSVLIATVLASITQ 318
VF + D+V W +LISG + G++++AL F K + S K DS ++ +V +
Sbjct: 429 LVFKSMEEKDMVAWGSLISGLCKNGKFKEALKVFGDMKDDDDSLKPDSDIMTSVTNACAG 488
Query: 319 MANVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSM 378
+ + G ++HG +++ GL +V V S+LID+YSKCG + VF M N++++NSM
Sbjct: 489 LEALRFGLQVHGSMIKTGLVLNVFVGSSLIDLYSKCGLPEMALKVFTSMSTENMVAWNSM 548
Query: 379 ILAYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGREL 429
I Y + + +F+ ML +G+ PD + +++L A + G+ L
Sbjct: 549 ISCYSRNNLPELSIDLFNLMLSQGIFPDSVSITSVLVAISSTASLLKGKSL 599
Score = 184 bits (467), Expect = 1e-46
Identities = 122/437 (27%), Positives = 212/437 (47%), Gaps = 15/437 (3%)
Query: 12 LTNVCKSLLRV---KQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDK----- 63
L C +L + K +H ++ DPF AT ++ +Y ++YA VFD
Sbjct: 66 LLKACSALTNLSYGKTIHGSVVVLGWRYDPFIATSLVNMYVKCGFLDYAVQVFDGWSQSQ 125
Query: 64 --TSTRSVFLWNSMIRAFAKARRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFG 121
S R V +WNSMI + K RRF + FR MLV +RPD ++ + + +F
Sbjct: 126 SGVSARDVTVWNSMIDGYFKFRRFKEGVGCFRRMLVFGVRPDAFSLSIVVSVMCKEGNFR 185
Query: 122 ML--RVVHGSAVSVGLGLDPICCSALVSAYSKLGVVHEARRVFDGIVE-PDLVLWNSLIS 178
+ +HG + L D +AL+ Y K G+ +A RVF I + ++VLWN +I
Sbjct: 186 REEGKQIHGFMLRNSLDTDSFLKTALIDMYFKFGLSIDAWRVFVEIEDKSNVVLWNVMIV 245
Query: 179 AYGGSGMWEIGIQMFSSMRLAGKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDS 238
+GGSG+ E + ++ + K + G LG + S G+++H K GL +
Sbjct: 246 GFGGSGICESSLDLYMLAKNNSVKLVSTSFTGALGACSQSENSGFGRQIHCDVVKMGLHN 305
Query: 239 DCHVGSLLVSMYSRCKCIDSAYRVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLN 298
D +V + L+SMYS+C + A VF + + L W+A+++ Y++ AL F +
Sbjct: 306 DPYVCTSLLSMYSKCGMVGEAETVFSCVVDKRLEIWNAMVAAYAENDYGYSALDLFGFMR 365
Query: 299 MKSKKLDSVLIATVLASITQMANVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLH 358
KS DS ++ V++ + + G +H + + ++S + SAL+ +YSKCG
Sbjct: 366 QKSVLPDSFTLSNVISCCSVLGLYNYGKSVHAELFKRPIQSTSTIESALLTLYSKCGCDP 425
Query: 359 LGTCVFRIMLERNIISYNSMILAYGLHGCASQAFTMFDEML--QKGLVPDEGTFSALLSA 416
VF+ M E++++++ S+I +G +A +F +M L PD +++ +A
Sbjct: 426 DAYLVFKSMEEKDMVAWGSLISGLCKNGKFKEALKVFGDMKDDDDSLKPDSDIMTSVTNA 485
Query: 417 CCHAGLVKDGRELFWRM 433
C ++ G ++ M
Sbjct: 486 CAGLEALRFGLQVHGSM 502
Score = 157 bits (398), Expect = 1e-38
Identities = 109/377 (28%), Positives = 186/377 (48%), Gaps = 19/377 (5%)
Query: 73 NSMIRAFAKARRFSNAISLFRTMLVDDIRP---DNYTYACAIRACADSFDFGMLRVVHGS 129
NS IRA + + A+ L+ D P +T+ ++AC+ + + +HGS
Sbjct: 28 NSGIRALIQKGEYLQALHLYSKH--DGSSPFWTSVFTFPSLLKACSALTNLSYGKTIHGS 85
Query: 130 AVSVGLGLDPICCSALVSAYSKLGVVHEARRVFD-------GIVEPDLVLWNSLISAYGG 182
V +G DP ++LV+ Y K G + A +VFD G+ D+ +WNS+I Y
Sbjct: 86 VVVLGWRYDPFIATSLVNMYVKCGFLDYAVQVFDGWSQSQSGVSARDVTVWNSMIDGYFK 145
Query: 183 SGMWEIGIQMFSSMRLAGKKPDGFTLAGLLGGIADSSLL--SIGQELHGLSQKSGLDSDC 240
++ G+ F M + G +PD F+L+ ++ + G+++HG ++ LD+D
Sbjct: 146 FRRFKEGVGCFRRMLVFGVRPDAFSLSIVVSVMCKEGNFRREEGKQIHGFMLRNSLDTDS 205
Query: 241 HVGSLLVSMYSRCKCIDSAYRVFCGIFN-PDLVTWSALISGYSQCGEYQKALLFFRKLNM 299
+ + L+ MY + A+RVF I + ++V W+ +I G+ G + +L +
Sbjct: 206 FLKTALIDMYFKFGLSIDAWRVFVEIEDKSNVVLWNVMIVGFGGSGICESSLDLYMLAKN 265
Query: 300 KSKKLDSVLIATVLASITQMANVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHL 359
S KL S L + +Q N G +IH V++ GL +D V ++L+ MYSKCG +
Sbjct: 266 NSVKLVSTSFTGALGACSQSENSGFGRQIHCDVVKMGLHNDPYVCTSLLSMYSKCGMVGE 325
Query: 360 GTCVFRIMLERNIISYNSMILAYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCH 419
VF ++++ + +N+M+ AY + A +F M QK ++PD T S ++S C
Sbjct: 326 AETVFSCVVDKRLEIWNAMVAAYAENDYGYSALDLFGFMRQKSVLPDSFTLSNVISCCSV 385
Query: 420 AGLVKDGR----ELFWR 432
GL G+ ELF R
Sbjct: 386 LGLYNYGKSVHAELFKR 402
Score = 63.5 bits (153), Expect = 3e-10
Identities = 58/222 (26%), Positives = 95/222 (42%), Gaps = 5/222 (2%)
Query: 18 SLLRVKQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMIR 77
SLL+ K LH L+ + D +I +Y YA ++F K +S+ WN MI
Sbjct: 592 SLLKGKSLHGYTLRLGIPSDTHLKNALIDMYVKCGFSKYAENIFKKMQHKSLITWNLMIY 651
Query: 78 AFAKARRFSNAISLFRTMLVDDIRPDNYTYACAIRACADS-FDFGMLRVVHGSAVSVGLG 136
+ A+SLF M PD+ T+ I AC S F + G+
Sbjct: 652 GYGSHGDCITALSLFDEMKKAGESPDDVTFLSLISACNHSGFVEEGKNIFEFMKQDYGIE 711
Query: 137 LDPICCSALVSAYSKLGVVHEARRVFDGI-VEPDLVLWNSLISAYGGSGMWEIGIQMFSS 195
+ + +V + G++ EA + +E D +W L+SA E+GI S+
Sbjct: 712 PNMEHYANMVDLLGRAGLLEEAYSFIKAMPIEADSSIWLCLLSASRTHHNVELGI--LSA 769
Query: 196 MRLAGKKPD-GFTLAGLLGGIADSSLLSIGQELHGLSQKSGL 236
+L +P+ G T L+ ++ L + +L GL ++ GL
Sbjct: 770 EKLLRMEPERGSTYVQLINLYMEAGLKNEAAKLLGLMKEKGL 811
>At5g65570 putative protein
Length = 738
Score = 294 bits (753), Expect = 8e-80
Identities = 183/545 (33%), Positives = 293/545 (53%), Gaps = 8/545 (1%)
Query: 10 SELTNVC---KSLLRVKQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTST 66
S+L C +S+ +K + A +LK+ + ++++ I+YA VFD S
Sbjct: 69 SQLLRQCIDERSISGIKTIQAHMLKSGFPAE-ISGSKLVDASLKCGDIDYARQVFDGMSE 127
Query: 67 RSVFLWNSMIRAFAKARRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVV 126
R + WNS+I K RR A+ ++R M+ +++ PD YT + +A +D +
Sbjct: 128 RHIVTWNSLIAYLIKHRRSKEAVEMYRLMITNNVLPDEYTLSSVFKAFSDLSLEKEAQRS 187
Query: 127 HGSAVSVGLGLDPICC-SALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGM 185
HG AV +GL + + SALV Y K G EA+ V D + E D+VL +LI Y G
Sbjct: 188 HGLAVILGLEVSNVFVGSALVDMYVKFGKTREAKLVLDRVEEKDVVLITALIVGYSQKGE 247
Query: 186 WEIGIQMFSSMRLAGKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSL 245
++ F SM + +P+ +T A +L + + G+ +HGL KSG +S +
Sbjct: 248 DTEAVKAFQSMLVEKVQPNEYTYASVLISCGNLKDIGNGKLIHGLMVKSGFESALASQTS 307
Query: 246 LVSMYSRCKCIDSAYRVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLD 305
L++MY RC +D + RVF I P+ V+W++LISG Q G + AL+ FRK+ S K +
Sbjct: 308 LLTMYLRCSLVDDSLRVFKCIEYPNQVSWTSLISGLVQNGREEMALIEFRKMMRDSIKPN 367
Query: 306 SVLIATVLASITQMANVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFR 365
S +++ L + +A G +IHG V ++G + D S LID+Y KCG + VF
Sbjct: 368 SFTLSSALRGCSNLAMFEEGRQIHGIVTKYGFDRDKYAGSGLIDLYGKCGCSDMARLVFD 427
Query: 366 IMLERNIISYNSMILAYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKD 425
+ E ++IS N+MI +Y +G +A +F+ M+ GL P++ T ++L AC ++ LV++
Sbjct: 428 TLSEVDVISLNTMIYSYAQNGFGREALDLFERMINLGLQPNDVTVLSVLLACNNSRLVEE 487
Query: 426 GRELFWRMKDEFNIKARPEHYVYMVKLLGGVGELEEAYNLTQSLPKPVDKAILGALLSCC 485
G ELF + + I +HY MV LLG G LEEA LT + P D + LLS C
Sbjct: 488 GCELFDSFRKD-KIMLTNDHYACMVDLLGRAGRLEEAEMLTTEVINP-DLVLWRTLLSAC 545
Query: 486 DSYGNSELAETVAQQIFKSNPADNVYRVMLSNIYAGDGRWDDVKKLRDKMVGGQ-KKMRG 544
+ E+AE + ++I + P D +++SN+YA G+W+ V +++ KM + KK
Sbjct: 546 KVHRKVEMAERITRKILEIEPGDEGTLILMSNLYASTGKWNRVIEMKSKMKDMKLKKNPA 605
Query: 545 VSWIE 549
+SW+E
Sbjct: 606 MSWVE 610
>At5g13230 putative protein
