

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140670.14 + phase: 0
(359 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
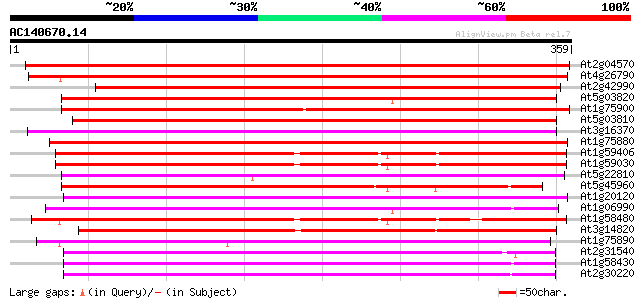
Score E
Sequences producing significant alignments: (bits) Value
At2g04570 putative GDSL-motif lipase/hydrolase 493 e-140
At4g26790 putative APG protein 428 e-120
At2g42990 putative GDSL-motif lipase/hydrolase 394 e-110
At5g03820 putative protein 280 1e-75
At1g75900 family II lipase EXL3 (EXL3) 278 3e-75
At5g03810 putative protein 274 6e-74
At3g16370 putative APG protein 273 1e-73
At1g75880 family II extracellular lipase 1 like protein (EXL1) 271 3e-73
At1g59406 proline-rich protein, putative 270 1e-72
At1g59030 proline-rich protein, putative 270 1e-72
At5g22810 GDSL-motif lipase/hydrolase-like protein 267 8e-72
At5g45960 GDSL-motif lipase/hydrolase-like protein 266 2e-71
At1g20120 anter-specific proline-rich like protein (APG precursor) 265 4e-71
At1g06990 unknown protein 262 2e-70
At1g58480 hypothetical protein 261 4e-70
At3g14820 hypothetical protein 259 2e-69
At1g75890 family II lipase EXL2 (EXL2) 259 2e-69
At2g31540 putative GDSL-motif lipase/hydrolase 256 1e-68
At1g58430 unknown protein 256 2e-68
At2g30220 putative GDSL-motif lipase/hydrolase 250 7e-67
>At2g04570 putative GDSL-motif lipase/hydrolase
Length = 350
Score = 493 bits (1270), Expect = e-140
Identities = 229/348 (65%), Positives = 281/348 (79%)
Query: 11 YTNNILCILLLQRLDLSLVKSAKVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLG 70
+ ++ IL L + ++ + K+PAIIVFGDSSVDAGNNN+I TVARSNF+PYGRDF+G
Sbjct: 3 HLKSLFTILFLIAMSSTVTFAGKIPAIIVFGDSSVDAGNNNYIPTVARSNFEPYGRDFVG 62
Query: 71 GKPTGRFSNGRIATDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDV 130
GKPTGRF NG+IATDF+SEA G+KP IPAYLDPS+NIS FATGV+FASAATGYDNATSDV
Sbjct: 63 GKPTGRFCNGKIATDFMSEALGLKPIIPAYLDPSYNISDFATGVTFASAATGYDNATSDV 122
Query: 131 LSVIPLWKQLEYYKEYQKKLGAYLGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQ 190
LSV+PLWKQLEYYKEYQ KL AY G+ + ETI +LY+IS+GTNDFLENY+ PGR+SQ
Sbjct: 123 LSVLPLWKQLEYYKEYQTKLKAYQGKDRGTETIESSLYLISIGTNDFLENYFAFPGRSSQ 182
Query: 191 YTPSEYQNFLAGIAQNFIHKLYDLGAKKISLGGLPPMGCLPLERTTNFAGGNDCVSNYNN 250
Y+ S YQ+FLAGIA+ F+ KL+ LGA+KISLGGLPPMGC+PLER TN G +CV YN+
Sbjct: 183 YSVSLYQDFLAGIAKEFVKKLHGLGARKISLGGLPPMGCMPLERATNIGTGGECVGRYND 242
Query: 251 IALEFNGKLNKLTTKLKKDLPGIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFE 310
IA++FN KL+K+ KL K+LPG LVFSNPY+ + ++K P +GF+V ACCATGMFE
Sbjct: 243 IAVQFNSKLDKMVEKLSKELPGSNLVFSNPYEPFMRIIKNPSSFGFEVVGAACCATGMFE 302
Query: 311 MGYACSRASLFSCMDASRYVFWDSFHPTEKTNGIVANYLVKNALAQFL 358
MGY C R + F+C +A +YVFWDSFHPT+KTN I+AN L+ + FL
Sbjct: 303 MGYGCQRNNPFTCTNADKYVFWDSFHPTQKTNHIMANALMNSTFPHFL 350
>At4g26790 putative APG protein
Length = 351
Score = 428 bits (1100), Expect = e-120
Identities = 205/347 (59%), Positives = 263/347 (75%), Gaps = 2/347 (0%)
Query: 13 NNILCILLLQRLDLSLVKS--AKVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLG 70
N +L LLL L + AK PA+IVFGDS+VD+GNNN ISTV +SNFQPYGRD+
Sbjct: 4 NRVLAFLLLAAQLLVKIPETCAKFPALIVFGDSTVDSGNNNQISTVLKSNFQPYGRDYFD 63
Query: 71 GKPTGRFSNGRIATDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDV 130
GK TGRFSNGRIA DFISE G+K +PAYLDP++NI+ FATGV FASA TG DNATS V
Sbjct: 64 GKATGRFSNGRIAPDFISEGLGLKNAVPAYLDPAYNIADFATGVCFASAGTGLDNATSAV 123
Query: 131 LSVIPLWKQLEYYKEYQKKLGAYLGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQ 190
LSV+PLWK++EYYKEYQ +L +YLGE+KA E I+++LY+IS+GTNDFLENYY +P + +
Sbjct: 124 LSVMPLWKEVEYYKEYQTRLRSYLGEEKANEIISESLYLISIGTNDFLENYYLLPRKLRK 183
