

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140031.2 + phase: 0 /pseudo
(484 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
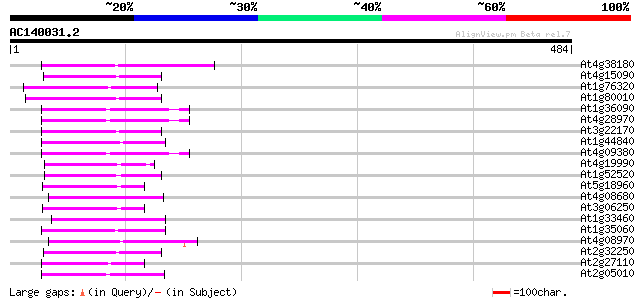
Score E
Sequences producing significant alignments: (bits) Value
At4g38180 unknown protein (At4g38180) 59 8e-09
At4g15090 unknown protein 57 3e-08
At1g76320 putative phytochrome A signaling protein 52 1e-06
At1g80010 hypothetical protein 50 2e-06
At1g36090 hypothetical protein 50 2e-06
At4g28970 putative protein 50 3e-06
At3g22170 far-red impaired response protein, putative 49 6e-06
At1g44840 hypothetical protein 49 8e-06
At4g09380 putative protein 47 2e-05
At4g19990 putative protein 47 2e-05
At1g52520 F6D8.26 46 4e-05
At5g18960 FAR1 - like protein 46 5e-05
At4g08680 putative MuDR-A-like transposon protein 46 5e-05
At3g06250 unknown protein 44 2e-04
At1g33460 mutator transposase MUDRA, putative 44 2e-04
At1g35060 hypothetical protein 44 3e-04
At4g08970 putative transposon protein 43 3e-04
At2g32250 Mutator-like transposase 43 3e-04
At2g27110 Mutator-like transposase 43 4e-04
At2g05010 Mutator-like transposase 43 4e-04
>At4g38180 unknown protein (At4g38180)
Length = 788
Score = 58.5 bits (140), Expect = 8e-09
Identities = 35/151 (23%), Positives = 72/151 (47%), Gaps = 4/151 (2%)
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
F +F + D+TY++N Y++P GV G F+ +E E +FVW+
Sbjct: 293 FTHFGDTVTFDTTYRSNRYRLPFAPFTGVNHHGQPILFGCAFIINETEASFVWLFNTWLA 352
Query: 88 LLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVKVRCI-LDCRYPMAKK 146
+++ + P + TD D + A+ +V P + C +H+ K + + + ++P +
Sbjct: 353 AMSA--HPPVSITTDHDAVIRAAIMHVFPGARHRFCKWHILKKCQEKLSHVFLKHPSFES 410
Query: 147 D-GKEVKHGDVVKKIMRVW*VIIESHPLKNY 176
D K V + V+ R W +++ + L+++
Sbjct: 411 DFHKCVNLTESVEDFERCWFSLLDKYELRDH 441
>At4g15090 unknown protein
Length = 768
Score = 56.6 bits (135), Expect = 3e-08
Identities = 32/102 (31%), Positives = 53/102 (51%), Gaps = 2/102 (1%)
Query: 30 NFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCKLL 89
+F V+ D+TY K+P+ +GV +G V E E FVW+++ + +
Sbjct: 212 SFNDVVSFDTTYVKFNDKLPLALFIGVNHHSQPMLLGCALVADESMETFVWLIKTWLRAM 271
Query: 90 TSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNV 131
R PKV++TD+D LM+AVS +LP + +HV + +
Sbjct: 272 GGR--APKVILTDQDKFLMSAVSELLPNTRHCFALWHVLEKI 311
>At1g76320 putative phytochrome A signaling protein
Length = 670
Score = 51.6 bits (122), Expect = 1e-06
Identities = 30/115 (26%), Positives = 58/115 (50%), Gaps = 2/115 (1%)
Query: 13 LLKTYFALIQHPLSFFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTH 72
LL+ F + + + +F V+ +++Y + YK+P+ VGV +G +
Sbjct: 191 LLRNVFWVDAKGIEDYKSFSDVVSFETSYFVSKYKVPLVLFVGVNHHVQPVLLGCGLLAD 250
Query: 73 EKEENFVWVLQMMCKLLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHV 127
+ +VW++Q L+ PKV++TD++ + A++ VLPE+ C +HV
Sbjct: 251 DTVYTYVWLMQSW--LVAMGGQKPKVMLTDQNNAIKAAIAAVLPETRHCYCLWHV 303
>At1g80010 hypothetical protein
Length = 696
Score = 50.4 bits (119), Expect = 2e-06
Identities = 28/118 (23%), Positives = 56/118 (46%), Gaps = 2/118 (1%)
Query: 14 LKTYFALIQHPLSFFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHE 73
L+ F + + +++F VL+ D+T +N Y++P+ VG+ T +G + +