Length = 822
Score = 293 bits (751), Expect = 1e-79
Identities = 164/526 (31%), Positives = 277/526 (52%), Gaps = 1/526 (0%)
Query: 25 LHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMIRAFAKARR 84
LH+ ++K + F +I Y+ ++ A VF+ + + +W ++ + +
Sbjct: 168 LHSPIVKLGYDSNAFVGAALINAYSVCGSVDSARTVFEGILCKDIVVWAGIVSCYVENGY 227
Query: 85 FSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGLGLDPICCSA 144
F +++ L M + P+NYT+ A++A F + VHG + LDP
Sbjct: 228 FEDSLKLLSCMRMAGFMPNNYTFDTALKASIGLGAFDFAKGVHGQILKTCYVLDPRVGVG 287
Query: 145 LVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSMRLAGKKPD 204
L+ Y++LG + +A +VF+ + + D+V W+ +I+ + +G + +F MR A P+
Sbjct: 288 LLQLYTQLGDMSDAFKVFNEMPKNDVVPWSFMIARFCQNGFCNEAVDLFIRMREAFVVPN 347
Query: 205 GFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDSAYRVFC 264
FTL+ +L G A +G++LHGL K G D D +V + L+ +Y++C+ +D+A ++F
Sbjct: 348 EFTLSSILNGCAIGKCSGLGEQLHGLVVKVGFDLDIYVSNALIDVYAKCEKMDTAVKLFA 407
Query: 265 GIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLASITQMANVLP 324
+ + + V+W+ +I GY GE KA FR+ + V ++ L + +A++
Sbjct: 408 ELSSKNEVSWNTVIVGYENLGEGGKAFSMFREALRNQVSVTEVTFSSALGACASLASMDL 467
Query: 325 GCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSMILAYGL 384
G ++HG ++ V VS++LIDMY+KCG + VF M ++ S+N++I Y
Sbjct: 468 GVQVHGLAIKTNNAKKVAVSNSLIDMYAKCGDIKFAQSVFNEMETIDVASWNALISGYST 527
Query: 385 HGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGRELFWRMKDEFNIKARPE 444
HG QA + D M + P+ TF +LS C +AGL+ G+E F M + I+ E
Sbjct: 528 HGLGRQALRILDIMKDRDCKPNGLTFLGVLSGCSNAGLIDQGQECFESMIRDHGIEPCLE 587
Query: 445 HYVYMVKLLGGVGELEEAYNLTQSLPKPVDKAILGALLSCCDSYGNSELAETVAQQIFKS 504
HY MV+LLG G+L++A L + +P I A+LS + N E A A++I K
Sbjct: 588 HYTCMVRLLGRSGQLDKAMKLIEGIPYEPSVMIWRAMLSASMNQNNEEFARRSAEEILKI 647
Query: 505 NPADNVYRVMLSNIYAGDGRWDDVKKLRDKMVG-GQKKMRGVSWIE 549
NP D V++SN+YAG +W +V +R M G KK G+SWIE
Sbjct: 648 NPKDEATYVLVSNMYAGAKQWANVASIRKSMKEMGVKKEPGLSWIE 693
Score = 122 bits (306), Expect = 5e-28
Identities = 92/361 (25%), Positives = 164/361 (44%), Gaps = 2/361 (0%)
Query: 23 KQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMIRAFAKA 82
K +H +LKT DP +++LY ++ A VF++ V W+ MI F +
Sbjct: 267 KGVHGQILKTCYVLDPRVGVGLLQLYTQLGDMSDAFKVFNEMPKNDVVPWSFMIARFCQN 326
Query: 83 RRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGLGLDPICC 142
+ A+ LF M + P+ +T + + CA G+ +HG V VG LD
Sbjct: 327 GFCNEAVDLFIRMREAFVVPNEFTLSSILNGCAIGKCSGLGEQLHGLVVKVGFDLDIYVS 386
Query: 143 SALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSMRLAGKK 202
+AL+ Y+K + A ++F + + V WN++I Y G MF
Sbjct: 387 NALIDVYAKCEKMDTAVKLFAELSSKNEVSWNTVIVGYENLGEGGKAFSMFREALRNQVS 446
Query: 203 PDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDSAYRV 262
T + LG A + + +G ++HGL+ K+ V + L+ MY++C I A V
Sbjct: 447 VTEVTFSSALGACASLASMDLGVQVHGLAIKTNNAKKVAVSNSLIDMYAKCGDIKFAQSV 506
Query: 263 FCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLASITQMANV 322
F + D+ +W+ALISGYS G ++AL + + K + + VL+ + +
Sbjct: 507 FNEMETIDVASWNALISGYSTHGLGRQALRILDIMKDRDCKPNGLTFLGVLSGCSNAGLI 566
Query: 323 LPGCEIHGYVLR-HGLESDVKVSSALIDMYSKCGFLHLGTCVFR-IMLERNIISYNSMIL 380
G E ++R HG+E ++ + ++ + + G L + I E +++ + +M+
Sbjct: 567 DQGQECFESMIRDHGIEPCLEHYTCMVRLLGRSGQLDKAMKLIEGIPYEPSVMIWRAMLS 626
Query: 381 A 381
A
Sbjct: 627 A 627
Score = 114 bits (285), Expect = 1e-25
Identities = 82/322 (25%), Positives = 147/322 (45%), Gaps = 6/322 (1%)
Query: 98 DDIRP--DNYTYACAIRACADSFDFGMLRVVHGSAVSVGLGLDPICCSALVSAYSKLGVV 155
D I P D++ Y +R C D + +H + G LD + L++AY K G
Sbjct: 41 DSIIPGLDSHAYGAMLRRCIQKNDPISAKAIHCDILKKGSCLDLFATNILLNAYVKAGFD 100
Query: 156 HEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSMRLAGKKPDGFTLAGLLGGI 215
+A +FD + E + V + +L Y + I ++S + G + + L
Sbjct: 101 KDALNLFDEMPERNNVSFVTLAQGYA----CQDPIGLYSRLHREGHELNPHVFTSFLKLF 156
Query: 216 ADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDSAYRVFCGIFNPDLVTWS 275
I LH K G DS+ VG+ L++ YS C +DSA VF GI D+V W+
Sbjct: 157 VSLDKAEICPWLHSPIVKLGYDSNAFVGAALINAYSVCGSVDSARTVFEGILCKDIVVWA 216
Query: 276 ALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLASITQMANVLPGCEIHGYVLRH 335
++S Y + G ++ +L + M ++ T L + + +HG +L+
Sbjct: 217 GIVSCYVENGYFEDSLKLLSCMRMAGFMPNNYTFDTALKASIGLGAFDFAKGVHGQILKT 276
Query: 336 GLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSMILAYGLHGCASQAFTMF 395
D +V L+ +Y++ G + VF M + +++ ++ MI + +G ++A +F
Sbjct: 277 CYVLDPRVGVGLLQLYTQLGDMSDAFKVFNEMPKNDVVPWSFMIARFCQNGFCNEAVDLF 336
Query: 396 DEMLQKGLVPDEGTFSALLSAC 417
M + +VP+E T S++L+ C
Sbjct: 337 IRMREAFVVPNEFTLSSILNGC 358
Score = 112 bits (280), Expect = 5e-25
Identities = 81/307 (26%), Positives = 151/307 (48%), Gaps = 10/307 (3%)
Query: 23 KQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMIRAFAKA 82
+QLH ++K D + + +I +YA ++ A +F + S+++ WN++I +
Sbjct: 368 EQLHGLVVKVGFDLDIYVSNALIDVYAKCEKMDTAVKLFAELSSKNEVSWNTVIVGYENL 427
Query: 83 RRFSNAISLFRTMLVDDIRPDNYTYACAIRACAD--SFDFGMLRVVHGSAVSVGLGLDPI 140
A S+FR L + + T++ A+ ACA S D G+ VHG A+
Sbjct: 428 GEGGKAFSMFREALRNQVSVTEVTFSSALGACASLASMDLGVQ--VHGLAIKTNNAKKVA 485
Query: 141 CCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSMRLAG 200
++L+ Y+K G + A+ VF+ + D+ WN+LIS Y G+ +++ M+
Sbjct: 486 VSNSLIDMYAKCGDIKFAQSVFNEMETIDVASWNALISGYSTHGLGRQALRILDIMKDRD 545
Query: 201 KKPDGFTLAGLLGGIADSSLLSIGQE-LHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDSA 259
KP+G T G+L G +++ L+ GQE + + G++ + +V + R +D A
Sbjct: 546 CKPNGLTFLGVLSGCSNAGLIDQGQECFESMIRDHGIEPCLEHYTCMVRLLGRSGQLDKA 605
Query: 260 YRVFCGI-FNPDLVTWSALIS-GYSQCGE--YQKALLFFRKLNMKSKKLDSVLIATVLAS 315
++ GI + P ++ W A++S +Q E +++ K+N K + VL++ + A
Sbjct: 606 MKLIEGIPYEPSVMIWRAMLSASMNQNNEEFARRSAEEILKINPKD-EATYVLVSNMYAG 664
Query: 316 ITQMANV 322
Q ANV
Sbjct: 665 AKQWANV 671
>At3g22150 hypothetical protein
Length = 820
Score = 286 bits (733), Expect = 2e-77
Identities = 181/549 (32%), Positives = 300/549 (53%), Gaps = 10/549 (1%)
Query: 14 NVCKSLLRVKQLHACLLKT--HLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFL 71
++ +S+ + + +LK KD F + I +YA I + VFD R++ +
Sbjct: 225 SISRSIKKANVFYGLMLKLGDEYVKDLFVVSSAISMYAELGDIESSRRVFDSCVERNIEV 284
Query: 72 WNSMIRAFAKARRFSNAISLFRTMLVD-DIRPDNYTYACAIRACADSFDFGMLRVVHGSA 130
WN+MI + + +I LF + +I D TY A A + + R HG
Sbjct: 285 WNTMIGVYVQNDCLVESIELFLEAIGSKEIVSDEVTYLLAASAVSALQQVELGRQFHGFV 344
Query: 131 VSVGLGLDPICCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGI 190
L + ++L+ YS+ G VH++ VF + E D+V WN++ISA+ +G+ + G+
Sbjct: 345 SKNFRELPIVIVNSLMVMYSRCGSVHKSFGVFLSMRERDVVSWNTMISAFVQNGLDDEGL 404
Query: 191 QMFSSMRLAGKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMY 250