Query: 191 YTPSEYQNFLAGIAQNFIHKLYDLGAKKISLGGLPPMGCLPLERTTNFAGGNDCVSNYNN 250
Y+ +EYQ FL GIA +F+ +Y LGA+K+SL GL P GCLPLERTT G+ C+ YN
Sbjct: 184 YSVNEYQYFLIGIAADFVTDIYRLGARKMSLSGLSPFGCLPLERTTQLFYGSKCIEEYNI 243
Query: 251 IALEFNGKLNKLTTKLKKDLPGIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFE 310
+A +FN K+ + +L +DL GI+LVFSNPYD++ ++ P +GF+ ACC TG +E
Sbjct: 244 VARDFNIKMEEKVFQLNRDLNGIQLVFSNPYDLVSEIIYHPEAFGFENVRSACCGTGYYE 303
Query: 311 MGYACSRASLFSCMDASRYVFWDSFHPTEKTNGIVANYLVKNALAQF 357
M Y C + + F+C DAS+YVFWDSFHPTEKTN IVAN+++K L++F
Sbjct: 304 MSYLCDKMNPFTCSDASKYVFWDSFHPTEKTNAIVANHVLKYDLSRF 350
>At2g42990 putative GDSL-motif lipase/hydrolase
Length = 303
Score = 394 bits (1011), Expect = e-110
Identities = 172/297 (57%), Positives = 238/297 (79%)
Query: 56 VARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEAFGIKPYIPAYLDPSFNISQFATGVS 115
+AR+NF+PYGRDF GG+ TGRF NGR+++DF SEA+G+KP +PAYLDPS+NIS FATGV
Sbjct: 1 MARANFEPYGRDFPGGRATGRFCNGRLSSDFTSEAYGLKPTVPAYLDPSYNISDFATGVC 60
Query: 116 FASAATGYDNATSDVLSVIPLWKQLEYYKEYQKKLGAYLGEKKAKETITKALYIISLGTN 175
FASA TGYDN+T+DVL VIPLWK++EY+KEYQ L AYLG ++A + I ++LYI+S+GTN
Sbjct: 61 FASAGTGYDNSTADVLGVIPLWKEVEYFKEYQSNLSAYLGHRRAAKIIRESLYIVSIGTN 120
Query: 176 DFLENYYTIPGRASQYTPSEYQNFLAGIAQNFIHKLYDLGAKKISLGGLPPMGCLPLERT 235
DFLENYYT+P R SQ++ S+YQ+FL IA+ F+ +Y LGA+K+S G+ PMGCLPLER
Sbjct: 121 DFLENYYTLPDRRSQFSISQYQDFLVEIAEVFLKDIYRLGARKMSFTGISPMGCLPLERV 180
Query: 236 TNFAGGNDCVSNYNNIALEFNGKLNKLTTKLKKDLPGIRLVFSNPYDVLLGVVKKPGQYG 295
TN C +YN++A++FNG+L +L TKL ++L GI++ F+NPYD++ +V KP YG
Sbjct: 181 TNLDDPFSCARSYNDLAVDFNGRLRRLVTKLNRELTGIKIYFANPYDIMWDIVTKPNLYG 240
Query: 296 FQVASMACCATGMFEMGYACSRASLFSCMDASRYVFWDSFHPTEKTNGIVANYLVKN 352
+++S ACC TG+FEMG+ C + + +C DA+++VFWD+FHPTE+TN IV+++ K+
Sbjct: 241 LEISSSACCGTGLFEMGFLCGQDNPLTCSDANKFVFWDAFHPTERTNQIVSDHFFKH 297
>At5g03820 putative protein
Length = 354
Score = 280 bits (715), Expect = 1e-75
Identities = 139/319 (43%), Positives = 200/319 (62%), Gaps = 2/319 (0%)
Query: 34 VPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEAFGI 93
VPA+I+ GDS VDAGNNN ++T+ ++NF PYGRDFL TGRFSNG++ATDF +E+ G
Sbjct: 28 VPALIIMGDSVVDAGNNNRLNTLIKANFPPYGRDFLAHNATGRFSNGKLATDFTAESLGF 87
Query: 94 KPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYYKEYQKKLGAY 153
Y YL N + TG +FAS A+GYD+ T+ + I L +QL+ YKEYQ K+
Sbjct: 88 TSYPVPYLSQEANGTNLLTGANFASGASGYDDGTAIFYNAITLNQQLKNYKEYQNKVTNI 147
Query: 154 LGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAGIAQNFIHKLYD 213
+G ++A + + A++++S G++DFL++YY P +TP +Y + L F+ LYD
Sbjct: 148 VGSERANKIFSGAIHLLSTGSSDFLQSYYINPILNRIFTPDQYSDRLMKPYSTFVQNLYD 207
Query: 214 LGAKKISLGGLPPMGCLPLERTTNFAGGND--CVSNYNNIALEFNGKLNKLTTKLKKDLP 271
LGA+KI + LPP+GCLP T GN+ CV N A+ FN KLN + L +LP
Sbjct: 208 LGARKIGVTTLPPLGCLPAAITLFGETGNNNTCVERLNQDAVSFNTKLNNTSMNLTNNLP 267
Query: 272 GIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCMDASRYVF 331
G++LV + Y+ LL + P + GF + ACC TG E + C+ S+ +C +A+ YVF
Sbjct: 268 GLKLVVFDIYNPLLNMAMNPVENGFFESRRACCGTGTVETSFLCNARSVGTCSNATNYVF 327
Query: 332 WDSFHPTEKTNGIVANYLV 350
WD FHP+E N ++AN L+
Sbjct: 328 WDGFHPSEAANRVIANNLL 346
>At1g75900 family II lipase EXL3 (EXL3)
Length = 364
Score = 278 bits (712), Expect = 3e-75
Identities = 130/325 (40%), Positives = 195/325 (60%), Gaps = 1/325 (0%)
Query: 34 VPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEAFGI 93
+PA+I FGDS VD G NN + TV + +F PYG +F G TGRF +GR+ D ++E GI
Sbjct: 41 IPAVIAFGDSIVDTGMNNNVKTVVKCDFLPYGINFQSGVATGRFCDGRVPADLLAEELGI 100
Query: 94 KPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYYKEYQKKLGAY 153
