

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140720.4 + phase: 0
(653 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
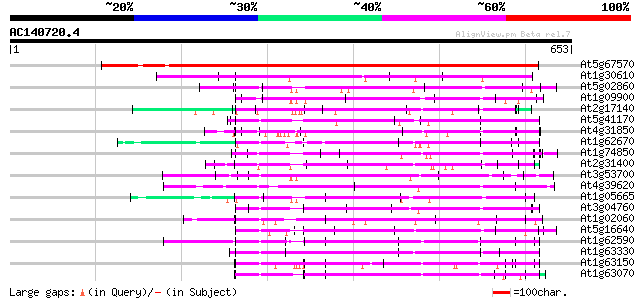
Score E
Sequences producing significant alignments: (bits) Value
At5g67570 putative protein 626 e-179
At1g30610 hypothetical protein 369 e-102
At5g02860 putative protein 124 1e-28
At1g09900 hypothetical protein 107 2e-23
At2g17140 hypothetical protein 106 4e-23
At5g41170 salt-inducible protein-like 102 7e-22
At4g31850 putative protein 100 3e-21
At1g62670 PPR-repeat protein 100 3e-21
At1g74850 hypothetical protein 98 2e-20
At2g31400 unknown protein 97 4e-20
At3g53700 putative protein 96 5e-20
At4g39620 putative protein 94 2e-19
At1g05665 putative protein 94 2e-19
At3g04760 unknown protein 92 7e-19
At1g02060 unknown protein 92 9e-19
At5g16640 putative protein 92 1e-18
At1g62590 putative membrane-associated salt-inducible protein (A... 92 1e-18
At1g63330 unknown protein 91 2e-18
At1g63150 unknown protein 91 2e-18
At1g63070 unknown protein 91 2e-18
>At5g67570 putative protein
Length = 505
Score = 626 bits (1614), Expect = e-179
Identities = 301/512 (58%), Positives = 396/512 (76%), Gaps = 11/512 (2%)
Query: 108 LIGKPWEGVEKVDFLERIKV--NYEHRGEKLKRESLIELKEMFRERKMDELKWVFEDDIE 165
++G PWEG+E+V E + E +LK+E+L ELK++ + +L+WV +DD++
Sbjct: 1 MVGNPWEGIERVKLKELVSGVRREEVSAGELKKENLKELKKILEK----DLRWVLDDDVD 56
Query: 166 INEVWFDENNNGKRKKTSKRSEVQVVRFLVDRLCDKEIRAKDWKFSRLMKLSGLSFTEGQ 225
+ E D+ + ++ R+E + VR LVDRL +EI K WKF R+M SGL FTE Q
Sbjct: 57 VEEFDLDKEFDPAKRW---RNEGEAVRVLVDRLSGREINEKHWKFVRMMNQSGLQFTEDQ 113
Query: 226 LMMIIEMLGVKRCWKQALSVVQWVYNSKDHRKFQSRFVYTKLLAVLGKARRPKEALQIFN 285
++ I++ LG K+ WKQA +VV WVY+ K + +SRFVYTKLL+VLG ARRP+EALQIFN
Sbjct: 114 MLKIVDRLGRKQSWKQASAVVHWVYSDKKRKHLRSRFVYTKLLSVLGFARRPQEALQIFN 173
Query: 286 MMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETL-KYMYRKNWDPTLEP 344
MLG+ ++YPDMAAYH IAVTLGQAGLLKELL ++E MRQKP L K + +KNWDP LEP
Sbjct: 174 QMLGDRQLYPDMAAYHCIAVTLGQAGLLKELLKVIERMRQKPTKLTKNLRQKNWDPVLEP 233
Query: 345 DVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELF 404
D+V+YNA+LNACVP+ QWK VSWVF ++RK+ L+PNGATYGLAMEVML+SG +D VH+ F
Sbjct: 234 DLVVYNAILNACVPTLQWKAVSWVFVELRKNGLRPNGATYGLAMEVMLESGKFDRVHDFF 293
Query: 405 EKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNC 464
KM+ +GE P+A+TYKV+VR W+EGK++EAV+AVRDME++GV+GT SVYYELACCLCN
Sbjct: 294 RKMKSSGEAPKAITYKVLVRALWREGKIEEAVEAVRDMEQKGVIGTGSVYYELACCLCNN 353
Query: 465 GRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDHCAPNVGTV 524
GRW DA LEV ++KRL + +PLE+TFTG+I +S++GGH+DDC+ IF+YM+D C PN+GT
Sbjct: 354 GRWCDAMLEVGRMKRLENCRPLEITFTGLIAASLNGGHVDDCMAIFQYMKDKCDPNIGTA 413
Query: 525 NTMLKVYSQNDMFSTAKVLFEEVKVAK-SDLRPDAYTYNLMLEASSRGHQWEYFEHVYKE 583
N MLKVY +NDMFS AK LFEE+ K + L P+ YTY+ MLEAS+R QWEYFEHVY+
Sbjct: 414 NMMLKVYGRNDMFSEAKELFEEIVSRKETHLVPNEYTYSFMLEASARSLQWEYFEHVYQT 473
Query: 584 MILSGYHLDQNKHLPLLVKASRAGKVDNILVI 615
M+LSGY +DQ KH +L++ASRAGKV N+ ++
Sbjct: 474 MVLSGYQMDQTKHASMLIEASRAGKVTNLCLV 505
>At1g30610 hypothetical protein
Length = 1006
Score = 369 bits (947), Expect = e-102
Identities = 195/477 (40%), Positives = 289/477 (59%), Gaps = 42/477 (8%)
Query: 172 DENNNGKRKKTSKRSEVQV-VRFLVDRLCDKEIRAKDWKFSRLMKLSGLSFTEGQLMMII 230
DE+++ K + R E++ + L L +I +W+FS+ ++ + + +T+ +M +I
Sbjct: 417 DESSDIVDKPATSRVEMEDRIEKLAKVLNGADINMPEWQFSKAIRSAKIRYTDYTVMRLI 476
Query: 231 EMLGVKRCWKQALSVVQWVYNSKDHRKFQSRFVYTKLLAVLGKARRPKEALQIFNMMLGN 290
LG W++ L V++W+ ++ + R +YT L VLGK+RRP EAL +F+ ML
Sbjct: 477 HFLGKLGNWRRVLQVIEWLQRQDRYKSNKIRIIYTTALNVLGKSRRPVEALNVFHAMLLQ 536
Query: 291 IRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKP-ETLKYMYRKNWDPTLEPDVVIY 349
I YPDM AY SIAVTLGQAG +KEL +++ MR P + K + WDP LEPDVV+Y
Sbjct: 537 ISSYPDMVAYRSIAVTLGQAGHIKELFYVIDTMRSPPKKKFKPTTLEKWDPRLEPDVVVY 596
Query: 350 NAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQR 409
NAVLNACV KQW+G WV QQ+++ KP+ TYGL MEVML Y+LVHE F KMQ+
Sbjct: 597 NAVLNACVQRKQWEGAFWVLQQLKQRGQKPSPVTYGLIMEVMLACEKYNLVHEFFRKMQK 656
Query: 410 NGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQD 469
+ +P AL Y+V+V T WKEGK DEAV V DME RG++G+A++YY+LA CLC+ GR +
Sbjct: 657 S-SIPNALAYRVLVNTLWKEGKSDEAVHTVEDMESRGIVGSAALYYDLARCLCSAGRCNE 715
Query: 470 A----------------------------TLEVEKIKRLPHAKPLEVTFTGMIRSSMDGG 501
+++KI R+ + KPL VT+TG+I++ +D G
Sbjct: 716 GLNMVNFVNPVVLKLIENLIYKADLVHTIQFQLKKICRVAN-KPLVVTYTGLIQACVDSG 774
Query: 502 HIDDCICIFEYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKV----------AK 551
+I + IF+ M+ C+PN+ T N MLK Y Q +F A+ LF+++ +
Sbjct: 775 NIKNAAYIFDQMKKVCSPNLVTCNIMLKAYLQGGLFEEARELFQKMSEDGNHIKNSSDFE 834
Query: 552 SDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGK 608
S + PD YT+N ML+ + +W+ F + Y+EM+ GYH + +HL ++++ASRAGK
Sbjct: 835 SRVLPDTYTFNTMLDTCAEQEKWDDFGYAYREMLRHGYHFNAKRHLRMVLEASRAGK 891
Score = 69.7 bits (169), Expect = 5e-12
Identities = 65/286 (22%), Positives = 110/286 (37%), Gaps = 28/286 (9%)
Query: 244 SVVQWVYNSKDHRKFQSRFVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSI 303
++V + + Y L+ L K + EA+ M + A Y+ +
Sbjct: 645 NLVHEFFRKMQKSSIPNALAYRVLVNTLWKEGKSDEAVHTVEDMESR-GIVGSAALYYDL 703
Query: 304 AVTLGQAGLLKELLNIVECMR--------------QKPETLKYMYRKNWDPTLEPDVVIY 349
A L AG E LN+V + T+++ +K +P VV Y
Sbjct: 704 ARCLCSAGRCNEGLNMVNFVNPVVLKLIENLIYKADLVHTIQFQLKKICRVANKPLVVTY 763
Query: 350 NAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQR 409
++ ACV S K +++F QM+K PN T + ++ LQ G ++ ELF+KM
Sbjct: 764 TGLIQACVDSGNIKNAAYIFDQMKKVC-SPNLVTCNIMLKAYLQGGLFEEARELFQKMSE 822
Query: 410 NGE------------VPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYEL 457
+G +P+ T+ M+ T ++ K D+ A R+M R G A + +
Sbjct: 823 DGNHIKNSSDFESRVLPDTYTFNTMLDTCAEQEKWDDFGYAYREMLRHGYHFNAKRHLRM 882
Query: 458 ACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHI 503
G+ + E ++R P + R G HI
Sbjct: 883 VLEASRAGKEEVMEATWEHMRRSNRIPPSPLIKERFFRKLEKGDHI 928
Score = 42.4 bits (98), Expect = 8e-04
Identities = 32/179 (17%), Positives = 76/179 (41%), Gaps = 4/179 (2%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
YT L+ + K A IF+ M P++ + + Q GL +E + + M
Sbjct: 763 YTGLIQACVDSGNIKNAAYIFDQMKKVCS--PNLVTCNIMLKAYLQGGLFEEARELFQKM 820
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
+ +K +++ + PD +N +L+ C ++W + +++M + N
Sbjct: 821 SEDGNHIKNS--SDFESRVLPDTYTFNTMLDTCAEQEKWDDFGYAYREMLRHGYHFNAKR 878
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDM 442
+ + ++G +++ +E M+R+ +P + K ++G A+ ++ D+
Sbjct: 879 HLRMVLEASRAGKEEVMEATWEHMRRSNRIPPSPLIKERFFRKLEKGDHISAISSLADL 937
>At5g02860 putative protein
Length = 819
Score = 124 bits (312), Expect = 1e-28
Identities = 88/399 (22%), Positives = 173/399 (43%), Gaps = 20/399 (5%)
Query: 222 TEGQLMMIIEMLGVKRCWKQALSVVQWVYNSKDHRKFQSRFVYTKLLAVLGKARRPKEAL 281
T +L+ ++ LG + + AL W KD++ V ++++LGK R A
Sbjct: 134 TSSELLAFLKGLGFHKKFDLALRAFDWFMKQKDYQSMLDNSVVAIIISMLGKEGRVSSAA 193
Query: 282 QIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNWDPT 341
+FN + + D+ +Y S+ +G +E +N+ ++K +
Sbjct: 194 NMFNGLQED-GFSLDVYSYTSLISAFANSGRYREAVNV--------------FKKMEEDG 238
Query: 342 LEPDVVIYNAVLNACVP-SKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLV 400
+P ++ YN +LN W ++ + ++M+ + P+ TY + + +
Sbjct: 239 CKPTLITYNVILNVFGKMGTPWNKITSLVEKMKSDGIAPDAYTYNTLITCCKRGSLHQEA 298
Query: 401 HELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACC 460
++FE+M+ G + +TY ++ + K + EA+K + +M G + Y L
Sbjct: 299 AQVFEEMKAAGFSYDKVTYNALLDVYGKSHRPKEAMKVLNEMVLNGFSPSIVTYNSLISA 358
Query: 461 LCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDH-CAP 519
G +A +E++ KP T+T ++ G ++ + IFE M++ C P
Sbjct: 359 YARDGMLDEA-MELKNQMAEKGTKPDVFTYTTLLSGFERAGKVESAMSIFEEMRNAGCKP 417
Query: 520 NVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEH 579
N+ T N +K+Y F+ +F+E+ V L PD T+N +L +
Sbjct: 418 NICTFNAFIKMYGNRGKFTEMMKIFDEINVC--GLSPDIVTWNTLLAVFGQNGMDSEVSG 475
Query: 580 VYKEMILSGYHLDQNKHLPLLVKASRAGKVDNILVIYIR 618
V+KEM +G+ ++ L+ SR G + + +Y R
Sbjct: 476 VFKEMKRAGFVPERETFNTLISAYSRCGSFEQAMTVYRR 514
Score = 92.0 bits (227), Expect = 9e-19
Identities = 93/415 (22%), Positives = 169/415 (40%), Gaps = 57/415 (13%)
Query: 261 RFVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIV 320
+ Y LL V GK+ RPKEA+++ N M+ N P + Y+S+ + G+L E + +
Sbjct: 314 KVTYNALLDVYGKSHRPKEAMKVLNEMVLN-GFSPSIVTYNSLISAYARDGMLDEAMELK 372
Query: 321 ECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPN 380
M +K +PDV Y +L+ + + + +F++MR + KPN
Sbjct: 373 NQMAEKGT--------------KPDVFTYTTLLSGFERAGKVESAMSIFEEMRNAGCKPN 418
Query: 381 GATY--------------------------GLAMEVML---------QSGNYDLVHELFE 405
T+ GL+ +++ Q+G V +F+
Sbjct: 419 ICTFNAFIKMYGNRGKFTEMMKIFDEINVCGLSPDIVTWNTLLAVFGQNGMDSEVSGVFK 478
Query: 406 KMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCG 465
+M+R G VPE T+ ++ + + G ++A+ R M GV S Y + L G
Sbjct: 479 EMKRAGFVPERETFNTLISAYSRCGSFEQAMTVYRRMLDAGVTPDLSTYNTVLAALARGG 538
Query: 466 RWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDH-CAPNVGTV 524
W+ + + +++ KP E+T+ ++ + +G I + E + P +
Sbjct: 539 MWEQSEKVLAEMED-GRCKPNELTYCSLLHAYANGKEIGLMHSLAEEVYSGVIEPRAVLL 597
Query: 525 NTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEM 584
T++ V S+ D+ A+ F E+K + PD T N M+ R V M
Sbjct: 598 KTLVLVCSKCDLLPEAERAFSELK--ERGFSPDITTLNSMVSIYGRRQMVAKANGVLDYM 655
Query: 585 ILSGYHLDQNKHLPLLVKASRA---GKVDNILVIYIRSQCLFYISAHSIISYKNC 636
G+ + L+ SR+ GK + IL + I +++ + Y C
Sbjct: 656 KERGFTPSMATYNSLMYMHSRSADFGKSEEILREILAKGIKPDIISYNTVIYAYC 710
Score = 73.6 bits (179), Expect = 3e-13
Identities = 76/359 (21%), Positives = 142/359 (39%), Gaps = 34/359 (9%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
+ + + G + E ++IF+ + + PD+ ++++ GQ G+ E+ + + M
Sbjct: 422 FNAFIKMYGNRGKFTEMMKIFDE-INVCGLSPDIVTWNTLLAVFGQNGMDSEVSGVFKEM 480
Query: 324 RQK---PETLKY------------------MYRKNWDPTLEPDVVIYNAVLNACVPSKQW 362
++ PE + +YR+ D + PD+ YN VL A W
Sbjct: 481 KRAGFVPERETFNTLISAYSRCGSFEQAMTVYRRMLDAGVTPDLSTYNTVLAALARGGMW 540
Query: 363 KGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVM 422
+ V +M KPN TY + L+H L E++ P A+ K +
Sbjct: 541 EQSEKVLAEMEDGRCKPNELTYCSLLHAYANGKEIGLMHSLAEEVYSGVIEPRAVLLKTL 600
Query: 423 VRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQ-----DATLEVEKI 477
V K + EA +A +++ RG + + GR Q + L+ K
Sbjct: 601 VLVCSKCDLLPEAERAFSELKERGFSPDITTLNSMVSIY---GRRQMVAKANGVLDYMKE 657
Query: 478 KRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDHCAPNVGTVNTMLKVYSQNDMF 537
+ + + M S D G ++ + E + P++ + NT++ Y +N
Sbjct: 658 RGFTPSMATYNSLMYMHSRSADFGKSEE--ILREILAKGIKPDIISYNTVIYAYCRNTRM 715
Query: 538 STAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKH 596
A +F E++ S + PD TYN + + + +E V + MI G +QN +
Sbjct: 716 RDASRIFSEMR--NSGIVPDVITYNTFIGSYAADSMFEEAIGVVRYMIKHGCRPNQNTY 772
Score = 57.4 bits (137), Expect = 2e-08
Identities = 37/190 (19%), Positives = 83/190 (43%), Gaps = 15/190 (7%)
Query: 295 PDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLN 354
PD+ +S+ G+ ++ + +++ M+++ T P + YN+++
Sbjct: 627 PDITTLNSMVSIYGRRQMVAKANGVLDYMKERGFT--------------PSMATYNSLMY 672
Query: 355 ACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVP 414
S + + +++ +KP+ +Y + ++ +F +M+ +G VP
Sbjct: 673 MHSRSADFGKSEEILREILAKGIKPDIISYNTVIYAYCRNTRMRDASRIFSEMRNSGIVP 732
Query: 415 EALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEV 474
+ +TY + ++ + +EA+ VR M + G + Y + C R +A L V
Sbjct: 733 DVITYNTFIGSYAADSMFEEAIGVVRYMIKHGCRPNQNTYNSIVDGYCKLNRKDEAKLFV 792
Query: 475 EKIKRL-PHA 483
E ++ L PHA
Sbjct: 793 EDLRNLDPHA 802
>At1g09900 hypothetical protein
Length = 598
Score = 107 bits (267), Expect = 2e-23
Identities = 89/347 (25%), Positives = 147/347 (41%), Gaps = 28/347 (8%)
Query: 271 LGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETL 330
LGK R+ + L+I L PD+ Y+ + +AG + L++++ M
Sbjct: 150 LGKTRKAAKILEI----LEGSGAVPDVITYNVMISGYCKAGEINNALSVLDRM------- 198
Query: 331 KYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEV 390
++ PDVV YN +L + S + K V +M + P+ TY + +E
Sbjct: 199 ----------SVSPDVVTYNTILRSLCDSGKLKQAMEVLDRMLQRDCYPDVITYTILIEA 248
Query: 391 MLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGT 450
+ +L ++M+ G P+ +TY V+V KEG++DEA+K + DM G
Sbjct: 249 TCRDSGVGHAMKLLDEMRDRGCTPDVVTYNVLVNGICKEGRLDEAIKFLNDMPSSGCQPN 308
Query: 451 ASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIF 510
+ + +C+ GRW DA + + R + P VTF +I G + I I
Sbjct: 309 VITHNIILRSMCSTGRWMDAEKLLADMLRKGFS-PSVVTFNILINFLCRKGLLGRAIDIL 367
Query: 511 EYMQDH-CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASS 569
E M H C PN + N +L + + A E ++ PD TYN ML A
Sbjct: 368 EKMPQHGCQPNSLSYNPLLHGFCKEKKMDRAIEYLE--RMVSRGCYPDIVTYNTMLTALC 425
Query: 570 RGHQWEYFEHVYKEMILSGYH---LDQNKHLPLLVKASRAGKVDNIL 613
+ + E + ++ G + N + L KA + GK +L
Sbjct: 426 KDGKVEDAVEILNQLSSKGCSPVLITYNTVIDGLAKAGKTGKAIKLL 472
Score = 89.0 bits (219), Expect = 8e-18
Identities = 83/385 (21%), Positives = 161/385 (41%), Gaps = 36/385 (9%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y +++ KA AL + + M V PD+ Y++I +L +G LK+ + +++ M
Sbjct: 175 YNVMISGYCKAGEINNALSVLDRM----SVSPDVVTYNTILRSLCDSGKLKQAMEVLDRM 230
Query: 324 RQK---PETLKY------------------MYRKNWDPTLEPDVVIYNAVLNACVPSKQW 362
Q+ P+ + Y + + D PDVV YN ++N +
Sbjct: 231 LQRDCYPDVITYTILIEATCRDSGVGHAMKLLDEMRDRGCTPDVVTYNVLVNGICKEGRL 290
Query: 363 KGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVM 422
M S +PN T+ + + M +G + +L M R G P +T+ ++
Sbjct: 291 DEAIKFLNDMPSSGCQPNVITHNIILRSMCSTGRWMDAEKLLADMLRKGFSPSVVTFNIL 350
Query: 423 VRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPH 482
+ ++G + A+ + M + G + Y L C + A +E ++R+
Sbjct: 351 INFLCRKGLLGRAIDILEKMPQHGCQPNSLSYNPLLHGFCKEKKMDRA---IEYLERMVS 407
