

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140034.13 + phase: 0 /partial
(433 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
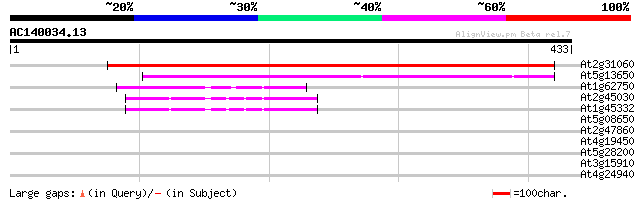
Score E
Sequences producing significant alignments: (bits) Value
At2g31060 putative GTP-binding protein 579 e-165
At5g13650 GTP-binding protein typA (tyrosine phosphorylated prot... 237 1e-62
At1g62750 elongation factor G, putative 43 3e-04
At2g45030 putative mitochondrial translation elongation factor G 42 5e-04
At1g45332 mitochondrial elongation factor, putative 42 5e-04
At5g08650 GTP-binding protein LepA homolog 33 0.40
At2g47860 unknown protein 32 0.68
At4g19450 unknown protein 31 1.2
At5g28200 putative protein 30 2.6
At3g15910 hypothetical protein 30 3.4
At4g24940 ubiquitin activating enzyme like protein 28 7.5
>At2g31060 putative GTP-binding protein
Length = 664
Score = 579 bits (1492), Expect = e-165
Identities = 284/345 (82%), Positives = 319/345 (92%)
Query: 76 LRNKDSGAEKIEDGKVVKVMKKKGTTMVVTDCAGAGDIISIAGLSSPSIGHTVTTVEIMS 135
LR DSG+EKIE+ KVVK+MKKKGTT+V D AGAGDII +AGL++PSIGHTV + E+ +
Sbjct: 294 LRKTDSGSEKIEEAKVVKLMKKKGTTIVSIDAAGAGDIICMAGLTAPSIGHTVASAEVTT 353
Query: 136 ALPTVELDPPTISMTFGVNDSPLAGRDGTHLTGGKIGDRLMAESETNLAINVLPGMSETF 195
ALPTVELDPPTISMTFGVNDSPLAG+DGTHLTGG+IGDRLMAE+ETNLAINV+PG+SE++
Sbjct: 354 ALPTVELDPPTISMTFGVNDSPLAGQDGTHLTGGRIGDRLMAEAETNLAINVIPGLSESY 413
Query: 196 EVQGRGELQLGILIENMRREGFELSVSPPKVMYKTDKGQKLEPIEEVTIEVNDEHVGFVM 255
EVQGRGELQLGILIENMRREGFELSVSPPKVMYKT+KGQKLEPIEEVTIE+NDEHVG VM
Sbjct: 414 EVQGRGELQLGILIENMRREGFELSVSPPKVMYKTEKGQKLEPIEEVTIEINDEHVGLVM 473
Query: 256 EALSHRRAEITDMGPVAGTVGRTRLCLTCPSRGLVGYRSVFSSETRGTGFMHRAFLKYEK 315
EALSHRRAE+ DMGPV G GRTRL LTCPSRGLVGYR VFSS+TRGTGFMHRAFL YEK
Sbjct: 474 EALSHRRAEVIDMGPVPGNEGRTRLSLTCPSRGLVGYRCVFSSDTRGTGFMHRAFLTYEK 533
Query: 316 FRGPLGNVRKGVLVSVGFGSITLHALMSLEARGTLFVSPGMEAYDGMIVGEHSRDTDLDV 375
+RGPLGNVRKGVLVS+ G+IT H+LMSLEARG LFVSPG+++YDGMI+GEHSR+TDLD+
Sbjct: 534 YRGPLGNVRKGVLVSMARGTITAHSLMSLEARGILFVSPGLDSYDGMIIGEHSRETDLDL 593
Query: 376 NPVRAKQLTNVRSVNKDDTVKLIPPRLMTLEEAIGYVASDELIEV 420
NPV+AK+LTN+RS KD+ VKL PPRLMTLEEAIGYVASDELIEV
Sbjct: 594 NPVKAKELTNIRSAGKDENVKLSPPRLMTLEEAIGYVASDELIEV 638
>At5g13650 GTP-binding protein typA (tyrosine phosphorylated protein
A)
Length = 609