K +PAYLDP+ TGVSFAS +GYD T +++VI L QL Y++EY +K+
Sbjct: 101 KSIVPAYLDPNLKSKDLLTGVSFASGGSGYDPITPKLVAVISLEDQLSYFEEYIEKVKNI 160
Query: 154 LGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAGIAQNFIHKLYD 213
+GE + + +L+++ G++D YYT+ R +Y Y ++ A F+ KLY
Sbjct: 161 VGEARKDFIVANSLFLLVAGSDDIANTYYTLRARP-EYDVDSYTTLMSDSASEFVTKLYG 219
Query: 214 LGAKKISLGGLPPMGCLPLERTTNFAGGNDCVSNYNNIALEFNGKLNKLTTKLKKDLPGI 273
G +++++ G PP+GC+P +RT DC NYN A FN KL+ L+K LPGI
Sbjct: 220 YGVRRVAVFGAPPIGCVPSQRTLGGGILRDCADNYNEAAKLFNSKLSPKLDSLRKTLPGI 279
Query: 274 RLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCMDASRYVFWD 333
+ ++ N YD L +++ P YGF+V++ CC TG E+ C++ + C D S +VFWD
Sbjct: 280 KPIYINIYDPLFDIIQNPANYGFEVSNKGCCGTGAIEVAVLCNKITSSVCPDVSTHVFWD 339
Query: 334 SFHPTEKTNGIVANYLVKNALAQFL 358
S+HPTEKT ++ + L+ + QF+
Sbjct: 340 SYHPTEKTYKVLVSLLINKFVNQFV 364
>At5g03810 putative protein
Length = 320
Score = 274 bits (700), Expect = 6e-74
Identities = 136/311 (43%), Positives = 194/311 (61%), Gaps = 1/311 (0%)
Query: 41 GDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEAFGIKPYIPAY 100
GDS VDAGNNN T+ ++NF PYGRDF+ TGRFSNG++ATDF +E G Y AY
Sbjct: 2 GDSVVDAGNNNHRITLVKANFPPYGRDFVAHSATGRFSNGKLATDFTAENLGFTSYPVAY 61
Query: 101 LDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYYKEYQKKLGAYLGEKKAK 160
L N + TG +FAS A+G+D+AT+ + I L +QL+ YKEYQ K+ +G+++A
Sbjct: 62 LSQEANETNLLTGANFASGASGFDDATAIFYNAITLSQQLKNYKEYQNKVTNIVGKERAN 121
Query: 161 ETITKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAGIAQNFIHKLYDLGAKKIS 220
E + A++++S G++DFL++YY P +TP +Y + L F+ LY LGA++I
Sbjct: 122 EIFSGAIHLLSTGSSDFLQSYYINPILNRIFTPDQYSDHLLRSYSTFVQNLYGLGARRIG 181
Query: 221 LGGLPPMGCLPLERTT-NFAGGNDCVSNYNNIALEFNGKLNKLTTKLKKDLPGIRLVFSN 279
+ LPP+GCLP T G N CV N A+ FN KLN + L +LPG++LV +
Sbjct: 182 VTTLPPLGCLPAAITLFGGVGNNMCVERLNQDAVSFNTKLNNTSINLTNNLPGLKLVVFD 241
Query: 280 PYDVLLGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCMDASRYVFWDSFHPTE 339
Y+ LL +V P +YGF + ACC TG E + C+ S+ +C +A+ YVFWD FHP+E
Sbjct: 242 IYNPLLNMVINPVEYGFFESRRACCGTGTMETSFLCNALSVGTCSNATNYVFWDGFHPSE 301
Query: 340 KTNGIVANYLV 350
N ++AN L+
Sbjct: 302 AANRVIANNLL 312
>At3g16370 putative APG protein
Length = 353
Score = 273 bits (698), Expect = 1e-73
Identities = 143/341 (41%), Positives = 202/341 (58%), Gaps = 2/341 (0%)
Query: 12 TNNILCILLLQRLDLSLVKSAK-VPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLG 70
T++ L + L+ L + + A+ VPAI+ FGDS VD GNNN++ T+ R+++ PYGRDF
Sbjct: 5 TSSFLLLTLVSTLSILQISFAQLVPAIMTFGDSVVDVGNNNYLPTLFRADYPPYGRDFAN 64
Query: 71 GKPTGRFSNGRIATDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDV 130
K TGRF NG++ATD +E G Y PAYL P + G +FASAA+GYD+ + +
Sbjct: 65 HKATGRFCNGKLATDITAETLGFTKYPPAYLSPEASGKNLLIGANFASAASGYDDKAALL 124
Query: 131 LSVIPLWKQLEYYKEYQKKLGAYLGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQ 190
IPL++Q+EY+KEY+ KL G KKA I A+ ++S G++DF++NYY P
Sbjct: 125 NHAIPLYQQVEYFKEYKSKLIKIAGSKKADSIIKGAICLLSAGSSDFVQNYYVNPLLYKV 184
Query: 191 YTPSEYQNFLAGIAQNFIHKLYDLGAKKISLGGLPPMGCLPLERTTNFAGGNDCVSNYNN 250
YT Y +FL FI ++Y +GA+KI + LPP GCLP RT CVS N
Sbjct: 185 YTVDAYGSFLIDNFSTFIKQVYAVGARKIGVTSLPPTGCLPAARTLFGFHEKGCVSRLNT 244
Query: 251 IALEFNGKLNKLTTKLKKDLPGIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFE 310
A FN KLN +KL+K +++V + Y L +V+ P + GF A+ CC TG E
Sbjct: 245 DAQNFNKKLNAAASKLQKQYSDLKIVVFDIYSPLYDLVQNPSKSGFTEATKGCCGTGTVE 304
Query: 311 -MGYACSRASLFSCMDASRYVFWDSFHPTEKTNGIVANYLV 350
C+ S +C +A++YVFWDS HP+E N I+A L+
Sbjct: 305 TTSLLCNPKSFGTCSNATQYVFWDSVHPSEAANEILATALI 345
>At1g75880 family II extracellular lipase 1 like protein (EXL1)