+ M+ G K D T+ LL ++ IG++ H + G+ + + S L+ MY
Sbjct: 405 MLVYEMQKQGFKIDYITVTALLSAASNLRNKEIGKQTHAFLIRQGIQFE-GMNSYLIDMY 463
Query: 251 SRCKCIDSAYRVF--CGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVL 308
S+ I + ++F G D TW+++ISGY+Q G +K L FRK+ ++ + ++V
Sbjct: 464 SKSGLIRISQKLFEGSGYAERDQATWNSMISGYTQNGHTEKTFLVFRKMLEQNIRPNAVT 523
Query: 309 IATVLASITQMANVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIML 368
+A++L + +Q+ +V G ++HG+ +R L+ +V V+SAL+DMYSK G + +F
Sbjct: 524 VASILPACSQIGSVDLGKQLHGFSIRQYLDQNVFVASALVDMYSKAGAIKYAEDMFSQTK 583
Query: 369 ERNIISYNSMILAYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGRE 428
ERN ++Y +MIL YG HG +A ++F M + G+ PD TF A+LSAC ++GL+ +G +
Sbjct: 584 ERNSVTYTTMILGYGQHGMGERAISLFLSMQESGIKPDAITFVAVLSACSYSGLIDEGLK 643
Query: 429 LFWRMKDEFNIKARPEHYVYMVKLLGGVGELEEAYNLTQSLPKPVDKAIL-GALLSCCDS 487
+F M++ +NI+ EHY + +LG VG + EAY + L + + A L G+LL C
Sbjct: 644 IFEEMREVYNIQPSSEHYCCITDMLGRVGRVNEAYEFVKGLGEEGNIAELWGSLLGSCKL 703
Query: 488 YGNSELAETVAQQIFKSNPADNV--YRVMLSNIYAGDGRWDDVKKLRDKM-VGGQKKMRG 544
+G ELAETV++++ K + N Y V+LSN+YA + +W V K+R M G KK G
Sbjct: 704 HGELELAETVSERLAKFDKGKNFSGYEVLLSNMYAEEQKWKSVDKVRRGMREKGLKKEVG 763
Query: 545 VSWIEGSYY 553
S IE + Y
Sbjct: 764 RSGIEIAGY 772
Score = 126 bits (317), Expect = 3e-29
Identities = 114/479 (23%), Positives = 205/479 (41%), Gaps = 34/479 (7%)
Query: 57 AHHVFDKTSTRSVFLWNSMIRAFAKARRFSNAISLFRTM--LVDDIRPDNYTYACAIRAC 114
A +FD + LWN++I F A+ + M D YTY+ ++AC
Sbjct: 58 ARQLFDAIPKPTTVLWNTIIIGFICNNLPHEALLFYSRMKKTAPFTNCDAYTYSSTLKAC 117
Query: 115 ADSFDFGMLRVVHGSAVSVGLGLDPICCSALVSAYSKLGVVHEA------RRVFDGIVEP 168
A++ + + VH + + ++L++ Y + R+VFD +
Sbjct: 118 AETKNLKAGKAVHCHLIRCLQNSSRVVHNSLMNMYVSCLNAPDCFEYDVVRKVFDNMRRK 177
Query: 169 DLVLWNSLISAYGGSGMWEIGIQMFSSMRLAGKKPDGFTLAGLLGGIADSSLLSIGQELH 228
++V WN+LIS Y +G + F M KP + + ++ S + +
Sbjct: 178 NVVAWNTLISWYVKTGRNAEACRQFGIMMRMEVKPSPVSFVNVFPAVSISRSIKKANVFY 237
Query: 229 GLSQKSGLD--SDCHVGSLLVSMYSRCKCIDSAYRVFCGIFNPDLVTWSALISGYSQCGE 286
GL K G + D V S +SMY+ I+S+ RVF ++ W+ +I Y Q
Sbjct: 238 GLMLKLGDEYVKDLFVVSSAISMYAELGDIESSRRVFDSCVERNIEVWNTMIGVYVQNDC 297
Query: 287 YQKAL-LFFRKLNMKSKKLDSVLIATVLASITQMANVLPGCEIHGYVLRHGLESDVKVSS 345
+++ LF + K D V ++++ + V G + HG+V ++ E + + +
Sbjct: 298 LVESIELFLEAIGSKEIVSDEVTYLLAASAVSALQQVELGRQFHGFVSKNFRELPIVIVN 357
Query: 346 ALIDMYSKCGFLHLGTCVFRIMLERNIISYNSMILAYGLHGCASQAFTMFDEMLQKGLVP 405
+L+ MYS+CG +H VF M ER+++S+N+MI A+ +G + + EM ++G
Sbjct: 358 SLMVMYSRCGSVHKSFGVFLSMRERDVVSWNTMISAFVQNGLDDEGLMLVYEMQKQGFKI 417
Query: 406 DEGTFSALLSAC-----------CHAGLVKDGRELFWRMKDEFNIKARPEHYVYMVKLLG 454
D T +ALLSA HA L++ G + ++ ++ KL
Sbjct: 418 DYITVTALLSAASNLRNKEIGKQTHAFLIRQGIQFEGMNSYLIDMYSKSGLIRISQKLFE 477
Query: 455 GVGELEEAYNLTQSLPKPVDKAILGALLSCCDSYGNSELAETVAQQIFKSNPADNVYRV 513
G G E D+A +++S G++E V +++ + N N V
Sbjct: 478 GSGYAER------------DQATWNSMISGYTQNGHTEKTFLVFRKMLEQNIRPNAVTV 524
>At5g39350 putative protein
Length = 677
Score = 286 bits (732), Expect = 2e-77
Identities = 165/530 (31%), Positives = 281/530 (52%), Gaps = 5/530 (0%)
Query: 25 LHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMIRAFAKARR 84
+H +L++ +D + ++ +Y + A VFD R V WN+MI + +
Sbjct: 139 VHGRILRSWFGRDKYVQNALLAMYMNFGKVEMARDVFDVMKNRDVISWNTMISGYYRNGY 198
Query: 85 FSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGLGLDPICCSA 144
++A+ +F M+ + + D+ T + C D M R VH LG +A
Sbjct: 199 MNDALMMFDWMVNESVDLDHATIVSMLPVCGHLKDLEMGRNVHKLVEEKRLGDKIEVKNA 258
Query: 145 LVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSMRLAGKKPD 204
LV+ Y K G + EAR VFD + D++ W +I+ Y G E +++ M+ G +P+
Sbjct: 259 LVNMYLKCGRMDEARFVFDRMERRDVITWTCMINGYTEDGDVENALELCRLMQFEGVRPN 318
Query: 205 GFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDSAYRVFC 264
T+A L+ D+ ++ G+ LHG + + + SD + + L+SMY++CK +D +RVF
Sbjct: 319 AVTIASLVSVCGDALKVNDGKCLHGWAVRQQVYSDIIIETSLISMYAKCKRVDLCFRVFS 378
Query: 265 GIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLASITQMANVLP 324
G WSA+I+G Q AL F+++ + + + + ++L + +A++
Sbjct: 379 GASKYHTGPWSAIIAGCVQNELVSDALGLFKRMRREDVEPNIATLNSLLPAYAALADLRQ 438
Query: 325 GCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERN----IISYNSMIL 380
IH Y+ + G S + ++ L+ +YSKCG L +F + E++ ++ + ++I
Sbjct: 439 AMNIHCYLTKTGFMSSLDAATGLVHVYSKCGTLESAHKIFNGIQEKHKSKDVVLWGALIS 498
Query: 381 AYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGRELFWRMKDEFNIK 440
YG+HG A +F EM++ G+ P+E TF++ L+AC H+GLV++G LF M + +
Sbjct: 499 GYGMHGDGHNALQVFMEMVRSGVTPNEITFTSALNACSHSGLVEEGLTLFRFMLEHYKTL 558
Query: 441 ARPEHYVYMVKLLGGVGELEEAYNLTQSLPKPVDKAILGALLSCCDSYGNSELAETVAQQ 500
AR HY +V LLG G L+EAYNL ++P + GALL+ C ++ N +L E A +
Sbjct: 559 ARSNHYTCIVDLLGRAGRLDEAYNLITTIPFEPTSTVWGALLAACVTHENVQLGEMAANK 618
Query: 501 IFKSNPADNVYRVMLSNIYAGDGRWDDVKKLRDKMVG-GQKKMRGVSWIE 549
+F+ P + V+L+NIYA GRW D++K+R M G +K G S IE
Sbjct: 619 LFELEPENTGNYVLLANIYAALGRWKDMEKVRSMMENVGLRKKPGHSTIE 668
Score = 174 bits (441), Expect = 1e-43
Identities = 128/498 (25%), Positives = 235/498 (46%), Gaps = 7/498 (1%)
Query: 17 KSLLRVKQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMI 76
+S+ + K LH ++ +T + YA HI YA +F++ S+ +N +I
Sbjct: 29 QSISKTKALHCHVITGGRVSGHILSTLSVT-YALCGHITYARKLFEEMPQSSLLSYNIVI 87
Query: 77 RAFAKARRFSNAISLFRTMLVDDIR--PDNYTYACAIRACADSFDFGMLRVVHGSAVSVG 134
R + + + +AIS+F M+ + ++ PD YTY +A + + VVHG +
Sbjct: 88 RMYVREGLYHDAISVFIRMVSEGVKCVPDGYTYPFVAKAAGELKSMKLGLVVHGRILRSW 147
Query: 135 LGLDPICCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFS 194
G D +AL++ Y G V AR VFD + D++ WN++IS Y +G + MF
Sbjct: 148 FGRDKYVQNALLAMYMNFGKVEMARDVFDVMKNRDVISWNTMISGYYRNGYMNDALMMFD 207
Query: 195 SMRLAGKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCK 254
M D T+ +L L +G+ +H L ++ L V + LV+MY +C
Sbjct: 208 WMVNESVDLDHATIVSMLPVCGHLKDLEMGRNVHKLVEEKRLGDKIEVKNALVNMYLKCG 267
Query: 255 CIDSAYRVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLA 314
+D A VF + D++TW+ +I+GY++ G+ + AL R + + + ++V IA++++
Sbjct: 268 RMDEARFVFDRMERRDVITWTCMINGYTEDGDVENALELCRLMQFEGVRPNAVTIASLVS 327
Query: 315 SITQMANVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIIS 374
V G +HG+ +R + SD+ + ++LI MY+KC + L VF + +
Sbjct: 328 VCGDALKVNDGKCLHGWAVRQQVYSDIIIETSLISMYAKCKRVDLCFRVFSGASKYHTGP 387
Query: 375 YNSMILAYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGRELFWRMK 434
++++I + S A +F M ++ + P+ T ++LL A ++ + +
Sbjct: 388 WSAIIAGCVQNELVSDALGLFKRMRREDVEPNIATLNSLLPAYAALADLRQAMNIHCYL- 446
Query: 435 DEFNIKARPEHYVYMVKLLGGVGELEEA---YNLTQSLPKPVDKAILGALLSCCDSYGNS 491
+ + + +V + G LE A +N Q K D + GAL+S +G+
Sbjct: 447 TKTGFMSSLDAATGLVHVYSKCGTLESAHKIFNGIQEKHKSKDVVLWGALISGYGMHGDG 506
