

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140772.6 + phase: 0 /partial
(40 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
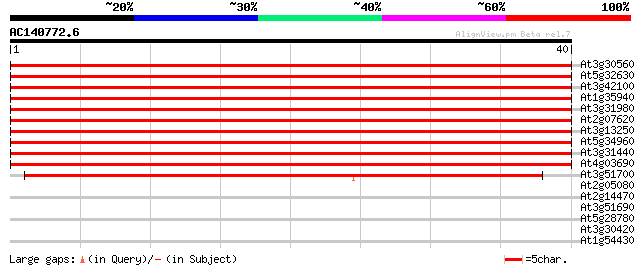
Score E
Sequences producing significant alignments: (bits) Value
At3g30560 hypothetical protein 50 2e-07
At5g32630 putative protein 49 5e-07
At3g42100 putative protein 47 1e-06
At1g35940 hypothetical protein 46 3e-06
At3g31980 hypothetical protein 45 5e-06
At2g07620 putative helicase 45 5e-06
At3g13250 hypothetical protein 45 7e-06
At5g34960 putative protein 45 9e-06
At3g31440 hypothetical protein 44 2e-05
At4g03690 hypothetical protein 41 1e-04
At3g51700 unknown protein 40 3e-04
At2g05080 putative helicase 36 0.003
At2g14470 pseudogene 35 0.006
At3g51690 putative protein 34 0.013
At5g28780 putative protein 33 0.028
At3g30420 hypothetical protein 31 0.11
At1g54430 hypothetical protein 31 0.11
>At3g30560 hypothetical protein
Length = 1473
Score = 50.1 bits (118), Expect = 2e-07
Identities = 21/40 (52%), Positives = 30/40 (74%)
Query: 1 LYVTISRVTSRGGLKILINDDDGDDTNVASSVIYREVFRN 40
LYV +SRV S+GGLK+LI D G N ++V+++E+FRN
Sbjct: 1433 LYVAMSRVKSKGGLKVLITDSKGKQKNETTNVVFKEIFRN 1472
>At5g32630 putative protein
Length = 856
Score = 48.9 bits (115), Expect = 5e-07
Identities = 22/40 (55%), Positives = 31/40 (77%)
Query: 1 LYVTISRVTSRGGLKILINDDDGDDTNVASSVIYREVFRN 40
LYV +SRVTS+ GLKILI D DGD ++V+++E+F+N
Sbjct: 815 LYVALSRVTSKSGLKILILDKDGDIQKQTTNVVFKELFQN 854
>At3g42100 putative protein
Length = 1752
Score = 47.4 bits (111), Expect = 1e-06
Identities = 22/40 (55%), Positives = 31/40 (77%)
Query: 1 LYVTISRVTSRGGLKILINDDDGDDTNVASSVIYREVFRN 40
LYV +SRVTS+ GLKILI D DG+ ++V+++EVF+N
Sbjct: 1711 LYVALSRVTSKKGLKILILDKDGNMQKQTTNVVFKEVFQN 1750
>At1g35940 hypothetical protein
Length = 1678
Score = 46.2 bits (108), Expect = 3e-06
Identities = 22/40 (55%), Positives = 30/40 (75%)
Query: 1 LYVTISRVTSRGGLKILINDDDGDDTNVASSVIYREVFRN 40
LYV +SRVTS+ GLKILI D DG ++V+++EVF+N
Sbjct: 1637 LYVALSRVTSKKGLKILILDKDGKLQKQTTNVVFKEVFQN 1676
>At3g31980 hypothetical protein
Length = 1099
Score = 45.4 bits (106), Expect = 5e-06
Identities = 19/40 (47%), Positives = 31/40 (77%)
Query: 1 LYVTISRVTSRGGLKILINDDDGDDTNVASSVIYREVFRN 40
LYV +SRVTS+ GLK+LI D +G+ + +V+++E+F+N
Sbjct: 1057 LYVALSRVTSKKGLKVLIVDKEGNTQSQTMNVVFKEIFQN 1096
>At2g07620 putative helicase
Length = 1241
Score = 45.4 bits (106), Expect = 5e-06
Identities = 19/40 (47%), Positives = 31/40 (77%)
Query: 1 LYVTISRVTSRGGLKILINDDDGDDTNVASSVIYREVFRN 40
LYV +SRVTS+ GLK+LI D +G+ + +V+++E+F+N
Sbjct: 1199 LYVALSRVTSKKGLKVLIVDKEGNTQSQTMNVVFKEIFQN 1238
>At3g13250 hypothetical protein
Length = 1419
Score = 45.1 bits (105), Expect = 7e-06
Identities = 21/40 (52%), Positives = 30/40 (74%)
Query: 1 LYVTISRVTSRGGLKILINDDDGDDTNVASSVIYREVFRN 40
LYV +SRVTS+ GLKILI D +G ++V+++EVF+N
Sbjct: 1379 LYVALSRVTSKTGLKILILDKEGKIQKQTTNVVFKEVFQN 1418
>At5g34960 putative protein
Length = 1033
Score = 44.7 bits (104), Expect = 9e-06
Identities = 21/40 (52%), Positives = 29/40 (72%)
Query: 1 LYVTISRVTSRGGLKILINDDDGDDTNVASSVIYREVFRN 40
LYV +SRVTS+ GLK LI D DG ++V+++EVF+N
Sbjct: 992 LYVALSRVTSKKGLKFLILDKDGKLQKQTTNVVFKEVFQN 1031