Length = 374
Score = 271 bits (694), Expect = 3e-73
Identities = 137/332 (41%), Positives = 201/332 (60%)
Query: 26 LSLVKSAKVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIATD 85
+ + K+ VPA+IVFGDS VDAGNN+ + T AR ++ PYG DF GG TGRFSNG++ D
Sbjct: 42 VKIPKNTTVPAVIVFGDSIVDAGNNDDMITEARCDYAPYGIDFDGGVATGRFSNGKVPGD 101
Query: 86 FISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYYKE 145
++E GIKP IPAY +P+ + TGV+FAS GY T+ + IPL +QL Y++E
Sbjct: 102 IVAEELGIKPNIPAYRNPNLKPEELLTGVTFASGGAGYVPLTTKIAGGIPLPQQLIYFEE 161
Query: 146 YQKKLGAYLGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAGIAQ 205
Y +KL +GEK+ K I +L+++ G+ND +++T+P YT + + +A A+
Sbjct: 162 YIEKLKQMVGEKRTKFIIKNSLFVVICGSNDIANDFFTLPPVRLHYTVASFTALMADNAR 221
Query: 206 NFIHKLYDLGAKKISLGGLPPMGCLPLERTTNFAGGNDCVSNYNNIALEFNGKLNKLTTK 265
+F LY GA++I + G PP+GC+P +RT DCV+ +N+ A FN KL+
Sbjct: 222 SFAQTLYGYGARRILVFGAPPIGCVPSQRTVAGGPTRDCVARFNDAAKLFNTKLSANIDV 281
Query: 266 LKKDLPGIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCMD 325
L + L +++ + Y LL ++ P QYGF+VA+ CC TG+ E+ C+ + C
Sbjct: 282 LSRTLQDPTIIYIDIYSPLLDLILNPHQYGFKVANKGCCGTGLIEVTALCNNYTASVCPI 341
Query: 326 ASRYVFWDSFHPTEKTNGIVANYLVKNALAQF 357
S YVFWDSFHPTEK I+ L+ L +F
Sbjct: 342 RSDYVFWDSFHPTEKAYRIIVAKLLDRYLNRF 373
>At1g59406 proline-rich protein, putative
Length = 349
Score = 270 bits (689), Expect = 1e-72
Identities = 128/329 (38%), Positives = 204/329 (61%), Gaps = 7/329 (2%)
Query: 30 KSAKVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISE 89
K+ +PA+IVFGDS +D GNNN + T+ + NF PYG+D+ GG TGRFS+GR+ +D I+E
Sbjct: 24 KNTTIPALIVFGDSIMDTGNNNNLPTLLKCNFPPYGKDYPGGFATGRFSDGRVPSDLIAE 83
Query: 90 AFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYYKEYQKK 149
G+ +PAY++P GV+FAS TGYD T+ ++SVI +W QL +KEY K
Sbjct: 84 KLGLAKTLPAYMNPYLKPEDLLKGVTFASGGTGYDPLTAKIMSVISVWDQLINFKEYISK 143
Query: 150 LGAYLGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAGIAQNFIH 209
+ + GE+KAK+ + + +++ +ND Y + +Y + Y NFLA A +F+
Sbjct: 144 IKRHFGEEKAKDILEHSFFLVVSSSNDLAHTYL---AQTHRYDRTSYANFLADSAVHFVR 200
Query: 210 KLYDLGAKKISLGGLPPMGCLPLERTTNFAG--GNDCVSNYNNIALEFNGKLNKLTTKLK 267
+L+ LGA+KI + P+GC+PL+RT F G C NN+A +FN +L+ L
Sbjct: 201 ELHKLGARKIGVFSAVPVGCVPLQRTV-FGGFFTRGCNQPLNNMAKQFNARLSPALDSLD 259
Query: 268 KDLPGIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCMDAS 327
K+L G+ +++ N YD L +++ P +YGF+VA CC G+ + Y C+ + F+C ++S
Sbjct: 260 KELDGV-ILYINVYDTLFDMIQHPKKYGFEVADRGCCGKGLLAISYLCNSLNPFTCSNSS 318
Query: 328 RYVFWDSFHPTEKTNGIVANYLVKNALAQ 356
Y+FWDS+HP+E+ ++ + L+ L++
Sbjct: 319 AYIFWDSYHPSERAYQVIVDNLLDKYLSK 347
>At1g59030 proline-rich protein, putative
Length = 349
Score = 270 bits (689), Expect = 1e-72
Identities = 128/329 (38%), Positives = 204/329 (61%), Gaps = 7/329 (2%)
Query: 30 KSAKVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISE 89
K+ +PA+IVFGDS +D GNNN + T+ + NF PYG+D+ GG TGRFS+GR+ +D I+E
Sbjct: 24 KNTTIPALIVFGDSIMDTGNNNNLPTLLKCNFPPYGKDYPGGFATGRFSDGRVPSDLIAE 83
Query: 90 AFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYYKEYQKK 149
G+ +PAY++P GV+FAS TGYD T+ ++SVI +W QL +KEY K
Sbjct: 84 KLGLAKTLPAYMNPYLKPEDLLKGVTFASGGTGYDPLTAKIMSVISVWDQLINFKEYISK 143
Query: 150 LGAYLGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAGIAQNFIH 209
+ + GE+KAK+ + + +++ +ND Y + +Y + Y NFLA A +F+
Sbjct: 144 IKRHFGEEKAKDILEHSFFLVVSSSNDLAHTYL---AQTHRYDRTSYANFLADSAVHFVR 200
Query: 210 KLYDLGAKKISLGGLPPMGCLPLERTTNFAG--GNDCVSNYNNIALEFNGKLNKLTTKLK 267
+L+ LGA+KI + P+GC+PL+RT F G C NN+A +FN +L+ L
Sbjct: 201 ELHKLGARKIGVFSAVPVGCVPLQRTV-FGGFFTRGCNQPLNNMAKQFNARLSPALDSLD 259