Query: 492 ELAETVAQQIFKSNPADN 509
A V ++ +S N
Sbjct: 507 HNALQVFMEMVRSGVTPN 524
>At1g03540 hypothetical protein
Length = 609
Score = 285 bits (730), Expect = 4e-77
Identities = 179/549 (32%), Positives = 287/549 (51%), Gaps = 11/549 (2%)
Query: 8 LHSELTNVCK---SLLRVKQLHACLLKTHLSKDPFYATQIIRLY-AFNNHINYAHHVFDK 63
L++ L C S + Q HA ++K+ L D ++ LY + VFD
Sbjct: 63 LYASLLQTCNKVFSFIHGIQFHAHVVKSGLETDRNVGNSLLSLYFKLGPGMRETRRVFDG 122
Query: 64 TSTRSVFLWNSMIRAFAKARRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGML 123
+ W SM+ + + A+ +F M+ + + +T + A++AC++ + +
Sbjct: 123 RFVKDAISWTSMMSGYVTGKEHVKALEVFVEMVSFGLDANEFTLSSAVKACSELGEVRLG 182
Query: 124 RVVHGSAVSVGLGLDPICCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGS 183
R HG ++ G + S L Y +ARRVFD + EPD++ W +++SA+ +
Sbjct: 183 RCFHGVVITHGFEWNHFISSTLAYLYGVNREPVDARRVFDEMPEPDVICWTAVLSAFSKN 242
Query: 184 GMWEIGIQMFSSM-RLAGKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHV 242
++E + +F +M R G PDG T +L + L G+E+HG +G+ S+ V
Sbjct: 243 DLYEEALGLFYAMHRGKGLVPDGSTFGTVLTACGNLRRLKQGKEIHGKLITNGIGSNVVV 302
Query: 243 GSLLVSMYSRCKCIDSAYRVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSK 302
S L+ MY +C + A +VF G+ + V+WSAL+ GY Q GE++KA+ FR++ K
Sbjct: 303 ESSLLDMYGKCGSVREARQVFNGMSKKNSVSWSALLGGYCQNGEHEKAIEIFREMEEK-- 360
Query: 303 KLDSVLIATVLASITQMANVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTC 362
D TVL + +A V G EIHG +R G +V V SALID+Y K G + +
Sbjct: 361 --DLYCFGTVLKACAGLAAVRLGKEIHGQYVRRGCFGNVIVESALIDLYGKSGCIDSASR 418
Query: 363 VFRIMLERNIISYNSMILAYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGL 422
V+ M RN+I++N+M+ A +G +A + F++M++KG+ PD +F A+L+AC H G+
Sbjct: 419 VYSKMSIRNMITWNAMLSALAQNGRGEEAVSFFNDMVKKGIKPDYISFIAILTACGHTGM 478
Query: 423 VKDGRELFWRMKDEFNIKARPEHYVYMVKLLGGVGELEEAYNLTQSLPKPVDKAILGALL 482
V +GR F M + IK EHY M+ LLG G EEA NL + D ++ G LL
Sbjct: 479 VDEGRNYFVLMAKSYGIKPGTEHYSCMIDLLGRAGLFEEAENLLERAECRNDASLWGVLL 538
Query: 483 SCCDSYGN-SELAETVAQQIFKSNPADNVYRVMLSNIYAGDGRWDDVKKLRDKMV-GGQK 540
C + + S +AE +A+++ + P ++ V+LSN+Y GR D +R MV G
Sbjct: 539 GPCAANADASRVAERIAKRMMELEPKYHMSYVLLSNMYKAIGRHGDALNIRKLMVRRGVA 598
Query: 541 KMRGVSWIE 549
K G SWI+
Sbjct: 599 KTVGQSWID 607
>At3g57430 putative protein
Length = 803
Score = 285 bits (729), Expect = 5e-77
Identities = 169/548 (30%), Positives = 289/548 (51%), Gaps = 16/548 (2%)
Query: 17 KSLLRVKQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMI 76
+ L+ KQ+HA L+ + F ++ +Y + + + R + WN+++
Sbjct: 129 EGLMMGKQVHAYGLRKG-ELNSFIINTLVAMYGKLGKLASSKVLLGSFGGRDLVTWNTVL 187
Query: 77 RAFAKARRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVG-L 135
+ + + A+ R M+++ + PD +T + + AC+ + +H A+ G L
Sbjct: 188 SSLCQNEQLLEALEYLREMVLEGVEPDEFTISSVLPACSHLEMLRTGKELHAYALKNGSL 247
Query: 136 GLDPICCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSS 195
+ SALV Y V RRVFDG+ + + LWN++I+ Y + + + +F
Sbjct: 248 DENSFVGSALVDMYCNCKQVLSGRRVFDGMFDRKIGLWNAMIAGYSQNEHDKEALLLFIG 307
Query: 196 MR-LAGKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCK 254
M AG + T+AG++ S S + +HG K GLD D V + L+ MYSR
Sbjct: 308 MEESAGLLANSTTMAGVVPACVRSGAFSRKEAIHGFVVKRGLDRDRFVQNTLMDMYSRLG 367
Query: 255 CIDSAYRVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKL---------- 304
ID A R+F + + DLVTW+ +I+GY ++ ALL K+ +K+
Sbjct: 368 KIDIAMRIFGKMEDRDLVTWNTMITGYVFSEHHEDALLLLHKMQNLERKVSKGASRVSLK 427
Query: 305 -DSVLIATVLASITQMANVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCV 363
+S+ + T+L S ++ + G EIH Y +++ L +DV V SAL+DMY+KCG L + V
Sbjct: 428 PNSITLMTILPSCAALSALAKGKEIHAYAIKNNLATDVAVGSALVDMYAKCGCLQMSRKV 487
Query: 364 FRIMLERNIISYNSMILAYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLV 423
F + ++N+I++N +I+AYG+HG +A + M+ +G+ P+E TF ++ +AC H+G+V
Sbjct: 488 FDQIPQKNVITWNVIIMAYGMHGNGQEAIDLLRMMMVQGVKPNEVTFISVFAACSHSGMV 547
Query: 424 KDGRELFWRMKDEFNIKARPEHYVYMVKLLGGVGELEEAYNLTQSLPKPVDKA-ILGALL 482
+G +F+ MK ++ ++ +HY +V LLG G ++EAY L +P+ +KA +LL
Sbjct: 548 DEGLRIFYVMKPDYGVEPSSDHYACVVDLLGRAGRIKEAYQLMNMMPRDFNKAGAWSSLL 607
Query: 483 SCCDSYGNSELAETVAQQIFKSNPADNVYRVMLSNIYAGDGRWDDVKKLRDKM-VGGQKK 541
+ N E+ E AQ + + P + V+L+NIY+ G WD ++R M G +K
Sbjct: 608 GASRIHNNLEIGEIAAQNLIQLEPNVASHYVLLANIYSSAGLWDKATEVRRNMKEQGVRK 667
Query: 542 MRGVSWIE 549
G SWIE
Sbjct: 668 EPGCSWIE 675
Score = 187 bits (476), Expect = 1e-47
Identities = 129/433 (29%), Positives = 217/433 (49%), Gaps = 36/433 (8%)
Query: 23 KQLHACLLKTHLSKDPF-YATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMIRAFAK 81
KQ+HA + K D A ++ LY + VFD+ S R+ WNS+I +
Sbjct: 30 KQIHAHVYKFGYGVDSVTVANTLVNLYRKCGDFGAVYKVFDRISERNQVSWNSLISSLCS 89
Query: 82 ARRFSNAISLFRTMLVDDIRPDNYTYACAIRACAD-SFDFGML--RVVHGSAVSVGLGLD 138
++ A+ FR ML +++ P ++T + AC++ G++ + VH + G L+
Sbjct: 90 FEKWEMALEAFRCMLDENVEPSSFTLVSVVTACSNLPMPEGLMMGKQVHAYGLRKG-ELN 148
Query: 139 PICCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSMRL 198
+ LV+ Y KLG + ++ + DLV WN+++S+ + ++ M L
Sbjct: 149 SFIINTLVAMYGKLGKLASSKVLLGSFGGRDLVTWNTVLSSLCQNEQLLEALEYLREMVL 208
Query: 199 AGKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSG-LDSDCHVGSLLVSMYSRCKCID 257
G +PD FT++ +L + +L G+ELH + K+G LD + VGS LV MY CK +
Sbjct: 209 EGVEPDEFTISSVLPACSHLEMLRTGKELHAYALKNGSLDENSFVGSALVDMYCNCKQVL 268
Query: 258 SAYRVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLASIT 317
S RVF G+F+ + W+A+I+GYSQ ++ALL F + M+ A +LA+ T
Sbjct: 269 SGRRVFDGMFDRKIGLWNAMIAGYSQNEHDKEALLLF--IGMEES-------AGLLANST 319
Query: 318 QMANVLPGC----------EIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIM 367
MA V+P C IHG+V++ GL+ D V + L+DMYS+ G + + +F M
Sbjct: 320 TMAGVVPACVRSGAFSRKEAIHGFVVKRGLDRDRFVQNTLMDMYSRLGKIDIAMRIFGKM 379
Query: 368 LERNIISYNSMILAYGLHGCASQAFTMFDEM------LQKG-----LVPDEGTFSALLSA 416
+R+++++N+MI Y A + +M + KG L P+ T +L +
Sbjct: 380 EDRDLVTWNTMITGYVFSEHHEDALLLLHKMQNLERKVSKGASRVSLKPNSITLMTILPS 439
Query: 417 CCHAGLVKDGREL 429
C + G+E+
Sbjct: 440 CAALSALAKGKEI 452
Score = 150 bits (378), Expect = 2e-36
Identities = 100/334 (29%), Positives = 175/334 (51%), Gaps = 9/334 (2%)
Query: 95 MLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGLGLDPI-CCSALVSAYSKLG 153
M+V I+PDNY + ++A AD D + + +H G G+D + + LV+ Y K G
Sbjct: 1 MIVLGIKPDNYAFPALLKAVADLQDMELGKQIHAHVYKFGYGVDSVTVANTLVNLYRKCG 60
Query: 154 VVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSMRLAGKKPDGFTLAGLLG 213
+VFD I E + V WNSLIS+ WE+ ++ F M +P FTL ++
Sbjct: 61 DFGAVYKVFDRISERNQVSWNSLISSLCSFEKWEMALEAFRCMLDENVEPSSFTLVSVVT 120
Query: 214 GIADSSL---LSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDSAYRVFCGIF-NP 269
++ + L +G+++H + G + + + + LV+MY + + S+ +V G F
