

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140549.19 - phase: 0
(373 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
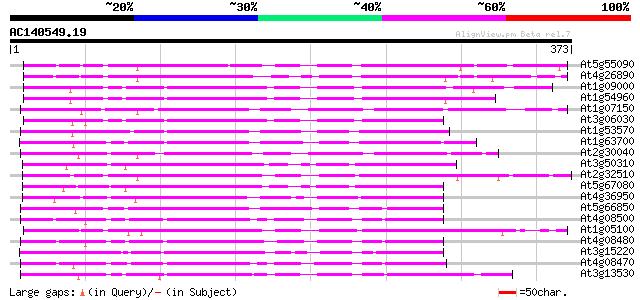
At5g55090 putative protein 202 3e-52
At4g26890 putative NPK1-related protein kinase 201 4e-52
At1g09000 NPK1-related protein kinase 1S (ANP1) 197 8e-51
At1g54960 NPK1-related protein kinase 2 (ANP2) 197 1e-50
At1g07150 hypothetical protein 196 1e-50
At3g06030 NPK1-related protein kinase 3 196 2e-50
At1g53570 MEK kinase MAP3Ka, putative 196 2e-50
At1g63700 putative protein kinase 195 3e-50
At2g30040 putative protein kinase 189 3e-48
At3g50310 protein kinase -like protein 184 5e-47
At2g32510 putative protein kinase 184 5e-47
At5g67080 protein kinase-like protein 183 1e-46
At4g36950 MAP3K-like protein kinase 181 6e-46
At5g66850 MAP protein kinase like protein 176 2e-44
At4g08500 MEKK1/MAP kinase kinase kinase 174 7e-44
At1g05100 putative NPK1-related MAP kinase 172 3e-43
At4g08480 putative mitogen-activated protein kinase 166 1e-41
At3g15220 putative MAP kinase 164 6e-41
At4g08470 putative mitogen-activated protein kinase 163 2e-40
At3g13530 MAP3K epsilon protein kinase 160 8e-40
>At5g55090 putative protein
Length = 448
Score = 202 bits (513), Expect = 3e-52
Identities = 135/373 (36%), Positives = 205/373 (54%), Gaps = 45/373 (12%)
Query: 10 WMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGRDALENEVNILNTLNSSPSPYIV 69
W++G ++G GS +V L + S G F VK A + L+ E +IL+ L+S PYIV
Sbjct: 6 WIRGPIIGRGSTATVSLGITNS-GDFFAVKSAEFSSSA-FLQREQSILSKLSS---PYIV 60
Query: 70 QCLGTDYDHQDNQL--HVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLYHLHQ 127
+ +G++ ++++L ++ MEY+SGGSL D+ GG L E ++R YTRQI+ GL +LH
Sbjct: 61 KYIGSNVTKENDKLMYNLLMEYVSGGSLHDLIKNSGGKLPEPLIRSYTRQILKGLMYLHD 120
Query: 128 HGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEVL 187
GIVHCD+K +NV++ K+ D GCAK V++N N+ S GTP +M+PEV
Sbjct: 121 QGIVHCDVKSQNVMIGGE-IAKIVDLGCAKTVEENENLEFS--------GTPAFMSPEVA 171
Query: 188 LMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMFKIACG 247
+ S AD+W+LGCTVIEMATG PW + +++ +AA++KI
Sbjct: 172 RGEEQS-----------FPADVWALGCTVIEMATGSSPWPE-----LNDVVAAIYKIGFT 215
Query: 248 DGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVSTTLTHHKH---YSASSP 304
P P+ S++G DFLR+CL +DPK+R T ELL HPFL + ++SSP
Sbjct: 216 GESPVIPVWLSEKGQDFLRKCLRKDPKQRWTVEELLQHPFLDEEDNDSDQTGNCLNSSSP 275
Query: 305 ASVLEVHQFEDTYDDDDDDDDSDDNDELISPAGGNHFFITKELLCPQGTTMWQLEDSASG 364
++VL+ +F D + +D+++ P + F+ + P D +SG
Sbjct: 276 STVLD-QRFWDLCETSRSRFIKEDHED---PFANSTNFLWDDDSLPGDRIKKLAGDESSG 331
Query: 365 ------NHWITIR 371
N WI +R
Sbjct: 332 EPDWETNGWIEVR 344
>At4g26890 putative NPK1-related protein kinase
Length = 444
Score = 201 bits (512), Expect = 4e-52
Identities = 141/376 (37%), Positives = 202/376 (53%), Gaps = 54/376 (14%)
Query: 10 WMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGRDALENEVNILNTLNSSPSPYIV 69
W +G ++G GS +V +A++ S+G LF VK A + L+ E +IL+TL+S P++V
Sbjct: 5 WTRGPIIGRGSTATVSIAIS-SSGELFAVKSADLSSS-SLLQKEQSILSTLSS---PHMV 59
Query: 70 QCLGTDYDHQDNQL--HVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLYHLHQ 127
+ +GT + N L ++ MEY+SGG+L D+ GG L E +R YTRQI++GL +LH+
Sbjct: 60 KYIGTGLTRESNGLVYNILMEYVSGGNLHDLIKNSGGKLPEPEIRSYTRQILNGLVYLHE 119
Query: 128 HGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEVL 187
GIVHCDLK NVL+ +G +K+AD GCAK V K+ GTP +MAPEV
Sbjct: 120 RGIVHCDLKSHNVLVEENGVLKIADMGCAKSVDKS-----------EFSGTPAFMAPEV- 167