Query: 268 KDLPGIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCMDAS 327
K+L G+ +++ N YD L +++ P +YGF+VA CC G+ + Y C+ + F+C ++S
Sbjct: 260 KELDGV-ILYINVYDTLFDMIQHPKKYGFEVADRGCCGKGLLAISYLCNSLNPFTCSNSS 318
Query: 328 RYVFWDSFHPTEKTNGIVANYLVKNALAQ 356
Y+FWDS+HP+E+ ++ + L+ L++
Sbjct: 319 AYIFWDSYHPSERAYQVIVDNLLDKYLSK 347
>At5g22810 GDSL-motif lipase/hydrolase-like protein
Length = 337
Score = 267 bits (682), Expect = 8e-72
Identities = 136/325 (41%), Positives = 190/325 (57%), Gaps = 3/325 (0%)
Query: 34 VPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEAFGI 93
VPAI +FGDS VD GNNN I T+ ++NF PYGRDF PTGRF NG++ATDF +E G
Sbjct: 10 VPAIFIFGDSVVDVGNNNDIYTIVKANFPPYGRDFTTHTPTGRFCNGKLATDFTAENLGF 69
Query: 94 KPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYYKEYQKKLGAY 153
K Y AYL G +FASAA+GY + T+ + S I L +QLE+YK+Y ++
Sbjct: 70 KSYPQAYLSKKAKGKNLLIGANFASAASGYYDGTAKLYSAISLPQQLEHYKDYISRIQEI 129
Query: 154 L---GEKKAKETITKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAGIAQNFIHK 210
A I+ +YI+S G++DF++NYY P +P E+ + L +FI
Sbjct: 130 ATSNNNSNASAIISNGIYIVSAGSSDFIQNYYINPLLYRDQSPDEFSDLLILSYSSFIQN 189
Query: 211 LYDLGAKKISLGGLPPMGCLPLERTTNFAGGNDCVSNYNNIALEFNGKLNKLTTKLKKDL 270
LY LGA++I + LPP+GCLP T C NN A+ FN KLN + LK++L
Sbjct: 190 LYSLGARRIGVTTLPPLGCLPAAITVVGPHEGGCSEKLNNDAISFNNKLNTTSQDLKRNL 249
Query: 271 PGIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCMDASRYV 330
G+ LV + Y L + +P ++GF A ACC TG+ E C+ S+ +C +A+ YV
Sbjct: 250 IGLNLVVFDIYQPLYDLATRPSEFGFAEARRACCGTGLLETSILCNPKSVGTCNNATEYV 309
Query: 331 FWDSFHPTEKTNGIVANYLVKNALA 355
FWD FHPTE N I+A+ L+ + ++
Sbjct: 310 FWDGFHPTEAANKILADNLLVSGIS 334
>At5g45960 GDSL-motif lipase/hydrolase-like protein
Length = 375
Score = 266 bits (679), Expect = 2e-71
Identities = 132/314 (42%), Positives = 199/314 (63%), Gaps = 8/314 (2%)
Query: 34 VPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEAFGI 93
V AI+VFGDS+VD GNNN+I TV + NF PYG DF PTGRF NGR+ TDFI+ G+
Sbjct: 45 VSAILVFGDSTVDPGNNNYIDTVFKCNFPPYGLDFRNKTPTGRFCNGRLVTDFIASYIGV 104
Query: 94 KPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYYKEYQKKLGAY 153
K +P YLDP+ I++ +GVSFASA +GYD T + +VI + QLEY++EY++KL
Sbjct: 105 KENVPPYLDPNLGINELISGVSFASAGSGYDPLTPTITNVIDIPTQLEYFREYKRKLEGK 164
Query: 154 LGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAGIAQNFIHKLYD 213
+G+++ ++ I +A++ +S GTNDF+ NY+TIP R +T YQ F+ + FI L+
Sbjct: 165 MGKQEMEKHIEEAMFCVSAGTNDFVINYFTIPIRRKTFTIEAYQQFVISNLKQFIQGLWK 224
Query: 214 LGAKKISLGGLPPMGCLPLERTTNFAG----GNDCVSNYNNIALEFNGKLNKLTTKLKKD 269
GA+KI++ GLPP+GCLP+ T F+G C+ ++ +A +N L K ++
Sbjct: 225 EGARKITVAGLPPIGCLPIV-ITLFSGEALTNRRCIDRFSTVATNYNFLLQKQLALMQVG 283
Query: 270 LP--GIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCMDAS 327
L G ++ + + Y+ + V++ P ++GF+ CC +G E + C+ S + C + S
Sbjct: 284 LAHLGSKIFYLDVYNPVYEVIRDPRKFGFEEVFSGCCGSGYLEASFLCNPKS-YVCPNTS 342
Query: 328 RYVFWDSFHPTEKT 341
YVF+DS HP+EKT
Sbjct: 343 AYVFFDSIHPSEKT 356
>At1g20120 anter-specific proline-rich like protein (APG precursor)
Length = 402
Score = 265 bits (676), Expect = 4e-71
Identities = 129/323 (39%), Positives = 191/323 (58%)
Query: 35 PAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEAFGIK 94
PAI FGDS +D GNN++I T+ ++NF PYG +F PTGRF NG+I +DFI++ G+K
Sbjct: 77 PAIFAFGDSILDTGNNDYILTLIKANFLPYGMNFPDKVPTGRFCNGKIPSDFIADYIGVK 136
Query: 95 PYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYYKEYQKKLGAYL 154
P +PAYL P TGVSFAS +GYD T V+S IP+ KQL Y++EY +K+ ++
Sbjct: 137 PVVPAYLRPGLTQEDLLTGVSFASGGSGYDPLTPIVVSAIPMSKQLTYFQEYIEKVKGFV 196