Sbjct: 121 ACSNLPMPEGLMMGKQVHAYGLRKG-ELNSFIINTLVAMYGKLGKLASS-KVLLGSFGGR 178
Query: 270 DLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLASITQMANVLPGCEIH 329
DLVTW+ ++S Q + +AL + R++ ++ + D I++VL + + + + G E+H
Sbjct: 179 DLVTWNTVLSSLCQNEQLLEALEYLREMVLEGVEPDEFTISSVLPACSHLEMLRTGKELH 238
Query: 330 GYVLRHG-LESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSMILAYGLHGCA 388
Y L++G L+ + V SAL+DMY C + G VF M +R I +N+MI Y +
Sbjct: 239 AYALKNGSLDENSFVGSALVDMYCNCKQVLSGRRVFDGMFDRKIGLWNAMIAGYSQNEHD 298
Query: 389 SQAFTMFDEMLQK-GLVPDEGTFSALLSACCHAG 421
+A +F M + GL+ + T + ++ AC +G
Sbjct: 299 KEALLLFIGMEESAGLLANSTTMAGVVPACVRSG 332
>At5g04780 selenium-binding protein-like
Length = 864
Score = 284 bits (727), Expect = 8e-77
Identities = 168/498 (33%), Positives = 277/498 (54%), Gaps = 22/498 (4%)
Query: 70 FLWNSMIRAFAKARRFSNAISLFRTMLV------DDIRPDNYTYACA--------IRACA 115
F N +IR RR SN SL R + V +++ P Y+ + ++ CA
Sbjct: 6 FTVNFLIRCKVLPRR-SNTSSLSRNISVLASYDQEEVSPGRYSNEFSNRNLVHEILQLCA 64
Query: 116 DSFDFGMLRVVHGSAVSVGLGLDPICCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNS 175
+ + HG + + L D + L++AYSK G V AR+VFDG++E LV WN+
Sbjct: 65 RNGAVMEAKACHGKIIRIDLEGDVTLLNVLINAYSKCGFVELARQVFDGMLERSLVSWNT 124
Query: 176 LISAYGGSGMWEIGIQMFSSMRLAGKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSG 235
+I Y + M + +F MR G K FT++ +L + ++LH LS K+
Sbjct: 125 MIGLYTRNRMESEALDIFLEMRNEGFKFSEFTISSVLSACGVNCDALECKKLHCLSVKTC 184
Query: 236 LDSDCHVGSLLVSMYSRCKCIDSAYRVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFR 295
+D + +VG+ L+ +Y++C I A +VF + + VTWS++++GY Q Y++ALL +R
Sbjct: 185 IDLNLYVGTALLDLYAKCGMIKDAVQVFESMQDKSSVTWSSMVAGYVQNKNYEEALLLYR 244
Query: 296 KLNMKSKKLDSVLIATVLASITQMANVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCG 355
+ S + + +++V+ + + +A ++ G ++H + + G S+V V+S+ +DMY+KCG
Sbjct: 245 RAQRMSLEQNQFTLSSVICACSNLAALIEGKQMHAVICKSGFGSNVFVASSAVDMYAKCG 304
Query: 356 FLHLGTCVFRIMLERNIISYNSMILAYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLS 415
L +F + E+N+ +N++I + H + +F++M Q G+ P+E TFS+LLS
Sbjct: 305 SLRESYIIFSEVQEKNLELWNTIISGFAKHARPKEVMILFEKMQQDGMHPNEVTFSSLLS 364
Query: 416 ACCHAGLVKDGRELFWRMKDEFNIKARPEHYVYMVKLLGGVGELEEAYNLTQSLPKPVDK 475
C H GLV++GR F M+ + + HY MV +LG G L EAY L +S+P
Sbjct: 365 VCGHTGLVEEGRRFFKLMRTTYGLSPNVVHYSCMVDILGRAGLLSEAYELIKSIPFDPTA 424
Query: 476 AILGALLSCCDSYGNSELAETVAQQIFKSNPADNVYRVMLSNIYAGDGRWDDVKK----L 531
+I G+LL+ C Y N ELAE A+++F+ P + V+LSNIYA + +W+++ K L
Sbjct: 425 SIWGSLLASCRVYKNLELAEVAAEKLFELEPENAGNHVLLSNIYAANKQWEEIAKSRKLL 484
Query: 532 RDKMVGGQKKMRGVSWIE 549
RD V KK+RG SWI+
Sbjct: 485 RDCDV---KKVRGKSWID 499
Score = 130 bits (328), Expect = 1e-30
Identities = 79/351 (22%), Positives = 169/351 (47%), Gaps = 4/351 (1%)
Query: 11 ELTNVCK---SLLRVKQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTR 67
E+ +C +++ K H +++ L D +I Y+ + A VFD R
Sbjct: 58 EILQLCARNGAVMEAKACHGKIIRIDLEGDVTLLNVLINAYSKCGFVELARQVFDGMLER 117
Query: 68 SVFLWNSMIRAFAKARRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVH 127
S+ WN+MI + + R S A+ +F M + + +T + + AC + D + +H
Sbjct: 118 SLVSWNTMIGLYTRNRMESEALDIFLEMRNEGFKFSEFTISSVLSACGVNCDALECKKLH 177
Query: 128 GSAVSVGLGLDPICCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWE 187
+V + L+ +AL+ Y+K G++ +A +VF+ + + V W+S+++ Y + +E
Sbjct: 178 CLSVKTCIDLNLYVGTALLDLYAKCGMIKDAVQVFESMQDKSSVTWSSMVAGYVQNKNYE 237
Query: 188 IGIQMFSSMRLAGKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLV 247
+ ++ + + + FTL+ ++ ++ + L G+++H + KSG S+ V S V
Sbjct: 238 EALLLYRRAQRMSLEQNQFTLSSVICACSNLAALIEGKQMHAVICKSGFGSNVFVASSAV 297
Query: 248 SMYSRCKCIDSAYRVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSV 307
MY++C + +Y +F + +L W+ +ISG+++ ++ ++ F K+ + V
Sbjct: 298 DMYAKCGSLRESYIIFSEVQEKNLELWNTIISGFAKHARPKEVMILFEKMQQDGMHPNEV 357
Query: 308 LIATVLASITQMANVLPGCEIHGYV-LRHGLESDVKVSSALIDMYSKCGFL 357
+++L+ V G + +GL +V S ++D+ + G L
Sbjct: 358 TFSSLLSVCGHTGLVEEGRRFFKLMRTTYGLSPNVVHYSCMVDILGRAGLL 408
>At4g13650 unknown protein
Length = 1024
Score = 283 bits (725), Expect = 1e-76
Identities = 166/533 (31%), Positives = 262/533 (49%), Gaps = 1/533 (0%)
Query: 18 SLLRVKQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMIR 77
+L R +QLHA K + + ++ LYA I A F +T +V LWN M+
Sbjct: 364 TLFRGQQLHAYTTKLGFASNNKIEGALLNLYAKCADIETALDYFLETEVENVVLWNVMLV 423
Query: 78 AFAKARRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGLGL 137
A+ N+ +FR M +++I P+ YTY ++ C D + +H + L
Sbjct: 424 AYGLLDDLRNSFRIFRQMQIEEIVPNQYTYPSILKTCIRLGDLELGEQIHSQIIKTNFQL 483
Query: 138 DPICCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSMR 197
+ CS L+ Y+KLG + A + D+V W ++I+ Y + + F M
Sbjct: 484 NAYVCSVLIDMYAKLGKLDTAWDILIRFAGKDVVSWTTMIAGYTQYNFDDKALTTFRQML 543
Query: 198 LAGKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCID 257
G + D L + A L GQ++H + SG SD + LV++YSRC I+
Sbjct: 544 DRGIRSDEVGLTNAVSACAGLQALKEGQQIHAQACVSGFSSDLPFQNALVTLYSRCGKIE 603
Query: 258 SAYRVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLASIT 317
+Y F D + W+AL+SG+ Q G ++AL F ++N + ++ + + + +
Sbjct: 604 ESYLAFEQTEAGDNIAWNALVSGFQQSGNNEEALRVFVRMNREGIDNNNFTFGSAVKAAS 663
Query: 318 QMANVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNS 377
+ AN+ G ++H + + G +S+ +V +ALI MY+KCG + F + +N +S+N+
Sbjct: 664 ETANMKQGKQVHAVITKTGYDSETEVCNALISMYAKCGSISDAEKQFLEVSTKNEVSWNA 723
Query: 378 MILAYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGRELFWRMKDEF 437
+I AY HG S+A FD+M+ + P+ T +LSAC H GLV G F M E+
Sbjct: 724 IINAYSKHGFGSEALDSFDQMIHSNVRPNHVTLVGVLSACSHIGLVDKGIAYFESMNSEY 783
Query: 438 NIKARPEHYVYMVKLLGGVGELEEAYNLTQSLPKPVDKAILGALLSCCDSYGNSELAETV 497
+ +PEHYV +V +L G L A Q +P D + LLS C + N E+ E
Sbjct: 784 GLSPKPEHYVCVVDMLTRAGLLSRAKEFIQEMPIKPDALVWRTLLSACVVHKNMEIGEFA 843
Query: 498 AQQIFKSNPADNVYRVMLSNIYAGDGRWDDVKKLRDKM-VGGQKKMRGVSWIE 549
A + + P D+ V+LSN+YA +WD R KM G KK G SWIE
Sbjct: 844 AHHLLELEPEDSATYVLLSNLYAVSKKWDARDLTRQKMKEKGVKKEPGQSWIE 896
Score = 215 bits (547), Expect = 6e-56
Identities = 129/463 (27%), Positives = 235/463 (49%), Gaps = 8/463 (1%)
Query: 7 WLHSELTNVCKSLLRVKQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTST 66
WL SL ++LH+ +LK L + + ++ Y F + A VFD+
Sbjct: 49 WLLEGCLKTNGSLDEGRKLHSQILKLGLDSNGCLSEKLFDFYLFKGDLYGAFKVFDEMPE 108
Query: 67 RSVFLWNSMIRAFAKARRFSNAISLFRTMLVDDIRPDNYTYACAIRAC-ADSFDFGMLRV 125
R++F WN MI+ A LF M+ +++ P+ T++ + AC S F ++
Sbjct: 109 RTIFTWNKMIKELASRNLIGEVFGLFVRMVSENVTPNEGTFSGVLEACRGGSVAFDVVEQ 168
Query: 126 VHGSAVSVGLGLDPICCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGM 185
+H + GL + C+ L+ YS+ G V ARRVFDG+ D W ++IS +