Sbjct: 272 LRNVFWIDARARAAYSHFGDVLLFDTTCLSNAYELPLVAFVGINHHGDTILLGCGLLADQ 331
Query: 74 KEENFVWVLQMMCKLLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNV 131
E +VW+ + + R P++ +T++ + AVS V P + HV N+
Sbjct: 332 SFETYVWLFRAWLTCMLGR--PPQIFITEQCKAMRTAVSEVFPRAHHRLSLTHVLHNI 387
>At1g36090 hypothetical protein
Length = 645
Score = 50.4 bits (119), Expect = 2e-06
Identities = 34/129 (26%), Positives = 62/129 (47%), Gaps = 11/129 (8%)
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
F V+V+D+T+ +Y + Y + F + EK+ +++W L+ K
Sbjct: 364 FRAMRKVIVVDATHLKTVYGGMLVIATAHDPNHHHYPLAFGIIDSEKDVSWIWFLE---K 420
Query: 88 LLTSRMNVPKVV-VTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVKVRCILDCRYPMAKK 146
L T +VP++V ++D+ ++ AV V P + C +H+ +N++ R +D K
Sbjct: 421 LKTVYSDVPRLVFISDRHQSIKKAVKTVYPNALHAACIWHLCQNMRDRVKID-------K 473
Query: 147 DGKEVKHGD 155
DG VK D
Sbjct: 474 DGAAVKFRD 482
>At4g28970 putative protein
Length = 914
Score = 50.1 bits (118), Expect = 3e-06
Identities = 34/129 (26%), Positives = 61/129 (46%), Gaps = 11/129 (8%)
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
F V+V+D+T+ +Y + Y + F + EK+ +++W L+ K
Sbjct: 404 FRAMRKVIVVDATHLKTVYGGMLVIATAQDPNHHHYPLAFGIIDSEKDVSWIWFLE---K 460
Query: 88 LLTSRMNVPKVV-VTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVKVRCILDCRYPMAKK 146
L T +VP +V ++D+ ++ AV V P + C +H+ +N++ R +D K
Sbjct: 461 LKTVYSDVPGLVFISDRHQSIKKAVKTVYPNALHAACIWHLCQNMRDRVTID-------K 513
Query: 147 DGKEVKHGD 155
DG VK D
Sbjct: 514 DGAAVKFRD 522
>At3g22170 far-red impaired response protein, putative
Length = 814
Score = 48.9 bits (115), Expect = 6e-06
Identities = 26/104 (25%), Positives = 50/104 (48%), Gaps = 2/104 (1%)
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
+ +F V+ +D+TY N YKMP+ VGV +G ++ E + W+++ +
Sbjct: 278 YGSFCDVVSLDTTYVRNKYKMPLAIFVGVNQHYQYMVLGCALISDESAATYSWLMETWLR 337
Query: 88 LLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNV 131
+ + PKV++T+ D + + V + P + +HV V
Sbjct: 338 AIGGQ--APKVLITELDVVMNSIVPEIFPNTRHCLFLWHVLMKV 379
>At1g44840 hypothetical protein
Length = 926
Score = 48.5 bits (114), Expect = 8e-06
Identities = 23/107 (21%), Positives = 53/107 (49%), Gaps = 2/107 (1%)
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
F + +LV+D T+ YK + G + Y +GF V E +E++ W + +
Sbjct: 568 FRHLRRILVVDGTHLKGKYKGVLLTSSGQDANFQVYPLGFAVVDSENDESWTWFFTKLER 627
Query: 88 LLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVKVR 134
++ + +++D+ ++++ AV V P++ C H+ +N++ +
Sbjct: 628 IIADSKTL--TILSDRHSSILVAVKRVFPQANHGACIIHLCRNIQTK 672
>At4g09380 putative protein
Length = 960
Score = 47.4 bits (111), Expect = 2e-05
Identities = 33/129 (25%), Positives = 60/129 (45%), Gaps = 11/129 (8%)
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
F V+V+D+T+ +Y + Y + F + EK+ +++W L+
Sbjct: 404 FRAMRKVIVVDATHLKTVYGGMLVIATAQDPNHHHYPLAFGIIDSEKDVSWIWFLE---N 460
Query: 88 LLTSRMNVPKVV-VTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVKVRCILDCRYPMAKK 146
L T +VP +V ++D+ ++ AV V P + C +H+ +N++ R +D K
Sbjct: 461 LKTVYSDVPGLVFISDRHQSIKKAVKTVYPNALHAACIWHLCQNMRDRVKID-------K 513
Query: 147 DGKEVKHGD 155
DG VK D
Sbjct: 514 DGAAVKFRD 522
>At4g19990 putative protein
Length = 672