Query: 188 LMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMFKIACG 247
+ +E R AD+W+LGCT+IEM TG PW + +++ +AAM+KI
Sbjct: 168 ------ARGEEQRF----PADVWALGCTMIEMMTGSSPWPE-----LNDVVAAMYKIGFS 212
Query: 248 DGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL----VSTTLTHHKHYSASS 303
P P S + DFL+ CL D K+R T ELL HPFL S T K+ SS
Sbjct: 213 GESPAIPAWISDKAKDFLKNCLKEDQKQRWTVEELLKHPFLDDDEESQTSDCLKN-KTSS 271
Query: 304 PASVLEVHQFEDTYDD--------DDDDDDSDDNDELISPAGGNHFFITKELLCPQGTTM 355
P++VL+ +F D+ + D +D ++ ++ L SPA E
Sbjct: 272 PSTVLD-QRFWDSCESSKSHLVSIDHEDPFAEYSESLDSPADRIEKLAGDEF---SSLLD 327
Query: 356 WQLEDSASGNHWITIR 371
W ED WI +R
Sbjct: 328 WDTEDDGG---WIQVR 340
>At1g09000 NPK1-related protein kinase 1S (ANP1)
Length = 666
Score = 197 bits (501), Expect = 8e-51
Identities = 131/371 (35%), Positives = 191/371 (51%), Gaps = 55/371 (14%)
Query: 10 WMKGKMVGCGSFGSVHLAMNKSTGGLFVVK----------KAHSEAGRDALENEVNILNT 59
W KG+++G G+FG+V++ MN +G L VK K ++A LE EV +L
Sbjct: 69 WRKGQLIGRGAFGTVYMGMNLDSGELLAVKQVLIAANFASKEKTQAHIQELEEEVKLLKN 128
Query: 60 LNSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIV 119
L+ P IV+ LGT +D+ L++ +E++ GGS++ + KFG E VVR YTRQ++
Sbjct: 129 LSH---PNIVRYLGTV--REDDTLNILLEFVPGGSISSLLEKFG-PFPESVVRTYTRQLL 182
Query: 120 HGLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTP 179
GL +LH H I+H D+K N+L+ + G +KLADFG +K+V + M + S GTP
Sbjct: 183 LGLEYLHNHAIMHRDIKGANILVDNKGCIKLADFGASKQVAELATMTGAKSM----KGTP 238
Query: 180 LWMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMA 239
WMAPEV+L +S +ADIWS+GCTVIEM TG+ PW +A
Sbjct: 239 YWMAPEVILQTGHS-----------FSADIWSVGCTVIEMVTGKAPWSQQ-----YKEVA 282
Query: 240 AMFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVSTTLTHHKHY 299
A+F I P P S + DFL +CL P R TA ELL HPF++ HK
Sbjct: 283 AIFFIGTTKSHPPIPDTLSSDAKDFLLKCLQEVPNLRPTASELLKHPFVMG----KHKES 338
Query: 300 SASSPASV---------LEVHQFEDTYDDDDDDDDSDDNDELISPAGGNHFFITKELLCP 350
+++ SV L+++ + T D DD N G ++ + +
Sbjct: 339 ASTDLGSVLNNLSTPLPLQINNTKSTPDSTCDDVGDMCN------FGSLNYSLVDPVKSI 392
Query: 351 QGTTMWQLEDS 361
Q +WQ D+
Sbjct: 393 QNKNLWQQNDN 403
>At1g54960 NPK1-related protein kinase 2 (ANP2)
Length = 651
Score = 197 bits (500), Expect = 1e-50
Identities = 123/332 (37%), Positives = 180/332 (54%), Gaps = 44/332 (13%)
Query: 10 WMKGKMVGCGSFGSVHLAMNKSTGGLFVVK----------KAHSEAGRDALENEVNILNT 59
W KG+++G G+FG+V++ MN +G L VK K ++A LE EV +L
Sbjct: 68 WRKGQLIGRGAFGTVYMGMNLDSGELLAVKQVLITSNCASKEKTQAHIQELEEEVKLLKN 127
Query: 60 LNSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIV 119
L+ P IV+ LGT +D L++ +E++ GGS++ + KFG + E VVR YT Q++
Sbjct: 128 LSH---PNIVRYLGTV--REDETLNILLEFVPGGSISSLLEKFG-AFPESVVRTYTNQLL 181
Query: 120 HGLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTP 179
GL +LH H I+H D+K N+L+ + G +KLADFG +K+V + ++ + S GTP
Sbjct: 182 LGLEYLHNHAIMHRDIKGANILVDNQGCIKLADFGASKQVAELATISGAKSM----KGTP 237
Query: 180 LWMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMA 239
WMAPEV+L +S +ADIWS+GCTVIEM TG+ PW +A
Sbjct: 238 YWMAPEVILQTGHS-----------FSADIWSVGCTVIEMVTGKAPWSQQ-----YKEIA 281
Query: 240 AMFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL-------VSTT 292
A+F I P P + S + DFL +CL ++P R TA ELL HPF+ S
Sbjct: 282 AIFHIGTTKSHPPIPDNISSDANDFLLKCLQQEPNLRPTASELLKHPFVTGKQKESASKD 341
Query: 293 LTHHKHYSASS-PASVLEVHQFEDTYDDDDDD 323
LT S S P+ + + ++ + DD D
Sbjct: 342 LTSFMDNSCSPLPSELTNITSYQTSTSDDVGD 373
