

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140721.10 + phase: 0 /pseudo
(329 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
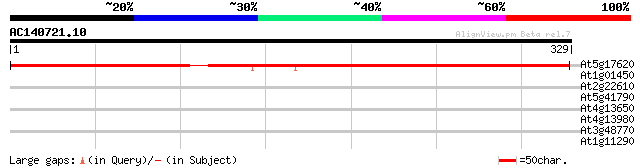
Score E
Sequences producing significant alignments: (bits) Value
At5g17620 unknown protein 402 e-112
At1g01450 hypothetical protein 29 3.1
At2g22610 putative kinesin heavy chain 29 4.0
At5g41790 myosin heavy chain-like protein 28 5.2
At4g13650 unknown protein 28 5.2
At4g13980 unknown protein 28 9.0
At3g48770 hypothetical protein 28 9.0
At1g11290 hypothetical protein 28 9.0
>At5g17620 unknown protein
Length = 329
Score = 402 bits (1032), Expect = e-112
Identities = 222/339 (65%), Positives = 257/339 (75%), Gaps = 21/339 (6%)
Query: 1 MASKQMEVIQKKLGMLNYPRANASAQSLLFAGMERYALFEWLFFRLLGDKSPFSQQNLQG 60
MA+KQME IQKKL +L+YPRANA AQSLLFAGMERYAL EWLFF+LLGDKSPFSQQNLQG
Sbjct: 1 MAAKQMEEIQKKLRLLSYPRANAPAQSLLFAGMERYALLEWLFFKLLGDKSPFSQQNLQG 60
Query: 61 DALDRDEETARIQYLAEIAKFLGITTTVDTDAIQTHLLNDDLFFFL*LW*CRYGIGINLV 120
DA RDEET RIQYLAEIAKFLGIT TVD +AIQ H +D L N+V
Sbjct: 61 DAGVRDEETVRIQYLAEIAKFLGITPTVDIEAIQGHGTYEDRMEML----------RNIV 110
Query: 121 CIFVSIVLTSR*PRTYT**IL--------LQKNRHKYFLKNVNCFPQMFRFSPY--IP*V 170
+ + + + + + + + + F + FP + +P V
Sbjct: 111 DLVEASLFSDNQEWSIDEQVAKDIQLIDAIAERQSLIFSEECKLFPADVQIQSIYPLPDV 170
Query: 171 AELESKLTEQSKILLNLQQKVDDLASKHAYNPDEEYTEVESQLRAHLESFLETARTFNMI 230
+ELE+KL+EQ+KIL NLQQKVDDLA+KHAYNPDEEYTEVESQLRA LESFLETAR FN I
Sbjct: 171 SELETKLSEQAKILSNLQQKVDDLAAKHAYNPDEEYTEVESQLRARLESFLETARAFNTI 230
Query: 231 YTKEIRPWTHMMEVPQLHGFGPAANRLLEAYKMLLKFLGNLQNLRDSHAALAFGSSET-S 289
YTKEIRPWTHMMEVPQLHGFGPAANRLLEAY MLLKFLGNL+NLRDSHAAL+ GSS T +
Sbjct: 231 YTKEIRPWTHMMEVPQLHGFGPAANRLLEAYNMLLKFLGNLKNLRDSHAALSIGSSGTVA 290
Query: 290 GGPSSVSRIISECESEMTVINRDLGILSASIAREHGEKM 328
G PSSV+RI+S+CE+ +TV+NRDLGILSASIARE GE++
Sbjct: 291 GEPSSVTRIVSDCEAALTVLNRDLGILSASIAREQGERL 329
>At1g01450 hypothetical protein
Length = 470
Score = 29.3 bits (64), Expect = 3.1
Identities = 16/52 (30%), Positives = 25/52 (47%)
Query: 15 MLNYPRANASAQSLLFAGMERYALFEWLFFRLLGDKSPFSQQNLQGDALDRD 66
+L + +A SL ++ F + F LL K PF +LQGD + R+
Sbjct: 207 VLEEQEQSGTAGSLKYSDKSDVYSFGMVSFELLTGKVPFEDSHLQGDKMSRN 258
>At2g22610 putative kinesin heavy chain
Length = 1068
Score = 28.9 bits (63), Expect = 4.0
Identities = 16/73 (21%), Positives = 40/73 (53%), Gaps = 1/73 (1%)
Query: 171 AELESKLTEQSKILLNLQQKVDDLASKHAYNPDEEYTEVESQLRAHLESFLETARTFNMI 230
A+L+ +L + +I NLQQKV +L K + + +Q LE+ L+ + +++
Sbjct: 806 AQLQERLKSRDEICSNLQQKVKELECKLRERHQSD-SAANNQKVKDLENNLKESEGSSLV 864
Query: 231 YTKEIRPWTHMME 243
+ ++++ + + ++
Sbjct: 865 WQQKVKDYENKLK 877
>At5g41790 myosin heavy chain-like protein
Length = 1305
Score = 28.5 bits (62), Expect = 5.2
Identities = 35/166 (21%), Positives = 68/166 (40%), Gaps = 25/166 (15%)
Query: 172 ELESKLTEQSKILLNLQQKVDDLASKHAYNPDEEYTEVES--QLRAHLESFLETARTFNM 229