Query: 483 --AKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDMFST 539
P VT+ M+ + G ++D + I + C+P + T NT++ ++
Sbjct: 408 RGCYPDIVTYNTMLTALCKDGKVEDAVEILNQLSSKGCSPVLITYNTVIDGLAKAGKTGK 467
Query: 540 AKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWE---YFEHVYKEMILSGYHLDQNKH 596
A L +E++ DL+PD TY+ ++ SR + + F H ++ M + + N
Sbjct: 468 AIKLLDEMRA--KDLKPDTITYSSLVGGLSREGKVDEAIKFFHEFERMGIRPNAVTFNSI 525
Query: 597 LPLLVKASRAGKVDNILVIYIRSQC 621
+ L K+ + + + LV I C
Sbjct: 526 MLGLCKSRQTDRAIDFLVFMINRGC 550
Score = 69.3 bits (168), Expect = 6e-12
Identities = 54/221 (24%), Positives = 94/221 (42%), Gaps = 22/221 (9%)
Query: 295 PDMAAYHSIAVTLGQAGLLKELLNIVECMRQ---KPETLKY-------MYRKNWDPTLE- 343
P + ++ + L + GLL ++I+E M Q +P +L Y K D +E
Sbjct: 342 PSVVTFNILINFLCRKGLLGRAIDILEKMPQHGCQPNSLSYNPLLHGFCKEKKMDRAIEY 401
Query: 344 ----------PDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQ 393
PD+V YN +L A + + + Q+ P TY ++ + +
Sbjct: 402 LERMVSRGCYPDIVTYNTMLTALCKDGKVEDAVEILNQLSSKGCSPVLITYNTVIDGLAK 461
Query: 394 SGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASV 453
+G +L ++M+ P+ +TY +V +EGKVDEA+K + ER G+ A
Sbjct: 462 AGKTGKAIKLLDEMRAKDLKPDTITYSSLVGGLSREGKVDEAIKFFHEFERMGIRPNAVT 521
Query: 454 YYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMI 494
+ + LC R D ++ KP E ++T +I
Sbjct: 522 FNSIMLGLCK-SRQTDRAIDFLVFMINRGCKPNETSYTILI 561
Score = 58.9 bits (141), Expect = 9e-09
Identities = 40/176 (22%), Positives = 80/176 (44%), Gaps = 7/176 (3%)
Query: 391 MLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGT 450
M+++G + + E M +G VP+ + ++R F + GK +A K + +E G +
Sbjct: 112 MVRTGELEEGFKFLENMVYHGNVPDIIPCTTLIRGFCRLGKTRKAAKILEILEGSGAVPD 171
Query: 451 ASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIF 510
Y + C G +A ++++ P VT+ ++RS D G + + +
Sbjct: 172 VITYNVMISGYCKAGEINNALSVLDRMS----VSPDVVTYNTILRSLCDSGKLKQAMEVL 227
Query: 511 EYM-QDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLML 565
+ M Q C P+V T +++ ++ A L +E++ PD TYN+++
Sbjct: 228 DRMLQRDCYPDVITYTILIEATCRDSGVGHAMKLLDEMR--DRGCTPDVVTYNVLV 281
>At2g17140 hypothetical protein
Length = 903
Score = 106 bits (265), Expect = 4e-23
Identities = 81/312 (25%), Positives = 143/312 (44%), Gaps = 13/312 (4%)
Query: 287 MLGNIRVYPDMAAYHSIAVTLG-QAGLLKELLNIVECMRQKPETLK----YMYRKNWDPT 341
+L +++ ++ H++ ++ Q L LL++V + K + ++ P
Sbjct: 48 ILVRAKMHEEIQELHNLILSSSIQKTKLSSLLSVVSIFAKSNHIDKAFPQFQLVRSRFPE 107
Query: 342 LEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVH 401
+P V +YN +L +C+ ++ + VSW+++ M + P T+ L + + S D
Sbjct: 108 NKPSVYLYNLLLESCIKERRVEFVSWLYKDMVLCGIAPQTYTFNLLIRALCDSSCVDAAR 167
Query: 402 ELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCL 461
ELF++M G P T+ ++VR + K G D+ ++ + ME GV+ +Y +
Sbjct: 168 ELFDEMPEKGCKPNEFTFGILVRGYCKAGLTDKGLELLNAMESFGVLPNKVIYNTIVSSF 227
Query: 462 CNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQ-----DH 516
C GR D+ VEK+ R P VTF I + G + D IF M+
Sbjct: 228 CREGRNDDSEKMVEKM-REEGLVPDIVTFNSRISALCKEGKVLDASRIFSDMELDEYLGL 286
Query: 517 CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEY 576
PN T N MLK + + + AK LFE ++ ++D +YN+ L+ R ++
Sbjct: 287 PRPNSITYNLMLKGFCKVGLLEDAKTLFESIR--ENDDLASLQSYNIWLQGLVRHGKFIE 344
Query: 577 FEHVYKEMILSG 588
E V K+M G
Sbjct: 345 AETVLKQMTDKG 356
Score = 92.0 bits (227), Expect = 9e-19
Identities = 104/496 (20%), Positives = 197/496 (38%), Gaps = 72/496 (14%)
Query: 144 LKEMFRERKMDELKWVFEDDIEINEVWFDENNNGKRKKTSKRSEVQVVRFLVDRLCDKEI 203
L+ +ER+++ + W+++D + N + S V R L D + +K
Sbjct: 119 LESCIKERRVEFVSWLYKDMVLCGIAPQTYTFNLLIRALCDSSCVDAARELFDEMPEKGC 178
Query: 204 RAKDWKFSRLMK---LSGLSFTEGQLMMIIEMLGV---KRCWKQALSVVQWVYNSKDHRK 257
+ ++ F L++ +GL+ +L+ +E GV K + +S + D K
Sbjct: 179 KPNEFTFGILVRGYCKAGLTDKGLELLNAMESFGVLPNKVIYNTIVSSFCREGRNDDSEK 238
Query: 258 FQSRF----------VYTKLLAVLGKARRPKEALQIFNMM-----LGNIRVYPDMAAYHS 302
+ + ++ L K + +A +IF+ M LG R P+ Y+
Sbjct: 239 MVEKMREEGLVPDIVTFNSRISALCKEGKVLDASRIFSDMELDEYLGLPR--PNSITYNL 296
Query: 303 IAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNW---------------------DPT 341
+ + GLL++ + E +R+ + W D
Sbjct: 297 MLKGFCKVGLLEDAKTLFESIRENDDLASLQSYNIWLQGLVRHGKFIEAETVLKQMTDKG 356
Query: 342 LEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVH 401
+ P + YN +++ + M+++ + P+ TYG + G D
Sbjct: 357 IGPSIYSYNILMDGLCKLGMLSDAKTIVGLMKRNGVCPDAVTYGCLLHGYCSVGKVDAAK 416
Query: 402 ELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCL 461
L ++M RN +P A T +++ + WK G++ EA + +R M +G Y L
Sbjct: 417 SLLQEMMRNNCLPNAYTCNILLHSLWKMGRISEAEELLRKMNEKG--------YGLDTVT 468
Query: 462 CN------CGRWQ-DATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQ 514
CN CG + D +E+ K R+ + L I G +DD + ++
Sbjct: 469 CNIIVDGLCGSGELDKAIEIVKGMRVHGSAALGNLGNSYI------GLVDDSL-----IE 517
Query: 515 DHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQW 574
++C P++ T +T+L + F+ AK LF E+ K L+PD+ YN+ + + +
Sbjct: 518 NNCLPDLITYSTLLNGLCKAGRFAEAKNLFAEMMGEK--LQPDSVAYNIFIHHFCKQGKI 575
Query: 575 EYFEHVYKEMILSGYH 590
V K+M G H
Sbjct: 576 SSAFRVLKDMEKKGCH 591
Score = 67.8 bits (164), Expect = 2e-11
Identities = 73/337 (21%), Positives = 134/337 (39%), Gaps = 56/337 (16%)
Query: 260 SRFVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNI 319
+ F + L+ KA + L++ N M + V P+ Y++I + + G + +
Sbjct: 181 NEFTFGILVRGYCKAGLTDKGLELLNAM-ESFGVLPNKVIYNTIVSSFCREGRNDDSEKM 239
Query: 320 VECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSL-- 377
VE MR++ L PD+V +N+ ++A + S +F M
Sbjct: 240 VEKMREEG--------------LVPDIVTFNSRISALCKEGKVLDASRIFSDMELDEYLG 285
Query: 378 --KPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEA 435
+PN TY L ++ + G + LFE ++ N ++ +Y + ++ + GK EA
Sbjct: 286 LPRPNSITYNLMLKGFCKVGLLEDAKTLFESIRENDDLASLQSYNIWLQGLVRHGKFIEA 345
Query: 436 VKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIR 495
++ M +G+ + Y L LC G DA V +KR
Sbjct: 346 ETVLKQMTDKGIGPSIYSYNILMDGLCKLGMLSDAKTIVGLMKR---------------- 389
Query: 496 SSMDGGHIDDCICIFEYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLR 555
+ +C P+ T +L Y AK L +E+ +++
Sbjct: 390 ---------NGVC----------PDAVTYGCLLHGYCSVGKVDAAKSLLQEMM--RNNCL 428
Query: 556 PDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLD 592
P+AYT N++L + + + E + ++M GY LD
Sbjct: 429 PNAYTCNILLHSLWKMGRISEAEELLRKMNEKGYGLD 465
Score = 57.8 bits (138), Expect = 2e-08
Identities = 39/182 (21%), Positives = 85/182 (46%), Gaps = 2/182 (1%)
Query: 365 VSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVR 424
+ V + +++ P+ TY + + ++G + LF +M P+++ Y + +
Sbjct: 508 IGLVDDSLIENNCLPDLITYSTLLNGLCKAGRFAEAKNLFAEMMGEKLQPDSVAYNIFIH 567
Query: 425 TFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAK 484
F K+GK+ A + ++DME++G + Y L L + + ++++K
Sbjct: 568 HFCKQGKISSAFRVLKDMEKKGCHKSLETYNSLILGLGIKNQIFEIHGLMDEMKE-KGIS 626
Query: 485 PLEVTFTGMIRSSMDGGHIDDCICIF-EYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVL 543
P T+ I+ +G ++D + E MQ + APNV + +++ + + F A+ +
Sbjct: 627 PNICTYNTAIQYLCEGEKVEDATNLLDEMMQKNIAPNVFSFKYLIEAFCKVPDFDMAQEV 686
Query: 544 FE 545
FE
Sbjct: 687 FE 688
Score = 52.8 bits (125), Expect = 6e-07
Identities = 48/264 (18%), Positives = 102/264 (38%), Gaps = 3/264 (1%)
Query: 344 PDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHEL 403
PD++ Y+ +LN + ++ +F +M L+P+ Y + + + G +
Sbjct: 522 PDLITYSTLLNGLCKAGRFAEAKNLFAEMMGEKLQPDSVAYNIFIHHFCKQGKISSAFRV 581
Query: 404 FEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCN 463
+ M++ G TY ++ + ++ E + +M+ +G+ Y LC
Sbjct: 582 LKDMEKKGCHKSLETYNSLILGLGIKNQIFEIHGLMDEMKEKGISPNICTYNTAIQYLCE 641
Query: 464 CGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDHCAPNVGT 523
+ +DAT ++++ + + P +F +I + D +FE C G
Sbjct: 642 GEKVEDATNLLDEMMQ-KNIAPNVFSFKYLIEAFCKVPDFDMAQEVFETAVSICGQKEGL 700
Query: 524 VNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKE 583
+ M A L E V +L + Y ++E+ + + E + +
Sbjct: 701 YSLMFNELLAAGQLLKATELLEAVLDRGFEL--GTFLYKDLVESLCKKDELEVASGILHK 758
Query: 584 MILSGYHLDQNKHLPLLVKASRAG 607
MI GY D +P++ + G
Sbjct: 759 MIDRGYGFDPAALMPVIDGLGKMG 782
Score = 42.7 bits (99), Expect = 6e-04
Identities = 46/220 (20%), Positives = 90/220 (40%), Gaps = 23/220 (10%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y+ LL L KA R EA +F M+G ++ PD AY+ + G + +++ M
Sbjct: 527 YSTLLNGLCKAGRFAEAKNLFAEMMGE-KLQPDSVAYNIFIHHFCKQGKISSAFRVLKDM 585
Query: 324 RQKP-----ETLKYMYR----KNW------------DPTLEPDVVIYNAVLNACVPSKQW 362
+K ET + KN + + P++ YN + ++
Sbjct: 586 EKKGCHKSLETYNSLILGLGIKNQIFEIHGLMDEMKEKGISPNICTYNTAIQYLCEGEKV 645
Query: 363 KGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVM 422
+ + + +M + ++ PN ++ +E + ++D+ E+FE E L Y +M
Sbjct: 646 EDATNLLDEMMQKNIAPNVFSFKYLIEAFCKVPDFDMAQEVFETAVSICGQKEGL-YSLM 704
Query: 423 VRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLC 462
G++ +A + + + RG +Y +L LC
Sbjct: 705 FNELLAAGQLLKATELLEAVLDRGFELGTFLYKDLVESLC 744
Score = 40.8 bits (94), Expect = 0.002
Identities = 34/155 (21%), Positives = 67/155 (42%), Gaps = 18/155 (11%)
Query: 295 PDMAAYHSIAVTLGQAGLLKELLNI-VECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVL 353
PD+ Y ++ L +AG E N+ E M +K L+PD V YN +
Sbjct: 522 PDLITYSTLLNGLCKAGRFAEAKNLFAEMMGEK---------------LQPDSVAYNIFI 566
Query: 354 NACVPSKQWKGVSWVFQQMRKSSLKPNGATY-GLAMEVMLQSGNYDLVHELFEKMQRNGE 412
+ + V + M K + TY L + + +++ ++ +H L ++M+ G
Sbjct: 567 HHFCKQGKISSAFRVLKDMEKKGCHKSLETYNSLILGLGIKNQIFE-IHGLMDEMKEKGI 625
Query: 413 VPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGV 447
P TY ++ + KV++A + +M ++ +
Sbjct: 626 SPNICTYNTAIQYLCEGEKVEDATNLLDEMMQKNI 660
>At5g41170 salt-inducible protein-like
Length = 527
Score = 102 bits (254), Expect = 7e-22
Identities = 86/358 (24%), Positives = 160/358 (44%), Gaps = 24/358 (6%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
+T L+ R +EA+ + N M+ + + PD+ Y +I +L + G + L++ + M
Sbjct: 145 FTSLINGFCLGNRMEEAMSMVNQMV-EMGIKPDVVMYTTIIDSLCKNGHVNYALSLFDQM 203
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
+N+ + PDVV+Y +++N S +W+ + + M K +KP+ T
Sbjct: 204 ------------ENYG--IRPDVVMYTSLVNGLCNSGRWRDADSLLRGMTKRKIKPDVIT 249
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
+ ++ ++ G + EL+ +M R P TY ++ F EG VDEA + ME
Sbjct: 250 FNALIDAFVKEGKFLDAEELYNEMIRMSIAPNIFTYTSLINGFCMEGCVDEARQMFYLME 309
Query: 444 RRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPL---EVTFTGMIRSSMDG 500
+G Y L C C + DA KI K L +T+T +I+
Sbjct: 310 TKGCFPDVVAYTSLINGFCKCKKVDDAM----KIFYEMSQKGLTGNTITYTTLIQGFGQV 365
Query: 501 GHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSD-LRPDA 558
G + +F +M PN+ T N +L N A ++FE+++ + D + P+
Sbjct: 366 GKPNVAQEVFSHMVSRGVPPNIRTYNVLLHCLCYNGKVKKALMIFEDMQKREMDGVAPNI 425
Query: 559 YTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVDNILVIY 616
+TYN++L + E V+++M + + ++ +AGKV N + ++
Sbjct: 426 WTYNVLLHGLCYNGKLEKALMVFEDMRKREMDIGIITYTIIIQGMCKAGKVKNAVNLF 483
Score = 74.7 bits (182), Expect = 1e-13
Identities = 67/289 (23%), Positives = 122/289 (42%), Gaps = 20/289 (6%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
+ L+ K + +A +++N M+ + + P++ Y S+ G + E + M
Sbjct: 250 FNALIDAFVKEGKFLDAEELYNEMI-RMSIAPNIFTYTSLINGFCMEGCVDEARQMFYLM 308
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
K PDVV Y +++N K+ +F +M + L N T
Sbjct: 309 ETKG--------------CFPDVVAYTSLINGFCKCKKVDDAMKIFYEMSQKGLTGNTIT 354
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
Y ++ Q G ++ E+F M G P TY V++ GKV +A+ DM+
Sbjct: 355 YTTLIQGFGQVGKPNVAQEVFSHMVSRGVPPNIRTYNVLLHCLCYNGKVKKALMIFEDMQ 414
Query: 444 RRGVMGTAS---VYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDG 500
+R + G A Y L LC G+ + A + E +++ + +T+T +I+
Sbjct: 415 KREMDGVAPNIWTYNVLLHGLCYNGKLEKALMVFEDMRKREMDIGI-ITYTIIIQGMCKA 473
Query: 501 GHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVK 548
G + + + +F + PNV T TM+ + + A VLF ++K
Sbjct: 474 GKVKNAVNLFCSLPSKGVKPNVVTYTTMISGLFREGLKHEAHVLFRKMK 522
Score = 72.8 bits (177), Expect = 6e-13
Identities = 75/365 (20%), Positives = 152/365 (41%), Gaps = 21/365 (5%)
Query: 254 DHRKFQSRFVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLL 313
+ R S +TKLL V+ K ++ F++++ M H +
Sbjct: 65 ESRPLPSIIDFTKLLNVIAKMKK-------FDVVINLCDHLQIMGVSHDLYTC------- 110
Query: 314 KELLNIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNA-CVPSKQWKGVSWVFQQM 372
LL C +P K EPD+V + +++N C+ ++ + +S V QM
Sbjct: 111 -NLLMNCFCQSSQPYLASSFLGKMMKLGFEPDIVTFTSLINGFCLGNRMEEAMSMV-NQM 168
Query: 373 RKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKV 432
+ +KP+ Y ++ + ++G+ + LF++M+ G P+ + Y +V G+
Sbjct: 169 VEMGIKPDVVMYTTIIDSLCKNGHVNYALSLFDQMENYGIRPDVVMYTSLVNGLCNSGRW 228
Query: 433 DEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTG 492
+A +R M +R + + L G++ DA ++ R+ A P T+T
Sbjct: 229 RDADSLLRGMTKRKIKPDVITFNALIDAFVKEGKFLDAEELYNEMIRMSIA-PNIFTYTS 287
Query: 493 MIRSSMDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAK 551
+I G +D+ +F M+ C P+V +++ + + A +F E +++
Sbjct: 288 LINGFCMEGCVDEARQMFYLMETKGCFPDVVAYTSLINGFCKCKKVDDAMKIFYE--MSQ 345
Query: 552 SDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVDN 611
L + TY +++ + + + V+ M+ G + + LL GKV
Sbjct: 346 KGLTGNTITYTTLIQGFGQVGKPNVAQEVFSHMVSRGVPPNIRTYNVLLHCLCYNGKVKK 405
Query: 612 ILVIY 616
L+I+
Sbjct: 406 ALMIF 410
Score = 61.2 bits (147), Expect = 2e-09
Identities = 46/184 (25%), Positives = 83/184 (45%), Gaps = 12/184 (6%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
YT L+ G+ +P A ++F+ M+ V P++ Y+ + L G +K+ L I E M
Sbjct: 355 YTTLIQGFGQVGKPNVAQEVFSHMVSR-GVPPNIRTYNVLLHCLCYNGKVKKALMIFEDM 413
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
+++ + P++ YN +L+ + + + VF+ MRK + T
Sbjct: 414 QKREMD-----------GVAPNIWTYNVLLHGLCYNGKLEKALMVFEDMRKREMDIGIIT 462