>At1g07150 hypothetical protein
Length = 499
Score = 196 bits (499), Expect = 1e-50
Identities = 126/372 (33%), Positives = 198/372 (52%), Gaps = 55/372 (14%)
Query: 8 SEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAG----RDALENEVNILNTLNSS 63
S W++G +G G FG+V A++K+ G +F VK ++LENE+++ +L
Sbjct: 21 SFWVRGACIGRGCFGAVSTAISKTNGEVFAVKSVDLATSLPTQSESLENEISVFRSLK-- 78
Query: 64 PSPYIVQCLGTDYDHQDNQL--HVFMEYMSGGSLADVSHKFGGSL-NEDVVRVYTRQIVH 120
P PYIV+ LG + ++++EY+ G +A SH+ GG + +E +++ YT +V
Sbjct: 79 PHPYIVKFLGDGVSKEGTTTFRNLYLEYLPNGDVA--SHRAGGKIEDETLLQRYTACLVS 136
Query: 121 GLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPL 180
L H+H G VHCD+K +N+L++ S VKLADFG A R+ + +I G+PL
Sbjct: 137 ALRHVHSQGFVHCDVKARNILVSQSSMVKLADFGSAFRI-------HTPRALITPRGSPL 189
Query: 181 WMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAA 240
WMAPEV+ R +D+WSLGCT+IEM TG+P W D + S+S
Sbjct: 190 WMAPEVI-----------RREYQGPESDVWSLGCTIIEMFTGKPAWEDHGIDSLS----- 233
Query: 241 MFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVSTTLTHHKHYS 300
+I+ D +P FP S+ G DFL +CL RDP +R + +LL HPFL + H+ +
Sbjct: 234 --RISFSDELPVFPSKLSEIGRDFLEKCLKRDPNQRWSCDQLLQHPFL---SQCHNSSPT 288
Query: 301 ASSPASVLE-VHQFEDTYDDDDDDDDSDDNDELISPAGGNHFFITKELLCPQGTTMWQLE 359
SSP VL+ V+ D +++++ S+ D K ++C TT +
Sbjct: 289 ESSPRCVLDWVNSGFDLEEEEEEVGRSEFED------------AAKAIICNLATTGGVIW 336
Query: 360 DSASGNHWITIR 371
+S + W+ +R
Sbjct: 337 ES---DGWVEVR 345
>At3g06030 NPK1-related protein kinase 3
Length = 651
Score = 196 bits (498), Expect = 2e-50
Identities = 115/289 (39%), Positives = 169/289 (57%), Gaps = 36/289 (12%)
Query: 10 WMKGKMVGCGSFGSVHLAMNKSTGGLFVVKK---AHSEAGRDA-------LENEVNILNT 59
W KG+++GCG+FG V++ MN +G L +K+ A S A ++ LE EV +L
Sbjct: 68 WRKGELIGCGAFGRVYMGMNLDSGELLAIKQVLIAPSSASKEKTQGHIRELEEEVQLLKN 127
Query: 60 LNSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIV 119
L+ P IV+ LGT + + L++ ME++ GGS++ + KFG S E V+ +YT+Q++
Sbjct: 128 LSH---PNIVRYLGTV--RESDSLNILMEFVPGGSISSLLEKFG-SFPEPVIIMYTKQLL 181
Query: 120 HGLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTP 179
GL +LH +GI+H D+K N+L+ + G ++LADFG +K+V + +N + S GTP
Sbjct: 182 LGLEYLHNNGIMHRDIKGANILVDNKGCIRLADFGASKKVVELATVNGAKSM----KGTP 237
Query: 180 LWMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMA 239
WMAPEV+L +S +ADIWS+GCTVIEMATG+PPW + A
Sbjct: 238 YWMAPEVILQTGHS-----------FSADIWSVGCTVIEMATGKPPWSEQ-----YQQFA 281
Query: 240 AMFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL 288
A+ I P P S E DFL +CL ++P R +A ELL HPF+
Sbjct: 282 AVLHIGRTKAHPPIPEDLSPEAKDFLMKCLHKEPSLRLSATELLQHPFV 330
>At1g53570 MEK kinase MAP3Ka, putative
Length = 609
Score = 196 bits (498), Expect = 2e-50
Identities = 113/289 (39%), Positives = 164/289 (56%), Gaps = 30/289 (10%)
Query: 8 SEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKK----AHSEAGRDALENEVNILNTLNSS 63
S W KGK +G G+FG V+L N G + +K+ + + ++ L+ +N LN
Sbjct: 212 STWKKGKFLGSGTFGQVYLGFNSEKGKMCAIKEVKVISDDQTSKECLKQLNQEINLLNQL 271
Query: 64 PSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLY 123
P IVQ G++ + L V++EY+SGGS+ + +G S E V++ YTRQI+ GL
Sbjct: 272 CHPNIVQYYGSELSEET--LSVYLEYVSGGSIHKLLKDYG-SFTEPVIQNYTRQILAGLA 328
Query: 124 HLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMA 183
+LH VH D+K N+L+ +G +KLADFG AK V + S +++ G+P WMA
Sbjct: 329 YLHGRNTVHRDIKGANILVDPNGEIKLADFGMAKHV-------TAFSTMLSFKGSPYWMA 381
Query: 184 PEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMFK 243