Query: 155 GEKKAKETITKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAGIAQNFIHKLYDL 214
G++KA+ I+K L I+ G++D YY Y Y +F+A A +F +LY+
Sbjct: 197 GKEKAEHIISKGLAIVVAGSDDLANTYYGEHLEEFLYDIDTYTSFMASSAASFAMQLYES 256
Query: 215 GAKKISLGGLPPMGCLPLERTTNFAGGNDCVSNYNNIALEFNGKLNKLTTKLKKDLPGIR 274
GAKKI G+ P+GC+P++RTT C N A FN KL+ +L K +
Sbjct: 257 GAKKIGFIGVSPIGCIPIQRTTRGGLKRKCADELNFAAQLFNSKLSTSLNELAKTMKNTT 316
Query: 275 LVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCMDASRYVFWDS 334
LV+ + Y +++ P +YGF CC TG+ E+G C++ + C + S ++FWDS
Sbjct: 317 LVYIDIYSSFNDMIQNPKKYGFDEIDRGCCGTGLLELGPLCNKYTSLLCKNVSSFMFWDS 376
Query: 335 FHPTEKTNGIVANYLVKNALAQF 357
+HPTE+ I++ V+N + F
Sbjct: 377 YHPTERAYKILSQKFVENDMGPF 399
>At1g06990 unknown protein
Length = 360
Score = 262 bits (670), Expect = 2e-70
Identities = 127/330 (38%), Positives = 197/330 (59%), Gaps = 3/330 (0%)
Query: 24 LDLSLVKSAKVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIA 83
++++ V + PAI+VFGDS++D GNNN+I T R+NF PYG +F G TGRFSNG++
Sbjct: 25 INVTNVNVSMFPAILVFGDSTIDTGNNNYIKTYIRANFPPYGCNFPGHNATGRFSNGKLI 84
Query: 84 TDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYY 143
DFI+ GIK +P +LDP + S TGV FASA +GYDN T S + + KQ +
Sbjct: 85 PDFIASLMGIKDTVPPFLDPHLSDSDIITGVCFASAGSGYDNLTDRATSTLSVDKQADML 144
Query: 144 KEYQKKLGAYLGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAGI 203
+ Y ++L +G++KA +++AL I+S GTNDF N Y P R + YQ+F+
Sbjct: 145 RSYVERLSQIVGDEKAASIVSEALVIVSSGTNDFNLNLYDTPSRRQKLGVDGYQSFILSN 204
Query: 204 AQNFIHKLYDLGAKKISLGGLPPMGCLPLERTTNFAGGND--CVSNYNNIALEFNGKLNK 261
NF+ +LYD+G +KI + GLPP+GCLP++ T N+ C+ N+ + EFN KL
Sbjct: 205 VHNFVQELYDIGCRKIMVLGLPPVGCLPIQMTMAMQKQNERRCIDKQNSDSQEFNQKLKN 264
Query: 262 LTTKLKKDLPGIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLF 321
T+++ +L G + + + Y L + P +YG + + CC TG E+ Y C+ +
Sbjct: 265 SLTEMQSNLTGSVIFYGDIYGALFDMATNPQRYGLKETTRGCCGTGEIELAYLCNALTRI 324
Query: 322 SCMDASRYVFWDSFHPTEKTNGIVANYLVK 351
C + ++Y+FWD HP++ +++ LV+
Sbjct: 325 -CPNPNQYLFWDDIHPSQIAYIVISLSLVE 353
>At1g58480 hypothetical protein
Length = 342
Score = 261 bits (667), Expect = 4e-70
Identities = 131/347 (37%), Positives = 211/347 (60%), Gaps = 17/347 (4%)
Query: 15 ILCILLLQRLDLSLVK---SAKVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGG 71
+L L+L ++ + VK +A +PA+IVFGDS +D GNNN + T+ + NF PYG+D+ GG
Sbjct: 6 LLLALVLIAVEANAVKQGINATIPALIVFGDSIMDTGNNNNLPTLLKCNFPPYGKDYPGG 65
Query: 72 KPTGRFSNGRIATDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVL 131
TGRFS+GR+ +D I+E G+ +PAY++ GV+FAS TGYD T+ ++
Sbjct: 66 DATGRFSDGRVPSDLIAEKLGLAKTLPAYMNSYLKPEDLLKGVTFASRGTGYDPLTAKIM 125
Query: 132 SVIPLWKQLEYYKEYQKKLGAYLGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQY 191
SVI +W QL Y+KEY K+ + GE+KAK+ + + +++ +ND Y +A +Y
Sbjct: 126 SVISVWDQLIYFKEYISKIKRHFGEEKAKDILEHSFFLVVSSSNDLAHTYL---AQAHRY 182
Query: 192 TPSEYQNFLAGIAQNFIHKLYDLGAKKISLGGLPPMGCLPLERTTNFAG--GNDCVSNYN 249
+ Y NFLA A +F+ +L+ LGA+KI + P+GC+PL+RT F G C N
Sbjct: 183 DRTSYANFLADSAVHFVRELHKLGARKIGVFSAVPVGCVPLQRTV-FGGFFTRGCNEPLN 241
Query: 250 NIALEFNGKLNKLTTKLKKDLPGIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMF 309
N+A +FN +L+ L K+L G+ +++ N YD L +++ P +YG CC G+
Sbjct: 242 NMAKQFNARLSPALDSLDKELDGV-ILYINVYDTLFDMIQHPKKYG-------CCGKGLL 293
Query: 310 EMGYACSRASLFSCMDASRYVFWDSFHPTEKTNGIVANYLVKNALAQ 356
+ Y C+ + F+C ++S Y+FWDS+HP+E+ ++ + L+ L++
Sbjct: 294 TISYLCNSLNPFTCSNSSSYIFWDSYHPSERAYQVIVDNLLDKYLSK 340
>At3g14820 hypothetical protein
Length = 311
Score = 259 bits (662), Expect = 2e-69
Identities = 130/306 (42%), Positives = 188/306 (60%), Gaps = 4/306 (1%)