Score = 237 bits (604), Expect = 1e-62
Identities = 136/319 (42%), Positives = 190/319 (58%), Gaps = 3/319 (0%)
Query: 103 VVTDCAGAGDIISIAGLSSPSIGHTVTTVEIMSALPTVELDPPTISMTFGVNDSPLAGRD 162
V TD AGDI ++ G+ + IG T+ LPT++++ PT+ M+F VN SP +GR+
Sbjct: 274 VPTDSVEAGDICAVCGIDNIQIGETIADKVHGKPLPTIKVEEPTVKMSFSVNTSPFSGRE 333
Query: 163 GTHLTGGKIGDRLMAESETNLAINVLPG-MSETFEVQGRGELQLGILIENMRREGFELSV 221
G ++T + DRL E E NLA+ V G ++TF V GRG L + ILIENMRREG+E V
Sbjct: 334 GKYVTSRNLRDRLNRELERNLAMKVEDGETADTFIVSGRGTLHITILIENMRREGYEFMV 393
Query: 222 SPPKVMYKTDKGQKLEPIEEVTIEVNDEHVGFVMEALSHRRAEITDMGPVAGTVGRTRLC 281
PPKV+ K + LEP E T+EV + H+G V+E L RR ++ DM V G+ G T L
Sbjct: 394 GPPKVINKRVNDKLLEPYEIATVEVPEAHMGPVVELLGKRRGQMFDMQGV-GSEGTTFLR 452
Query: 282 LTCPSRGLVGYRSVFSSETRGTGFMHRAFLKYEKFRGPLGNVRKGVLVSVGFGSITLHAL 341
P+RGL+G R+ + +RGT ++ F Y + G + G LV+ G+ T +AL
Sbjct: 453 YKIPTRGLLGLRNAILTASRGTAILNTVFDSYGPWAGDISTRDLGSLVAFEDGTSTSYAL 512
Query: 342 MSLEARGTLFVSPGMEAYDGMIVGEHSRDTDLDVNPVRAKQLTNVRSVNKDDTVKLIPPR 401
S + RG +FV G++ Y G IVG H R DL +N + K TN+RS NKD TV L P
Sbjct: 513 ASAQERGQMFVGSGVDVYKGQIVGIHQRPGDLGLNICKKKAATNIRS-NKDVTVILDTPL 571
Query: 402 LMTLEEAIGYVASDELIEV 420
+L++ I Y+ DEL+EV
Sbjct: 572 TYSLDDCIEYIEEDELVEV 590
>At1g62750 elongation factor G, putative
Length = 783
Score = 43.1 bits (100), Expect = 3e-04
Identities = 36/148 (24%), Positives = 69/148 (46%), Gaps = 9/148 (6%)
Query: 83 AEKIEDGKVVKVMKKKGTTMVVTDCAGAGDIISIAGLSSPSIGHTVTTVEIMSALPTVEL 142
A K + ++ ++++ + A GDII++AGL G T++ E L ++
Sbjct: 431 ANKGKKERIGRLLEMHANSREDVKVALTGDIIALAGLKDTITGETLSDPENPVVLERMDF 490
Query: 143 DPPTISMTFGVNDSPLAGRDGTHLTGGKIGDRLMAESETNLAINVLPGMSETFEVQGRGE 202
P I + P D + G I +A+ + + + M++T ++G GE
Sbjct: 491 PDPVIKVAI----EPKTKADIDKMATGLI---KLAQEDPSFHFSRDEEMNQTV-IEGMGE 542
Query: 203 LQLGILIENMRRE-GFELSVSPPKVMYK 229
L L I+++ ++RE E +V P+V Y+
Sbjct: 543 LHLEIIVDRLKREFKVEANVGAPQVNYR 570
Score = 31.2 bits (69), Expect = 1.2
Identities = 22/79 (27%), Positives = 33/79 (40%), Gaps = 2/79 (2%)
Query: 236 LEPIEEVTIEVNDEHVGFVMEALSHRRAEITDMGPVAGTVGRTRLCLTCPSRGLVGYRSV 295
LEPI V + +EH+G V+ L+ RR +I G G G + P + Y S
Sbjct: 688 LEPIMRVEVVTPEEHLGDVIGDLNSRRGQINSFGDKPG--GLKVVDSLVPLAEMFQYVST 745
Query: 296 FSSETRGTGFMHRAFLKYE 314
T+G K++
Sbjct: 746 LRGMTKGRASYTMQLAKFD 764