PEV++ +N + A DIWSLGCT++EMAT +PPW S +AA+FK
Sbjct: 382 PEVVMSQNGYT----------HAVDIWSLGCTILEMATSKPPW------SQFEGVAAIFK 425
Query: 244 IACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVSTT 292
I P+ P H S + +F+R CL R+P R TA +LL HPFL +TT
Sbjct: 426 IGNSKDTPEIPDHLSNDAKNFIRLCLQRNPTVRPTASQLLEHPFLRNTT 474
>At1g63700 putative protein kinase
Length = 883
Score = 195 bits (496), Expect = 3e-50
Identities = 116/311 (37%), Positives = 172/311 (55%), Gaps = 37/311 (11%)
Query: 7 GSEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKA-------HSEAGRDALENEVNILNT 59
GS W KG+++G GSFG V+L N +G + +K+ S L E+++L+
Sbjct: 397 GSRWKKGRLLGMGSFGHVYLGFNSESGEMCAMKEVTLCSDDPKSRESAQQLGQEISVLSR 456
Query: 60 LNSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIV 119
L IVQ G++ D++L++++EY+SGGS+ + ++G E+ +R YT+QI+
Sbjct: 457 LRHQN---IVQYYGSET--VDDKLYIYLEYVSGGSIYKLLQEYG-QFGENAIRNYTQQIL 510
Query: 120 HGLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTP 179
GL +LH VH D+K N+L+ G VK+ADFG AK + + S ++ G+P
Sbjct: 511 SGLAYLHAKNTVHRDIKGANILVDPHGRVKVADFGMAKHI-------TAQSGPLSFKGSP 563
Query: 180 LWMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMA 239
WMAPEV+ N S++ A DIWSLGCTV+EMAT +PPW S +
Sbjct: 564 YWMAPEVIKNSNGSNL----------AVDIWSLGCTVLEMATTKPPW------SQYEGVP 607
Query: 240 AMFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVSTTLTHHKHY 299
AMFKI +P P H S+EG DF+R+CL R+P R TA +LL+H F V + +
Sbjct: 608 AMFKIGNSKELPDIPDHLSEEGKDFVRKCLQRNPANRPTAAQLLDHAF-VRNVMPMERPI 666
Query: 300 SASSPASVLEV 310
+ PA + V
Sbjct: 667 VSGEPAEAMNV 677
>At2g30040 putative protein kinase
Length = 463
Score = 189 bits (479), Expect = 3e-48
Identities = 122/325 (37%), Positives = 179/325 (54%), Gaps = 43/325 (13%)
Query: 8 SEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHSE----AGRDALENEVNILNTLNSS 63
S W++G VG G FG+V A++K GGLF VK + ++LENE+ IL ++ S
Sbjct: 15 SSWIRGSCVGRGCFGTVSKALSKIDGGLFAVKSIDLATCLPSQAESLENEIVILRSMKSH 74
Query: 64 PSPYIVQCLGTDYDHQDNQL--HVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHG 121
P+ IV+ LG D + ++ +EY G +A+ GG +NE ++R Y +V
Sbjct: 75 PN--IVRFLGDDVSKEGTASFRNLHLEYSPEGDVAN-----GGIVNETLLRRYVWCLVSA 127
Query: 122 LYHLHQHGIVHCDLKCKNVLLASSG-NVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPL 180
L H+H +GIVHCD+K KNVL+ + G +VKLADFG A +K S+ ++ G+PL
Sbjct: 128 LSHVHSNGIVHCDVKSKNVLVFNGGSSVKLADFGSAVEFEK-------STIHVSPRGSPL 180
Query: 181 WMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAA 240
WMAPEV+ R +D+WSLGCTVIEM TG+P W D S+S
Sbjct: 181 WMAPEVV-----------RREYQGPESDVWSLGCTVIEMLTGKPAWEDHGFDSLS----- 224
Query: 241 MFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVSTTLTHHKHYS 300
+I + +P P+ S+ G DFL +CL RD +R + +LL HPFL H ++
Sbjct: 225 --RIGFSNDLPFIPVGLSELGRDFLEKCLKRDRSQRWSCDQLLQHPFLCQD--HHDSFFT 280
Query: 301 ASSPASVLEVHQFEDTYDDDDDDDD 325
SSP VL+ E +D++++ D+
Sbjct: 281 ESSPRCVLDWVNSE--FDEEEESDE 303
>At3g50310 protein kinase -like protein
Length = 342
Score = 184 bits (468), Expect = 5e-47
Identities = 109/295 (36%), Positives = 163/295 (54%), Gaps = 25/295 (8%)
Query: 9 EWMKGKMVGCGSFGSVHLAMNKSTGGLF---VVKKAHSEAGRDALENEVNILNTLNSSPS 65
EW++G+ +G G+F +V A G F + K+ G +L NE ++L++L P
Sbjct: 2 EWVRGETIGFGTFSTVSTATKSRNSGDFPALIAVKSTDAYGAASLSNEKSVLDSLGDCPE 61
Query: 66 PYIVQCLGTD--YDHQDNQLHVFMEYMSGGSLADVSHKFGGS-LNEDVVRVYTRQIVHGL 122
I++C G D ++ + ++ +EY S GSLA K GG L E VR +T ++ GL
Sbjct: 62 --IIRCYGEDSTVENGEEMHNLLLEYASRGSLASYMKKLGGEGLPESTVRRHTGSVLRGL 119