Query: 45 VDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEAFGIKPYIPAYLDPS 104
+D GNNN I T+ +SNF PYGRDF G PTGRFS+G++ +D I+E+ GI +P YL +
Sbjct: 1 MDTGNNNDIPTLLKSNFPPYGRDFPGAIPTGRFSDGKVPSDIIAESLGIAKTLPPYLGSN 60
Query: 105 FNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYYKEYQKKLGAYLGEKKAKETIT 164
GV FAS +GYD TS +LSV+ + QL+Y++EY K+ + GE+K K +
Sbjct: 61 LKPHDLLKGVIFASGGSGYDPLTSTLLSVVSMSDQLKYFQEYLAKIKQHFGEEKVKFILE 120
Query: 165 KALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAGIAQNFIHKLYDLGAKKISLGGL 224
K+++++ +ND E Y+ R+ +Y + Y +L +A FI +L +LGAK I L
Sbjct: 121 KSVFLVVSSSNDLAETYWV---RSVEYDRNSYAEYLVELASEFIKELSELGAKNIGLFSG 177
Query: 225 PPMGCLPLERTTNFAGGNDCVSNYNNIALEFNGKLNKLTTKLKKDLPGIRLVFSNPYDVL 284
P+GCLP +RT C NN+AL FN KL+ LKK+LP RL+F + YD L
Sbjct: 178 VPVGCLPAQRTLFGGFERKCYEKLNNMALHFNSKLSSSLDTLKKELPS-RLIFIDVYDTL 236
Query: 285 LGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCMDASRYVFWDSFHPTEKTNGI 344
L ++K P YGF+VA CC TG E+ C++ + F+C DAS +VF+DS+HP+EK I
Sbjct: 237 LDIIKNPTNYGFKVADKGCCGTGKIELMELCNKFTPFTCSDASTHVFFDSYHPSEKAYQI 296
Query: 345 VANYLV 350
+ + L+
Sbjct: 297 ITHKLL 302
>At1g75890 family II lipase EXL2 (EXL2)
Length = 379
Score = 259 bits (661), Expect = 2e-69
Identities = 137/346 (39%), Positives = 203/346 (58%), Gaps = 17/346 (4%)
Query: 18 ILLLQRLDLSLVK---SAKVPAIIVFGDSSVDAGNNNFI-STVARSNFQPYGRDFLGGKP 73
+LL + +LVK + PAIIVFGDS VDAGNN+ I +T+AR N+ PYG DF GG P
Sbjct: 26 VLLCKTSTNALVKQPPNETTPAIIVFGDSIVDAGNNDDIMTTLARCNYPPYGIDFDGGIP 85
Query: 74 TGRFSNGRIATDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSV 133
TGRF NG++ATDFI+ FGIKP IPAY +P+ TGV+FAS GY T+ + +
Sbjct: 86 TGRFCNGKVATDFIAGKFGIKPSIPAYRNPNLKPEDLLTGVTFASGGAGYVPFTTQLSTY 145
Query: 134 IPLWK-------------QLEYYKEYQKKLGAYLGEKKAKETITKALYIISLGTNDFLEN 180
+ ++K QL+ ++EY +K+ +GE++ K I +L+++ G+ND
Sbjct: 146 LFIYKPLLFLKGGIALSQQLKLFEEYVEKMKKMVGEERTKLIIKNSLFMVICGSNDITNT 205
Query: 181 YYTIPGRASQYTPSEYQNFLAGIAQNFIHKLYDLGAKKISLGGLPPMGCLPLERTTNFAG 240
Y+ +P QY + + +A A++F KL++ GA++I + G PP+GC+P +RT
Sbjct: 206 YFGLPSVQQQYDVASFTTLMADNARSFAQKLHEYGARRIQVFGAPPVGCVPSQRTLAGGP 265
Query: 241 GNDCVSNYNNIALEFNGKLNKLTTKLKKDLPGIRLVFSNPYDVLLGVVKKPGQYGFQVAS 300
+CV +N+ +N KL L + L +++ + YD LL ++ P QYGF+V
Sbjct: 266 TRNCVVRFNDATKLYNVKLAANLGSLSRTLGDKTIIYVDIYDSLLDIILDPRQYGFKVVD 325
Query: 301 MACCATGMFEMGYACSRASLFSCMDASRYVFWDSFHPTEKTNGIVA 346
CC TG+ E+ C+ + C + YVFWDSFHPTEKT I+A
Sbjct: 326 KGCCGTGLIEVALLCNNFAADVCPNRDEYVFWDSFHPTEKTYRIMA 371
>At2g31540 putative GDSL-motif lipase/hydrolase
Length = 360
Score = 256 bits (654), Expect = 1e-68
Identities = 126/320 (39%), Positives = 193/320 (59%), Gaps = 8/320 (2%)
Query: 35 PAIIVFGDSSVDAGNNNF-ISTVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEAFGI 93
PAI++FGDS+VD GNNN+ + T+ R+ PYG D GK GRFSNG++ +D I+ I
Sbjct: 34 PAILIFGDSTVDTGNNNYPLPTIFRAEHFPYGMDLPDGKANGRFSNGKLISDIIATKLNI 93
Query: 94 KPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYYKEYQKKLGAY 153
K +IP +L P+ + TGV FASA GYD+ TS I + +Q +K Y +L
Sbjct: 94 KEFIPPFLQPNLSDQDILTGVCFASAGAGYDDLTSLSTQAIRVSEQPNMFKSYIARLKGI 153
Query: 154 LGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQYT-PSEYQNFLAGIAQNFIHKLY 212
+G+KKA E I A ++S G NDF+ NYY IP R +Y S YQ+F+ +NF+ +LY
Sbjct: 154 VGDKKAMEIINNAFVVVSAGPNDFILNYYEIPSRRLEYPFISGYQDFILKRLENFVRELY 213
Query: 213 DLGAKKISLGGLPPMGCLPLERTTNFAG-GNDCVSNYNNIALEFNGKLNKLTTKLKKDLP 271
LG + + +GGLPPMGCLP+ T F C+ ++N ++ +N KL L +++ LP
Sbjct: 214 SLGVRNVLVGGLPPMGCLPIHMTAKFRNIFRFCLEHHNKDSVLYNEKLQNLLPQIEASLP 273
Query: 272 GIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLFS--CMDASRY 329