>At2g45030 putative mitochondrial translation elongation factor G
Length = 754
Score = 42.4 bits (98), Expect = 5e-04
Identities = 38/149 (25%), Positives = 68/149 (45%), Gaps = 10/149 (6%)
Query: 90 KVVKVMKKKGTTMVVTDCAGAGDIISIAGLSSPSIGHTVTTVEIMSALPTVELDPPTISM 149
KV ++++ M A AG I+++ G+ S G T T + + ++ + P +S+
Sbjct: 407 KVPRLVRMHSNDMEDIQEAHAGQIVAVFGIECAS-GDTFTDGSVKYTMTSMNVPEPVMSL 465
Query: 150 TFGVNDSPLAGRDGTHLTGGKIGDRLMAESETNLAINVLPGMSETFEVQGRGELQLGILI 209
P++ G + K +R E T + + P +T + G GEL L I +
Sbjct: 466 AV----QPVSKDSGGQFS--KALNRFQKEDPT-FRVGLDPESGQTI-ISGMGELHLDIYV 517
Query: 210 ENMRRE-GFELSVSPPKVMYKTDKGQKLE 237
E MRRE + +V P+V ++ Q+ E
Sbjct: 518 ERMRREYKVDATVGKPRVNFRETITQRAE 546
>At1g45332 mitochondrial elongation factor, putative
Length = 754
Score = 42.4 bits (98), Expect = 5e-04
Identities = 38/149 (25%), Positives = 68/149 (45%), Gaps = 10/149 (6%)
Query: 90 KVVKVMKKKGTTMVVTDCAGAGDIISIAGLSSPSIGHTVTTVEIMSALPTVELDPPTISM 149
KV ++++ M A AG I+++ G+ S G T T + + ++ + P +S+
Sbjct: 407 KVPRLVRMHSNDMEDIQEAHAGQIVAVFGIECAS-GDTFTDGSVKYTMTSMNVPEPVMSL 465
Query: 150 TFGVNDSPLAGRDGTHLTGGKIGDRLMAESETNLAINVLPGMSETFEVQGRGELQLGILI 209
P++ G + K +R E T + + P +T + G GEL L I +
Sbjct: 466 AV----QPVSKDSGGQFS--KALNRFQKEDPT-FRVGLDPESGQTI-ISGMGELHLDIYV 517
Query: 210 ENMRRE-GFELSVSPPKVMYKTDKGQKLE 237
E MRRE + +V P+V ++ Q+ E
Sbjct: 518 ERMRREYKVDATVGKPRVNFRETITQRAE 546
>At5g08650 GTP-binding protein LepA homolog
Length = 675
Score = 32.7 bits (73), Expect = 0.40
Identities = 32/137 (23%), Positives = 55/137 (39%), Gaps = 24/137 (17%)
Query: 201 GELQLGILIENMRRE-GFELSVSPPKVMYKT-----------------DKGQKL---EPI 239
G L + I+ E + RE L + P V+Y+ D GQ+ EP
Sbjct: 419 GLLHMEIVQERLEREYNLNLITTAPSVVYRVNSVNGDTTLCSNPSRLPDPGQRKSVEEPY 478
Query: 240 EEVTIEVNDEHVGFVMEALSHRRAEITDMGPVAGTVGRTRLCLTCPSRGLVG-YRSVFSS 298
++ + +++G +ME RR E +M +A R + P +VG + S
Sbjct: 479 VKIELLTPKDYIGALMELAQERRGEFKEMKYIA--ENRASILYELPLAEMVGDFFDQLKS 536
Query: 299 ETRGTGFMHRAFLKYEK 315
T+G M + + Y +
Sbjct: 537 RTKGYASMEYSVIGYRE 553
>At2g47860 unknown protein
Length = 635
Score = 32.0 bits (71), Expect = 0.68
Identities = 25/81 (30%), Positives = 41/81 (49%), Gaps = 11/81 (13%)
Query: 121 SPSIGHTVT-TVEIMSALPTVELDPPTISMTFGVNDSPLAGRDGTHLTGGKIGDRLMAES 179
S SI HT V +++ T+E+ S TF ++ PL R G +I ++
Sbjct: 23 SSSIRHTPQWPVSDVTSDLTIEVG----SATFSLHKFPLVSRSG------RIRKLVLESK 72