Query: 123 YHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWM 182
H+H G HCD+K N+LL + G+VK+ADFG A RV ++ + S + GTPL+M
Sbjct: 120 RHIHAKGFAHCDIKLANILLFNDGSVKIADFGLAMRVDGDLTALRKS---VEIRGTPLYM 176
Query: 183 APEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMF 242
APE +ND +AAD+W+LGC V+EM +G+ W + S+ M+ +
Sbjct: 177 APE--------CVNDNEY---GSAADVWALGCAVVEMFSGKTAW---SVKEGSHFMSLLI 222
Query: 243 KIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVSTTLTHHK 297
+I GD +P+ P S+EG DFL +C V+DP KR TA LLNH F+ H+
Sbjct: 223 RIGVGDELPKIPEMLSEEGKDFLSKCFVKDPAKRWTAEMLLNHSFVTIDLEDDHR 277
>At2g32510 putative protein kinase
Length = 372
Score = 184 bits (468), Expect = 5e-47
Identities = 123/377 (32%), Positives = 193/377 (50%), Gaps = 42/377 (11%)
Query: 9 EWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGR-DALENEVNILNTLNSSPSPY 67
EW +G+++G GS +V+ A ++ + VK SE R + L+ E IL++L+S PY
Sbjct: 2 EWTRGRILGRGSTATVYAAAGHNSDEILAVKS--SEVHRSEFLQREAKILSSLSS---PY 56
Query: 68 IVQCLGTDYDHQDNQL---HVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLYH 124
++ G++ + N + ++ MEY G+L D + K GG ++E V YTR I+ GL +
Sbjct: 57 VIGYRGSETKRESNGVVMYNLLMEYAPYGTLTDAAAKDGGRVDETRVVKYTRDILKGLEY 116
Query: 125 LHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAP 184
+H GIVHCD+K NV+++ G K+ADFGCAKRV GTP +MAP
Sbjct: 117 IHSKGIVHCDVKGSNVVISEKGEAKIADFGCAKRVDPVFESPVM--------GTPAFMAP 168
Query: 185 EVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMFKI 244
EV + +DIW++GCT+IEM TG PPW D S +P++ ++++
Sbjct: 169 EVARGEKQGK-----------ESDIWAVGCTMIEMVTGSPPWTKAD--SREDPVSVLYRV 215
Query: 245 ACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVSTTLTHH---KHYSA 301
P+ P ++E DFL +CL R+ +R TA +LLNHPFL + +
Sbjct: 216 GYSSETPELPCLLAEEAKDFLEKCLKREANERWTATQLLNHPFLTTKPDIEPVLVPGLIS 275
Query: 302 SSPASVLEVHQFEDTYDDDDDD-----DDSDDNDELISPAGGNHFFITKELLCPQGTTMW 356
+SP SV + + ++++++ DS D D L G + L C G
Sbjct: 276 NSPTSVTDQTFWRSVEEEEEEETEEIQKDSRDLDRL--SLWGCYSERIGRLKCVGGLDGT 333
Query: 357 QLEDSASGNHWITIRSR 373
+ + G WI +R+R
Sbjct: 334 RCD--MEGGDWIMVRAR 348
>At5g67080 protein kinase-like protein
Length = 344
Score = 183 bits (465), Expect = 1e-46
Identities = 112/288 (38%), Positives = 159/288 (54%), Gaps = 28/288 (9%)
Query: 9 EWMKGKMVGCGSFGSVHLAMNKSTGG-----LFVVKKAHSEAGRDALENEVNILNTLNSS 63
EW++G+ +G G+F +V LA + L VK A S G +L NE ++L+ L
Sbjct: 2 EWIRGETIGYGTFSTVSLATRSNNDSGEFPPLMAVKSADSY-GAASLANEKSVLDNLGDD 60
Query: 64 PSPYIVQCLGTD--YDHQDNQLHVFMEYMSGGSLADVSHKFGGS-LNEDVVRVYTRQIVH 120
+ IV+C G D ++ + ++F+EY S GSL K G + E VR +T ++
Sbjct: 61 CNE-IVRCFGEDRTVENGEEMHNLFLEYASRGSLESYLKKLAGEGVPESTVRRHTGSVLR 119
Query: 121 GLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPL 180
GL H+H +G HCDLK N+LL G VK+ADFG AKR+ +N + GTPL
Sbjct: 120 GLRHIHANGFAHCDLKLGNILLFGDGAVKIADFGLAKRIGDLTALNYG----VQIRGTPL 175
Query: 181 WMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAA 240
+MAPE S+ND + D+W+LGC V+EM +G+ W L SN M+
Sbjct: 176 YMAPE--------SVNDNEY---GSEGDVWALGCVVVEMFSGKTAW---SLKEGSNFMSL 221
Query: 241 MFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL 288
+ +I GD +P P S++G DFL +C V+DPKKR TA LLNHPF+
Sbjct: 222 LLRIGVGDEVPMIPEELSEQGRDFLSKCFVKDPKKRWTAEMLLNHPFV 269
>At4g36950 MAP3K-like protein kinase
Length = 799
Score = 181 bits (459), Expect = 6e-46
Identities = 109/287 (37%), Positives = 166/287 (56%), Gaps = 30/287 (10%)
Query: 9 EWMKGKMVGCGSFGSVHLAM-----NKSTGGLFVVKKAHSEAGRDALENEVNILNTLNSS 63