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
Y + ++ M ++G LF + G P +TY M+ ++EG EA R M+
Sbjct: 463 YTIIIQGMCKAGKVKNAVNLFCSLPSKGVKPNVVTYTTMISGLFREGLKHEAHVLFRKMK 522
Query: 444 RRGV 447
GV
Sbjct: 523 EDGV 526
Score = 51.2 bits (121), Expect = 2e-06
Identities = 47/224 (20%), Positives = 90/224 (39%), Gaps = 18/224 (8%)
Query: 258 FQSRFVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELL 317
F YT L+ K ++ +A++IF M + + Y ++ GQ G
Sbjct: 314 FPDVVAYTSLINGFCKCKKVDDAMKIFYEM-SQKGLTGNTITYTTLIQGFGQVG------ 366
Query: 318 NIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSL 377
KP + ++ + P++ YN +L+ + + K +F+ M+K +
Sbjct: 367 --------KPNVAQEVFSHMVSRGVPPNIRTYNVLLHCLCYNGKVKKALMIFEDMQKREM 418
Query: 378 K---PNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDE 434
PN TY + + + +G + +FE M++ +TY ++++ K GKV
Sbjct: 419 DGVAPNIWTYNVLLHGLCYNGKLEKALMVFEDMRKREMDIGIITYTIIIQGMCKAGKVKN 478
Query: 435 AVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIK 478
AV + +GV Y + L G +A + K+K
Sbjct: 479 AVNLFCSLPSKGVKPNVVTYTTMISGLFREGLKHEAHVLFRKMK 522
>At4g31850 putative protein
Length = 1112
Score = 100 bits (248), Expect = 3e-21
Identities = 105/422 (24%), Positives = 182/422 (42%), Gaps = 45/422 (10%)
Query: 227 MMIIEMLGVKRCWKQALSVVQWVYNSKDHRKFQ---SRFVYTKLLAVLGKARRPKEALQI 283
+ I + L VK KQA Y + R+F + + Y L+ +L K+R EA+++
Sbjct: 157 LTIFKSLSVKGGLKQA------PYALRKMREFGFVLNAYSYNGLIHLLLKSRFCTEAMEV 210
Query: 284 FN-MMLGNIRVYPDMAAYHSIAVTLGQA-------GLLKELLNIVECMRQKPETLKY--- 332
+ M+L R P + Y S+ V LG+ GLLKE+ E + KP +
Sbjct: 211 YRRMILEGFR--PSLQTYSSLMVGLGKRRDIDSVMGLLKEM----ETLGLKPNVYTFTIC 264
Query: 333 ---------------MYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSL 377
+ ++ D PDVV Y +++A +++ VF++M+
Sbjct: 265 IRVLGRAGKINEAYEILKRMDDEGCGPDVVTYTVLIDALCTARKLDCAKEVFEKMKTGRH 324
Query: 378 KPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVK 437
KP+ TY ++ + + D V + + +M+++G VP+ +T+ ++V K G EA
Sbjct: 325 KPDRVTYITLLDRFSDNRDLDSVKQFWSEMEKDGHVPDVVTFTILVDALCKAGNFGEAFD 384
Query: 438 AVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSS 497
+ M +G++ Y L C L R DA LE+ KP T+ I
Sbjct: 385 TLDVMRDQGILPNLHTYNTLICGLLRVHRLDDA-LELFGNMESLGVKPTAYTYIVFIDYY 443
Query: 498 MDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRP 556
G + FE M+ APN+ N L ++ AK +F +K L P
Sbjct: 444 GKSGDSVSALETFEKMKTKGIAPNIVACNASLYSLAKAGRDREAKQIFYGLK--DIGLVP 501
Query: 557 DAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVDNILVIY 616
D+ TYN+M++ S+ + + + EM+ +G D L+ +A +VD ++
Sbjct: 502 DSVTYNMMMKCYSKVGEIDEAIKLLSEMMENGCEPDVIVVNSLINTLYKADRVDEAWKMF 561
Query: 617 IR 618
+R
Sbjct: 562 MR 563
Score = 83.6 bits (205), Expect = 3e-16
Identities = 70/329 (21%), Positives = 138/329 (41%), Gaps = 21/329 (6%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y L+ L +A + A +F + + + PD+A Y+ + G++G + EL +
Sbjct: 788 YNLLIGGLLEADMIEIAQDVF-LQVKSTGCIPDVATYNFLLDAYGKSGKIDELFEL---- 842
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWK-GVSWVFQQMRKSSLKPNGA 382
Y++ E + + +N V++ V + + + M P
Sbjct: 843 ----------YKEMSTHECEANTITHNIVISGLVKAGNVDDALDLYYDLMSDRDFSPTAC 892
Query: 383 TYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDM 442
TYG ++ + +SG +LFE M G P Y +++ F K G+ D A + M
Sbjct: 893 TYGPLIDGLSKSGRLYEAKQLFEGMLDYGCRPNCAIYNILINGFGKAGEADAACALFKRM 952
Query: 443 ERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGH 502
+ GV Y L CLC GR + +++K P V + +I
Sbjct: 953 VKEGVRPDLKTYSVLVDCLCMVGRVDEGLHYFKELKE-SGLNPDVVCYNLIINGLGKSHR 1011
Query: 503 IDDCICIFEYMQDH--CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYT 560
+++ + +F M+ P++ T N+++ M A ++ E++ ++ L P+ +T
Sbjct: 1012 LEEALVLFNEMKTSRGITPDLYTYNSLILNLGIAGMVEEAGKIYNEIQ--RAGLEPNVFT 1069
Query: 561 YNLMLEASSRGHQWEYFEHVYKEMILSGY 589
+N ++ S + E+ VY+ M+ G+
Sbjct: 1070 FNALIRGYSLSGKPEHAYAVYQTMVTGGF 1098
Score = 83.6 bits (205), Expect = 3e-16
Identities = 83/349 (23%), Positives = 146/349 (41%), Gaps = 26/349 (7%)
Query: 262 FVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLL---KELLN 318
+ +T + VLG+A + EA +I M + PD+ Y + L A L KE+
Sbjct: 259 YTFTICIRVLGRAGKINEAYEILKRM-DDEGCGPDVVTYTVLIDALCTARKLDCAKEVFE 317
Query: 319 IVECMRQKPETLKYM--------------YRKNWDPTLE----PDVVIYNAVLNACVPSK 360
++ R KP+ + Y+ ++ W + PDVV + +++A +
Sbjct: 318 KMKTGRHKPDRVTYITLLDRFSDNRDLDSVKQFWSEMEKDGHVPDVVTFTILVDALCKAG 377
Query: 361 QWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYK 420
+ MR + PN TY + +L+ D ELF M+ G P A TY
Sbjct: 378 NFGEAFDTLDVMRDQGILPNLHTYNTLICGLLRVHRLDDALELFGNMESLGVKPTAYTYI 437
Query: 421 VMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRL 480
V + + K G A++ M+ +G+ L GR ++A +K +
Sbjct: 438 VFIDYYGKSGDSVSALETFEKMKTKGIAPNIVACNASLYSLAKAGRDREAKQIFYGLKDI 497
Query: 481 PHAKPLEVTFTGMIRSSMDGGHIDDCICIF-EYMQDHCAPNVGTVNTMLKVYSQNDMFST 539
P VT+ M++ G ID+ I + E M++ C P+V VN+++ + D
Sbjct: 498 GLV-PDSVTYNMMMKCYSKVGEIDEAIKLLSEMMENGCEPDVIVVNSLINTLYKADRVDE 556
Query: 540 AKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSG 588
A +F +K K L+P TYN +L + + + +++ M+ G
Sbjct: 557 AWKMFMRMKEMK--LKPTVVTYNTLLAGLGKNGKIQEAIELFEGMVQKG 603
Score = 75.1 bits (183), Expect = 1e-13
Identities = 78/382 (20%), Positives = 153/382 (39%), Gaps = 52/382 (13%)
Query: 271 LGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETL 330
L KA R +EA QIF L +I + PD Y+ + + G + E + ++ M +
Sbjct: 478 LAKAGRDREAKQIF-YGLKDIGLVPDSVTYNMMMKCYSKVGEIDEAIKLLSEMMENG--- 533
Query: 331 KYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEV 390
EPDV++ N+++N + + +F +M++ LKP TY +
Sbjct: 534 -----------CEPDVIVVNSLINTLYKADRVDEAWKMFMRMKEMKLKPTVVTYNTLLAG 582
Query: 391 MLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGT 450
+ ++G ELFE M + G P +T+ + K +V A+K + M G +
Sbjct: 583 LGKNGKIQEAIELFEGMVQKGCPPNTITFNTLFDCLCKNDEVTLALKMLFKMMDMGCVPD 642
Query: 451 ASVYYELACCLCNCGRWQDATLEVEKIKRLPHA--------------------------- 483
Y + L G+ ++A ++K+L +
Sbjct: 643 VFTYNTIIFGLVKNGQVKEAMCFFHQMKKLVYPDFVTLCTLLPGVVKASLIEDAYKIITN 702
Query: 484 -------KPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDH--CAPNVGTVNTMLKVYSQN 534
+P + + +I S + ID+ + E + + C + +++ ++
Sbjct: 703 FLYNCADQPANLFWEDLIGSILAEAGIDNAVSFSERLVANGICRDGDSILVPIIRYSCKH 762
Query: 535 DMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQN 594
+ S A+ LFE+ ++P TYNL++ E + V+ ++ +G D
Sbjct: 763 NNVSGARTLFEKF-TKDLGVQPKLPTYNLLIGGLLEADMIEIAQDVFLQVKSTGCIPDVA 821
Query: 595 KHLPLLVKASRAGKVDNILVIY 616
+ LL ++GK+D + +Y
Sbjct: 822 TYNFLLDAYGKSGKIDELFELY 843
Score = 74.7 bits (182), Expect = 1e-13
Identities = 69/296 (23%), Positives = 119/296 (39%), Gaps = 37/296 (12%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y L+ L + R +AL++F M ++ V P Y G++G L E M
Sbjct: 401 YNTLICGLLRVHRLDDALELFGNM-ESLGVKPTAYTYIVFIDYYGKSGDSVSALETFEKM 459
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
+ K + P++V NA L + + + + +F ++ L P+ T
Sbjct: 460 KTKG--------------IAPNIVACNASLYSLAKAGRDREAKQIFYGLKDIGLVPDSVT 505
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
Y + M+ + G D +L +M NG P+ + ++ T +K +VDEA K M+
Sbjct: 506 YNMMMKCYSKVGEIDEAIKLLSEMMENGCEPDVIVVNSLINTLYKADRVDEAWKMFMRMK 565
Query: 444 RRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHI 503
+ T Y L L G+ Q+A +E+ + P +TF +
Sbjct: 566 EMKLKPTVVTYNTLLAGLGKNGKIQEA-IELFEGMVQKGCPPNTITFNTLF--------- 615
Query: 504 DDCIC-----------IFEYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVK 548
DC+C +F+ M C P+V T NT++ +N A F ++K
Sbjct: 616 -DCLCKNDEVTLALKMLFKMMDMGCVPDVFTYNTIIFGLVKNGQVKEAMCFFHQMK 670
Score = 70.9 bits (172), Expect = 2e-12
Identities = 66/281 (23%), Positives = 120/281 (42%), Gaps = 6/281 (2%)
Query: 339 DPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYD 398
D ++P + YN ++ + + + VF Q++ + P+ ATY ++ +SG D
Sbjct: 778 DLGVQPKLPTYNLLIGGLLEADMIEIAQDVFLQVKSTGCIPDVATYNFLLDAYGKSGKID 837
Query: 399 LVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRD-MERRGVMGTASVYYEL 457
+ EL+++M + +T+ +++ K G VD+A+ D M R TA Y L
Sbjct: 838 ELFELYKEMSTHECEANTITHNIVISGLVKAGNVDDALDLYYDLMSDRDFSPTACTYGPL 897
Query: 458 ACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYM-QDH 516
L GR +A E + +P + +I G D +F+ M ++
Sbjct: 898 IDGLSKSGRLYEAKQLFEGMLDY-GCRPNCAIYNILINGFGKAGEADAACALFKRMVKEG 956
Query: 517 CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEY 576
P++ T + ++ F+E+K +S L PD YNL++ + H+ E
Sbjct: 957 VRPDLKTYSVLVDCLCMVGRVDEGLHYFKELK--ESGLNPDVVCYNLIINGLGKSHRLEE 1014
Query: 577 FEHVYKEMILS-GYHLDQNKHLPLLVKASRAGKVDNILVIY 616
++ EM S G D + L++ AG V+ IY
Sbjct: 1015 ALVLFNEMKTSRGITPDLYTYNSLILNLGIAGMVEEAGKIY 1055
Score = 56.2 bits (134), Expect = 6e-08
Identities = 53/217 (24%), Positives = 96/217 (43%), Gaps = 25/217 (11%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAG-------LLKEL 316
Y L+ L K+ R EA Q+F ML + P+ A Y+ + G+AG L K +
Sbjct: 894 YGPLIDGLSKSGRLYEAKQLFEGML-DYGCRPNCAIYNILINGFGKAGEADAACALFKRM 952
Query: 317 LN------------IVECM---RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQ 361
+ +V+C+ + E L Y +++ + L PDVV YN ++N S +
Sbjct: 953 VKEGVRPDLKTYSVLVDCLCMVGRVDEGLHY-FKELKESGLNPDVVCYNLIINGLGKSHR 1011
Query: 362 WKGVSWVFQQMRKS-SLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYK 420
+ +F +M+ S + P+ TY + + +G + +++ ++QR G P T+
Sbjct: 1012 LEEALVLFNEMKTSRGITPDLYTYNSLILNLGIAGMVEEAGKIYNEIQRAGLEPNVFTFN 1071
Query: 421 VMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYEL 457
++R + GK + A + M G Y +L
Sbjct: 1072 ALIRGYSLSGKPEHAYAVYQTMVTGGFSPNTGTYEQL 1108
Score = 52.4 bits (124), Expect = 8e-07
Identities = 60/306 (19%), Positives = 114/306 (36%), Gaps = 45/306 (14%)
Query: 291 IRVYPD----MAAYHSIAVTLGQAGLLKELLNIVECMRQ--KPETLKYMYRKNWDPTLEP 344
++ +PD + + S+A L + ++E +R K E + Y++ ++
Sbjct: 92 LKSFPDTDSSFSYFKSVAGNLNLVHTTETCNYMLEALRVDGKLEEMAYVFDLMQKRIIKR 151
Query: 345 DVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELF 404
D Y + + K + ++MR+ N +Y + ++L+S E++
Sbjct: 152 DTNTYLTIFKSLSVKGGLKQAPYALRKMREFGFVLNAYSYNGLIHLLLKSRFCTEAMEVY 211
Query: 405 EKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNC 464
+M G P TY ++ K +D + +++ME G+
Sbjct: 212 RRMILEGFRPSLQTYSSLMVGLGKRRDIDSVMGLLKEMETLGL----------------- 254
Query: 465 GRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDH-CAPNVGT 523
KP TFT IR G I++ I + M D C P+V T
Sbjct: 255 -------------------KPNVYTFTICIRVLGRAGKINEAYEILKRMDDEGCGPDVVT 295
Query: 524 VNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKE 583
++ AK +FE++K + +PD TY +L+ S + + + E
Sbjct: 296 YTVLIDALCTARKLDCAKEVFEKMKTGRH--KPDRVTYITLLDRFSDNRDLDSVKQFWSE 353
Query: 584 MILSGY 589
M G+
Sbjct: 354 MEKDGH 359
>At1g62670 PPR-repeat protein
Length = 630
Score = 100 bits (248), Expect = 3e-21
Identities = 70/311 (22%), Positives = 135/311 (42%), Gaps = 41/311 (13%)
Query: 343 EPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHE 402
+P+ V +N +++ + + +M +P+ TYG+ + + + G+ DL
Sbjct: 183 QPNTVTFNTLIHGLFLHNKASEAMALIDRMVAKGCQPDLVTYGVVVNGLCKRGDTDLAFN 242
Query: 403 LFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLC 462
L KM++ P L Y ++ K +D+A+ ++ME +G+ Y L CLC
Sbjct: 243 LLNKMEQGKLEPGVLIYNTIIDGLCKYKHMDDALNLFKEMETKGIRPNVVTYSSLISCLC 302
Query: 463 NCGRWQDAT-------------------------------LEVEK-----IKRLPHAKPL 486
N GRW DA+ +E EK +KR P
Sbjct: 303 NYGRWSDASRLLSDMIERKINPDVFTFSALIDAFVKEGKLVEAEKLYDEMVKR--SIDPS 360
Query: 487 EVTFTGMIRSSMDGGHIDDCICIFEYM-QDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFE 545
VT++ +I +D+ +FE+M HC P+V T NT++K + + +F
Sbjct: 361 IVTYSSLINGFCMHDRLDEAKQMFEFMVSKHCFPDVVTYNTLIKGFCKYKRVEEGMEVFR 420
Query: 546 EVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASR 605
E +++ L + TYN++++ + + + ++KEM+ G + + LL +
Sbjct: 421 E--MSQRGLVGNTVTYNILIQGLFQAGDCDMAQEIFKEMVSDGVPPNIMTYNTLLDGLCK 478
Query: 606 AGKVDNILVIY 616
GK++ +V++
Sbjct: 479 NGKLEKAMVVF 489
Score = 87.0 bits (214), Expect = 3e-17
Identities = 71/323 (21%), Positives = 136/323 (41%), Gaps = 19/323 (5%)
Query: 263 VYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVEC 322
+Y ++ L K + +AL +F M + P++ Y S+ L G + ++
Sbjct: 258 IYNTIIDGLCKYKHMDDALNLFKEMETK-GIRPNVVTYSSLISCLCNYGRWSDASRLLSD 316
Query: 323 MRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGA 382
M ++ + PDV ++A+++A V + ++ +M K S+ P+
Sbjct: 317 MIERK--------------INPDVFTFSALIDAFVKEGKLVEAEKLYDEMVKRSIDPSIV 362
Query: 383 TYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDM 442
TY + D ++FE M P+ +TY +++ F K +V+E ++ R+M
Sbjct: 363 TYSSLINGFCMHDRLDEAKQMFEFMVSKHCFPDVVTYNTLIKGFCKYKRVEEGMEVFREM 422
Query: 443 ERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGH 502
+RG++G Y L L G D E+ K P +T+ ++ G
Sbjct: 423 SQRGLVGNTVTYNILIQGLFQAGDC-DMAQEIFKEMVSDGVPPNIMTYNTLLDGLCKNGK 481
Query: 503 IDDCICIFEYMQ-DHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTY 561
++ + +FEY+Q P + T N M++ + LF ++ ++PD Y
Sbjct: 482 LEKAMVVFEYLQRSKMEPTIYTYNIMIEGMCKAGKVEDGWDLF--CNLSLKGVKPDVVAY 539
Query: 562 NLMLEASSRGHQWEYFEHVYKEM 584
N M+ R E + ++KEM
Sbjct: 540 NTMISGFCRKGSKEEADALFKEM 562
Score = 86.7 bits (213), Expect = 4e-17
Identities = 113/521 (21%), Positives = 197/521 (37%), Gaps = 82/521 (15%)
Query: 126 KVNYEHRGEKLKRESLIELKEMFRERKMDELKWVFEDDIEINEVWFDENNNGKRKKTSKR 185
K +Y++R EKL R L ELK +D+ +F + ++ + +K
Sbjct: 43 KTSYDYR-EKLSRNGLSELK-------LDDAVALFGEMVKSRPFPSIIEFSKLLSAIAKM 94
Query: 186 SEVQVVRFLVDRLCDKEIRAKDWKFSRLMKLSGLSFTEGQLMMIIEMLG--VKRCWKQAL 243
++ VV L +++ + I + +S L+ QL + + +LG +K ++ +
Sbjct: 95 NKFDVVISLGEQMQNLGIPHNHYTYSILINCF---CRRSQLPLALAVLGKMMKLGYEPNI 151
Query: 244 SVVQWVYNSKDHRKFQSRFV-----------------YTKLLAVLGKARRPKEALQIFNM 286
+ + N H K S V + L+ L + EA+ + +
Sbjct: 152 VTLSSLLNGYCHSKRISEAVALVDQMFVTGYQPNTVTFNTLIHGLFLHNKASEAMALIDR 211