Score = 47.0 bits (110), Expect = 2e-05
Identities = 29/96 (30%), Positives = 47/96 (48%), Gaps = 5/96 (5%)
Query: 31 FPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFE-FVTHEKEENFVWVLQMMCKLL 89
F V+ +D+T+ N YK+P+ GV +GF +T E + FVW+ + K +
Sbjct: 203 FSDVVSIDTTFIKNEYKLPLVAFTGVNHHGQFLLLGFGLLLTDESKSGFVWLFRAWLKAM 262
Query: 90 TSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYF 125
P+V++T D L AV V P S +C++
Sbjct: 263 HGCR--PRVILTKHDQMLKEAVLEVFPSS--RHCFY 294
>At1g52520 F6D8.26
Length = 703
Score = 46.2 bits (108), Expect = 4e-05
Identities = 25/101 (24%), Positives = 49/101 (47%), Gaps = 3/101 (2%)
Query: 31 FPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCKLLT 90
F V+ +DS+Y + +++P+ GV T + F+ E E++ W+L++ ++
Sbjct: 294 FGDVIFIDSSYISGKFEIPLVTFTGVNHHGKTTLLSCGFLAGETMESYHWLLKVWLSVMK 353
Query: 91 SRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNV 131
P+ +VTD+ L A+S V P S H+ + +
Sbjct: 354 RS---PQTIVTDRCKPLEAAISQVFPRSHQRFSLTHIMRKI 391
>At5g18960 FAR1 - like protein
Length = 788
Score = 45.8 bits (107), Expect = 5e-05
Identities = 23/88 (26%), Positives = 42/88 (47%), Gaps = 2/88 (2%)
Query: 29 NNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCKL 88
+ F +V D++Y+ Y +P I+G +G V E +E F+W+ Q +
Sbjct: 394 SQFGDSVVFDTSYRKGSYSVPFATIIGFNHHRQPVLLGCAMVADESKEAFLWLFQTWLRA 453
Query: 89 LTSRMNVPKVVVTDKDTTLMNAVSNVLP 116
++ R P+ +V D+D + A+ V P
Sbjct: 454 MSGRR--PRSIVADQDLPIQQALVQVFP 479
>At4g08680 putative MuDR-A-like transposon protein
Length = 761
Score = 45.8 bits (107), Expect = 5e-05
Identities = 25/99 (25%), Positives = 48/99 (48%)
Query: 34 VLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCKLLTSRM 93
++ +D T+ + K + VG + Y + + V E EN++W +Q + K L
Sbjct: 305 IIGLDGTFLKVVVKGVLLTAVGHDPNNQIYPIAWAVVQSENAENWLWFVQQIKKDLNLED 364
Query: 94 NVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVK 132
V+++D+ L++AV LP + C H+ +N+K
Sbjct: 365 GSRFVILSDRSKGLLSAVKQELPNAEHRMCVKHIVENLK 403
>At3g06250 unknown protein
Length = 764
Score = 44.3 bits (103), Expect = 2e-04
Identities = 23/88 (26%), Positives = 41/88 (46%), Gaps = 2/88 (2%)
Query: 29 NNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCKL 88
+ F +V D++Y+ Y +P +G +G V E +E F W+ Q +
Sbjct: 370 SQFGDAVVFDTSYRKGDYSVPFATFIGFNHHRQPVLLGGALVADESKEAFSWLFQTWLRA 429
Query: 89 LTSRMNVPKVVVTDKDTTLMNAVSNVLP 116
++ R P+ +V D+D + AV+ V P
Sbjct: 430 MSGRR--PRSMVADQDLPIQQAVAQVFP 455
>At1g33460 mutator transposase MUDRA, putative
Length = 826
Score = 44.3 bits (103), Expect = 2e-04
Identities = 25/98 (25%), Positives = 47/98 (47%)
Query: 37 MDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCKLLTSRMNVP 96
+D T+ + K + + + + Y V + V E EN++W L + L +
Sbjct: 374 IDGTFLKHAVKGCLLTAIAHDANNQIYPVAWATVQFENAENWLWFLNQLKHDLELKDGSG 433
Query: 97 KVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVKVR 134
VV++D+ +++AV N LP + C H+ +N+K R
Sbjct: 434 YVVISDRCKGIISAVKNALPNAEHRPCVKHIVENLKKR 471
>At1g35060 hypothetical protein
Length = 873
Score = 43.5 bits (101), Expect = 3e-04
Identities = 23/107 (21%), Positives = 48/107 (44%), Gaps = 2/107 (1%)
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
F + VLV+D T+ YK + G + Y + F V E ++ + W + +