EW++ + +G GSF +V LA +K+ L VK + AL NE ++L+ L
Sbjct: 2 EWIRRETIGHGSFSTVSLATTSGSSSKAFPSLMAVKSSGVVCSA-ALRNERDVLDDLGDC 60
Query: 64 PSPYIVQCLGTDYDHQDNQ--LHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHG 121
IV+C G ++ + ++F+EY SGGSLAD G +L E VR +TR IV G
Sbjct: 61 SE--IVRCFGEGRTVENGEEIYNLFLEYASGGSLADRIKSSGEALPEFEVRRFTRSIVKG 118
Query: 122 LYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLW 181
L H+H +G HCD+K +NVL+ G+VK++DFG AKR +S + GTPL+
Sbjct: 119 LCHIHGNGFTHCDIKLENVLVFGDGDVKISDFGLAKR--------RSGEVCVEIRGTPLY 170
Query: 182 MAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAM 241
MAPE + N+ + ADIW+LGC+V+EM++G+ W +D + ++N M+ +
Sbjct: 171 MAPESV---NHGEFE--------SPADIWALGCSVVEMSSGKTAWCLEDGV-MNNVMSLL 218
Query: 242 FKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL 288
+I GD +P+ P+ S+EG DF+ +C V++ +R TA LL+HPFL
Sbjct: 219 VRIGSGDEVPRIPVELSEEGKDFVSKCFVKNAAERWTAEMLLDHPFL 265
>At5g66850 MAP protein kinase like protein
Length = 533
Score = 176 bits (445), Expect = 2e-44
Identities = 112/288 (38%), Positives = 154/288 (52%), Gaps = 31/288 (10%)
Query: 8 SEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAH-------SEAGRDALENEVNILNTL 60
S+W KGK++G G+FGSV++A N TG L +K+ S LE E+ +L+ L
Sbjct: 161 SQWKKGKLIGRGTFGSVYVASNSETGALCAMKEVELFPDDPKSAECIKQLEQEIKLLSNL 220
Query: 61 NSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVH 120
P IVQ G++ +++ +++EY+ GS+ G++ E VVR +TR I+
Sbjct: 221 QH---PNIVQYFGSET--VEDRFFIYLEYVHPGSINKYIRDHCGTMTESVVRNFTRHILS 275
Query: 121 GLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPL 180
GL +LH VH D+K N+L+ +SG VKLADFG AK + ++ G+P
Sbjct: 276 GLAYLHNKKTVHRDIKGANLLVDASGVVKLADFGMAKHL-------TGQRADLSLKGSPY 328
Query: 181 WMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAA 240
WMAPE++ N + A DIWSLGCT+IEM TG+PPW S AA
Sbjct: 329 WMAPELMQAVMQKDSNPDLAF----AVDIWSLGCTIIEMFTGKPPW------SEFEGAAA 378
Query: 241 MFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL 288
MFK+ P P S EG DFLR C R+P +R TA LL H FL
Sbjct: 379 MFKVMRDS--PPIPESMSPEGKDFLRLCFQRNPAERPTASMLLEHRFL 424
>At4g08500 MEKK1/MAP kinase kinase kinase
Length = 608
Score = 174 bits (441), Expect = 7e-44
Identities = 113/288 (39%), Positives = 164/288 (56%), Gaps = 38/288 (13%)
Query: 8 SEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHS-EAGRDA------LENEVNILNTL 60
+ W KG+++G GSFGSV+ ++ G F VK+ + G A LE E+ +L+ L
Sbjct: 331 TSWQKGQLLGRGSFGSVYEGIS-GDGDFFAVKEVSLLDQGSQAQECIQQLEGEIKLLSQL 389
Query: 61 NSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVH 120
IV+ GT D + L++F+E ++ GSL + ++ L + VV +YTRQI+
Sbjct: 390 QHQN---IVRYRGTAKD--GSNLYIFLELVTQGSLLKLYQRY--QLRDSVVSLYTRQILD 442
Query: 121 GLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPL 180
GL +LH G +H D+KC N+L+ ++G VKLADFG AK K N SC GTP
Sbjct: 443 GLKYLHDKGFIHRDIKCANILVDANGAVKLADFGLAKVSK----FNDIKSC----KGTPF 494
Query: 181 WMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAA 240
WMAPEV+ K++ + ADIWSLGCTV+EM TG+ P+ D P+ A
Sbjct: 495 WMAPEVINRKDSDGYG--------SPADIWSLGCTVLEMCTGQIPYSD------LEPVQA 540
Query: 241 MFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL 288
+F+I G +P+ P S + F+ +CL +P++R TA ELLNHPF+
Sbjct: 541 LFRIGRGT-LPEVPDTLSLDARLFILKCLKVNPEERPTAAELLNHPFV 587
>At1g05100 putative NPK1-related MAP kinase
Length = 339
Score = 172 bits (436), Expect = 3e-43
Identities = 120/376 (31%), Positives = 185/376 (48%), Gaps = 55/376 (14%)