G + ++++ Y+ ++ +++ P +YGF+ CC TG E + C ++FS C + S +
Sbjct: 274 GSKFLYADVYNPMMEMIQNPSKYGFKETKRGCCGTGFLETSFMC---NVFSPVCQNRSEF 330
Query: 330 VFWDSFHPTEKTNGIVANYL 349
+F+DS HP+E T ++ N L
Sbjct: 331 LFFDSIHPSEATYNVIGNLL 350
>At1g58430 unknown protein
Length = 360
Score = 256 bits (653), Expect = 2e-68
Identities = 127/318 (39%), Positives = 191/318 (59%), Gaps = 4/318 (1%)
Query: 35 PAIIVFGDSSVDAGNNNFIS-TVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEAFGI 93
PAI++FGDS+VD GNNN+ S T+ R+ PYG D P GRFSNG+I +D I+ I
Sbjct: 34 PAILIFGDSTVDTGNNNYPSQTIFRAKHVPYGIDLPNHSPNGRFSNGKIFSDIIATKLNI 93
Query: 94 KPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYYKEYQKKLGAY 153
K ++P +L P+ + TGV FASA GYD+ TS I + +Q +K Y +L +
Sbjct: 94 KQFVPPFLQPNLTDQEIVTGVCFASAGAGYDDQTSLTTQAIRVSEQPNMFKSYIARLKSI 153
Query: 154 LGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQY-TPSEYQNFLAGIAQNFIHKLY 212
+G+KKA + I AL ++S G NDF+ NYY +P Y + S+YQ+F+ NF+ +LY
Sbjct: 154 VGDKKAMKIINNALVVVSAGPNDFILNYYEVPSWRRMYPSISDYQDFVLSRLNNFVKELY 213
Query: 213 DLGAKKISLGGLPPMGCLPLERTTNFAGG-NDCVSNYNNIALEFNGKLNKLTTKLKKDLP 271
LG +KI +GGLPPMGCLP++ T F C+ N ++ +N KL KL + + L
Sbjct: 214 SLGCRKILVGGLPPMGCLPIQMTAQFRNVLRFCLEQENRDSVLYNQKLQKLLPQTQASLT 273
Query: 272 GIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCMDASRYVF 331
G ++++S+ YD ++ +++ P +YGF+ + CC TG E + C+ S C + S ++F
Sbjct: 274 GSKILYSDVYDPMMEMLQNPSKYGFKETTRGCCGTGFLETSFMCNAYSSM-CQNRSEFLF 332
Query: 332 WDSFHPTEKTNGIVANYL 349
+DS HP+E T + N L
Sbjct: 333 FDSIHPSEATYNYIGNVL 350
>At2g30220 putative GDSL-motif lipase/hydrolase
Length = 358
Score = 250 bits (639), Expect = 7e-67
Identities = 127/318 (39%), Positives = 188/318 (58%), Gaps = 4/318 (1%)
Query: 35 PAIIVFGDSSVDAGNNNFIS-TVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEAFGI 93
PAI++FGDS+ D GNNN+ S V ++N PYG D G + GRFSNG++ +D IS I
Sbjct: 32 PAILIFGDSTADTGNNNYYSQAVFKANHLPYGVDLPGHEANGRFSNGKLISDVISTKLNI 91
Query: 94 KPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYYKEYQKKLGAY 153
K ++P +L P+ + TGV FASA GYD+ TS IP+ +Q +K Y +L
Sbjct: 92 KEFVPPFLQPNISDQDIVTGVCFASAGAGYDDETSLSSKAIPVSQQPSMFKNYIARLKGI 151
Query: 154 LGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQY-TPSEYQNFLAGIAQNFIHKLY 212
+G+KKA E I AL +IS G NDF+ N+Y IP R +Y T YQ+F+ F+ +LY
Sbjct: 152 VGDKKAMEIINNALVVISAGPNDFILNFYDIPIRRLEYPTIYGYQDFVLKRLDGFVRELY 211
Query: 213 DLGAKKISLGGLPPMGCLPLERTTNFAG-GNDCVSNYNNIALEFNGKLNKLTTKLKKDLP 271
LG + I +GGLPPMGCLP++ T CV N ++ +N KL K +++ LP
Sbjct: 212 SLGCRNILVGGLPPMGCLPIQLTAKLRTILGICVEQENKDSILYNQKLVKKLPEIQASLP 271
Query: 272 GIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCMDASRYVF 331
G + +++N YD ++ +++ P +YGF+ CC TG E + C+ S +C + S ++F
Sbjct: 272 GSKFLYANVYDPVMDMIRNPSKYGFKETKKGCCGTGYLETSFLCTSLSK-TCPNHSDHLF 330
Query: 332 WDSFHPTEKTNGIVANYL 349
WDS HP+E + N++
Sbjct: 331 WDSIHPSEAAYKYLGNFI 348
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.138 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,061,196
Number of Sequences: 26719
Number of extensions: 344193
Number of successful extensions: 1229
Number of sequences better than 10.0: 113
Number of HSP's better than 10.0 without gapping: 110
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 759
Number of HSP's gapped (non-prelim): 123
length of query: 359
length of database: 11,318,596
effective HSP length: 100
effective length of query: 259
effective length of database: 8,646,696
effective search space: 2239494264
effective search space used: 2239494264
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC140670.14