Sbjct: 169 IHARILYQGLRDSTVVCNPLIDLYSRNGFVDLARRVFDGLRLKDHSSWVAMISGLSKNEC 228
Query: 186 WEIGIQMFSSMRLAGKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSL 245
I++F M + G P + + +L L IG++LHGL K G SD +V +
Sbjct: 229 EAEAIRLFCDMYVLGIMPTPYAFSSVLSACKKIESLEIGEQLHGLVLKLGFSSDTYVCNA 288
Query: 246 LVSMYSRCKCIDSAYRVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLD 305
LVS+Y + SA +F + D VT++ LI+G SQCG +KA+ F+++++ + D
Sbjct: 289 LVSLYFHLGNLISAEHIFSNMSQRDAVTYNTLINGLSQCGYGEKAMELFKRMHLDGLEPD 348
Query: 306 SVLIATVLASITQMANVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFR 365
S +A+++ + + + G ++H Y + G S+ K+ AL+++Y+KC + F
Sbjct: 349 SNTLASLVVACSADGTLFRGQQLHAYTTKLGFASNNKIEGALLNLYAKCADIETALDYFL 408
Query: 366 IMLERNIISYNSMILAYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKD 425
N++ +N M++AYGL +F +F +M + +VP++ T+ ++L C G ++
Sbjct: 409 ETEVENVVLWNVMLVAYGLLDDLRNSFRIFRQMQIEEIVPNQYTYPSILKTCIRLGDLEL 468
Query: 426 GRELFWR-MKDEFNIKARPEHYV--YMVKLLGGVGELEEAYNL 465
G ++ + +K F + A YV ++ + +G+L+ A+++
Sbjct: 469 GEQIHSQIIKTNFQLNA----YVCSVLIDMYAKLGKLDTAWDI 507
Score = 189 bits (479), Expect = 5e-48
Identities = 126/473 (26%), Positives = 223/473 (46%), Gaps = 2/473 (0%)
Query: 22 VKQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMIRAFAK 81
V+Q+HA +L L +I LY+ N ++ A VFD + W +MI +K
Sbjct: 166 VEQIHARILYQGLRDSTVVCNPLIDLYSRNGFVDLARRVFDGLRLKDHSSWVAMISGLSK 225
Query: 82 ARRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGLGLDPIC 141
+ AI LF M V I P Y ++ + AC + +HG + +G D
Sbjct: 226 NECEAEAIRLFCDMYVLGIMPTPYAFSSVLSACKKIESLEIGEQLHGLVLKLGFSSDTYV 285
Query: 142 CSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSMRLAGK 201
C+ALVS Y LG + A +F + + D V +N+LI+ G E +++F M L G
Sbjct: 286 CNALVSLYFHLGNLISAEHIFSNMSQRDAVTYNTLINGLSQCGYGEKAMELFKRMHLDGL 345
Query: 202 KPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDSAYR 261
+PD TLA L+ + L GQ+LH + K G S+ + L+++Y++C I++A
Sbjct: 346 EPDSNTLASLVVACSADGTLFRGQQLHAYTTKLGFASNNKIEGALLNLYAKCADIETALD 405
Query: 262 VFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLASITQMAN 321
F ++V W+ ++ Y + + + FR++ ++ + ++L + ++ +
Sbjct: 406 YFLETEVENVVLWNVMLVAYGLLDDLRNSFRIFRQMQIEEIVPNQYTYPSILKTCIRLGD 465
Query: 322 VLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSMILA 381
+ G +IH +++ + + V S LIDMY+K G L + ++++S+ +MI
Sbjct: 466 LELGEQIHSQIIKTNFQLNAYVCSVLIDMYAKLGKLDTAWDILIRFAGKDVVSWTTMIAG 525
Query: 382 YGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGRELFWRMKDEFNIKA 441
Y + +A T F +ML +G+ DE + +SAC +K+G+++ +
Sbjct: 526 YTQYNFDDKALTTFRQMLDRGIRSDEVGLTNAVSACAGLQALKEGQQIHAQACVSGFSSD 585
Query: 442 RPEHYVYMVKLLGGVGELEEAYNLTQSLPKPVDKAILGALLSCCDSYGNSELA 494
P +V L G++EE+Y L + D AL+S GN+E A
Sbjct: 586 LPFQNA-LVTLYSRCGKIEESY-LAFEQTEAGDNIAWNALVSGFQQSGNNEEA 636
Score = 95.1 bits (235), Expect = 9e-20
Identities = 61/265 (23%), Positives = 120/265 (45%), Gaps = 2/265 (0%)
Query: 17 KSLLRVKQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMI 76
++L +Q+HA + S D + ++ LY+ I ++ F++T WN+++
Sbjct: 565 QALKEGQQIHAQACVSGFSSDLPFQNALVTLYSRCGKIEESYLAFEQTEAGDNIAWNALV 624
Query: 77 RAFAKARRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGLG 136
F ++ A+ +F M + I +N+T+ A++A +++ + + VH G
Sbjct: 625 SGFQQSGNNEEALRVFVRMNREGIDNNNFTFGSAVKAASETANMKQGKQVHAVITKTGYD 684
Query: 137 LDPICCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSM 196
+ C+AL+S Y+K G + +A + F + + V WN++I+AY G + F M
Sbjct: 685 SETEVCNALISMYAKCGSISDAEKQFLEVSTKNEVSWNAIINAYSKHGFGSEALDSFDQM 744
Query: 197 RLAGKKPDGFTLAGLLGGIADSSLLSIG-QELHGLSQKSGLDSDCHVGSLLVSMYSRCKC 255
+ +P+ TL G+L + L+ G ++ + GL +V M +R
Sbjct: 745 IHSNVRPNHVTLVGVLSACSHIGLVDKGIAYFESMNSEYGLSPKPEHYVCVVDMLTRAGL 804
Query: 256 IDSAYRVFCGI-FNPDLVTWSALIS 279
+ A + PD + W L+S
Sbjct: 805 LSRAKEFIQEMPIKPDALVWRTLLS 829
>At1g50270 hypothetical protein
Length = 596
Score = 283 bits (725), Expect = 1e-76
Identities = 167/537 (31%), Positives = 277/537 (51%), Gaps = 8/537 (1%)
Query: 20 LRVKQLHACLLKT---HLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMI 76
L +KQ+H LL + + +D F + + R YA + + T S+ LW+S+I
Sbjct: 15 LHLKQIHCLLLTSPIFYTRRDLFLSRLLRRCCTAATQFRYARRLLCQLQTLSIQLWDSLI 74
Query: 77 RAFAKARRFSNAISL--FRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVG 134
F+ + +S +R M + + P +T+ ++A D + H V G
Sbjct: 75 GHFSGGITLNRRLSFLAYRHMRRNGVIPSRHTFPPLLKAVFKLRDSNPFQF-HAHIVKFG 133
Query: 135 LGLDPICCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFS 194
L DP ++L+S YS G+ A R+FDG + D+V W ++I + +G + F
Sbjct: 134 LDSDPFVRNSLISGYSSSGLFDFASRLFDGAEDKDVVTWTAMIDGFVRNGSASEAMVYFV 193
Query: 195 SMRLAGKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSG-LDSDCHVGSLLVSMYSRC 253
M+ G + T+ +L + G+ +HGL ++G + D +GS LV MY +C
Sbjct: 194 EMKKTGVAANEMTVVSVLKAAGKVEDVRFGRSVHGLYLETGRVKCDVFIGSSLVDMYGKC 253
Query: 254 KCIDSAYRVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVL 313
C D A +VF + + ++VTW+ALI+GY Q + K +L F ++ + +++VL
Sbjct: 254 SCYDDAQKVFDEMPSRNVVTWTALIAGYVQSRCFDKGMLVFEEMLKSDVAPNEKTLSSVL 313
Query: 314 ASITQMANVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNII 373
++ + + G +H Y++++ +E + + LID+Y KCG L VF + E+N+
Sbjct: 314 SACAHVGALHRGRRVHCYMIKNSIEINTTAGTTLIDLYVKCGCLEEAILVFERLHEKNVY 373
Query: 374 SYNSMILAYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGRELFWRM 433
++ +MI + HG A AF +F ML + P+E TF A+LSAC H GLV++GR LF M
Sbjct: 374 TWTAMINGFAAHGYARDAFDLFYTMLSSHVSPNEVTFMAVLSACAHGGLVEEGRRLFLSM 433
Query: 434 KDEFNIKARPEHYVYMVKLLGGVGELEEAYNLTQSLPKPVDKAILGALLSCCDSYGNSEL 493
K FN++ + +HY MV L G G LEEA L + +P + GAL C + + EL
Sbjct: 434 KGRFNMEPKADHYACMVDLFGRKGLLEEAKALIERMPMEPTNVVWGALFGSCLLHKDYEL 493
Query: 494 AETVAQQIFKSNPADNVYRVMLSNIYAGDGRWDDVKKLRDKMVGGQ-KKMRGVSWIE 549
+ A ++ K P+ + +L+N+Y+ WD+V ++R +M Q K G SWIE
Sbjct: 494 GKYAASRVIKLQPSHSGRYTLLANLYSESQNWDEVARVRKQMKDQQVVKSPGFSWIE 550
Score = 60.1 bits (144), Expect = 3e-09
Identities = 44/177 (24%), Positives = 78/177 (43%), Gaps = 28/177 (15%)
Query: 18 SLLRVKQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMIR 77
+L R +++H ++K + + T +I LY + A VF++ ++V+ W +MI
Sbjct: 321 ALHRGRRVHCYMIKNSIEINTTAGTTLIDLYVKCGCLEEAILVFERLHEKNVYTWTAMIN 380
Query: 78 AFAKARRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGLGL 137
FA +A LF TML + P+ T+ + ACA HG V G L
Sbjct: 381 GFAAHGYARDAFDLFYTMLSSHVSPNEVTFMAVLSACA-----------HGGLVEEGRRL 429
Query: 138 --------------DPICCSALVSAYSKLGVVHEARRVFDGI-VEPDLVLWNSLISA 179
D C +V + + G++ EA+ + + + +EP V+W +L +
Sbjct: 430 FLSMKGRFNMEPKADHYAC--MVDLFGRKGLLEEAKALIERMPMEPTNVVWGALFGS 484
>At1g18485 hypothetical protein
Length = 970
Score = 283 bits (725), Expect = 1e-76