Query: 287 MLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNWDPTLEPDV 346
M+ PD+ Y + L + G N++ M Q LEP V
Sbjct: 212 MVAK-GCQPDLVTYGVVVNGLCKRGDTDLAFNLLNKMEQGK--------------LEPGV 256
Query: 347 VIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEK 406
+IYN +++ K +F++M ++PN TY + + G + L
Sbjct: 257 LIYNTIIDGLCKYKHMDDALNLFKEMETKGIRPNVVTYSSLISCLCNYGRWSDASRLLSD 316
Query: 407 MQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGR 466
M P+ T+ ++ F KEGK+ EA K +M +R + + Y L C R
Sbjct: 317 MIERKINPDVFTFSALIDAFVKEGKLVEAEKLYDEMVKRSIDPSIVTYSSLINGFCMHDR 376
Query: 467 WQDATLEVE----------------------KIKRLPHAKPL------------EVTFTG 492
+A E K KR+ + VT+
Sbjct: 377 LDEAKQMFEFMVSKHCFPDVVTYNTLIKGFCKYKRVEEGMEVFREMSQRGLVGNTVTYNI 436
Query: 493 MIRSSMDGGHIDDCICIF-EYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAK 551
+I+ G D IF E + D PN+ T NT+L +N A V+FE ++ +
Sbjct: 437 LIQGLFQAGDCDMAQEIFKEMVSDGVPPNIMTYNTLLDGLCKNGKLEKAMVVFEYLQ--R 494
Query: 552 SDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLD 592
S + P YTYN+M+E + + E ++ + L G D
Sbjct: 495 SKMEPTIYTYNIMIEGMCKAGKVEDGWDLFCNLSLKGVKPD 535
Score = 82.8 bits (203), Expect = 5e-16
Identities = 66/307 (21%), Positives = 135/307 (43%), Gaps = 24/307 (7%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVE-- 321
Y+ L++ L R +A ++ + M+ ++ PD+ + ++ + G L E + +
Sbjct: 294 YSSLISCLCNYGRWSDASRLLSDMIER-KINPDVFTFSALIDAFVKEGKLVEAEKLYDEM 352
Query: 322 -------------------CMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQW 362
CM + + K M+ PDVV YN ++ K+
Sbjct: 353 VKRSIDPSIVTYSSLINGFCMHDRLDEAKQMFEFMVSKHCFPDVVTYNTLIKGFCKYKRV 412
Query: 363 KGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVM 422
+ VF++M + L N TY + ++ + Q+G+ D+ E+F++M +G P +TY +
Sbjct: 413 EEGMEVFREMSQRGLVGNTVTYNILIQGLFQAGDCDMAQEIFKEMVSDGVPPNIMTYNTL 472
Query: 423 VRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPH 482
+ K GK+++A+ ++R + T Y + +C G+ +D ++ L
Sbjct: 473 LDGLCKNGKLEKAMVVFEYLQRSKMEPTIYTYNIMIEGMCKAGKVEDG-WDLFCNLSLKG 531
Query: 483 AKPLEVTFTGMIRSSMDGGHIDDCICIFEYM-QDHCAPNVGTVNTMLKVYSQNDMFSTAK 541
KP V + MI G ++ +F+ M +D PN G NT+++ ++ +
Sbjct: 532 VKPDVVAYNTMISGFCRKGSKEEADALFKEMKEDGTLPNSGCYNTLIRARLRDGDREASA 591
Query: 542 VLFEEVK 548
L +E++
Sbjct: 592 ELIKEMR 598
Score = 50.4 bits (119), Expect = 3e-06
Identities = 43/191 (22%), Positives = 87/191 (45%), Gaps = 19/191 (9%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y L+ L +A A +IF M+ + V P++ Y+++ L + G L++ + + E
Sbjct: 434 YNILIQGLFQAGDCDMAQEIFKEMVSD-GVPPNIMTYNTLLDGLCKNGKLEKAMVVFE-- 490
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNA-CVPSKQWKGVSW-VFQQMRKSSLKPNG 381
Y+ R +PT + YN ++ C K G W +F + +KP+
Sbjct: 491 --------YLQRSKMEPT----IYTYNIMIEGMCKAGKVEDG--WDLFCNLSLKGVKPDV 536
Query: 382 ATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRD 441
Y + + G+ + LF++M+ +G +P + Y ++R ++G + + + +++
Sbjct: 537 VAYNTMISGFCRKGSKEEADALFKEMKEDGTLPNSGCYNTLIRARLRDGDREASAELIKE 596
Query: 442 MERRGVMGTAS 452
M G G AS
Sbjct: 597 MRSCGFAGDAS 607
>At1g74850 hypothetical protein
Length = 862
Score = 97.8 bits (242), Expect = 2e-20
Identities = 77/377 (20%), Positives = 157/377 (41%), Gaps = 61/377 (16%)
Query: 263 VYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVEC 322
+YT ++++LG+ + L++F+ M V + +Y ++ G+ G + L +++
Sbjct: 143 IYTIMISLLGREGLLDKCLEVFDEMPSQ-GVSRSVFSYTALINAYGRNGRYETSLELLDR 201
Query: 323 MRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACV-PSKQWKGVSWVFQQMRKSSLKPNG 381
M+ + + P ++ YN V+NAC W+G+ +F +MR ++P+
Sbjct: 202 MKNE--------------KISPSILTYNTVINACARGGLDWEGLLGLFAEMRHEGIQPDI 247
Query: 382 ATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRD 441
TY + G D +F M G VP+ TY +V TF K ++++ + +
Sbjct: 248 VTYNTLLSACAIRGLGDEAEMVFRTMNDGGIVPDLTTYSHLVETFGKLRRLEKVCDLLGE 307
Query: 442 MERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGG 501
M G + P ++ ++ + G
Sbjct: 308 MASGGSL------------------------------------PDITSYNVLLEAYAKSG 331
Query: 502 HIDDCICIFEYMQ-DHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYT 560
I + + +F MQ C PN T + +L ++ Q+ + + LF E+K + +D PDA T
Sbjct: 332 SIKEAMGVFHQMQAAGCTPNANTYSVLLNLFGQSGRYDDVRQLFLEMKSSNTD--PDAAT 389
Query: 561 YNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVDNILVIYIRSQ 620
YN+++E G ++ ++ +M+ D + ++ + G ++ I
Sbjct: 390 YNILIEVFGEGGYFKEVVTLFHDMVEENIEPDMETYEGIIFACGKGGLHEDARKI----- 444
Query: 621 CLFYISAHSIISYKNCY 637
L Y++A+ I+ Y
Sbjct: 445 -LQYMTANDIVPSSKAY 460
Score = 92.0 bits (227), Expect = 9e-19
Identities = 78/348 (22%), Positives = 143/348 (40%), Gaps = 19/348 (5%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y LL+ EA +F M + + PD+ Y + T G+ L+++ +++ M
Sbjct: 250 YNTLLSACAIRGLGDEAEMVFRTM-NDGGIVPDLTTYSHLVETFGKLRRLEKVCDLLGEM 308
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
PD+ YN +L A S K VF QM+ + PN T
Sbjct: 309 ASGGSL--------------PDITSYNVLLEAYAKSGSIKEAMGVFHQMQAAGCTPNANT 354
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
Y + + + QSG YD V +LF +M+ + P+A TY +++ F + G E V DM
Sbjct: 355 YSVLLNLFGQSGRYDDVRQLFLEMKSSNTDPDAATYNILIEVFGEGGYFKEVVTLFHDMV 414
Query: 444 RRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHI 503
+ YE C G + ++ + P +TG+I +
Sbjct: 415 EENIEPDMET-YEGIIFACGKGGLHEDARKILQYMTANDIVPSSKAYTGVIEAFGQAALY 473
Query: 504 DDCICIFEYMQD-HCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYN 562
++ + F M + P++ T +++L +++ + ++ + ++ S + + T+N
Sbjct: 474 EEALVAFNTMHEVGSNPSIETFHSLLYSFARGGLVKESEAILS--RLVDSGIPRNRDTFN 531
Query: 563 LMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVD 610
+EA +G ++E Y +M S D+ +L S A VD
Sbjct: 532 AQIEAYKQGGKFEEAVKTYVDMEKSRCDPDERTLEAVLSVYSFARLVD 579
Score = 89.4 bits (220), Expect = 6e-18
Identities = 76/353 (21%), Positives = 152/353 (42%), Gaps = 20/353 (5%)
Query: 259 QSRFVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKE-LL 317
+S F YT L+ G+ R + +L++ + M N ++ P + Y+++ + GL E LL
Sbjct: 174 RSVFSYTALINAYGRNGRYETSLELLDRMK-NEKISPSILTYNTVINACARGGLDWEGLL 232
Query: 318 NIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSL 377
+ MR + ++PD+V YN +L+AC VF+ M +
Sbjct: 233 GLFAEMRHEG--------------IQPDIVTYNTLLSACAIRGLGDEAEMVFRTMNDGGI 278
Query: 378 KPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVK 437
P+ TY +E + + V +L +M G +P+ +Y V++ + K G + EA+
Sbjct: 279 VPDLTTYSHLVETFGKLRRLEKVCDLLGEMASGGSLPDITSYNVLLEAYAKSGSIKEAMG 338
Query: 438 AVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSS 497
M+ G A+ Y L GR+ D ++K + P T+ +I
Sbjct: 339 VFHQMQAAGCTPNANTYSVLLNLFGQSGRYDDVRQLFLEMKS-SNTDPDAATYNILIEVF 397
Query: 498 MDGGHIDDCICIF-EYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRP 556
+GG+ + + +F + ++++ P++ T ++ + + A+ + + + +D+ P
Sbjct: 398 GEGGYFKEVVTLFHDMVEENIEPDMETYEGIIFACGKGGLHEDARKILQ--YMTANDIVP 455
Query: 557 DAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKV 609
+ Y ++EA + +E + M G + LL +R G V
Sbjct: 456 SSKAYTGVIEAFGQAALYEEALVAFNTMHEVGSNPSIETFHSLLYSFARGGLV 508
Score = 84.0 bits (206), Expect = 2e-16
Identities = 78/357 (21%), Positives = 144/357 (39%), Gaps = 52/357 (14%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y+ L+ GK RR ++ + M + PD+ +Y+ + ++G +KE + + M
Sbjct: 285 YSHLVETFGKLRRLEKVCDLLGEMASGGSL-PDITSYNVLLEAYAKSGSIKEAMGVFHQM 343
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
+ T P+ Y+ +LN S ++ V +F +M+ S+ P+ AT
Sbjct: 344 QAAGCT--------------PNANTYSVLLNLFGQSGRYDDVRQLFLEMKSSNTDPDAAT 389
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
Y + +EV + G + V LF M P+ TY+ ++ K G ++A K ++ M
Sbjct: 390 YNILIEVFGEGGYFKEVVTLFHDMVEENIEPDMETYEGIIFACGKGGLHEDARKILQYMT 449
Query: 444 RRGVMGTASVY------------YELACCLCNCGRWQDATLEVEKIKRLPHA-------K 484
++ ++ Y YE A N + +E L ++ K
Sbjct: 450 ANDIVPSSKAYTGVIEAFGQAALYEEALVAFNTMHEVGSNPSIETFHSLLYSFARGGLVK 509
Query: 485 PLEV---------------TFTGMIRSSMDGGHIDDCICIFEYMQ-DHCAPNVGTVNTML 528
E TF I + GG ++ + + M+ C P+ T+ +L
Sbjct: 510 ESEAILSRLVDSGIPRNRDTFNAQIEAYKQGGKFEEAVKTYVDMEKSRCDPDERTLEAVL 569
Query: 529 KVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMI 585
VYS + + FEE+K SD+ P Y +ML + +W+ + +EM+
Sbjct: 570 SVYSFARLVDECREQFEEMKA--SDILPSIMCYCMMLAVYGKTERWDDVNELLEEML 624
Score = 57.0 bits (136), Expect = 3e-08
Identities = 62/340 (18%), Positives = 132/340 (38%), Gaps = 52/340 (15%)
Query: 278 KEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKN 337
+ +L++F M I P+ Y + LG+ GLL + L + + M +
Sbjct: 122 QRSLRLFKYMQRQIWCKPNEHIYTIMISLLGREGLLDKCLEVFDEMPSQG---------- 171
Query: 338 WDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSG-N 396
+ V Y A++NA + +++ + +M+ + P+ TY + + G +
Sbjct: 172 ----VSRSVFSYTALINAYGRNGRYETSLELLDRMKNEKISPSILTYNTVINACARGGLD 227
Query: 397 YDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYE 456
++ + LF +M+ G P+ +TY ++ G DEA R M G++ + Y
Sbjct: 228 WEGLLGLFAEMRHEGIQPDIVTYNTLLSACAIRGLGDEAEMVFRTMNDGGIVPDLTTYSH 287
Query: 457 LACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDH 516
L VE +L + + C + E
Sbjct: 288 L----------------VETFGKLRRLEKV-------------------CDLLGEMASGG 312
Query: 517 CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEY 576
P++ + N +L+ Y+++ A +F +++ A P+A TY+++L + +++
Sbjct: 313 SLPDITSYNVLLEAYAKSGSIKEAMGVFHQMQAA--GCTPNANTYSVLLNLFGQSGRYDD 370
Query: 577 FEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVDNILVIY 616
++ EM S D + L+ G ++ ++
Sbjct: 371 VRQLFLEMKSSNTDPDAATYNILIEVFGEGGYFKEVVTLF 410
Score = 55.8 bits (133), Expect = 7e-08
Identities = 48/258 (18%), Positives = 113/258 (43%), Gaps = 4/258 (1%)
Query: 362 WKGVSWVFQQMRKSS-LKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYK 420
W+ +F+ M++ KPN Y + + ++ + G D E+F++M G +Y
Sbjct: 121 WQRSLRLFKYMQRQIWCKPNEHIYTIMISLLGREGLLDKCLEVFDEMPSQGVSRSVFSYT 180
Query: 421 VMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRL 480
++ + + G+ + +++ + M+ + + Y + G + L + R
Sbjct: 181 ALINAYGRNGRYETSLELLDRMKNEKISPSILTYNTVINACARGGLDWEGLLGLFAEMRH 240
Query: 481 PHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQD-HCAPNVGTVNTMLKVYSQNDMFST 539
+P VT+ ++ + G D+ +F M D P++ T + +++ + +
Sbjct: 241 EGIQPDIVTYNTLLSACAIRGLGDEAEMVFRTMNDGGIVPDLTTYSHLVETFGKLRRLEK 300
Query: 540 AKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPL 599
L E +A PD +YN++LEA ++ + V+ +M +G + N + L
Sbjct: 301 VCDLLGE--MASGGSLPDITSYNVLLEAYAKSGSIKEAMGVFHQMQAAGCTPNANTYSVL 358
Query: 600 LVKASRAGKVDNILVIYI 617
L ++G+ D++ +++
Sbjct: 359 LNLFGQSGRYDDVRQLFL 376
Score = 50.1 bits (118), Expect = 4e-06
Identities = 49/240 (20%), Positives = 105/240 (43%), Gaps = 6/240 (2%)
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEV-PEALTYKVMVRTFWKEGKVDEAVKAVRDM 442
+ L + G++ LF+ MQR P Y +M+ +EG +D+ ++ +M
Sbjct: 108 FALVFKEFAGRGDWQRSLRLFKYMQRQIWCKPNEHIYTIMISLLGREGLLDKCLEVFDEM 167
Query: 443 ERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGG- 501
+GV + Y L GR++ + ++++K P +T+ +I + GG
Sbjct: 168 PSQGVSRSVFSYTALINAYGRNGRYETSLELLDRMKN-EKISPSILTYNTVINACARGGL 226
Query: 502 HIDDCICIFEYMQ-DHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYT 560
+ + +F M+ + P++ T NT+L + + A+++F + + PD T
Sbjct: 227 DWEGLLGLFAEMRHEGIQPDIVTYNTLLSACAIRGLGDEAEMVFRTMN--DGGIVPDLTT 284
Query: 561 YNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVDNILVIYIRSQ 620
Y+ ++E + + E + EM G D + LL +++G + + ++ + Q
Sbjct: 285 YSHLVETFGKLRRLEKVCDLLGEMASGGSLPDITSYNVLLEAYAKSGSIKEAMGVFHQMQ 344
>At2g31400 unknown protein
Length = 918
Score = 96.7 bits (239), Expect = 4e-20
Identities = 67/308 (21%), Positives = 142/308 (45%), Gaps = 20/308 (6%)
Query: 262 FVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGL-LKELLNIV 320
+ ++ L++ G++ +EA+ +FN M + P++ Y+++ G+ G+ K++
Sbjct: 269 YAFSALISAYGRSGLHEEAISVFNSMK-EYGLRPNLVTYNAVIDACGKGGMEFKQVAKFF 327
Query: 321 ECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPN 380
+ M++ ++PD + +N++L C W+ +F +M ++ +
Sbjct: 328 DEMQRNG--------------VQPDRITFNSLLAVCSRGGLWEAARNLFDEMTNRRIEQD 373
Query: 381 GATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVR 440
+Y ++ + + G DL E+ +M +P ++Y ++ F K G+ DEA+
Sbjct: 374 VFSYNTLLDAICKGGQMDLAFEILAQMPVKRIMPNVVSYSTVIDGFAKAGRFDEALNLFG 433
Query: 441 DMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDG 500
+M G+ Y L GR ++A L++ + K VT+ ++
Sbjct: 434 EMRYLGIALDRVSYNTLLSIYTKVGRSEEA-LDILREMASVGIKKDVVTYNALLGGYGKQ 492
Query: 501 GHIDDCICIF-EYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAY 559
G D+ +F E ++H PN+ T +T++ YS+ ++ A +F E K A LR D
Sbjct: 493 GKYDEVKKVFTEMKREHVLPNLLTYSTLIDGYSKGGLYKEAMEIFREFKSA--GLRADVV 550
Query: 560 TYNLMLEA 567
Y+ +++A
Sbjct: 551 LYSALIDA 558
Score = 82.8 bits (203), Expect = 5e-16
Identities = 76/370 (20%), Positives = 167/370 (44%), Gaps = 29/370 (7%)
Query: 229 IIEMLGVKRCWKQALSVVQWVYNSKDHRKFQSRFVYTKLLAVLGKARRPKEALQIFNMML 288
II LG + +A+ ++ ++ RK + + + +++ LG+ + A +IF
Sbjct: 202 IIRELGNRNECDKAVGFYEFAVK-RERRKNEQGKLASAMISTLGRYGKVTIAKRIFETAF 260
Query: 289 ----GNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNWDPTLEP 344
GN VY A+ ++ G++GL +E +++ M++ L P
Sbjct: 261 AGGYGNT-VY----AFSALISAYGRSGLHEEAISVFNSMKEYG--------------LRP 301
Query: 345 DVVIYNAVLNACVPS-KQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHEL 403
++V YNAV++AC ++K V+ F +M+++ ++P+ T+ + V + G ++ L
Sbjct: 302 NLVTYNAVIDACGKGGMEFKQVAKFFDEMQRNGVQPDRITFNSLLAVCSRGGLWEAARNL 361
Query: 404 FEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCN 463
F++M + +Y ++ K G++D A + + M + +M Y +
Sbjct: 362 FDEMTNRRIEQDVFSYNTLLDAICKGGQMDLAFEILAQMPVKRIMPNVVSYSTVIDGFAK 421
Query: 464 CGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQD-HCAPNVG 522
GR+ +A +++ L A V++ ++ G ++ + I M +V
Sbjct: 422 AGRFDEALNLFGEMRYLGIALD-RVSYNTLLSIYTKVGRSEEALDILREMASVGIKKDVV 480
Query: 523 TVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYK 582