Query: 10 WMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGRDALENEVNILNTLNSSPSPYIV 69
W +GK +G GS +V A +G VK A + L+ E IL++LNS PY++
Sbjct: 3 WTRGKTLGRGSTATVSAATCHESGETLAVKSAEFHRS-EFLQREAKILSSLNS---PYVI 58
Query: 70 QCLGTDYD----HQDNQLHVF---MEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGL 122
G + H + + + MEY G+L DV+ K GG ++E V YTRQI+ GL
Sbjct: 59 GYRGCEITREPFHNNGEATTYSLLMEYAPYGTLTDVATKNGGFIDEARVVKYTRQILLGL 118
Query: 123 YHLHQH-GIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLW 181
++H GI HCD+K NVL+ +G K+ADFGCAK V+ + GTP +
Sbjct: 119 EYIHNSKGIAHCDIKGSNVLVGENGEAKIADFGCAKWVEPEITEPVR--------GTPAF 170
Query: 182 MAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAM 241
MAPE + +DIW++GCTVIEM TG PW+ D ++P++ +
Sbjct: 171 MAPEAARGERQGK-----------ESDIWAVGCTVIEMVTGSQPWIGADF---TDPVSVL 216
Query: 242 FKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVSTTLTHHKHYSA 301
+++ +P+ P +++ DFL +CL ++ +R TA +LLNHPFLV+
Sbjct: 217 YRVGYLGELPELPCSLTEQAKDFLGKCLKKEATERWTASQLLNHPFLVNKEPELVTGLVT 276
Query: 302 SSPASVLEVHQFEDTYDDDDDDDDS------DDNDELISPAGGNHFFITKELLCPQGTTM 355
+SP SV + + ++ +D S D+ ++S G H + +
Sbjct: 277 NSPTSVTDQMFWRSVEEEVSEDRSSWWECHEDERIGVLSWIG--HVVV---------EST 325
Query: 356 WQLEDSASGNHWITIR 371
W L+ G WIT+R
Sbjct: 326 WDLD----GEDWITVR 337
>At4g08480 putative mitogen-activated protein kinase
Length = 773
Score = 166 bits (421), Expect = 1e-41
Identities = 105/288 (36%), Positives = 164/288 (56%), Gaps = 38/288 (13%)
Query: 8 SEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHS-EAGRDA------LENEVNILNTL 60
+ W KG+++ GSFGSV+ A+++ G F VK+ + G A LE E+ +L+ L
Sbjct: 499 TSWQKGQLLRQGSFGSVYEAISED-GDFFAVKEVSLLDQGSQAQECIQQLEGEIALLSQL 557
Query: 61 NSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVH 120
I++ GTD D + L++F+E ++ GSL ++ ++ + + ++ +YT+QI+
Sbjct: 558 EHQN---ILRYRGTDKD--GSNLYIFLELVTQGSLLELYRRY--QIRDSLISLYTKQILD 610
Query: 121 GLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPL 180
GL +LH G +H D+KC +L+ ++G VKLADFG AK K N ++ T
Sbjct: 611 GLKYLHHKGFIHRDIKCATILVDANGTVKLADFGLAKVSKLNDIKSRKE--------TLF 662
Query: 181 WMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAA 240
WMAPEV+ K+N + ADIWSLGCTV+EM TG+ P+ D P+ A
Sbjct: 663 WMAPEVINRKDNDGYR--------SPADIWSLGCTVLEMCTGQIPYSD------LEPVEA 708
Query: 241 MFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL 288
+F+I G +P+ P S + F+ +CL +P++R TA ELLNHPF+
Sbjct: 709 LFRIRRGT-LPEVPDTLSLDARHFILKCLKLNPEERPTATELLNHPFV 755
>At3g15220 putative MAP kinase
Length = 690
Score = 164 bits (416), Expect = 6e-41
Identities = 100/282 (35%), Positives = 158/282 (55%), Gaps = 26/282 (9%)
Query: 7 GSEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGRDALENEVNILNTLNSSPSP 66
G+ + + +++G GSFG V+ A +K +K E D +E+ ++ L+ P
Sbjct: 12 GARFSQIELIGRGSFGDVYKAFDKDLNKEVAIKVIDLEESEDEIEDIQKEISVLSQCRCP 71
Query: 67 YIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLYHLH 126
YI + G+ Y HQ +L + MEYM+GGS+AD+ L+E + TR ++H + +LH
Sbjct: 72 YITEYYGS-YLHQ-TKLWIIMEYMAGGSVADLLQS-NNPLDETSIACITRDLLHAVEYLH 128
Query: 127 QHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEV 186
G +H D+K N+LL+ +G+VK+ADFG + ++ + ++ K+ GTP WMAPEV
Sbjct: 129 NEGKIHRDIKAANILLSENGDVKVADFGVSAQLTRTISRRKTFV------GTPFWMAPEV 182
Query: 187 LLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMFKIAC 246
+ +N+ N++ ADIWSLG TVIEMA G PP D +PM +F I
Sbjct: 183 I--QNSEGYNEK--------ADIWSLGITVIEMAKGEPPLAD------LHPMRVLF-IIP 225
