

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140030.1 + phase: 0
(336 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
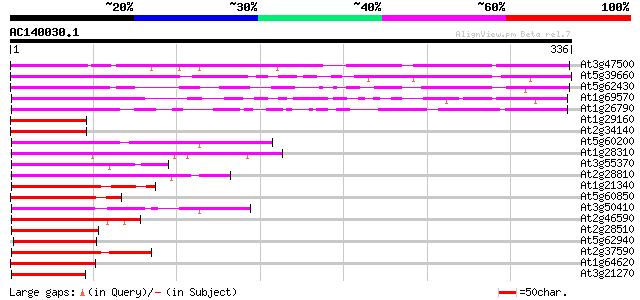
Score E
Sequences producing significant alignments: (bits) Value
At3g47500 H-protein promoter binding factor-2a 246 1e-65
At5g39660 unknown protein 213 9e-56
At5g62430 H-protein promoter binding factor-like protein 189 2e-48
At1g69570 H-protein promoter binding factor-2b, putative 145 2e-35
At1g26790 H-protein promoter binding factor-2b, putative 129 2e-30
At1g29160 ascorbate oxidase promoter-binding protein, putative 99 2e-21
At2g34140 putative DOF zinc finger protein 97 9e-21
At5g60200 zinc finger protein - like 91 7e-19
At1g28310 zinc finger like protein OBP3 91 1e-18
At3g55370 zinc finger protein OBP3 88 8e-18
At2g28810 putative DOF zinc finger protein 88 8e-18
At1g21340 DNA-binding protein, putative 87 1e-17
At5g60850 zinc finger protein OBP4 - like 87 1e-17
At3g50410 DNA binding protein 87 1e-17
At2g46590 DOF zinc finger like protein 87 1e-17
At2g28510 DOF zinc finger like protein 87 1e-17
At5g62940 Dof zinc finger protein - like 87 2e-17
At2g37590 putative DOF zinc finger protein 87 2e-17
At1g64620 zinc finger protein, putative 87 2e-17
At3g21270 Dof zinc finger protein 86 2e-17
>At3g47500 H-protein promoter binding factor-2a
Length = 448
Score = 246 bits (628), Expect = 1e-65
Identities = 156/354 (44%), Positives = 194/354 (54%), Gaps = 43/354 (12%)
Query: 1 MDTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTSHYRHLIIPE 60
M+TKFCY+NNYN+NQPRHFCK CQRYWTAGGTMRNVPVGAGRRKNK+ S+SHYRH+ I E
Sbjct: 118 METKFCYYNNYNINQPRHFCKACQRYWTAGGTMRNVPVGAGRRKNKS-SSSHYRHITISE 176
Query: 61 GAKLNSPNGIHSLGNGAAVLTFD---SNPPLCNTMASVLNIAE--RTQNCVSNGFHH--- 112
L + L VL+F + M V+ + E + N N FH
Sbjct: 177 A--LEAARLDPGLQANTRVLSFGLEAQQQHVAAPMTPVMKLQEDQKVSNGARNRFHGLAD 234
Query: 113 ---ASSYGGENDHSIGVSVTASNS-SERKSHTSTNGLVDKGVEGFPPQLQH----IPSPF 164
+ +D S G SVT SN+ S +S + +V+ + + IP
Sbjct: 235 QRLVARVENGDDCSSGSSVTTSNNHSVDESRAQSGSVVEAQMNNNNNNNMNGYACIPGVP 294
Query: 165 LPYTWNSAMPPPTFC-PPNYPLAFYTPVTPPAYWGCMPPPWNIPCISP-GSASVNESDSA 222
PYTWN AMPPP F PP YP+ FY P W IP + P S+S +
Sbjct: 295 WPYTWNPAMPPPGFYPPPGYPMPFY-------------PYWTIPMLPPHQSSSPISQKCS 341
Query: 223 HSSVPTLGKHSRDGNIITSVNSPKEKPETEHNSTESSVLIPKTLRFDDPSEAAKSSLWSK 282
+++ PTLGKH RD S K+ ETE VL+PKTLR DDP+EAAKSS+W+
Sbjct: 342 NTNSPTLGKHPRD------EGSSKKDNETERKQKAGCVLVPKTLRIDDPNEAAKSSIWTT 395
Query: 283 LGINNDKANSLNGGGMFNGFQSKGKDMNH-SVGTSPLMYANPAALSRSRTFHEE 335
LGI N+ GGMF GF K K N+ SP++ ANPAALSRS FHE+
Sbjct: 396 LGIKNEA--MCKAGGMFKGFDHKTKMYNNDKAENSPVLSANPAALSRSHNFHEQ 447
>At5g39660 unknown protein
Length = 457
Score = 213 bits (543), Expect = 9e-56
Identities = 139/351 (39%), Positives = 177/351 (49%), Gaps = 54/351 (15%)