T N +L Y + + K +F E+K + + P+ TY+ +++ S+G ++ +++
Sbjct: 481 TYNALLGGYGKQGKYDEVKKVFTEMK--REHVLPNLLTYSTLIDGYSKGGLYKEAMEIFR 538
Query: 583 EMILSGYHLD 592
E +G D
Sbjct: 539 EFKSAGLRAD 548
Score = 68.2 bits (165), Expect = 1e-11
Identities = 76/354 (21%), Positives = 132/354 (36%), Gaps = 48/354 (13%)
Query: 239 WKQALSVVQWVYNSKDHRKFQSRFVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMA 298
W+ A ++ + N R Q F Y LL + K + A +I M R+ P++
Sbjct: 355 WEAARNLFDEMTN---RRIEQDVFSYNTLLDAICKGGQMDLAFEILAQMPVK-RIMPNVV 410
Query: 299 AYHSIAVTLGQAGLLKELLNIVECMRQ---------------------KPETLKYMYRKN 337
+Y ++ +AG E LN+ MR + E + R+
Sbjct: 411 SYSTVIDGFAKAGRFDEALNLFGEMRYLGIALDRVSYNTLLSIYTKVGRSEEALDILREM 470
Query: 338 WDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNY 397
++ DVV YNA+L ++ V VF +M++ + PN TY ++ + G Y
Sbjct: 471 ASVGIKKDVVTYNALLGGYGKQGKYDEVKKVFTEMKREHVLPNLLTYSTLIDGYSKGGLY 530
Query: 398 DLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYEL 457
E+F + + G + + Y ++ K G V AV + +M + G+ Y +
Sbjct: 531 KEAMEIFREFKSAGLRADVVLYSALIDALCKNGLVGSAVSLIDEMTKEGISPNVVTYNSI 590
Query: 458 ACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFT-----------GMIRSSMDGGHIDDC 506
D + + LP + T G + + + DC
Sbjct: 591 IDAFGRSAT-MDRSADYSNGGSLPFSSSALSALTETEGNRVIQLFGQLTTESNNRTTKDC 649
Query: 507 -------ICIFEYM----QDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKV 549
CI E Q PNV T + +L S+ + F A +L EE+++
Sbjct: 650 EEGMQELSCILEVFRKMHQLEIKPNVVTFSAILNACSRCNSFEDASMLLEELRL 703
Score = 54.3 bits (129), Expect = 2e-07
Identities = 57/272 (20%), Positives = 110/272 (39%), Gaps = 38/272 (13%)
Query: 346 VVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSG-NYDLVHELF 404
V ++A+++A S + VF M++ L+PN TY ++ + G + V + F
Sbjct: 268 VYAFSALISAYGRSGLHEEAISVFNSMKEYGLRPNLVTYNAVIDACGKGGMEFKQVAKFF 327
Query: 405 EKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNC 464
++MQRNG P+ +T+ ++ + G + A +M R + Y L +C
Sbjct: 328 DEMQRNGVQPDRITFNSLLAVCSRGGLWEAARNLFDEMTNRRIEQDVFSYNTLLDAICKG 387
Query: 465 GRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDHCAPNVGTV 524
G+ L E + ++P + + PNV +
Sbjct: 388 GQMD---LAFEILAQMPVKRIM--------------------------------PNVVSY 412
Query: 525 NTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEM 584
+T++ +++ F A LF E++ L D +YN +L ++ + E + +EM
Sbjct: 413 STVIDGFAKAGRFDEALNLFGEMRYLGIAL--DRVSYNTLLSIYTKVGRSEEALDILREM 470
Query: 585 ILSGYHLDQNKHLPLLVKASRAGKVDNILVIY 616
G D + LL + GK D + ++
Sbjct: 471 ASVGIKKDVVTYNALLGGYGKQGKYDEVKKVF 502
Score = 47.4 bits (111), Expect = 3e-05
Identities = 51/226 (22%), Positives = 98/226 (42%), Gaps = 19/226 (8%)
Query: 394 SGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVK----AVRDMERRGVMG 449
SG+ ++ H L + + TY ++R + D+AV AV+ R+ G
Sbjct: 176 SGDDEMFHSLMLSFESKLCGSDDCTY--IIRELGNRNECDKAVGFYEFAVKRERRKNEQG 233
Query: 450 TASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVT---FTGMIRSSMDGGHIDDC 506
+ + + GR+ T+ ++I A T F+ +I + G ++
Sbjct: 234 KLA-----SAMISTLGRYGKVTI-AKRIFETAFAGGYGNTVYAFSALISAYGRSGLHEEA 287
Query: 507 ICIFEYMQDH-CAPNVGTVNTMLKVYSQNDM-FSTAKVLFEEVKVAKSDLRPDAYTYNLM 564
I +F M+++ PN+ T N ++ + M F F+E++ ++ ++PD T+N +
Sbjct: 288 ISVFNSMKEYGLRPNLVTYNAVIDACGKGGMEFKQVAKFFDEMQ--RNGVQPDRITFNSL 345
Query: 565 LEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVD 610
L SRG WE +++ EM D + LL + G++D
Sbjct: 346 LAVCSRGGLWEAARNLFDEMTNRRIEQDVFSYNTLLDAICKGGQMD 391
Score = 40.4 bits (93), Expect = 0.003
Identities = 26/103 (25%), Positives = 47/103 (45%), Gaps = 10/103 (9%)
Query: 333 MYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAM---- 388
++RK ++P+VV ++A+LNAC ++ S + +++R K G +GL M
Sbjct: 662 VFRKMHQLEIKPNVVTFSAILNACSRCNSFEDASMLLEELRLFDNKVYGVVHGLLMGQRE 721
Query: 389 EVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGK 431
V LQ+ LF+K+ + Y + W G+
Sbjct: 722 NVWLQA------QSLFDKVNEMDGSTASAFYNALTDMLWHFGQ 758
>At3g53700 putative protein
Length = 754
Score = 96.3 bits (238), Expect = 5e-20
Identities = 85/381 (22%), Positives = 168/381 (43%), Gaps = 34/381 (8%)
Query: 240 KQALSVVQWVYNSKDHRKFQSRFVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAA 299
+ AL+ +Q + N F ++ + L+ L KA K A++I ++ML PD+
Sbjct: 276 EDALNFIQEMSNQDGF--FPDQYTFNTLVNGLCKAGHVKHAIEIMDVMLQE-GYDPDVYT 332
Query: 300 YHSIAVTLGQAGLLKELLNIVECMRQK---PETLKY------------------MYRKNW 338
Y+S+ L + G +KE + +++ M + P T+ Y + R
Sbjct: 333 YNSVISGLCKLGEVKEAVEVLDQMITRDCSPNTVTYNTLISTLCKENQVEEATELARVLT 392
Query: 339 DPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYD 398
+ PDV +N+++ ++ + +F++MR +P+ TY + ++ + G D
Sbjct: 393 SKGILPDVCTFNSLIQGLCLTRNHRVAMELFEEMRSKGCEPDEFTYNMLIDSLCSKGKLD 452
Query: 399 LVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELA 458
+ ++M+ +G +TY ++ F K K EA + +ME GV + Y L
Sbjct: 453 EALNMLKQMELSGCARSVITYNTLIDGFCKANKTREAEEIFDEMEVHGVSRNSVTYNTLI 512
Query: 459 CCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDH-C 517
LC R +DA ++++ + KP + T+ ++ GG I I + M + C
Sbjct: 513 DGLCKSRRVEDAAQLMDQM-IMEGQKPDKYTYNSLLTHFCRGGDIKKAADIVQAMTSNGC 571
Query: 518 APNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYF 577
P++ T T++ + A L +++ +L P A YN +++ R +
Sbjct: 572 EPDIVTYGTLISGLCKAGRVEVASKLLRSIQMKGINLTPHA--YNPVIQGLFRKRKTTEA 629
Query: 578 EHVYKEMILSGYHLDQNKHLP 598
++++EM L+QN+ P
Sbjct: 630 INLFREM------LEQNEAPP 644
Score = 77.4 bits (189), Expect = 2e-14
Identities = 72/333 (21%), Positives = 137/333 (40%), Gaps = 25/333 (7%)
Query: 266 KLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQ 325
KLL L AL++FN+ P+ A Y I + LG++G ++ I+E M+
Sbjct: 52 KLLDSLRSQPDDSAALRLFNLASKKPNFSPEPALYEEILLRLGRSGSFDDMKKILEDMKS 111
Query: 326 ----------------------KPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWK 363
+ E L + + L+PD YN +LN V K
Sbjct: 112 SRCEMGTSTFLILIESYAQFELQDEILSVVDWMIDEFGLKPDTHFYNRMLNLLVDGNSLK 171
Query: 364 GVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMV 423
V +M +KP+ +T+ + ++ + ++ + E M G VP+ T+ ++
Sbjct: 172 LVEISHAKMSVWGIKPDVSTFNVLIKALCRAHQLRPAILMLEDMPSYGLVPDEKTFTTVM 231
Query: 424 RTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHA 483
+ + +EG +D A++ M G + + C GR +DA ++++
Sbjct: 232 QGYIEEGDLDGALRIREQMVEFGCSWSNVSVNVIVHGFCKEGRVEDALNFIQEMSNQDGF 291
Query: 484 KPLEVTFTGMIRSSMDGGHIDDCICIFEYM-QDHCAPNVGTVNTMLKVYSQNDMFSTAKV 542
P + TF ++ GH+ I I + M Q+ P+V T N+++ + A
Sbjct: 292 FPDQYTFNTLVNGLCKAGHVKHAIEIMDVMLQEGYDPDVYTYNSVISGLCKLGEVKEAVE 351
Query: 543 LFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWE 575
+ ++ + D P+ TYN ++ + +Q E
Sbjct: 352 VLDQ--MITRDCSPNTVTYNTLISTLCKENQVE 382
Score = 76.3 bits (186), Expect = 5e-14
Identities = 79/357 (22%), Positives = 146/357 (40%), Gaps = 21/357 (5%)
Query: 279 EALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNW 338
E L + + M+ + PD Y+ + L LK +VE K W
Sbjct: 136 EILSVVDWMIDEFGLKPDTHFYNRMLNLLVDGNSLK----LVEISHAKMSV--------W 183
Query: 339 DPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYD 398
++PDV +N ++ A + Q + + + M L P+ T+ M+ ++ G+ D
Sbjct: 184 G--IKPDVSTFNVLIKALCRAHQLRPAILMLEDMPSYGLVPDEKTFTTVMQGYIEEGDLD 241
Query: 399 LVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERR-GVMGTASVYYEL 457
+ E+M G ++ V+V F KEG+V++A+ +++M + G + L
Sbjct: 242 GALRIREQMVEFGCSWSNVSVNVIVHGFCKEGRVEDALNFIQEMSNQDGFFPDQYTFNTL 301
Query: 458 ACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYM-QDH 516
LC G + A +E+ + P T+ +I G + + + + + M
Sbjct: 302 VNGLCKAGHVKHA-IEIMDVMLQEGYDPDVYTYNSVISGLCKLGEVKEAVEVLDQMITRD 360
Query: 517 CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEY 576
C+PN T NT++ + + A L V +K L PD T+N +++
Sbjct: 361 CSPNTVTYNTLISTLCKENQVEEATEL-ARVLTSKGIL-PDVCTFNSLIQGLCLTRNHRV 418
Query: 577 FEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVDNILVIYIRSQCLFYISAHSIISY 633
+++EM G D+ + L+ GK+D L + Q A S+I+Y
Sbjct: 419 AMELFEEMRSKGCEPDEFTYNMLIDSLCSKGKLDEAL--NMLKQMELSGCARSVITY 473
Score = 49.7 bits (117), Expect = 5e-06
Identities = 69/328 (21%), Positives = 135/328 (41%), Gaps = 25/328 (7%)
Query: 179 RKKTSKRSEVQVVRF--LVDRLCDKEIRAKDWKFSRLMKLSGLSFTEGQLMMIIEMLGVK 236
R TSK V F L+ LC + M+ G E M+I+ L K
Sbjct: 389 RVLTSKGILPDVCTFNSLIQGLCLTRNHRVAMELFEEMRSKGCEPDEFTYNMLIDSLCSK 448
Query: 237 RCWKQALSVVQWVYNSKDHRKFQSRFVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPD 296
+AL++++ + S R S Y L+ KA + +EA +IF+ M + V +
Sbjct: 449 GKLDEALNMLKQMELSGCAR---SVITYNTLIDGFCKANKTREAEEIFDEMEVH-GVSRN 504
Query: 297 MAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNAC 356
Y+++ L ++ +++ +++ M + + +PD YN++L
Sbjct: 505 SVTYNTLIDGLCKSRRVEDAAQLMDQMIMEGQ--------------KPDKYTYNSLLTHF 550
Query: 357 VPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEA 416
K + + Q M + +P+ TYG + + ++G ++ +L +Q G
Sbjct: 551 CRGGDIKKAADIVQAMTSNGCEPDIVTYGTLISGLCKAGRVEVASKLLRSIQMKGINLTP 610
Query: 417 LTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELAC-CLCNCG----RWQDAT 471
Y +++ +++ K EA+ R+M + +V Y + LCN G D
Sbjct: 611 HAYNPVIQGLFRKRKTTEAINLFREMLEQNEAPPDAVSYRIVFRGLCNGGGPIREAVDFL 670
Query: 472 LEVEKIKRLPHAKPLEVTFTGMIRSSMD 499
+E+ + +P L + G++ SM+
Sbjct: 671 VELLEKGFVPEFSSLYMLAEGLLTLSME 698
>At4g39620 putative protein
Length = 563
Score = 94.4 bits (233), Expect = 2e-19
Identities = 88/411 (21%), Positives = 182/411 (43%), Gaps = 24/411 (5%)
Query: 180 KKTSKRSEVQVVRFLVDRLCDKEIRAKDW-KFSRLMKLSGLSFTEGQLMMIIEMLGVKRC 238
+++++R +VR L+ R+ D+E K K+ ++++ ++ E LG
Sbjct: 60 RESAERENRVLVRSLMSRISDREPLVKTLDKYVKVVRCD-------HCFLLFEELGKSDK 112
Query: 239 WKQALSVVQWVYNSKDHRKFQSRFVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMA 298
W Q L V +W+ K VY+KL++V+GK + + A+ +F+ M N PD +
Sbjct: 113 WLQCLEVFRWM--QKQRWYIPDNGVYSKLISVMGKKGQTRMAMWLFSEMK-NSGCRPDAS 169
Query: 299 AYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVP 358
Y++ L+ L+ + + + Y+ + +P+VV YN +L A
Sbjct: 170 VYNA---------LITAHLHTRDKAKALEKVRGYLDKMKGIERCQPNVVTYNILLRAFAQ 220
Query: 359 SKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALT 418
S + V+ +F+ + S + P+ T+ M+ ++G + + +M+ N P+ +T
Sbjct: 221 SGKVDQVNALFKDLDMSPVSPDVYTFNGVMDAYGKNGMIKEMEAVLTRMRSNECKPDIIT 280
Query: 419 YKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIK 478
+ V++ ++ K+ + ++ + + + R T + + A +K+
Sbjct: 281 FNVLIDSYGKKQEFEKMEQTFKSLMRSKEKPTLPTFNSMIINYGKARMIDKAEWVFKKMN 340
Query: 479 RLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYM-QDHCAPNVGTVNTMLKVYSQNDMF 537
+ + P +T+ MI G + IFE + + T+N ML+VY +N ++
Sbjct: 341 DMNYI-PSFITYECMIMMYGYCGSVSRAREIFEEVGESDRVLKASTLNAMLEVYCRNGLY 399
Query: 538 STAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSG 588
A LF + + PDA TY + +A ++ E + + K+M G
Sbjct: 400 IEADKLFHNASAFR--VHPDASTYKFLYKAYTKADMKEQVQILMKKMEKDG 448
Score = 63.5 bits (153), Expect = 3e-10
Identities = 49/239 (20%), Positives = 109/239 (45%), Gaps = 10/239 (4%)
Query: 402 ELFEKMQRNG-EVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACC 460
E+F MQ+ +P+ Y ++ K+G+ A+ +M+ G ASVY L
Sbjct: 118 EVFRWMQKQRWYIPDNGVYSKLISVMGKKGQTRMAMWLFSEMKNSGCRPDASVYNALITA 177
Query: 461 LCNCGRWQDATLEV----EKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQ-D 515
+ A +V +K+K + +P VT+ ++R+ G +D +F+ +
Sbjct: 178 HLHTRDKAKALEKVRGYLDKMKGIERCQPNVVTYNILLRAFAQSGKVDQVNALFKDLDMS 237
Query: 516 HCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWE 575
+P+V T N ++ Y +N M + + ++ ++ +PD T+N+++++ + ++E
Sbjct: 238 PVSPDVYTFNGVMDAYGKNGMIKEMEAVLTRMR--SNECKPDIITFNVLIDSYGKKQEFE 295
Query: 576 YFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVDNILVIYIRSQCLFYISAHSIISYK 634
E +K ++ S +++ +A +D ++ + + YI S I+Y+
Sbjct: 296 KMEQTFKSLMRSKEKPTLPTFNSMIINYGKARMIDKAEWVFKKMNDMNYIP--SFITYE 352
Score = 40.0 bits (92), Expect = 0.004
Identities = 32/193 (16%), Positives = 80/193 (40%), Gaps = 15/193 (7%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
+ L+ GK + ++ Q F ++ + + P + ++S+ + G+A ++
Sbjct: 281 FNVLIDSYGKKQEFEKMEQTFKSLMRS-KEKPTLPTFNSMIINYGKARMI---------- 329
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
+ +++++K D P + Y ++ +F+++ +S +T
Sbjct: 330 ----DKAEWVFKKMNDMNYIPSFITYECMIMMYGYCGSVSRAREIFEEVGESDRVLKAST 385
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
+EV ++G Y +LF P+A TYK + + + K ++ ++ ME
Sbjct: 386 LNAMLEVYCRNGLYIEADKLFHNASAFRVHPDASTYKFLYKAYTKADMKEQVQILMKKME 445
Query: 444 RRGVMGTASVYYE 456
+ G++ + E
Sbjct: 446 KDGIVPNKRFFLE 458
>At1g05665 putative protein
Length = 732
Score = 94.4 bits (233), Expect = 2e-19
Identities = 78/320 (24%), Positives = 140/320 (43%), Gaps = 26/320 (8%)
Query: 296 DMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNA 355
++A+Y+ + + Q G +KE +++ M K T PDV+ Y+ V+N
Sbjct: 236 NVASYNIVIHFVCQLGRIKEAHHLLLLMELKGYT--------------PDVISYSTVVNG 281
Query: 356 CVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPE 415
+ V + + M++ LKPN YG + ++ + E F +M R G +P+
Sbjct: 282 YCRFGELDKVWKLIEVMKRKGLKPNSYIYGSIIGLLCRICKLAEAEEAFSEMIRQGILPD 341
Query: 416 ALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVE 475
+ Y ++ F K G + A K +M R + Y + C G +E
Sbjct: 342 TVVYTTLIDGFCKRGDIRAASKFFYEMHSRDITPDVLTYTAIISGFCQIG----DMVEAG 397
Query: 476 KIKRLPHAKPLE---VTFTGMIRSSMDGGHIDDCICIFEYM-QDHCAPNVGTVNTMLK-V 530
K+ K LE VTFT +I GH+ D + +M Q C+PNV T T++ +
Sbjct: 398 KLFHEMFCKGLEPDSVTFTELINGYCKAGHMKDAFRVHNHMIQAGCSPNVVTYTTLIDGL 457
Query: 531 YSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYH 590
+ D+ S ++L E K+ L+P+ +TYN ++ + E + E +G +
Sbjct: 458 CKEGDLDSANELLHEMWKIG---LQPNIFTYNSIVNGLCKSGNIEEAVKLVGEFEAAGLN 514
Query: 591 LDQNKHLPLLVKASRAGKVD 610
D + L+ ++G++D
Sbjct: 515 ADTVTYTTLMDAYCKSGEMD 534
Score = 88.6 bits (218), Expect = 1e-17
Identities = 76/361 (21%), Positives = 148/361 (40%), Gaps = 26/361 (7%)