Sbjct: 519 FKHLRRVLVVDGTHLKGKYKGVLLTASGQDANFQVYPLAFAVVDSENDDAWTWFFTKLER 578
Query: 88 LLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVKVR 134
++ N +++D+ ++ V V P++ C H+ +N++ R
Sbjct: 579 IIAD--NNTLTILSDRHESIKVGVKKVFPQAHHGACIIHLCRNIQAR 623
>At4g08970 putative transposon protein
Length = 597
Score = 43.1 bits (100), Expect = 3e-04
Identities = 29/132 (21%), Positives = 63/132 (46%), Gaps = 5/132 (3%)
Query: 34 VLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCKLLTSRM 93
V+++D T + +K + + + + + F V E E + W L+ + +L
Sbjct: 458 VVLIDGTAIKHKFKGVLLTASMQDANFMVFPIAFGIVDSESEPAWSWFLRQLTTILPDAA 517
Query: 94 NVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVKVRCILDCRYPMAKKDGK---E 150
+V V+V+D+ ++ A+ V E+F C H+++NV+++ + +K + E
Sbjct: 518 DV--VIVSDRHRSIYAAMGQVYLEAFHGACAVHIERNVRLKFPKKGVSNLVRKAARAFNE 575
Query: 151 VKHGDVVKKIMR 162
+G + K+I R
Sbjct: 576 TNYGGLYKEIER 587
>At2g32250 Mutator-like transposase
Length = 684
Score = 43.1 bits (100), Expect = 3e-04
Identities = 24/102 (23%), Positives = 47/102 (45%), Gaps = 2/102 (1%)
Query: 30 NFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCKLL 89
+F V++ D+ Y N Y++P +GV+ +G + E + W+ + K +
Sbjct: 215 SFSDVVLFDTFYVRNGYRIPFAPFIGVSHHRQYVLLGCALIGEVSESTYSWLFRTWLKAV 274
Query: 90 TSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNV 131
+ P V++TD+D L + V V P+ + C + V +
Sbjct: 275 GGQ--APGVMITDQDKLLSDIVVEVFPDVRHIFCLWSVLSKI 314
>At2g27110 Mutator-like transposase
Length = 851
Score = 42.7 bits (99), Expect = 4e-04
Identities = 21/89 (23%), Positives = 39/89 (43%), Gaps = 2/89 (2%)
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
+ +F + +D+ Y+ N +++P GV G + E + +F+W+ +
Sbjct: 240 YTHFGDTVTLDTRYRCNQFRVPFAPFTGVNHHGQAILFGCALILDESDTSFIWLFKTF-- 297
Query: 88 LLTSRMNVPKVVVTDKDTTLMNAVSNVLP 116
L R P +VTD+D + A V P
Sbjct: 298 LTAMRDQPPVSLVTDQDRAIQIAAGQVFP 326
>At2g05010 Mutator-like transposase
Length = 731
Score = 42.7 bits (99), Expect = 4e-04
Identities = 26/107 (24%), Positives = 52/107 (48%), Gaps = 4/107 (3%)
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
F V+V+D+T+ +Y + Y + F + EK+ +++W L+ K
Sbjct: 403 FREMRKVIVVDATHLKTVYGGMLVIATTQDPNHHHYPLAFGIIDSEKDVSWIWFLE---K 459
Query: 88 LLTSRMNVPKVV-VTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVKV 133
L T NVP +V ++D+ + AV V + +C +H+ +N+++
Sbjct: 460 LKTVYPNVPGLVFISDRHQNIKKAVKMVYLNALHASCIWHLSQNMRL 506
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.342 0.150 0.517
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,522,230
Number of Sequences: 26719
Number of extensions: 435255
Number of successful extensions: 1609
Number of sequences better than 10.0: 64
Number of HSP's better than 10.0 without gapping: 27
Number of HSP's successfully gapped in prelim test: 37
Number of HSP's that attempted gapping in prelim test: 1560
Number of HSP's gapped (non-prelim): 75
length of query: 484
length of database: 11,318,596
effective HSP length: 103
effective length of query: 381
effective length of database: 8,566,539
effective search space: 3263851359
effective search space used: 3263851359
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 62 (28.5 bits)
Medicago: description of AC140031.2