Identities = 162/531 (30%), Positives = 284/531 (52%), Gaps = 4/531 (0%)
Query: 23 KQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMIRAFAKA 82
K +H +K L K+ ++ +Y+ I A +F + ++V WN+M+ F+
Sbjct: 312 KGVHGWAVKLRLDKELVLNNALMDMYSKCGCITNAQMIFKMNNNKNVVSWNTMVGGFSAE 371
Query: 83 RRFSNAISLFRTMLV--DDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGLGLDPI 140
+ R ML +D++ D T A+ C L+ +H ++ + +
Sbjct: 372 GDTHGTFDVLRQMLAGGEDVKADEVTILNAVPVCFHESFLPSLKELHCYSLKQEFVYNEL 431
Query: 141 CCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSMRLAG 200
+A V++Y+K G + A+RVF GI + WN+LI + S + + M+++G
Sbjct: 432 VANAFVASYAKCGSLSYAQRVFHGIRSKTVNSWNALIGGHAQSNDPRLSLDAHLQMKISG 491
Query: 201 KKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDSAY 260
PD FT+ LL + L +G+E+HG ++ L+ D V ++S+Y C + +
Sbjct: 492 LLPDSFTVCSLLSACSKLKSLRLGKEVHGFIIRNWLERDLFVYLSVLSLYIHCGELCTVQ 551
Query: 261 RVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLASITQMA 320
+F + + LV+W+ +I+GY Q G +AL FR++ + +L + + V + + +
Sbjct: 552 ALFDAMEDKSLVSWNTVITGYLQNGFPDRALGVFRQMVLYGIQLCGISMMPVFGACSLLP 611
Query: 321 NVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSMIL 380
++ G E H Y L+H LE D ++ +LIDMY+K G + + VF + E++ S+N+MI+
Sbjct: 612 SLRLGREAHAYALKHLLEDDAFIACSLIDMYAKNGSITQSSKVFNGLKEKSTASWNAMIM 671
Query: 381 AYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGRELFWRMKDEFNIK 440
YG+HG A +A +F+EM + G PD+ TF +L+AC H+GL+ +G +MK F +K
Sbjct: 672 GYGIHGLAKEAIKLFEEMQRTGHNPDDLTFLGVLTACNHSGLIHEGLRYLDQMKSSFGLK 731
Query: 441 ARPEHYVYMVKLLGGVGELEEAYN-LTQSLPKPVDKAILGALLSCCDSYGNSELAETVAQ 499
+HY ++ +LG G+L++A + + + + D I +LLS C + N E+ E VA
Sbjct: 732 PNLKHYACVIDMLGRAGQLDKALRVVAEEMSEEADVGIWKSLLSSCRIHQNLEMGEKVAA 791
Query: 500 QIFKSNPADNVYRVMLSNIYAGDGRWDDVKKLRDKM-VGGQKKMRGVSWIE 549
++F+ P V+LSN+YAG G+W+DV+K+R +M +K G SWIE
Sbjct: 792 KLFELEPEKPENYVLLSNLYAGLGKWEDVRKVRQRMNEMSLRKDAGCSWIE 842
Score = 181 bits (458), Expect = 1e-45
Identities = 123/409 (30%), Positives = 203/409 (49%), Gaps = 15/409 (3%)
Query: 32 THLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMIRAFAKARRFSNAISL 91
T L D T+II +YA + + VFD ++++F WN++I ++++ + +
Sbjct: 114 TRLRNDDVLCTRIITMYAMCGSPDDSRFVFDALRSKNLFQWNAVISSYSRNELYDEVLET 173
Query: 92 FRTML-VDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGLGLDPICCSALVSAYS 150
F M+ D+ PD++TY C I+ACA D G+ VHG V GL D +ALVS Y
Sbjct: 174 FIEMISTTDLLPDHFTYPCVIKACAGMSDVGIGLAVHGLVVKTGLVEDVFVGNALVSFYG 233
Query: 151 KLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSMRL----AGKKPDGF 206
G V +A ++FD + E +LV WNS+I + +G E + M PD
Sbjct: 234 THGFVTDALQLFDIMPERNLVSWNSMIRVFSDNGFSEESFLLLGEMMEENGDGAFMPDVA 293
Query: 207 TLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDSAYRVFCGI 266
TL +L A + +G+ +HG + K LD + + + L+ MYS+C CI +A +F
Sbjct: 294 TLVTVLPVCAREREIGLGKGVHGWAVKLRLDKELVLNNALMDMYSKCGCITNAQMIFKMN 353
Query: 267 FNPDLVTWSALISGYSQCGEYQKALLFFRKL--NMKSKKLDSVLIATVLASITQMANVLP 324
N ++V+W+ ++ G+S G+ R++ + K D V I + + + LP
Sbjct: 354 NNKNVVSWNTMVGGFSAEGDTHGTFDVLRQMLAGGEDVKADEVTILNAV-PVCFHESFLP 412
Query: 325 GC-EIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSMILAYG 383
E+H Y L+ + V++A + Y+KCG L VF + + + S+N++I G
Sbjct: 413 SLKELHCYSLKQEFVYNELVANAFVASYAKCGSLSYAQRVFHGIRSKTVNSWNALI---G 469
Query: 384 LHGCASQAFTMFDEMLQ---KGLVPDEGTFSALLSACCHAGLVKDGREL 429
H ++ D LQ GL+PD T +LLSAC ++ G+E+
Sbjct: 470 GHAQSNDPRLSLDAHLQMKISGLLPDSFTVCSLLSACSKLKSLRLGKEV 518
Score = 160 bits (405), Expect = 2e-39
Identities = 105/410 (25%), Positives = 201/410 (48%), Gaps = 6/410 (1%)
Query: 25 LHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMIRAFAKARR 84
+H ++KT L +D F ++ Y + + A +FD R++ WNSMIR F+
Sbjct: 209 VHGLVVKTGLVEDVFVGNALVSFYGTHGFVTDALQLFDIMPERNLVSWNSMIRVFSDNGF 268
Query: 85 FSNAISLFRTMLVDD----IRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGLGLDPI 140
+ L M+ ++ PD T + CA + G+ + VHG AV + L + +
Sbjct: 269 SEESFLLLGEMMEENGDGAFMPDVATLVTVLPVCAREREIGLGKGVHGWAVKLRLDKELV 328
Query: 141 CCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSMRLAG 200
+AL+ YSK G + A+ +F ++V WN+++ + G + M G
Sbjct: 329 LNNALMDMYSKCGCITNAQMIFKMNNNKNVVSWNTMVGGFSAEGDTHGTFDVLRQMLAGG 388
Query: 201 K--KPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDS 258
+ K D T+ + S L +ELH S K + V + V+ Y++C +
Sbjct: 389 EDVKADEVTILNAVPVCFHESFLPSLKELHCYSLKQEFVYNELVANAFVASYAKCGSLSY 448
Query: 259 AYRVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLASITQ 318
A RVF GI + + +W+ALI G++Q + + +L ++ + DS + ++L++ ++
Sbjct: 449 AQRVFHGIRSKTVNSWNALIGGHAQSNDPRLSLDAHLQMKISGLLPDSFTVCSLLSACSK 508
Query: 319 MANVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSM 378
+ ++ G E+HG+++R+ LE D+ V +++ +Y CG L +F M +++++S+N++
Sbjct: 509 LKSLRLGKEVHGFIIRNWLERDLFVYLSVLSLYIHCGELCTVQALFDAMEDKSLVSWNTV 568
Query: 379 ILAYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGRE 428
I Y +G +A +F +M+ G+ + + AC ++ GRE
Sbjct: 569 ITGYLQNGFPDRALGVFRQMVLYGIQLCGISMMPVFGACSLLPSLRLGRE 618
Score = 132 bits (332), Expect = 5e-31
Identities = 90/336 (26%), Positives = 168/336 (49%), Gaps = 9/336 (2%)
Query: 111 IRACADSFDFGMLRVVHGSAV-SVGLGLDPICCSALVSAYSKLGVVHEARRVFDGIVEPD 169
++A D M R +H S L D + C+ +++ Y+ G ++R VFD + +
Sbjct: 91 LQASGKRKDIEMGRKIHQLVSGSTRLRNDDVLCTRIITMYAMCGSPDDSRFVFDALRSKN 150
Query: 170 LVLWNSLISAYGGSGMWEIGIQMFSSM-RLAGKKPDGFTLAGLLGGIADSSLLSIGQELH 228
L WN++IS+Y + +++ ++ F M PD FT ++ A S + IG +H
Sbjct: 151 LFQWNAVISSYSRNELYDEVLETFIEMISTTDLLPDHFTYPCVIKACAGMSDVGIGLAVH 210
Query: 229 GLSQKSGLDSDCHVGSLLVSMYSRCKCIDSAYRVFCGIFNPDLVTWSALISGYSQCGEYQ 288
GL K+GL D VG+ LVS Y + A ++F + +LV+W+++I +S G +
Sbjct: 211 GLVVKTGLVEDVFVGNALVSFYGTHGFVTDALQLFDIMPERNLVSWNSMIRVFSDNGFSE 270
Query: 289 KALLFFRKLNMKSK----KLDSVLIATVLASITQMANVLPGCEIHGYVLRHGLESDVKVS 344
++ L ++ ++ D + TVL + + G +HG+ ++ L+ ++ ++
Sbjct: 271 ESFLLLGEMMEENGDGAFMPDVATLVTVLPVCAREREIGLGKGVHGWAVKLRLDKELVLN 330
Query: 345 SALIDMYSKCGFLHLGTCVFRIMLERNIISYNSMILAYGLHGCASQAFTMFDEMLQKG-- 402
+AL+DMYSKCG + +F++ +N++S+N+M+ + G F + +ML G
Sbjct: 331 NALMDMYSKCGCITNAQMIFKMNNNKNVVSWNTMVGGFSAEGDTHGTFDVLRQMLAGGED 390
Query: 403 LVPDEGTFSALLSACCHAGLVKDGRELF-WRMKDEF 437
+ DE T + C H + +EL + +K EF
Sbjct: 391 VKADEVTILNAVPVCFHESFLPSLKELHCYSLKQEF 426
Score = 125 bits (314), Expect = 6e-29
Identities = 101/450 (22%), Positives = 197/450 (43%), Gaps = 38/450 (8%)
Query: 22 VKQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMIRAFAK 81
+K+LH LK + A + YA ++YA VF +++V WN++I A+
Sbjct: 414 LKELHCYSLKQEFVYNELVANAFVASYAKCGSLSYAQRVFHGIRSKTVNSWNALIGGHAQ 473
Query: 82 ARRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGLGLDPIC 141
+ ++ M + + PD++T + AC+ + + VHG + L D