Query: 262 FVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVE 321
++Y ++ +L + + EA + F+ M+ + PD Y ++ + G ++
Sbjct: 308 YIYGSIIGLLCRICKLAEAEEAFSEMIRQ-GILPDTVVYTTLIDGFCKRGDIRAASKFFY 366
Query: 322 CMRQK---PETLKY------------------MYRKNWDPTLEPDVVIYNAVLNACVPSK 360
M + P+ L Y ++ + + LEPD V + ++N +
Sbjct: 367 EMHSRDITPDVLTYTAIISGFCQIGDMVEAGKLFHEMFCKGLEPDSVTFTELINGYCKAG 426
Query: 361 QWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYK 420
K V M ++ PN TY ++ + + G+ D +EL +M + G P TY
Sbjct: 427 HMKDAFRVHNHMIQAGCSPNVVTYTTLIDGLCKEGDLDSANELLHEMWKIGLQPNIFTYN 486
Query: 421 VMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRL 480
+V K G ++EAVK V + E G+ Y L C G D E+ K
Sbjct: 487 SIVNGLCKSGNIEEAVKLVGEFEAAGLNADTVTYTTLMDAYCKSGE-MDKAQEILKEMLG 545
Query: 481 PHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYM-QDHCAPNVGTVNTMLKVYSQNDMFST 539
+P VTF ++ G ++D + +M APN T N+++K Y +
Sbjct: 546 KGLQPTIVTFNVLMNGFCLHGMLEDGEKLLNWMLAKGIAPNATTFNSLVKQYCIRNNLKA 605
Query: 540 AKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPL 599
A ++++ + + PD TY +++ + + +++EM G+ + + + L
Sbjct: 606 ATAIYKD--MCSRGVGPDGKTYENLVKGHCKARNMKEAWFLFQEMKGKGFSVSVSTYSVL 663
Query: 600 L 600
+
Sbjct: 664 I 664
Score = 62.4 bits (150), Expect = 8e-10
Identities = 57/250 (22%), Positives = 108/250 (42%), Gaps = 14/250 (5%)
Query: 368 VFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFW 427
VF++ + + N A+Y + + + Q G H L M+ G P+ ++Y +V +
Sbjct: 224 VFREFPEVGVCWNVASYNIVIHFVCQLGRIKEAHHLLLLMELKGYTPDVISYSTVVNGYC 283
Query: 428 KEGKVDEAVKAVRDMERRGVMGTASVY---YELACCLCNCGRWQDATLEVEKIKRLPHAK 484
+ G++D+ K + M+R+G+ + +Y L C +C ++A E+ + LP
Sbjct: 284 RFGELDKVWKLIEVMKRKGLKPNSYIYGSIIGLLCRICKLAEAEEAFSEMIRQGILPDT- 342
Query: 485 PLEVTFTGMIRSSMDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQ-NDMFSTAKV 542
V +T +I G I F M P+V T ++ + Q DM K+
Sbjct: 343 ---VVYTTLIDGFCKRGDIRAASKFFYEMHSRDITPDVLTYTAIISGFCQIGDMVEAGKL 399
Query: 543 LFEEVKVAKSDLRPDAYTYNLMLEASSR-GHQWEYFEHVYKEMILSGYHLDQNKHLPLLV 601
E + L PD+ T+ ++ + GH + F V+ MI +G + + L+
Sbjct: 400 FHE---MFCKGLEPDSVTFTELINGYCKAGHMKDAF-RVHNHMIQAGCSPNVVTYTTLID 455
Query: 602 KASRAGKVDN 611
+ G +D+
Sbjct: 456 GLCKEGDLDS 465
Score = 47.0 bits (110), Expect = 3e-05
Identities = 67/335 (20%), Positives = 123/335 (36%), Gaps = 53/335 (15%)
Query: 141 LIELKEMFRERKMDELKWVFEDDIEINEVWFDENNNGKRKKTSKRSEVQVVRFLVDRLCD 200
++E ++F E +F +E + V F E NG K + +V ++ C
Sbjct: 393 MVEAGKLFHE--------MFCKGLEPDSVTFTELINGYCKAGHMKDAFRVHNHMIQAGCS 444
Query: 201 KEIRAKDWKFSRLMKLSGLSFTEGQLMMIIEMLGVKRCWKQALSVVQWVYNS-------- 252
+ L K EG L E+L WK L + YNS
Sbjct: 445 PNVVTYTTLIDGLCK-------EGDLDSANELL--HEMWKIGLQPNIFTYNSIVNGLCKS 495
Query: 253 ---KDHRKFQSRF----------VYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAA 299
++ K F YT L+ K+ +A +I MLG + P +
Sbjct: 496 GNIEEAVKLVGEFEAAGLNADTVTYTTLMDAYCKSGEMDKAQEILKEMLGK-GLQPTIVT 554
Query: 300 YHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPS 359
++ + G+L++ + L +M K + P+ +N+++
Sbjct: 555 FNVLMNGFCLHGMLED----------GEKLLNWMLAKG----IAPNATTFNSLVKQYCIR 600
Query: 360 KQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTY 419
K + +++ M + P+G TY ++ ++ N LF++M+ G TY
Sbjct: 601 NNLKAATAIYKDMCSRGVGPDGKTYENLVKGHCKARNMKEAWFLFQEMKGKGFSVSVSTY 660
Query: 420 KVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVY 454
V+++ F K K EA + M R G+ ++
Sbjct: 661 SVLIKGFLKRKKFLEAREVFDQMRREGLAADKEIF 695
>At3g04760 unknown protein
Length = 602
Score = 92.4 bits (228), Expect = 7e-19
Identities = 77/333 (23%), Positives = 142/333 (42%), Gaps = 25/333 (7%)
Query: 279 EALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNW 338
EAL++ + ML + PDM Y++I + + G++ +V + K
Sbjct: 246 EALKLMDEMLSR-GLKPDMFTYNTIIRGMCKEGMVDRAFEMVRNLELKG----------- 293
Query: 339 DPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYD 398
EPDV+ YN +L A + +W+ + +M PN TY + + + + G +
Sbjct: 294 ---CEPDVISYNILLRALLNQGKWEEGEKLMTKMFSEKCDPNVVTYSILITTLCRDGKIE 350
Query: 399 LVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELA 458
L + M+ G P+A +Y ++ F +EG++D A++ + M G + Y +
Sbjct: 351 EAMNLLKLMKEKGLTPDAYSYDPLIAAFCREGRLDVAIEFLETMISDGCLPDIVNYNTVL 410
Query: 459 CCLCNCGRWQDATLEV----EKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQ 514
LC G+ D LE+ ++ P++ F+ + S G I I E M
Sbjct: 411 ATLCKNGK-ADQALEIFGKLGEVGCSPNSSSYNTMFSALWSS---GDKIRALHMILEMMS 466
Query: 515 DHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQW 574
+ P+ T N+M+ + M A L V + + P TYN++L + H+
Sbjct: 467 NGIDPDEITYNSMISCLCREGMVDEAFELL--VDMRSCEFHPSVVTYNIVLLGFCKAHRI 524
Query: 575 EYFEHVYKEMILSGYHLDQNKHLPLLVKASRAG 607
E +V + M+ +G ++ + L+ AG
Sbjct: 525 EDAINVLESMVGNGCRPNETTYTVLIEGIGFAG 557
Score = 90.5 bits (223), Expect = 3e-18
Identities = 65/275 (23%), Positives = 124/275 (44%), Gaps = 4/275 (1%)
Query: 343 EPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHE 402
+PDV YNA++N + + V +MR P+ TY + + + G DL +
Sbjct: 155 QPDVFAYNALINGFCKMNRIDDATRVLDRMRSKDFSPDTVTYNIMIGSLCSRGKLDLALK 214
Query: 403 LFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLC 462
+ ++ + P +TY +++ EG VDEA+K + +M RG+ Y + +C
Sbjct: 215 VLNQLLSDNCQPTVITYTILIEATMLEGGVDEALKLMDEMLSRGLKPDMFTYNTIIRGMC 274
Query: 463 NCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYM-QDHCAPNV 521
G D E+ + L +P +++ ++R+ ++ G ++ + M + C PNV
Sbjct: 275 KEG-MVDRAFEMVRNLELKGCEPDVISYNILLRALLNQGKWEEGEKLMTKMFSEKCDPNV 333
Query: 522 GTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVY 581
T + ++ ++ A L + +K + L PDAY+Y+ ++ A R + +
Sbjct: 334 VTYSILITTLCRDGKIEEAMNLLKLMK--EKGLTPDAYSYDPLIAAFCREGRLDVAIEFL 391
Query: 582 KEMILSGYHLDQNKHLPLLVKASRAGKVDNILVIY 616
+ MI G D + +L + GK D L I+
Sbjct: 392 ETMISDGCLPDIVNYNTVLATLCKNGKADQALEIF 426
Score = 57.8 bits (138), Expect = 2e-08
Identities = 52/217 (23%), Positives = 91/217 (40%), Gaps = 5/217 (2%)
Query: 393 QSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTAS 452
+SGNY L E M R G P+ + +++ F+ + +AV+ + +E+ G
Sbjct: 101 RSGNYIESLHLLETMVRKGYNPDVILCTKLIKGFFTLRNIPKAVRVMEILEKFG-QPDVF 159
Query: 453 VYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIF-E 511
Y L C R DAT ++++ R P VT+ MI S G +D + + +
Sbjct: 160 AYNALINGFCKMNRIDDATRVLDRM-RSKDFSPDTVTYNIMIGSLCSRGKLDLALKVLNQ 218
Query: 512 YMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRG 571
+ D+C P V T +++ A L +E + L+PD +TYN ++ +
Sbjct: 219 LLSDNCQPTVITYTILIEATMLEGGVDEALKLMDE--MLSRGLKPDMFTYNTIIRGMCKE 276
Query: 572 HQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGK 608
+ + + + L G D + LL GK
Sbjct: 277 GMVDRAFEMVRNLELKGCEPDVISYNILLRALLNQGK 313
Score = 48.1 bits (113), Expect = 2e-05
Identities = 41/182 (22%), Positives = 76/182 (41%), Gaps = 17/182 (9%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAG-LLKELLNIVEC 322
Y +LA L K + +AL+IF LG + P+ ++Y+++ L +G ++ L I+E
Sbjct: 406 YNTVLATLCKNGKADQALEIFGK-LGEVGCSPNSSSYNTMFSALWSSGDKIRALHMILEM 464
Query: 323 MRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGA 382
M ++PD + YN++++ + MR P+
Sbjct: 465 MSNG---------------IDPDEITYNSMISCLCREGMVDEAFELLVDMRSCEFHPSVV 509
Query: 383 TYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDM 442
TY + + ++ + + E M NG P TY V++ G EA++ D+
Sbjct: 510 TYNIVLLGFCKAHRIEDAINVLESMVGNGCRPNETTYTVLIEGIGFAGYRAEAMELANDL 569
Query: 443 ER 444
R
Sbjct: 570 VR 571
>At1g02060 unknown protein
Length = 710
Score = 92.0 bits (227), Expect = 9e-19
Identities = 59/236 (25%), Positives = 111/236 (47%), Gaps = 3/236 (1%)
Query: 368 VFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRN-GEVPEALTYKVMVRTF 426
+FQ M++ + P+ T+ + ++L+ G + H+LF++M+R G P++ T+ ++ F
Sbjct: 160 LFQTMKQMGISPSVLTFNSLLSILLKRGRTGMAHDLFDEMRRTYGVTPDSYTFNTLINGF 219
Query: 427 WKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDA-TLEVEKIKRLPHAKP 485
K VDEA + +DME Y + LC G+ + A + +K+ P
Sbjct: 220 CKNSMVDEAFRIFKDMELYHCNPDVVTYNTIIDGLCRAGKVKIAHNVLSGMLKKATDVHP 279
Query: 486 LEVTFTGMIRSSMDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDMFSTAKVLF 544
V++T ++R ID+ + +F M PN T NT++K S+ + K +
Sbjct: 280 NVVSYTTLVRGYCMKQEIDEAVLVFHDMLSRGLKPNAVTYNTLIKGLSEAHRYDEIKDIL 339
Query: 545 EEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLL 600
A + PDA T+N++++A + V++EM+ H D + L+
Sbjct: 340 IGGNDAFTTFAPDACTFNILIKAHCDAGHLDAAMKVFQEMLNMKLHPDSASYSVLI 395
Score = 89.4 bits (220), Expect = 6e-18
Identities = 93/434 (21%), Positives = 183/434 (41%), Gaps = 40/434 (9%)
Query: 203 IRAKDWKFSRLMKLSGLSFTEGQLMMIIEMLGVKRCWKQALSVVQWVYN-SKDHRKFQSR 261
+R DW ++ G S E +++E LG R A + + + S K Q R
Sbjct: 85 LRFFDWVSNK-----GFSHKEQSFFLMLEFLGRARNLNVARNFLFSIERRSNGCVKLQDR 139
Query: 262 FVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTL---GQAGLLKELLN 318
+ + L+ G A +E++++F M + + P + ++S+ L G+ G+ +L +
Sbjct: 140 Y-FNSLIRSYGNAGLFQESVKLFQTMK-QMGISPSVLTFNSLLSILLKRGRTGMAHDLFD 197
Query: 319 IVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLK 378
+ R+ + T PD +N ++N + +F+ M
Sbjct: 198 EM--------------RRTYGVT--PDSYTFNTLINGFCKNSMVDEAFRIFKDMELYHCN 241
Query: 379 PNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGE--VPEALTYKVMVRTFWKEGKVDEAV 436
P+ TY ++ + ++G + H + M + P ++Y +VR + + ++DEAV
Sbjct: 242 PDVVTYNTIIDGLCRAGKVKIAHNVLSGMLKKATDVHPNVVSYTTLVRGYCMKQEIDEAV 301
Query: 437 KAVRDMERRGVMGTASVYYELACCLCNCGRWQD-ATLEVEKIKRLPHAKPLEVTFTGMIR 495
DM RG+ A Y L L R+ + + + P TF +I+
Sbjct: 302 LVFHDMLSRGLKPNAVTYNTLIKGLSEAHRYDEIKDILIGGNDAFTTFAPDACTFNILIK 361
Query: 496 SSMDGGHIDDCICIFEYMQD-HCAPNVGTVNTMLKVYSQNDMFSTAKVLF-----EEVKV 549
+ D GH+D + +F+ M + P+ + + +++ + F A+ LF +EV +
Sbjct: 362 AHCDAGHLDAAMKVFQEMLNMKLHPDSASYSVLIRTLCMRNEFDRAETLFNELFEKEVLL 421
Query: 550 AKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKV 609
K + +P A YN M E + + E V+++++ G D + L+ R GK
Sbjct: 422 GKDECKPLAAAYNPMFEYLCANGKTKQAEKVFRQLMKRGVQ-DPPSYKTLITGHCREGKF 480
Query: 610 D---NILVIYIRSQ 620
+LV+ +R +
Sbjct: 481 KPAYELLVLMLRRE 494
Score = 63.9 bits (154), Expect = 3e-10
Identities = 65/316 (20%), Positives = 122/316 (38%), Gaps = 29/316 (9%)
Query: 262 FVYTKLLAVLGKARRPKEALQIFNMMLGNIRVY---PDMAAYHSIAVTLGQAGLLKELLN 318
+ + L+ K EA +IF M +Y PD+ Y++I L +AG +K N
Sbjct: 210 YTFNTLINGFCKNSMVDEAFRIFKDM----ELYHCNPDVVTYNTIIDGLCRAGKVKIAHN 265
Query: 319 IVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLK 378
++ M +K + P+VV Y ++ ++ VF M LK
Sbjct: 266 VLSGMLKKATDV------------HPNVVSYTTLVRGYCMKQEIDEAVLVFHDMLSRGLK 313
Query: 379 PNGATYGLAMEVMLQSGNYDLVHELF--EKMQRNGEVPEALTYKVMVRTFWKEGKVDEAV 436
PN TY ++ + ++ YD + ++ P+A T+ ++++ G +D A+
Sbjct: 314 PNAVTYNTLIKGLSEAHRYDEIKDILIGGNDAFTTFAPDACTFNILIKAHCDAGHLDAAM 373
Query: 437 KAVRDMERRGVMGTASVYYELACCLCNCGRWQDA------TLEVEKIKRLPHAKPLEVTF 490
K ++M + ++ Y L LC + A E E + KPL +
Sbjct: 374 KVFQEMLNMKLHPDSASYSVLIRTLCMRNEFDRAETLFNELFEKEVLLGKDECKPLAAAY 433
Query: 491 TGMIRSSMDGGHIDDCICIFEYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVA 550
M G +F + + + T++ + + F A L V +
Sbjct: 434 NPMFEYLCANGKTKQAEKVFRQLMKRGVQDPPSYKTLITGHCREGKFKPAYELL--VLML 491
Query: 551 KSDLRPDAYTYNLMLE 566
+ + PD TY L+++
Sbjct: 492 RREFVPDLETYELLID 507
Score = 56.6 bits (135), Expect = 4e-08
Identities = 59/242 (24%), Positives = 102/242 (41%), Gaps = 24/242 (9%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
YT L+ + EA+ +F+ ML + P+ Y+++ L +A E+ +I+
Sbjct: 284 YTTLVRGYCMKQEIDEAVLVFHDMLSR-GLKPNAVTYNTLIKGLSEAHRYDEIKDIL--- 339
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
+ + T PD +N ++ A + VFQ+M L P+ A+
Sbjct: 340 ---------IGGNDAFTTFAPDACTFNILIKAHCDAGHLDAAMKVFQEMLNMKLHPDSAS 390
Query: 384 YGLAMEVMLQSGNYD----LVHELFEKMQRNGE---VPEALTYKVMVRTFWKEGKVDEAV 436
Y + + + +D L +ELFEK G+ P A Y M GK +A
Sbjct: 391 YSVLIRTLCMRNEFDRAETLFNELFEKEVLLGKDECKPLAAAYNPMFEYLCANGKTKQAE 450
Query: 437 KAVRDMERRGVMGTASVYYELACCLCNCGRWQDA-TLEVEKIKR--LPHAKPLEVTFTGM 493
K R + +RGV S Y L C G+++ A L V ++R +P + E+ G+
Sbjct: 451 KVFRQLMKRGVQDPPS-YKTLITGHCREGKFKPAYELLVLMLRREFVPDLETYELLIDGL 509
Query: 494 IR 495
++
Sbjct: 510 LK 511
Score = 35.8 bits (81), Expect = 0.077
Identities = 60/347 (17%), Positives = 134/347 (38%), Gaps = 63/347 (18%)
Query: 280 ALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNWD 339
A+++F ML N++++PD A+Y + TL CMR + + + ++ + ++
Sbjct: 372 AMKVFQEML-NMKLHPDSASYSVLIRTL--------------CMRNEFDRAETLFNELFE 416
Query: 340 PTL-------EPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLK-------------- 378
+ +P YN + + + K VF+Q+ K ++
Sbjct: 417 KEVLLGKDECKPLAAAYNPMFEYLCANGKTKQAEKVFRQLMKRGVQDPPSYKTLITGHCR 476
Query: 379 --------------------PNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALT 418
P+ TY L ++ +L+ G L H+ ++M R+ +P A T
Sbjct: 477 EGKFKPAYELLVLMLRREFVPDLETYELLIDGLLKIGEALLAHDTLQRMLRSSYLPVATT 536
Query: 419 YKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIK 478
+ ++ K +E+ V M + + + ++ L + + + A L I
Sbjct: 537 FHSVLAELAKRKFANESFCLVTLMLEKRIRQNIDLSTQVVRLLFSSAQKEKAFL----IV 592
Query: 479 RLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEY-MQDHCAPNVGTVNTMLKVYSQNDMF 537
RL + V ++ + + D + + ++ ++ T NT+++ ++
Sbjct: 593 RLLYDNGYLVKMEELLGYLCENRKLLDAHTLVLFCLEKSQMVDIDTCNTVIEGLCKHKRH 652
Query: 538 STAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEM 584
S A L+ E+ + + + ++ A +WE + V K M
Sbjct: 653 SEAFSLYNELVELGNHQQLSCHV--VLRNALEAAGKWEELQFVSKRM 697
>At5g16640 putative protein
Length = 504
Score = 91.7 bits (226), Expect = 1e-18
Identities = 70/304 (23%), Positives = 134/304 (44%), Gaps = 19/304 (6%)
Query: 263 VYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVEC 322
+Y ++ L K+++ AL + N M + + PD+ Y+S+ L +G + +V C
Sbjct: 188 IYNTIIDGLCKSKQVDNALDLLNRMEKD-GIGPDVVTYNSLISGLCSSGRWSDATRMVSC 246
Query: 323 MRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGA 382
M ++ + PDV +NA+++ACV + +++M + SL P+