Query: 247 GDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL 288
+ PQ HFS++ +F+ CL + P +R +A EL+ H F+
Sbjct: 226 RETPPQLDEHFSRQVKEFVSLCLKKAPAERPSAKELIKHRFI 267
>At4g08470 putative mitogen-activated protein kinase
Length = 560
Score = 163 bits (412), Expect = 2e-40
Identities = 108/290 (37%), Positives = 165/290 (56%), Gaps = 38/290 (13%)
Query: 8 SEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKA-------HSEAGRDALENEVNILNTL 60
+ W+KG+++G GS+ SV+ A+++ G F VK+ ++ LE E+ +L+ L
Sbjct: 301 TSWLKGQLLGRGSYASVYEAISED-GDFFAVKEVSLLDKGIQAQECIQQLEGEIALLSQL 359
Query: 61 NSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVH 120
IV+ GT D ++L++F+E ++ GS+ + ++ L+ VV +YTRQI+
Sbjct: 360 QHQN---IVRYRGTAKDV--SKLYIFLELVTQGSVQKLYERY--QLSYTVVSLYTRQILA 412
Query: 121 GLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPL 180
GL +LH G VH D+KC N+L+ ++G VKLADFG A+ K N SC GT
Sbjct: 413 GLNYLHDKGFVHRDIKCANMLVDANGTVKLADFGLAEASK----FNDIMSC----KGTLF 464
Query: 181 WMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAA 240
WMAPEV+ K++ + + ADIWSLGCTV+EM TG+ P+ D P+ A
Sbjct: 465 WMAPEVINRKDSDG--------NGSPADIWSLGCTVLEMCTGQIPYSD------LKPIQA 510
Query: 241 MFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVS 290
FKI G +P P S + F+ CL +P++R TA ELL+HPF+++
Sbjct: 511 AFKIGRGT-LPDVPDTLSLDARHFILTCLKVNPEERPTAAELLHHPFVIN 559
>At3g13530 MAP3K epsilon protein kinase
Length = 1368
Score = 160 bits (406), Expect = 8e-40
Identities = 110/333 (33%), Positives = 170/333 (51%), Gaps = 40/333 (12%)
Query: 8 SEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHSE----AGRDALENEVNILNTLNSS 63
+++M G +G G++G V+ ++ G +K+ E + + E+++L LN
Sbjct: 18 NKYMLGDEIGKGAYGRVYKGLDLENGDFVAIKQVSLENIVQEDLNTIMQEIDLLKNLNHK 77
Query: 64 PSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADV--SHKFGGSLNEDVVRVYTRQIVHG 121
IV+ LG+ LH+ +EY+ GSLA++ +KFG E +V VY Q++ G
Sbjct: 78 N---IVKYLGSS--KTKTHLHIILEYVENGSLANIIKPNKFG-PFPESLVAVYIAQVLEG 131
Query: 122 LYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLW 181
L +LH+ G++H D+K N+L G VKLADFG A ++ + ++N S GTP W
Sbjct: 132 LVYLHEQGVIHRDIKGANILTTKEGLVKLADFGVATKLNE-ADVNTHSVV-----GTPYW 185
Query: 182 MAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAM 241
MAPEV+ M AA+DIWS+GCTVIE+ T PP+ D PM A+
Sbjct: 186 MAPEVIEMSGVC-----------AASDIWSVGCTVIELLTCVPPYYD------LQPMPAL 228
Query: 242 FKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVSTTLTHHKHYSA 301
F+I D P P S + DFLR+C +D ++R A LL+HP++ ++
Sbjct: 229 FRIVQDDN-PPIPDSLSPDITDFLRQCFKKDSRQRPDAKTLLSHPWIRNSRRALQSSLRH 287
Query: 302 SSPASVLEVHQFEDTYDDDDDDDDSDDNDELIS 334
S ++ E T + DD+ S D E +S
Sbjct: 288 SGTIKYMK----EATASSEKDDEGSQDAAESLS 316
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.135 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,578,058
Number of Sequences: 26719
Number of extensions: 443904
Number of successful extensions: 7180
Number of sequences better than 10.0: 1085
Number of HSP's better than 10.0 without gapping: 900
Number of HSP's successfully gapped in prelim test: 187
Number of HSP's that attempted gapping in prelim test: 3319
Number of HSP's gapped (non-prelim): 1899
length of query: 373
length of database: 11,318,596
effective HSP length: 101
effective length of query: 272
effective length of database: 8,619,977
effective search space: 2344633744
effective search space used: 2344633744
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC140549.19