E+ KL + + L ++ +L +H E + V+S Q A ++ L+ A
Sbjct: 369 EIMDKLEQAQNTIKELMDELGELKDRHKEKESELSSLVKSADQQVADMKQSLDNAEEEKK 428
Query: 230 IYTKEIRPWTHMMEVPQLHGFGPAANRLLEAYKMLLKFLGNLQNLRDSHAALAFGSS--- 286
+ ++ I +N + EA K + + + + L++SH +
Sbjct: 429 MLSQRILD---------------ISNEIQEAQKTIQEHMSESEQLKESHGVKERELTGLR 473
Query: 287 ---ETSGGPSSVSRIISECESEMTVINRDLGILSASIAREHGEKMS 329
ET SS +SE E+++ ++ + + LSAS+ EK S
Sbjct: 474 DIHETHQRESSTR--LSELETQLKLLEQRVVDLSASLNAAEEEKKS 517
>At4g13650 unknown protein
Length = 1024
Score = 28.5 bits (62), Expect = 5.2
Identities = 20/93 (21%), Positives = 39/93 (41%), Gaps = 8/93 (8%)
Query: 189 QKVDDLASKHAYNPDEEYTEVESQLRAHLESFLETARTFNMIYTKEIRPWTHMMEVPQLH 248
++V + +K Y+ + E + A S + + F + TK W ++ H
Sbjct: 672 KQVHAVITKTGYDSETEVCNALISMYAKCGSISDAEKQFLEVSTKNEVSWNAIINAYSKH 731
Query: 249 GFGPAANRLLEAYKMLLKFLGNLQNLRDSHAAL 281
GFG A L+++ ++ N+R +H L
Sbjct: 732 GFGSEA---LDSFDQMIH-----SNVRPNHVTL 756
>At4g13980 unknown protein
Length = 466
Score = 27.7 bits (60), Expect = 9.0
Identities = 35/154 (22%), Positives = 61/154 (38%), Gaps = 15/154 (9%)
Query: 176 KLTEQSKILLNLQQKVDDLASKHAYNPDEEYTEVESQLRAHLESFLETARTFNMIYTKEI 235
K ++K+L QQKV +KH + E+ + + L +FLETA N + K
Sbjct: 145 KAAIEAKLLKFKQQKV---VAKHQFEEMTEHVDDMENRQKKLLNFLETAIR-NPTFVKNF 200
Query: 236 RPWTHMMEVPQLHGFGPAANRLLEAYKMLLKFLGNLQNLRDSHAALAFGSSETSGGPSSV 295
+++ + RL E + + DSH + GSS G
Sbjct: 201 GKKVEQLDISAYN----KKRRLPEVEQ-------SKPPSEDSHLDNSSGSSRRESGNIFH 249
Query: 296 SRIISECESEMTVINRDLGILSASIAREHGEKMS 329
++ E++ + D+ ++S SI + E S
Sbjct: 250 QNFSNKLRLELSPADSDMNMVSHSIQSSNEEGAS 283
>At3g48770 hypothetical protein
Length = 1899
Score = 27.7 bits (60), Expect = 9.0
Identities = 22/77 (28%), Positives = 35/77 (44%), Gaps = 5/77 (6%)
Query: 250 FGPAANRLLEAYKMLLKFLGNLQNLRDSHAALAFGSSETSGGPS-SVSRIISECESEMTV 308
FG N +E Y L K+ +N SH AF S G + +++SE S + V
Sbjct: 1587 FGVRINPSIEDYCELWKYWEKTKNRLSSHECCAFWSFVVRHGDTVKAEKLLSESFSRLPV 1646
Query: 309 ----INRDLGILSASIA 321
N + G++ +SI+
Sbjct: 1647 HSPDCNNNDGVMLSSIS 1663
>At1g11290 hypothetical protein
Length = 809
Score = 27.7 bits (60), Expect = 9.0
Identities = 12/33 (36%), Positives = 18/33 (54%)
Query: 227 FNMIYTKEIRPWTHMMEVPQLHGFGPAANRLLE 259
F+M+ + + W M++ HGFG AA L E
Sbjct: 496 FDMMSERHVTTWNAMIDGYGTHGFGKAALELFE 528
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.332 0.143 0.430
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,315,402
Number of Sequences: 26719
Number of extensions: 236264
Number of successful extensions: 798
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 790
Number of HSP's gapped (non-prelim): 12
length of query: 329
length of database: 11,318,596
effective HSP length: 100
effective length of query: 229
effective length of database: 8,646,696
effective search space: 1980093384
effective search space used: 1980093384
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC140721.10