Sbjct: 247 MTKRE--------------IYPDVFTFNALIDACVKEGRVSEAEEFYEEMIRRSLDPDIV 292
Query: 383 TYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDM 442
TY L + + D E+F M G P+ +TY +++ + K KV+ +K +M
Sbjct: 293 TYSLLIYGLCMYSRLDEAEEMFGFMVSKGCFPDVVTYSILINGYCKSKKVEHGMKLFCEM 352
Query: 443 ERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGH 502
+RGV+ Y L C G+ A E+ + P +T+ ++ D G
Sbjct: 353 SQRGVVRNTVTYTILIQGYCRAGKLNVAE-EIFRRMVFCGVHPNIITYNVLLHGLCDNGK 411
Query: 503 IDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTY 561
I+ + I MQ + ++ T N +++ + + A ++ + L PD +TY
Sbjct: 412 IEKALVILADMQKNGMDADIVTYNIIIRGMCKAGEVADAWDIYCSLNC--QGLMPDIWTY 469
Query: 562 NLML 565
M+
Sbjct: 470 TTMM 473
Score = 75.5 bits (184), Expect = 9e-14
Identities = 60/274 (21%), Positives = 120/274 (42%), Gaps = 4/274 (1%)
Query: 343 EPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHE 402
EP +V + ++LN + ++F QM KPN Y ++ + +S D +
Sbjct: 148 EPSIVTFGSLLNGFCRGDRVYDALYMFDQMVGMGYKPNVVIYNTIIDGLCKSKQVDNALD 207
Query: 403 LFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLC 462
L +M+++G P+ +TY ++ G+ +A + V M +R + + L
Sbjct: 208 LLNRMEKDGIGPDVVTYNSLISGLCSSGRWSDATRMVSCMTKREIYPDVFTFNALIDACV 267
Query: 463 NCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYM-QDHCAPNV 521
GR +A E++ R P VT++ +I +D+ +F +M C P+V
Sbjct: 268 KEGRVSEAEEFYEEMIR-RSLDPDIVTYSLLIYGLCMYSRLDEAEEMFGFMVSKGCFPDV 326
Query: 522 GTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVY 581
T + ++ Y ++ LF E +++ + + TY ++++ R + E ++
Sbjct: 327 VTYSILINGYCKSKKVEHGMKLFCE--MSQRGVVRNTVTYTILIQGYCRAGKLNVAEEIF 384
Query: 582 KEMILSGYHLDQNKHLPLLVKASRAGKVDNILVI 615
+ M+ G H + + LL GK++ LVI
Sbjct: 385 RRMVFCGVHPNIITYNVLLHGLCDNGKIEKALVI 418
Score = 70.5 bits (171), Expect = 3e-12
Identities = 64/291 (21%), Positives = 120/291 (40%), Gaps = 4/291 (1%)
Query: 332 YMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVM 391
YM+ + +P+VVIYN +++ SKQ + +M K + P+ TY + +
Sbjct: 172 YMFDQMVGMGYKPNVVIYNTIIDGLCKSKQVDNALDLLNRMEKDGIGPDVVTYNSLISGL 231
Query: 392 LQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTA 451
SG + + M + P+ T+ ++ KEG+V EA + +M RR +
Sbjct: 232 CSSGRWSDATRMVSCMTKREIYPDVFTFNALIDACVKEGRVSEAEEFYEEMIRRSLDPDI 291
Query: 452 SVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIF- 510
Y L LC R +A E+ P VT++ +I ++ + +F
Sbjct: 292 VTYSLLIYGLCMYSRLDEAE-EMFGFMVSKGCFPDVVTYSILINGYCKSKKVEHGMKLFC 350
Query: 511 EYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSR 570
E Q N T +++ Y + + A+ +F ++ + P+ TYN++L
Sbjct: 351 EMSQRGVVRNTVTYTILIQGYCRAGKLNVAEEIFR--RMVFCGVHPNIITYNVLLHGLCD 408
Query: 571 GHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVDNILVIYIRSQC 621
+ E + +M +G D + ++ +AG+V + IY C
Sbjct: 409 NGKIEKALVILADMQKNGMDADIVTYNIIIRGMCKAGEVADAWDIYCSLNC 459
Score = 52.4 bits (124), Expect = 8e-07
Identities = 53/262 (20%), Positives = 110/262 (41%), Gaps = 7/262 (2%)
Query: 379 PNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKA 438
P+ A + + + + YD+V L+E+MQ G T +++ F + ++ A+
Sbjct: 79 PSIADFSRLLSAISKMKKYDVVIYLWEQMQMLGIPHNLCTCNILLNCFCRCSQLSLALSF 138
Query: 439 VRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSM 498
+ M + G + + L C R DA +++ + + KP V + +I
Sbjct: 139 LGKMIKLGHEPSIVTFGSLLNGFCRGDRVYDALYMFDQMVGMGY-KPNVVIYNTIIDGLC 197
Query: 499 DGGHIDDCICIFEYMQ-DHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPD 557
+D+ + + M+ D P+V T N+++ + +S A + + K ++ PD
Sbjct: 198 KSKQVDNALDLLNRMEKDGIGPDVVTYNSLISGLCSSGRWSDATRMVS--CMTKREIYPD 255
Query: 558 AYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPL---LVKASRAGKVDNILV 614
+T+N +++A + + E Y+EMI D + L L SR + + +
Sbjct: 256 VFTFNALIDACVKEGRVSEAEEFYEEMIRRSLDPDIVTYSLLIYGLCMYSRLDEAEEMFG 315
Query: 615 IYIRSQCLFYISAHSIISYKNC 636
+ C + +SI+ C
Sbjct: 316 FMVSKGCFPDVVTYSILINGYC 337
Score = 45.1 bits (105), Expect = 1e-04
Identities = 38/206 (18%), Positives = 85/206 (40%), Gaps = 22/206 (10%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYH----------------SIAVTL 307
Y+ L+ L R EA ++F M+ +PD+ Y + +
Sbjct: 294 YSLLIYGLCMYSRLDEAEEMFGFMVSK-GCFPDVVTYSILINGYCKSKKVEHGMKLFCEM 352
Query: 308 GQAGLLKELLNIV-----ECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQW 362
Q G+++ + C K + ++R+ + P+++ YN +L+ + +
Sbjct: 353 SQRGVVRNTVTYTILIQGYCRAGKLNVAEEIFRRMVFCGVHPNIITYNVLLHGLCDNGKI 412
Query: 363 KGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVM 422
+ + M+K+ + + TY + + M ++G +++ + G +P+ TY M
Sbjct: 413 EKALVILADMQKNGMDADIVTYNIIIRGMCKAGEVADAWDIYCSLNCQGLMPDIWTYTTM 472
Query: 423 VRTFWKEGKVDEAVKAVRDMERRGVM 448
+ +K+G EA R M+ G++
Sbjct: 473 MLGLYKKGLRREADALFRKMKEDGIL 498
Score = 32.3 bits (72), Expect = 0.85
Identities = 32/151 (21%), Positives = 64/151 (42%), Gaps = 15/151 (9%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
YT L+ +A + A +IF M+ V+P++ Y+ + L G +++ L I+ M
Sbjct: 364 YTILIQGYCRAGKLNVAEEIFRRMVF-CGVHPNIITYNVLLHGLCDNGKIEKALVILADM 422
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
++ ++ D+V YN ++ + + ++ + L P+ T
Sbjct: 423 QKNG--------------MDADIVTYNIIIRGMCKAGEVADAWDIYCSLNCQGLMPDIWT 468
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVP 414
Y M + + G LF KM+ +G +P
Sbjct: 469 YTTMMLGLYKKGLRREADALFRKMKEDGILP 499
>At1g62590 putative membrane-associated salt-inducible protein
(AL021637); similar to EST gb|AA728420
Length = 634
Score = 91.7 bits (226), Expect = 1e-18
Identities = 63/275 (22%), Positives = 125/275 (44%), Gaps = 6/275 (2%)
Query: 344 PDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHEL 403
PD + + +++ + + +M + +PN TYG+ + + + G+ DL L
Sbjct: 188 PDTITFTTLIHGLFLHNKASEAVALVDRMVQRGCQPNLVTYGVVVNGLCKRGDTDLALNL 247
Query: 404 FEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCN 463
KM+ + + + ++ + K VD+A+ ++ME +G+ Y L CLC+
Sbjct: 248 LNKMEAAKIEADVVIFNTIIDSLCKYRHVDDALNLFKEMETKGIRPNVVTYSSLISCLCS 307
Query: 464 CGRWQDAT-LEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYM-QDHCAPNV 521
GRW DA+ L + I++ P VTF +I + + G + +++ M + P++
Sbjct: 308 YGRWSDASQLLSDMIEK--KINPNLVTFNALIDAFVKEGKFVEAEKLYDDMIKRSIDPDI 365
Query: 522 GTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVY 581
T N+++ + +D AK +FE + D PD TYN +++ + + E ++
Sbjct: 366 FTYNSLVNGFCMHDRLDKAKQMFE--FMVSKDCFPDVVTYNTLIKGFCKSKRVEDGTELF 423
Query: 582 KEMILSGYHLDQNKHLPLLVKASRAGKVDNILVIY 616
+EM G D + L+ G DN ++
Sbjct: 424 REMSHRGLVGDTVTYTTLIQGLFHDGDCDNAQKVF 458
Score = 77.0 bits (188), Expect = 3e-14
Identities = 74/372 (19%), Positives = 155/372 (40%), Gaps = 22/372 (5%)
Query: 180 KKTSKRSEVQVVRF--LVDRLCDKEIRAKDWKFSRLMKLSGLSFTEGQLMMIIEMLGVKR 237
K + + E VV F ++D LC + M+ G+ +I L
Sbjct: 250 KMEAAKIEADVVIFNTIIDSLCKYRHVDDALNLFKEMETKGIRPNVVTYSSLISCLCSYG 309
Query: 238 CWKQALSVVQWVYNSKDHRKFQSRFVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDM 297
W A ++ + K + + + L+ K + EA ++++ M+ + PD+
Sbjct: 310 RWSDASQLLSDMIEKKINPNLVT---FNALIDAFVKEGKFVEAEKLYDDMIKR-SIDPDI 365
Query: 298 AAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACV 357
Y+S+ CM + + K M+ PDVV YN ++
Sbjct: 366 FTYNSLVNGF--------------CMHDRLDKAKQMFEFMVSKDCFPDVVTYNTLIKGFC 411
Query: 358 PSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEAL 417
SK+ + + +F++M L + TY ++ + G+ D ++F++M +G P+ +
Sbjct: 412 KSKRVEDGTELFREMSHRGLVGDTVTYTTLIQGLFHDGDCDNAQKVFKQMVSDGVPPDIM 471
Query: 418 TYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKI 477
TY +++ GK+++A++ M++ + +Y + +C G+ D ++
Sbjct: 472 TYSILLDGLCNNGKLEKALEVFDYMQKSEIKLDIYIYTTMIEGMCKAGKVDDG-WDLFCS 530
Query: 478 KRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYM-QDHCAPNVGTVNTMLKVYSQNDM 536
L KP VT+ MI + + + + M +D PN GT NT+++ + ++
Sbjct: 531 LSLKGVKPNVVTYNTMISGLCSKRLLQEAYALLKKMKEDGPLPNSGTYNTLIRAHLRDGD 590
Query: 537 FSTAKVLFEEVK 548
+ + L E++
Sbjct: 591 KAASAELIREMR 602
Score = 73.9 bits (180), Expect = 3e-13
Identities = 63/291 (21%), Positives = 118/291 (39%), Gaps = 4/291 (1%)
Query: 322 CMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNG 381
C R + + K +E DVVI+N ++++ + +F++M ++PN
Sbjct: 236 CKRGDTDLALNLLNKMEAAKIEADVVIFNTIIDSLCKYRHVDDALNLFKEMETKGIRPNV 295
Query: 382 ATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRD 441
TY + + G + +L M P +T+ ++ F KEGK EA K D
Sbjct: 296 VTYSSLISCLCSYGRWSDASQLLSDMIEKKINPNLVTFNALIDAFVKEGKFVEAEKLYDD 355
Query: 442 MERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGG 501
M +R + Y L C R D ++ + P VT+ +I+
Sbjct: 356 MIKRSIDPDIFTYNSLVNGFCMHDR-LDKAKQMFEFMVSKDCFPDVVTYNTLIKGFCKSK 414
Query: 502 HIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYT 560
++D +F M + T T+++ + A+ +F++ + + PD T
Sbjct: 415 RVEDGTELFREMSHRGLVGDTVTYTTLIQGLFHDGDCDNAQKVFKQ--MVSDGVPPDIMT 472
Query: 561 YNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVDN 611
Y+++L+ + E V+ M S LD + ++ +AGKVD+
Sbjct: 473 YSILLDGLCNNGKLEKALEVFDYMQKSEIKLDIYIYTTMIEGMCKAGKVDD 523
Score = 54.7 bits (130), Expect = 2e-07
Identities = 47/222 (21%), Positives = 96/222 (43%), Gaps = 4/222 (1%)
Query: 368 VFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFW 427
+F M KS P+ + + + + +D+V L EKMQR V TY +++ F
Sbjct: 72 LFGGMVKSRPLPSIVEFNKLLSAIAKMKKFDVVISLGEKMQRLEIVHGLYTYNILINCFC 131
Query: 428 KEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLE 487
+ ++ A+ + M + G + L C+ R DA V+++ + + +P
Sbjct: 132 RRSQISLALALLGKMMKLGYEPSIVTLSSLLNGYCHGKRISDAVALVDQMVEMGY-RPDT 190
Query: 488 VTFTGMIRSSMDGGHIDDCICIFEYM-QDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEE 546
+TFT +I + + + + M Q C PN+ T ++ + A L +
Sbjct: 191 ITFTTLIHGLFLHNKASEAVALVDRMVQRGCQPNLVTYGVVVNGLCKRGDTDLALNLLNK 250
Query: 547 VKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSG 588
++ AK + D +N ++++ + + +++KEM G
Sbjct: 251 MEAAK--IEADVVIFNTIIDSLCKYRHVDDALNLFKEMETKG 290
Score = 50.1 bits (118), Expect = 4e-06
Identities = 46/191 (24%), Positives = 82/191 (42%), Gaps = 19/191 (9%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
YT L+ L A ++F M+ + V PD+ Y + L G L++ L + + M
Sbjct: 438 YTTLIQGLFHDGDCDNAQKVFKQMVSD-GVPPDIMTYSILLDGLCNNGKLEKALEVFDYM 496
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNA-CVPSKQWKGVSW-VFQQMRKSSLKPNG 381
QK E ++ D+ IY ++ C K G W +F + +KPN
Sbjct: 497 -QKSE-------------IKLDIYIYTTMIEGMCKAGKVDDG--WDLFCSLSLKGVKPNV 540
Query: 382 ATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRD 441
TY + + + L +KM+ +G +P + TY ++R ++G + + +R+
Sbjct: 541 VTYNTMISGLCSKRLLQEAYALLKKMKEDGPLPNSGTYNTLIRAHLRDGDKAASAELIRE 600
Query: 442 MERRGVMGTAS 452
M +G AS
Sbjct: 601 MRSCRFVGDAS 611
>At1g63330 unknown protein
Length = 558
Score = 91.3 bits (225), Expect = 2e-18
Identities = 83/363 (22%), Positives = 155/363 (41%), Gaps = 19/363 (5%)
Query: 256 RKFQSRFVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKE 315
R S F + KLL+ + K + K L I +LG + Y VTL
Sbjct: 37 RPLPSIFEFNKLLSAIAKMK--KFDLVISLALLGKMM----KLGYEPSIVTLSS------ 84
Query: 316 LLNIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKS 375
LLN C ++ + + + PD + + +++ + + +M +
Sbjct: 85 LLNGY-CHGKRISDAVALVDQMVEMGYRPDTITFTTLIHGLFLHNKASEAVALVDRMVQR 143
Query: 376 SLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEA 435
+PN TYG+ + + + G+ DL L KM+ + + + ++ + K VD+A
Sbjct: 144 GCQPNLVTYGVVVNGLCKRGDIDLAFNLLNKMEAAKIEADVVIFNTIIDSLCKYRHVDDA 203
Query: 436 VKAVRDMERRGVMGTASVYYELACCLCNCGRWQDAT-LEVEKIKRLPHAKPLEVTFTGMI 494
+ ++ME +G+ Y L CLC+ GRW DA+ L + I++ P VTF +I
Sbjct: 204 LNLFKEMETKGIRPNVVTYSSLISCLCSYGRWSDASQLLSDMIEK--KINPNLVTFNALI 261
Query: 495 RSSMDGGHIDDCICIFEYM-QDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSD 553
+ + G + + + M + P++ T N+++ + +D AK +FE + D
Sbjct: 262 DAFVKEGKFVEAEKLHDDMIKRSIDPDIFTYNSLINGFCMHDRLDKAKQMFE--FMVSKD 319
Query: 554 LRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVDNIL 613
PD TYN +++ + + E +++EM G D + L+ G DN
Sbjct: 320 CFPDLDTYNTLIKGFCKSKRVEDGTELFREMSHRGLVGDTVTYTTLIQGLFHDGDCDNAQ 379
Query: 614 VIY 616
++
Sbjct: 380 KVF 382
Score = 80.9 bits (198), Expect = 2e-15
Identities = 72/308 (23%), Positives = 131/308 (42%), Gaps = 19/308 (6%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y+ L++ L R +A Q+ + M+ ++ P++ ++++ + G E + + M
Sbjct: 222 YSSLISCLCSYGRWSDASQLLSDMIEK-KINPNLVTFNALIDAFVKEGKFVEAEKLHDDM 280
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
++ +++PD+ YN+++N + +F+ M P+ T
Sbjct: 281 IKR--------------SIDPDIFTYNSLINGFCMHDRLDKAKQMFEFMVSKDCFPDLDT 326
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
Y ++ +S + ELF +M G V + +TY +++ + +G D A K + M
Sbjct: 327 YNTLIKGFCKSKRVEDGTELFREMSHRGLVGDTVTYTTLIQGLFHDGDCDNAQKVFKQMV 386
Query: 444 RRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHI 503
GV Y L LCN G+ + A LEV + K +T MI G +
Sbjct: 387 SDGVPPDIMTYSILLDGLCNNGKLEKA-LEVFDYMQKSEIKLDIYIYTTMIEGMCKAGKV 445
Query: 504 DDCICIF-EYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYN 562
DD +F PNV T NTM+ + A L +++K + PD+ TYN
Sbjct: 446 DDGWDLFCSLSLKGVKPNVVTYNTMISGLCSKRLLQEAYALLKKMK--EDGPLPDSGTYN 503
Query: 563 LMLEASSR 570
++ A R
Sbjct: 504 TLIRAHLR 511
Score = 70.5 bits (171), Expect = 3e-12
Identities = 47/228 (20%), Positives = 105/228 (45%), Gaps = 2/228 (0%)
Query: 322 CMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNG 381
CM + + K M+ PD+ YN ++ SK+ + + +F++M L +
Sbjct: 300 CMHDRLDKAKQMFEFMVSKDCFPDLDTYNTLIKGFCKSKRVEDGTELFREMSHRGLVGDT 359
Query: 382 ATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRD 441
TY ++ + G+ D ++F++M +G P+ +TY +++ GK+++A++
Sbjct: 360 VTYTTLIQGLFHDGDCDNAQKVFKQMVSDGVPPDIMTYSILLDGLCNNGKLEKALEVFDY 419
Query: 442 MERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGG 501
M++ + +Y + +C G+ D ++ L KP VT+ MI
Sbjct: 420 MQKSEIKLDIYIYTTMIEGMCKAGKVDDG-WDLFCSLSLKGVKPNVVTYNTMISGLCSKR 478
Query: 502 HIDDCICIFEYM-QDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVK 548
