

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140034.3 + phase: 0
(199 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
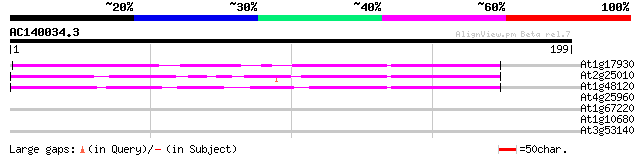
At1g17930 unknown protein 56 1e-08
At2g25010 unknown protein 52 2e-07
At1g48120 serine/threonine phosphatase PP7, putative 47 7e-06
At4g25960 P-glycoprotein-2 (pgp2) 29 1.4
At1g67220 hypothetical protein 29 1.4
At1g10680 putative P-glycoprotein-2 emb|CAA71277 29 1.4
At3g53140 caffeic acid O-methyltransferase - like protein 27 7.1
>At1g17930 unknown protein
Length = 478
Score = 55.8 bits (133), Expect = 1e-08
Identities = 43/173 (24%), Positives = 77/173 (43%), Gaps = 22/173 (12%)
Query: 2 PMGEMTVTLDDFACLTHLPIEGRMLDYEKKMPKHEGAALLMTYLGVSQHEAEKICNQEYG 61
P GEMT+TLD+ + + L ++G+ + K+ + L + + E G
Sbjct: 84 PCGEMTITLDEVSLILGLAVDGKPVVGVKEKDEDPSQVCLRLLGKLPKGELS-------G 136
Query: 62 GYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRWYLMYLVGCLLFDDRSN 121
++ L++ + A + +E+E Y R YL+Y+VG +F
Sbjct: 137 NRVTAKWLKESFAECPKGATM-------KEIE-------YHTRAYLIYIVGSTIFATTDP 182
Query: 122 KRIELIYLTTMADGYAGMRNYSWGGMTLAYLYGELADACRPGHRALGGSVTLL 174
+I + YL D + Y+WG LA+LY ++ +A + +GG +TLL
Sbjct: 183 SKISVDYLILFED-FEKAGEYAWGAAALAFLYRQIGNASQRSQSIIGGCLTLL 234
>At2g25010 unknown protein
Length = 509
Score = 52.0 bits (123), Expect = 2e-07
Identities = 48/175 (27%), Positives = 80/175 (45%), Gaps = 21/175 (12%)
Query: 1 MPMGEMTVTLDDFACLTHLPIEGRMLDYEKKMPKHEGAALLMTYLGVSQHEAEKICNQEY 60
+P+GEMT+TLD+ A + L I+G + K G + M G + N+E
Sbjct: 92 LPLGEMTITLDEVALVLGLEIDGDPIVGSK-----VGDEVAMDMCGRLLGKLPSAANKE- 145
Query: 61 GGYISYPRLRDFYTSYLGRANVLAGTEDPEELE-ELARVRTYCVRWYLMYLVGCLLFDDR 119
++ R++ ++L R +E PE+ ++ + T R YL+YL+G +F
Sbjct: 146 ---VNCSRVK---LNWLKR----TFSECPEDASFDVVKCHT---RAYLLYLIGSTIFATT 192
Query: 120 SNKRIELIYLTTMADGYAGMRNYSWGGMTLAYLYGELADACRPGHRALGGSVTLL 174
++ + YL D + Y+WG LA LY L +A + G +TLL
Sbjct: 193 DGDKVSVKYLPLFED-FDQAGRYAWGAAALACLYRALGNASLKSQSNICGCLTLL 246
>At1g48120 serine/threonine phosphatase PP7, putative
Length = 1338
Score = 47.0 bits (110), Expect = 7e-06
Identities = 45/174 (25%), Positives = 74/174 (41%), Gaps = 23/174 (13%)
Query: 1 MPMGEMTVTLDDFACLTHLPIEGRMLDYEKKMPKHEGAALLMTYLGVSQHEAEKICNQEY 60
+P GE+TVTL D L L ++G + K + A L LG + +
Sbjct: 101 LPAGEITVTLQDVNILLGLRVDGPAVTGSTK---YNWADLCEDLLGHRPGPKDL-----H 152
Query: 61 GGYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRWYLMYLVGCLLFDDRS 120
G ++S LR+ + + DP+E+ R + ++ L+ L+ D+S
Sbjct: 153 GSHVSLAWLRENFRNL---------PADPDEVTLKCHTRAF-----VLALMSGFLYGDKS 198
Query: 121 NKRIELIYLTTMADGYAGMRNYSWGGMTLAYLYGELADACRPGHRALGGSVTLL 174
+ L +L + D + + SWG TLA LY EL A + + G + LL
Sbjct: 199 KHDVALTFLPLLRD-FDEVAKLSWGSATLALLYRELCRASKRTVSTICGPLVLL 251