>At3g31440 hypothetical protein
Length = 536
Score = 43.9 bits (102), Expect = 2e-05
Identities = 20/40 (50%), Positives = 30/40 (75%)
Query: 1 LYVTISRVTSRGGLKILINDDDGDDTNVASSVIYREVFRN 40
LYV +SRVTS+ GLKILI D +G ++++++EVF+N
Sbjct: 495 LYVALSRVTSKKGLKILILDKNGKLQKQTTNIVFKEVFQN 534
>At4g03690 hypothetical protein
Length = 570
Score = 41.2 bits (95), Expect = 1e-04
Identities = 18/40 (45%), Positives = 27/40 (67%)
Query: 1 LYVTISRVTSRGGLKILINDDDGDDTNVASSVIYREVFRN 40
LYV +SRV S+ LK+LI D G ++VI++E+F+N
Sbjct: 529 LYVAMSRVKSKARLKVLITDSKGKQKKETTNVIFKEIFQN 568
>At3g51700 unknown protein
Length = 344
Score = 39.7 bits (91), Expect = 3e-04
Identities = 19/38 (50%), Positives = 26/38 (68%), Gaps = 1/38 (2%)
Query: 2 YVTISRVTSRGGLKILINDDDG-DDTNVASSVIYREVF 38
YV IS+V SR GLK+LI D DG D +V+++E+F
Sbjct: 295 YVAISKVKSRAGLKVLITDKDGKPDQEETKNVVFKELF 332
>At2g05080 putative helicase
Length = 1219
Score = 36.2 bits (82), Expect = 0.003
Identities = 17/23 (73%), Positives = 20/23 (86%)
Query: 1 LYVTISRVTSRGGLKILINDDDG 23
LYV ISRVTS+ GLKILI +D+G
Sbjct: 1158 LYVAISRVTSKSGLKILIVNDEG 1180
>At2g14470 pseudogene
Length = 1265
Score = 35.4 bits (80), Expect = 0.006
Identities = 17/23 (73%), Positives = 19/23 (81%)
Query: 1 LYVTISRVTSRGGLKILINDDDG 23
LYV +SRVTS+ GLKILI D DG
Sbjct: 1241 LYVALSRVTSKKGLKILILDKDG 1263
>At3g51690 putative protein
Length = 374
Score = 34.3 bits (77), Expect = 0.013
Identities = 16/38 (42%), Positives = 23/38 (60%)
Query: 1 LYVTISRVTSRGGLKILINDDDGDDTNVASSVIYREVF 38
++V IS+V SR GLK+LI D DG+ A + + F
Sbjct: 267 MFVAISKVKSRAGLKVLITDKDGNPQEEAKNYPFTLAF 304
>At5g28780 putative protein
Length = 337
Score = 33.1 bits (74), Expect = 0.028
Identities = 17/40 (42%), Positives = 25/40 (62%), Gaps = 2/40 (5%)
Query: 1 LYVTISRVTSRGGLKILINDDDGDDTNVASSVIYREVFRN 40
LYV +SRVTS GL IL DD +D +++Y+E + +
Sbjct: 292 LYVALSRVTSPIGLTILHGDDQKNDE--VKNIVYKEFYND 329
>At3g30420 hypothetical protein
Length = 837
Score = 31.2 bits (69), Expect = 0.11
Identities = 16/38 (42%), Positives = 24/38 (63%), Gaps = 1/38 (2%)
Query: 1 LYVTISRVTSRGGLKILINDDDGDDTNVASSVIYREVF 38
LYV +SRVTS GL +L + + ++++YREVF
Sbjct: 790 LYVALSRVTSPKGLTVL-DTSKKKEGKYVTNIVYREVF 826
>At1g54430 hypothetical protein
Length = 1639
Score = 31.2 bits (69), Expect = 0.11
Identities = 16/38 (42%), Positives = 24/38 (63%), Gaps = 1/38 (2%)
Query: 1 LYVTISRVTSRGGLKILINDDDGDDTNVASSVIYREVF 38
LYV +SRVTS GL +L + + ++++YREVF
Sbjct: 1592 LYVALSRVTSPKGLTVL-DTSKKKEGKYVTNIVYREVF 1628
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.139 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 885,662
Number of Sequences: 26719
Number of extensions: 21366
Number of successful extensions: 101
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 15
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 86
Number of HSP's gapped (non-prelim): 17
length of query: 40
length of database: 11,318,596
effective HSP length: 16
effective length of query: 24
effective length of database: 10,891,092
effective search space: 261386208
effective search space used: 261386208
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 53 (25.0 bits)
Medicago: description of AC140772.6