Sbjct: 474 SNDPRLSLDAHLQMKISGLLPDSFTVCSLLSACSKLKSLRLGKEVHGFIIRNWLERDLFV 533
Query: 142 CSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSMRLAGK 201
+++S Y G + + +FD + + LV WN++I+ Y +G + + +F M L G
Sbjct: 534 YLSVLSLYIHCGELCTVQALFDAMEDKSLVSWNTVITGYLQNGFPDRALGVFRQMVLYGI 593
Query: 202 KPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRCKCIDSAYR 261
+ G ++ + G + L +G+E H + K L+ D + L+ MY++ I + +
Sbjct: 594 QLCGISMMPVFGACSLLPSLRLGREAHAYALKHLLEDDAFIACSLIDMYAKNGSITQSSK 653
Query: 262 VFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLASITQMAN 321
VF G+ +W+A+I GY G ++A+ F ++ D + VL +
Sbjct: 654 VFNGLKEKSTASWNAMIMGYGIHGLAKEAIKLFEEMQRTGHNPDDLTFLGVLTACNHSGL 713
Query: 322 VLPGCE-IHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSMIL 380
+ G + GL+ ++K + +IDM + G L R++ E
Sbjct: 714 IHEGLRYLDQMKSSFGLKPNLKHYACVIDMLGRAGQLDK---ALRVVAE----------- 759
Query: 381 AYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGRELFWRMKDEFNIK 440
EM ++ D G + +LLS+C ++ G ++ ++ F ++
Sbjct: 760 ----------------EMSEEA---DVGIWKSLLSSCRIHQNLEMGEKVAAKL---FELE 797
Query: 441 -ARPEHYVYMVKLLGGVGELEEAYNLTQSL 469
+PE+YV + L G+G+ E+ + Q +
Sbjct: 798 PEKPENYVLLSNLYAGLGKWEDVRKVRQRM 827
Score = 98.6 bits (244), Expect = 8e-21
Identities = 67/266 (25%), Positives = 121/266 (45%), Gaps = 3/266 (1%)
Query: 17 KSLLRVKQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMI 76
KSL K++H +++ L +D F ++ LY + +FD +S+ WN++I
Sbjct: 510 KSLRLGKEVHGFIIRNWLERDLFVYLSVLSLYIHCGELCTVQALFDAMEDKSLVSWNTVI 569
Query: 77 RAFAKARRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGLG 136
+ + A+ +FR M++ I+ + AC+ + R H A+ L
Sbjct: 570 TGYLQNGFPDRALGVFRQMVLYGIQLCGISMMPVFGACSLLPSLRLGREAHAYALKHLLE 629
Query: 137 LDPICCSALVSAYSKLGVVHEARRVFDGIVEPDLVLWNSLISAYGGSGMWEIGIQMFSSM 196
D +L+ Y+K G + ++ +VF+G+ E WN++I YG G+ + I++F M
Sbjct: 630 DDAFIACSLIDMYAKNGSITQSSKVFNGLKEKSTASWNAMIMGYGIHGLAKEAIKLFEEM 689
Query: 197 RLAGKKPDGFTLAGLLGGIADSSLLSIG-QELHGLSQKSGLDSDCHVGSLLVSMYSRCKC 255
+ G PD T G+L S L+ G + L + GL + + ++ M R
Sbjct: 690 QRTGHNPDDLTFLGVLTACNHSGLIHEGLRYLDQMKSSFGLKPNLKHYACVIDMLGRAGQ 749
Query: 256 IDSAYRVFCGIFN--PDLVTWSALIS 279
+D A RV + D+ W +L+S
Sbjct: 750 LDKALRVVAEEMSEEADVGIWKSLLS 775
>At3g49710 putative protein
Length = 721
Score = 283 bits (724), Expect = 2e-76
Identities = 183/550 (33%), Positives = 280/550 (50%), Gaps = 10/550 (1%)
Query: 7 WLHSELTNVCKSLLRVKQLHACLLKTHLSKDPFYATQIIRLYAFNNHINYAHHVFDKTST 66
+L + N+ R+ A T + F I++ YA ++ I+ A +FD+
Sbjct: 44 YLSNHFVNLYSKCGRLSYARAAFYSTE-EPNVFSYNVIVKAYAKDSKIHIARQLFDEIPQ 102
Query: 67 RSVFLWNSMIRAFAKARRFSNAISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVV 126
+N++I +A AR A+ LF+ M D +T + I AC D D +++ +
Sbjct: 103 PDTVSYNTLISGYADARETFAAMVLFKRMRKLGFEVDGFTLSGLIAACCDRVD--LIKQL 160
Query: 127 HGSAVSVGLGLDPICCSALVSAYSKLGVVHEARRVFDGIVE-PDLVLWNSLISAYGGSGM 185
H +VS G +A V+ YSK G++ EA VF G+ E D V WNS+I AYG
Sbjct: 161 HCFSVSGGFDSYSSVNNAFVTYYSKGGLLREAVSVFYGMDELRDEVSWNSMIVAYGQHKE 220
Query: 186 WEIGIQMFSSMRLAGKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSL 245
+ ++ M G K D FTLA +L + L G++ HG K+G + HVGS
Sbjct: 221 GAKALALYKEMIFKGFKIDMFTLASVLNALTSLDHLIGGRQFHGKLIKAGFHQNSHVGSG 280
Query: 246 LVSMYSRCKCIDSAY---RVFCGIFNPDLVTWSALISGYSQCGEY-QKALLFFRKLNMKS 301
L+ YS+C D Y +VF I +PDLV W+ +ISGYS E ++A+ FR++
Sbjct: 281 LIDFYSKCGGCDGMYDSEKVFQEILSPDLVVWNTMISGYSMNEELSEEAVKSFRQMQRIG 340
Query: 302 KKLDSVLIATVLASITQMANVLPGCEIHGYVLRHGLESD-VKVSSALIDMYSKCGFLHLG 360
+ D V ++ + +++ +IHG ++ + S+ + V++ALI +Y K G L
Sbjct: 341 HRPDDCSFVCVTSACSNLSSPSQCKQIHGLAIKSHIPSNRISVNNALISLYYKSGNLQDA 400
Query: 361 TCVFRIMLERNIISYNSMILAYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHA 420
VF M E N +S+N MI Y HG ++A ++ ML G+ P++ TF A+LSAC H
Sbjct: 401 RWVFDRMPELNAVSFNCMIKGYAQHGHGTEALLLYQRMLDSGIAPNKITFVAVLSACAHC 460
Query: 421 GLVKDGRELFWRMKDEFNIKARPEHYVYMVKLLGGVGELEEAYNLTQSLPKPVDKAILGA 480
G V +G+E F MK+ F I+ EHY M+ LLG G+LEEA ++P A
Sbjct: 461 GKVDEGQEYFNTMKETFKIEPEAEHYSCMIDLLGRAGKLEEAERFIDAMPYKPGSVAWAA 520
Query: 481 LLSCCDSYGNSELAETVAQQIFKSNPADNVYRVMLSNIYAGDGRWDDVKKLRDKMVGGQ- 539
LL C + N LAE A ++ P VML+N+YA +W+++ +R M G +
Sbjct: 521 LLGACRKHKNMALAERAANELMVMQPLAATPYVMLANMYADARKWEEMASVRKSMRGKRI 580
Query: 540 KKMRGVSWIE 549
+K G SWIE
Sbjct: 581 RKKPGCSWIE 590
Score = 89.4 bits (220), Expect = 5e-18
Identities = 67/255 (26%), Positives = 124/255 (48%), Gaps = 35/255 (13%)
Query: 211 LLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSMYSRC----------------- 253
LL +A+ L + G+ LH L KS + S ++ + V++YS+C
Sbjct: 15 LLKSVAERDLFT-GKSLHALYVKSIVASSTYLSNHFVNLYSKCGRLSYARAAFYSTEEPN 73
Query: 254 --------------KCIDSAYRVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNM 299
I A ++F I PD V+++ LISGY+ E A++ F+++
Sbjct: 74 VFSYNVIVKAYAKDSKIHIARQLFDEIPQPDTVSYNTLISGYADARETFAAMVLFKRMRK 133
Query: 300 KSKKLDSVLIATVLASITQMANVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHL 359
++D ++ ++A+ +++ ++H + + G +S V++A + YSK G L
Sbjct: 134 LGFEVDGFTLSGLIAACCDRVDLIK--QLHCFSVSGGFDSYSSVNNAFVTYYSKGGLLRE 191
Query: 360 GTCVFRIMLE-RNIISYNSMILAYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACC 418
VF M E R+ +S+NSMI+AYG H ++A ++ EM+ KG D T +++L+A
Sbjct: 192 AVSVFYGMDELRDEVSWNSMIVAYGQHKEGAKALALYKEMIFKGFKIDMFTLASVLNALT 251
Query: 419 HAGLVKDGRELFWRM 433
+ GR+ ++
Sbjct: 252 SLDHLIGGRQFHGKL 266
Score = 49.3 bits (116), Expect = 6e-06
Identities = 32/114 (28%), Positives = 53/114 (46%), Gaps = 4/114 (3%)
Query: 321 NVLPGCEIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSMIL 380
++ G +H ++ + S +S+ +++YSKCG L F E N+ SYN ++
Sbjct: 23 DLFTGKSLHALYVKSIVASSTYLSNHFVNLYSKCGRLSYARAAFYSTEEPNVFSYNVIVK 82
Query: 381 AYGLHGCASQAFTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGRELFWRMK 434
AY A +FDE+ Q PD +++ L+S A LF RM+
Sbjct: 83 AYAKDSKIHIARQLFDEIPQ----PDTVSYNTLISGYADARETFAAMVLFKRMR 132
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.138 0.425
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,168,717
Number of Sequences: 26719
Number of extensions: 514782
Number of successful extensions: 9843
Number of sequences better than 10.0: 448
Number of HSP's better than 10.0 without gapping: 424
Number of HSP's successfully gapped in prelim test: 24
Number of HSP's that attempted gapping in prelim test: 1452
Number of HSP's gapped (non-prelim): 2323
length of query: 553
length of database: 11,318,596
effective HSP length: 104
effective length of query: 449
effective length of database: 8,539,820
effective search space: 3834379180
effective search space used: 3834379180
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC140849.2