Query: 180 ETNLAINVLPGMSETFEVQGR 200
+TNL + +PG SE+FE+ +
Sbjct: 73 DTNLNLAAVPGGSESFELAAK 93
>At4g19450 unknown protein
Length = 572
Score = 31.2 bits (69), Expect = 1.2
Identities = 27/107 (25%), Positives = 54/107 (50%), Gaps = 15/107 (14%)
Query: 326 GVLVSVGFGSITLHALMSLEARGTLFVSPG-MEAYDGMIVGEHSRD------TDLDVNPV 378
G++++ + + T+H LE G + V P +E + GM+ E +R+ D+ NPV
Sbjct: 253 GLVIARNWYNRTIHTSFRLEGSGFILVDPDELELHKGMLAHEANREGYQLLSDDVVQNPV 312
Query: 379 RAKQLTNVRSVNKDDTVKLIPPRLMTLE--EAIGYVASDELIEVRSN 423
++ +V ++D+ + +L+T + E +G S L+ RS+
Sbjct: 313 KSV------AVEEEDSDESCCKKLITRDQLEGLGIEHSLSLLLTRSD 353
>At5g28200 putative protein
Length = 528
Score = 30.0 bits (66), Expect = 2.6
Identities = 19/53 (35%), Positives = 27/53 (50%), Gaps = 3/53 (5%)
Query: 344 LEARGTLFVSPGME--AYDGMIVGEHSRDTDLDVN-PVRAKQLTNVRSVNKDD 393
LE +G +S + A + I GEHS+D D +N P +NV + N DD
Sbjct: 306 LETQGKTMISTLTDWFAKNSFIEGEHSKDPDCGINRPKEPVPASNVGASNSDD 358
>At3g15910 hypothetical protein
Length = 307
Score = 29.6 bits (65), Expect = 3.4
Identities = 13/46 (28%), Positives = 20/46 (43%)
Query: 6 WCLIYLRTWVLQRNNLTSLFFMLLLKKDGHPPHIPRILLPMPKICH 51
W Y+R + L+ S + + K P RI+ P P +CH
Sbjct: 182 WIQDYMRQYNLETTKQMSFLMAVFMDKRAPPETRKRIIFPHPLLCH 227
>At4g24940 ubiquitin activating enzyme like protein
Length = 322
Score = 28.5 bits (62), Expect = 7.5
Identities = 22/74 (29%), Positives = 36/74 (47%), Gaps = 4/74 (5%)
Query: 308 RAFLKYEKFRGPLGNVRKGVLVSVGFGSITL--HALMSLEARGTLFVSPGME-AYDGMIV 364
+A + +G + K ++++ G GS+TL L ++EA F+ P E Y G V
Sbjct: 31 KAHILVSGIKGTVAEFCKNIVLA-GVGSVTLMDDRLANMEALNANFLIPPDENVYSGKTV 89
Query: 365 GEHSRDTDLDVNPV 378
E D+ D NP+
Sbjct: 90 AEICSDSLKDFNPM 103
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.138 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,602,650
Number of Sequences: 26719
Number of extensions: 410113
Number of successful extensions: 865
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 855
Number of HSP's gapped (non-prelim): 13
length of query: 433
length of database: 11,318,596
effective HSP length: 102
effective length of query: 331
effective length of database: 8,593,258
effective search space: 2844368398
effective search space used: 2844368398
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC140034.13