>At4g25960 P-glycoprotein-2 (pgp2)
Length = 1233
Score = 29.3 bits (64), Expect = 1.4
Identities = 24/72 (33%), Positives = 36/72 (49%), Gaps = 14/72 (19%)
Query: 40 LLMTY-LGVSQHEAEKICNQEYGGYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARV 98
+L TY L +S H +EK+ Q YGG D +YL +AN+LAG E + + V
Sbjct: 818 VLATYPLVISGHISEKLFMQGYGG--------DLNKAYL-KANMLAG----ESVSNIRTV 864
Query: 99 RTYCVRWYLMYL 110
+C ++ L
Sbjct: 865 AAFCAEEKILEL 876
>At1g67220 hypothetical protein
Length = 1357
Score = 29.3 bits (64), Expect = 1.4
Identities = 30/139 (21%), Positives = 56/139 (39%), Gaps = 27/139 (19%)
Query: 62 GYISYPRLRDFYTSY-------LGRANVLAGTEDPEELEELARVRTYCVRWYLMYLVGCL 114
GY+ Y +LR F TSY +G+ ++ ++ + ++R +WY+ L
Sbjct: 932 GYLEYCKLRGFTTSYIWACPPKIGQDYIMYSHPKTQQTPDTKKLR----KWYVSML---- 983
Query: 115 LFDDRSNKRIELIYLTTMADGYAGMRNYSWGGMTLAYLYGE---------LADACRPGHR 165
++ ++ ++ +T + D + L Y G + D R G+
Sbjct: 984 ---QKAAEQRVVMNVTNLYDRFFDSTEEYMTAARLPYFEGSFWSNRAEIMIQDIEREGNN 1040
Query: 166 ALGGSVTLLTVRKLKFYIY 184
L V LL+ RK+K Y
Sbjct: 1041 ELQKKVKLLSRRKVKTMSY 1059
>At1g10680 putative P-glycoprotein-2 emb|CAA71277
Length = 1227
Score = 29.3 bits (64), Expect = 1.4
Identities = 34/120 (28%), Positives = 56/120 (46%), Gaps = 21/120 (17%)
Query: 40 LLMTY-LGVSQHEAEKICNQEYGGYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARV 98
+L TY L +S H +EKI Q YGG +S +YL +AN+LAG E + + V
Sbjct: 810 VLATYPLIISGHISEKIFMQGYGGNLS--------KAYL-KANMLAG----ESISNIRTV 856
Query: 99 RTYCVRWYLMYLVGCLLFD--DRSNKRIELIYLTTMADGYAGMRNYSWGGMTLAYLYGEL 156
+C ++ L L + +RS +R ++ + Y + + + LA YG +
Sbjct: 857 VAFCAEEKVLDLYSKELLEPSERSFRRGQMAGIL-----YGVSQFFIFSSYGLALWYGSI 911
>At3g53140 caffeic acid O-methyltransferase - like protein
Length = 359
Score = 26.9 bits (58), Expect = 7.1
Identities = 21/74 (28%), Positives = 30/74 (40%), Gaps = 13/74 (17%)
Query: 15 CLTHLPIEGRMLDYEKKMPKH-----------EGAALLMT-YLGVSQHEAEKICNQEYGG 62
C LP+ G+++ E +PK EG +MT Y +H E+ E G
Sbjct: 278 CYNALPVGGKLIACEPVLPKETDESHRTRALLEGDIFVMTIYRTKGKHRTEEEF-IELGL 336
Query: 63 YISYPRLRDFYTSY 76
+P R FY Y
Sbjct: 337 SAGFPTFRPFYIDY 350
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.325 0.142 0.440
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,513,188
Number of Sequences: 26719
Number of extensions: 184366
Number of successful extensions: 481
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 473
Number of HSP's gapped (non-prelim): 8
length of query: 199
length of database: 11,318,596
effective HSP length: 94
effective length of query: 105
effective length of database: 8,807,010
effective search space: 924736050
effective search space used: 924736050
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC140034.3