+ + + + M +D P+ GT NT+++ + ++ + + L E++
Sbjct: 479 LLQEAYALLKKMKEDGPLPDSGTYNTLIRAHLRDGDKAASAELIREMR 526
Score = 50.8 bits (120), Expect = 2e-06
Identities = 46/191 (24%), Positives = 83/191 (43%), Gaps = 19/191 (9%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
YT L+ L A ++F M+ + V PD+ Y + L G L++ L + + M
Sbjct: 362 YTTLIQGLFHDGDCDNAQKVFKQMVSD-GVPPDIMTYSILLDGLCNNGKLEKALEVFDYM 420
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNA-CVPSKQWKGVSW-VFQQMRKSSLKPNG 381
QK E ++ D+ IY ++ C K G W +F + +KPN
Sbjct: 421 -QKSE-------------IKLDIYIYTTMIEGMCKAGKVDDG--WDLFCSLSLKGVKPNV 464
Query: 382 ATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRD 441
TY + + + L +KM+ +G +P++ TY ++R ++G + + +R+
Sbjct: 465 VTYNTMISGLCSKRLLQEAYALLKKMKEDGPLPDSGTYNTLIRAHLRDGDKAASAELIRE 524
Query: 442 MERRGVMGTAS 452
M +G AS
Sbjct: 525 MRSCRFVGDAS 535
>At1g63150 unknown protein
Length = 629
Score = 91.3 bits (225), Expect = 2e-18
Identities = 78/344 (22%), Positives = 144/344 (41%), Gaps = 34/344 (9%)
Query: 262 FVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTL---GQAGLLKELLN 318
F +T L+ L + EA+ + + M+ PD+ Y ++ L G L LLN
Sbjct: 189 FTFTTLIHGLFLHNKASEAVALVDQMVQR-GCQPDLVTYGTVVNGLCKRGDIDLALNLLN 247
Query: 319 IVECMRQKPETL----------KY--------MYRKNWDPTLEPDVVIYNAVLNACVPSK 360
+E R K + KY ++ + + P+VV YN+++N
Sbjct: 248 KMEAARIKANVVIFNTIIDSLCKYRHVEVAVDLFTEMETKGIRPNVVTYNSLINCLCNYG 307
Query: 361 QWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYK 420
+W S + M + + PN T+ ++ + G +L E+M + P+ +TY
Sbjct: 308 RWSDASRLLSNMLEKKINPNVVTFNALIDAFFKEGKLVEAEKLHEEMIQRSIDPDTITYN 367
Query: 421 VMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRL 480
+++ F ++DEA + + M + + Y L C C R +D VE + +
Sbjct: 368 LLINGFCMHNRLDEAKQMFKFMVSKDCLPNIQTYNTLINGFCKCKRVEDG---VELFREM 424
Query: 481 PHAKPL--EVTFTGMIRSSMDGGHIDDCICIFEYMQDHCAP-NVGTVNTMLKVYSQNDMF 537
+ VT+T +I+ G D +F+ M + P ++ T + +L
Sbjct: 425 SQRGLVGNTVTYTTIIQGFFQAGDCDSAQMVFKQMVSNRVPTDIMTYSILLHGLCSYGKL 484
Query: 538 STAKVLFEEVKVAKSDLRPDAYTYNLMLE----ASSRGHQWEYF 577
TA V+F+ ++ KS++ + + YN M+E A G W+ F
Sbjct: 485 DTALVIFKYLQ--KSEMELNIFIYNTMIEGMCKAGKVGEAWDLF 526
Score = 79.0 bits (193), Expect = 8e-15
Identities = 59/275 (21%), Positives = 117/275 (42%), Gaps = 4/275 (1%)
Query: 343 EPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHE 402
+PD + +++ + + QM + +P+ TYG + + + G+ DL
Sbjct: 185 KPDTFTFTTLIHGLFLHNKASEAVALVDQMVQRGCQPDLVTYGTVVNGLCKRGDIDLALN 244
Query: 403 LFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLC 462
L KM+ + + ++ + K V+ AV +ME +G+ Y L CLC
Sbjct: 245 LLNKMEAARIKANVVIFNTIIDSLCKYRHVEVAVDLFTEMETKGIRPNVVTYNSLINCLC 304
Query: 463 NCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYM-QDHCAPNV 521
N GRW DA+ + + P VTF +I + G + + + E M Q P+
Sbjct: 305 NYGRWSDASRLLSNMLE-KKINPNVVTFNALIDAFFKEGKLVEAEKLHEEMIQRSIDPDT 363
Query: 522 GTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVY 581
T N ++ + ++ AK +F+ + D P+ TYN ++ + + E ++
Sbjct: 364 ITYNLLINGFCMHNRLDEAKQMFK--FMVSKDCLPNIQTYNTLINGFCKCKRVEDGVELF 421
Query: 582 KEMILSGYHLDQNKHLPLLVKASRAGKVDNILVIY 616
+EM G + + ++ +AG D+ +++
Sbjct: 422 REMSQRGLVGNTVTYTTIIQGFFQAGDCDSAQMVF 456
Score = 77.4 bits (189), Expect = 2e-14
Identities = 77/382 (20%), Positives = 157/382 (40%), Gaps = 54/382 (14%)
Query: 263 VYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVEC 322
++ ++ L K R + A+ +F M + P++ Y+S+ L G + ++
Sbjct: 260 IFNTIIDSLCKYRHVEVAVDLFTEMETK-GIRPNVVTYNSLINCLCNYGRWSDASRLLSN 318
Query: 323 MRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGA 382
M +K + P+VV +NA+++A + + ++M + S+ P+
Sbjct: 319 MLEKK--------------INPNVVTFNALIDAFFKEGKLVEAEKLHEEMIQRSIDPDTI 364
Query: 383 TYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDM 442
TY L + D ++F+ M +P TY ++ F K +V++ V+ R+M
Sbjct: 365 TYNLLINGFCMHNRLDEAKQMFKFMVSKDCLPNIQTYNTLINGFCKCKRVEDGVELFREM 424
Query: 443 ERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEV-TFTGMIRSSMDGG 501
+RG++G Y + G A + +++ + + P ++ T++ ++ G
Sbjct: 425 SQRGLVGNTVTYTTIIQGFFQAGDCDSAQMVFKQM--VSNRVPTDIMTYSILLHGLCSYG 482
Query: 502 HIDDCICIFEYMQDH-----------------------------CA----PNVGTVNTML 528
+D + IF+Y+Q C+ P+V T NTM+
Sbjct: 483 KLDTALVIFKYLQKSEMELNIFIYNTMIEGMCKAGKVGEAWDLFCSLSIKPDVVTYNTMI 542
Query: 529 KVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSG 588
+ A LF ++K + P++ TYN ++ A+ R + KEM SG
Sbjct: 543 SGLCSKRLLQEADDLFRKMK--EDGTLPNSGTYNTLIRANLRDCDRAASAELIKEMRSSG 600
Query: 589 YHLDQNKHLPLLVKASRAGKVD 610
+ D + + L+ G++D
Sbjct: 601 FVGDAST-ISLVTNMLHDGRLD 621
Score = 50.8 bits (120), Expect = 2e-06
Identities = 44/222 (19%), Positives = 93/222 (41%), Gaps = 4/222 (1%)
Query: 368 VFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFW 427
+F M KS P+ + + + + ++LV L E+MQ G + TY + + F
Sbjct: 70 LFGDMVKSRPFPSIVEFNKLLSAVAKMNKFELVISLGEQMQTLGISHDLYTYSIFINCFC 129
Query: 428 KEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLE 487
+ ++ A+ + M + G L C+ R DA V+++ + + KP
Sbjct: 130 RRSQLSLALAVLAKMMKLGYEPDIVTLSSLLNGYCHSKRISDAVALVDQMVEMGY-KPDT 188
Query: 488 VTFTGMIRSSMDGGHIDDCICIFEYM-QDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEE 546
TFT +I + + + + M Q C P++ T T++ + A L +
Sbjct: 189 FTFTTLIHGLFLHNKASEAVALVDQMVQRGCQPDLVTYGTVVNGLCKRGDIDLALNLLNK 248
Query: 547 VKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSG 588
++ A+ ++ + +N ++++ + E ++ EM G
Sbjct: 249 MEAAR--IKANVVIFNTIIDSLCKYRHVEVAVDLFTEMETKG 288
Score = 49.7 bits (117), Expect = 5e-06
Identities = 56/280 (20%), Positives = 106/280 (37%), Gaps = 43/280 (15%)
Query: 344 PDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHEL 403
P +V +N +L+A +++ V + +QM+ + + TY + + + L +
Sbjct: 81 PSIVEFNKLLSAVAKMNKFELVISLGEQMQTLGISHDLYTYSIFINCFCRRSQLSLALAV 140
Query: 404 FEKMQR-----------------------------------NGEVPEALTYKVMVRTFWK 428
KM + G P+ T+ ++ +
Sbjct: 141 LAKMMKLGYEPDIVTLSSLLNGYCHSKRISDAVALVDQMVEMGYKPDTFTFTTLIHGLFL 200
Query: 429 EGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHA--KPL 486
K EAV V M +RG Y + LC G D L + + ++ A K
Sbjct: 201 HNKASEAVALVDQMVQRGCQPDLVTYGTVVNGLCKRG---DIDLALNLLNKMEAARIKAN 257
Query: 487 EVTFTGMIRSSMDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDMFSTAKVLFE 545
V F +I S H++ + +F M+ PNV T N+++ +S A L
Sbjct: 258 VVIFNTIIDSLCKYRHVEVAVDLFTEMETKGIRPNVVTYNSLINCLCNYGRWSDASRLLS 317
Query: 546 EVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMI 585
+ K + P+ T+N +++A + + E +++EMI
Sbjct: 318 NMLEKK--INPNVVTFNALIDAFFKEGKLVEAEKLHEEMI 355
Score = 47.4 bits (111), Expect = 3e-05
Identities = 43/190 (22%), Positives = 84/190 (43%), Gaps = 20/190 (10%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
YT ++ +A A +F M+ N RV D+ Y + L G L L I + +
Sbjct: 436 YTTIIQGFFQAGDCDSAQMVFKQMVSN-RVPTDIMTYSILLHGLCSYGKLDTALVIFKYL 494
Query: 324 RQKPETLK-YMYR-------------KNWDP----TLEPDVVIYNAVLNACVPSKQWKGV 365
++ L ++Y + WD +++PDVV YN +++ + +
Sbjct: 495 QKSEMELNIFIYNTMIEGMCKAGKVGEAWDLFCSLSIKPDVVTYNTMISGLCSKRLLQEA 554
Query: 366 SWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRT 425
+F++M++ PN TY + L+ + EL ++M+ +G V +A T +V
Sbjct: 555 DDLFRKMKEDGTLPNSGTYNTLIRANLRDCDRAASAELIKEMRSSGFVGDASTIS-LVTN 613
Query: 426 FWKEGKVDEA 435
+G++D++
Sbjct: 614 MLHDGRLDKS 623
Score = 28.9 bits (63), Expect = 9.4
Identities = 20/91 (21%), Positives = 41/91 (44%), Gaps = 3/91 (3%)
Query: 503 IDDCICIF-EYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTY 561
+DD + +F + ++ P++ N +L ++ + F L E+++ + D YTY
Sbjct: 64 VDDAVDLFGDMVKSRPFPSIVEFNKLLSAVAKMNKFELVISLGEQMQTL--GISHDLYTY 121
Query: 562 NLMLEASSRGHQWEYFEHVYKEMILSGYHLD 592
++ + R Q V +M+ GY D
Sbjct: 122 SIFINCFCRRSQLSLALAVLAKMMKLGYEPD 152
>At1g63070 unknown protein
Length = 590
Score = 91.3 bits (225), Expect = 2e-18
Identities = 62/276 (22%), Positives = 120/276 (43%), Gaps = 5/276 (1%)
Query: 343 EPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHE 402
+PD V + +++ + + ++M +P+ TYG + + + G DL
Sbjct: 177 QPDTVTFTTLVHGLFQHNKASEAVALVERMVVKGCQPDLVTYGAVINGLCKRGEPDLALN 236
Query: 403 LFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLC 462
L KM++ + + Y ++ K +D+A ME +G+ Y L CLC
Sbjct: 237 LLNKMEKGKIEADVVIYNTIIDGLCKYKHMDDAFDLFNKMETKGIKPDVFTYNPLISCLC 296
Query: 463 NCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYM--QDHCAPN 520
N GRW DA+ + + + P V F +I + + G + + +++ M HC P+
Sbjct: 297 NYGRWSDASRLLSDMLE-KNINPDLVFFNALIDAFVKEGKLVEAEKLYDEMVKSKHCFPD 355
Query: 521 VGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHV 580
V NT++K + + +F E +++ L + TY ++ + + + V
Sbjct: 356 VVAYNTLIKGFCKYKRVEEGMEVFRE--MSQRGLVGNTVTYTTLIHGFFQARDCDNAQMV 413
Query: 581 YKEMILSGYHLDQNKHLPLLVKASRAGKVDNILVIY 616
+K+M+ G H D + LL G V+ LV++
Sbjct: 414 FKQMVSDGVHPDIMTYNILLDGLCNNGNVETALVVF 449
Score = 79.3 bits (194), Expect = 6e-15
Identities = 75/320 (23%), Positives = 129/320 (39%), Gaps = 24/320 (7%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
+T L+ L + + EA+ + M+ PD+ Y ++ L C
Sbjct: 183 FTTLVHGLFQHNKASEAVALVERMVVK-GCQPDLVTYGAVINGL--------------CK 227
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
R +P+ + K +E DVVIYN +++ K +F +M +KP+ T
Sbjct: 228 RGEPDLALNLLNKMEKGKIEADVVIYNTIIDGLCKYKHMDDAFDLFNKMETKGIKPDVFT 287
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDM- 442
Y + + G + L M P+ + + ++ F KEGK+ EA K +M
Sbjct: 288 YNPLISCLCNYGRWSDASRLLSDMLEKNINPDLVFFNALIDAFVKEGKLVEAEKLYDEMV 347
Query: 443 ERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGH 502
+ + Y L C R ++ +EV + VT+T +I
Sbjct: 348 KSKHCFPDVVAYNTLIKGFCKYKRVEEG-MEVFREMSQRGLVGNTVTYTTLIHGFFQARD 406
Query: 503 IDDCICIFEYM-QDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTY 561
D+ +F+ M D P++ T N +L N TA V+FE ++ K D++ D TY
Sbjct: 407 CDNAQMVFKQMVSDGVHPDIMTYNILLDGLCNNGNVETALVVFEYMQ--KRDMKLDIVTY 464
Query: 562 NLMLEASSRGHQ----WEYF 577
M+EA + + W+ F
Sbjct: 465 TTMIEALCKAGKVEDGWDLF 484
Score = 78.2 bits (191), Expect = 1e-14
Identities = 69/304 (22%), Positives = 132/304 (42%), Gaps = 18/304 (5%)
Query: 262 FVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVE 321
F Y L++ L R +A ++ + ML + PD+ ++++ + G L E + +
Sbjct: 286 FTYNPLISCLCNYGRWSDASRLLSDMLEK-NINPDLVFFNALIDAFVKEGKLVEAEKLYD 344
Query: 322 CMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNG 381
M + PDVV YN ++ K+ + VF++M + L N
Sbjct: 345 EMVKSKHCF-------------PDVVAYNTLIKGFCKYKRVEEGMEVFREMSQRGLVGNT 391
Query: 382 ATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRD 441
TY + Q+ + D +F++M +G P+ +TY +++ G V+ A+
Sbjct: 392 VTYTTLIHGFFQARDCDNAQMVFKQMVSDGVHPDIMTYNILLDGLCNNGNVETALVVFEY 451
Query: 442 MERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGG 501
M++R + Y + LC G+ +D ++ L KP VT+T M+ G
Sbjct: 452 MQKRDMKLDIVTYTTMIEALCKAGKVEDG-WDLFCSLSLKGVKPNVVTYTTMMSGFCRKG 510
Query: 502 HIDDCICIF-EYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYT 560
++ +F E +D PN GT NT+++ ++ + + L +E++ DA T
Sbjct: 511 LKEEADALFVEMKEDGPLPNSGTYNTLIRARLRDGDEAASAELIKEMR--SCGFAGDAST 568
Query: 561 YNLM 564
+ L+
Sbjct: 569 FGLV 572
Score = 52.4 bits (124), Expect = 8e-07
Identities = 55/284 (19%), Positives = 111/284 (38%), Gaps = 7/284 (2%)
Query: 344 PDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHEL 403
P +V ++ +L+A ++ V + +QM+ + N TY + + + L +
Sbjct: 73 PSIVEFSKLLSAIAKMNKFDLVISLGEQMQNLGISHNLYTYSIFINYFCRRSQLSLALAI 132
Query: 404 FEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCN 463
KM + G P +T ++ F ++ EAV V M G + L L
Sbjct: 133 LGKMMKLGYGPSIVTLNSLLNGFCHGNRISEAVALVDQMVEMGYQPDTVTFTTLVHGLFQ 192
Query: 464 CGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQ-DHCAPNVG 522
+ +A VE++ + +P VT+ +I G D + + M+ +V
Sbjct: 193 HNKASEAVALVERMV-VKGCQPDLVTYGAVINGLCKRGEPDLALNLLNKMEKGKIEADVV 251
Query: 523 TVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYK 582
NT++ + A LF K+ ++PD +TYN ++ +W +
Sbjct: 252 IYNTIIDGLCKYKHMDDAFDLFN--KMETKGIKPDVFTYNPLISCLCNYGRWSDASRLLS 309
Query: 583 EMILSGYHLDQ---NKHLPLLVKASRAGKVDNILVIYIRSQCLF 623
+M+ + D N + VK + + + + ++S+ F
Sbjct: 310 DMLEKNINPDLVFFNALIDAFVKEGKLVEAEKLYDEMVKSKHCF 353
Score = 49.3 bits (116), Expect = 7e-06
Identities = 46/234 (19%), Positives = 93/234 (39%), Gaps = 4/234 (1%)
Query: 368 VFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFW 427
+F M KS P+ + + + + +DLV L E+MQ G TY + + F
Sbjct: 62 LFGDMVKSRPFPSIVEFSKLLSAIAKMNKFDLVISLGEQMQNLGISHNLYTYSIFINYFC 121
Query: 428 KEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLE 487
+ ++ A+ + M + G + L C+ R +A V+++ + + +P
Sbjct: 122 RRSQLSLALAILGKMMKLGYGPSIVTLNSLLNGFCHGNRISEAVALVDQMVEMGY-QPDT 180
Query: 488 VTFTGMIRSSMDGGHIDDCICIFEYM-QDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEE 546
VTFT ++ + + + E M C P++ T ++ + A L
Sbjct: 181 VTFTTLVHGLFQHNKASEAVALVERMVVKGCQPDLVTYGAVINGLCKRGEPDLALNLLN- 239
Query: 547 VKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLL 600
K+ K + D YN +++ + + ++ +M G D + PL+
Sbjct: 240 -KMEKGKIEADVVIYNTIIDGLCKYKHMDDAFDLFNKMETKGIKPDVFTYNPLI 292
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.136 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,663,490
Number of Sequences: 26719
Number of extensions: 643020
Number of successful extensions: 6740
Number of sequences better than 10.0: 485
Number of HSP's better than 10.0 without gapping: 331
Number of HSP's successfully gapped in prelim test: 155
Number of HSP's that attempted gapping in prelim test: 2118
Number of HSP's gapped (non-prelim): 2326
length of query: 653
length of database: 11,318,596
effective HSP length: 106
effective length of query: 547
effective length of database: 8,486,382
effective search space: 4642050954
effective search space used: 4642050954
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC140720.4