Query: 1 MDTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTSHYRHLIIPE 60
M+TKFCY+NNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNK+ ++ + RH+ I
Sbjct: 146 METKFCYYNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKSPASHYNRHVSITS 205
Query: 61 GAKLNSPNGIH-SLGNGAAVLTFDSNPPLCNTMASVLNIAERTQNCVSNGFHHASSYGGE 119
+ NGA +LTF S+ LC +MAS LN+ E++ + E
Sbjct: 206 AEAMQKVARTDLQHPNGANLLTFGSDSVLCESMASGLNLVEKS-------LLKTQTVLQE 258
Query: 120 NDHSIGVSVTASNSSERKSHTSTNGLVDKGVEGFPPQLQHIPSPFLPYTWNSAMPPPTFC 179
+ + ++V + ++E S V FP P P PY WN
Sbjct: 259 PNEGLKITVPLNQTNEEAGTVSPL----PKVPCFPG-----PPPTWPYAWNGV---SWTI 306
Query: 180 PPNYPLAFYTPVTPPAYWGCMPPPWNIPCISPGS-------ASVNESDSAHSSVPTLGKH 232
P YP PPAYW C P +SPG+ N ++ + PTLGKH
Sbjct: 307 LPFYP--------PPAYWSC-------PGVSPGAWNSFTWMPQPNSPSGSNPNSPTLGKH 351
Query: 233 SRDGNIIT--SVNSPKEKPETEHNSTESSVLIPKTLRFDDPSEAAKSSLWSKLGINNDKA 290
SRD N + E E + E + +PKTLR DDP EAAKSS+W LGI D+
Sbjct: 352 SRDENAAEPGTAFDETESLGREKSKPERCLWVPKTLRIDDPEEAAKSSIWETLGIKKDE- 410
Query: 291 NSLNGGGMFNGFQSKGKDMN-----HSVGTSPLMYANPAALSRSRTFHEET 336
F F+S K+ + G P + ANPAALSRS FHE +
Sbjct: 411 ----NADTFGAFRSSTKEKSSLSEGRLPGRRPELQANPAALSRSANFHESS 457
>At5g62430 H-protein promoter binding factor-like protein
Length = 298
Score = 189 bits (479), Expect = 2e-48
Identities = 133/339 (39%), Positives = 161/339 (47%), Gaps = 107/339 (31%)
Query: 1 MDTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLS-TSHYRHLIIP 59
M+TKFCY+NNYNVNQPRHFCK CQRYWT+GGTMR+VP+GAGRRKNKN S TSHY H+ I
Sbjct: 62 METKFCYYNNYNVNQPRHFCKACQRYWTSGGTMRSVPIGAGRRKNKNNSPTSHYHHVTIS 121
Query: 60 EGAKLNSPNGIHSLGNGAAVLTFDSNPPLCNTMASVLNIAERTQNCVSNGFHHASSYGGE 119
E N P SLG+ Q SN F +
Sbjct: 122 E---TNGPVLSFSLGD--------------------------DQKVSSNRFGNQK----- 147
Query: 120 NDHSIGVSVTASNSSERKSHTSTNGLVDKGVEGFPPQLQHIPSPFLPYTWNSAMPPPTFC 179
+ N+ ER ++ ++NG L P PYTWN
Sbjct: 148 ------LVARIENNDERSNNNTSNG------------LNCFPGVSWPYTWNP-------- 181
Query: 180 PPNYPLAFYTPVTPPAYWGCMPPPWNIPCISPGSASVNESDSAHSSVPTLGKHSRDGNII 239
AFY P Y P W++P +S S + S TLGKHSRD
Sbjct: 182 ------AFY-----PVY-----PYWSMPVLS--------SPVSSSPTSTLGKHSRD---- 213
Query: 240 TSVNSPKEKPETEHNSTESSVLIPKTLRFDDPSEAAKSSLWSKLGINNDKANSLNGGGMF 299
E + SVL+PKTLR DDP+EAAKSS+W+ LGI N+ MF
Sbjct: 214 -------EDETVKQKQRNGSVLVPKTLRIDDPNEAAKSSIWTTLGIKNEV--------MF 258
Query: 300 NGFQSKGK---DMNHSVGTSPLMYANPAALSRSRTFHEE 335
NGF SK + TS ++ ANPAALSRS FHE+
Sbjct: 259 NGFGSKKEVKLSNKEETETSLVLCANPAALSRSINFHEQ 297
>At1g69570 H-protein promoter binding factor-2b, putative
Length = 399
Score = 145 bits (367), Expect = 2e-35
Identities = 117/341 (34%), Positives = 158/341 (46%), Gaps = 94/341 (27%)
Query: 2 DTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTSHYRHLIIPEG 61
+TKFCY+NNYNVNQPR+FC+ CQRYWTAGG+MRNVPVG+GRRKNK +S++ + E
Sbjct: 141 NTKFCYYNNYNVNQPRYFCRNCQRYWTAGGSMRNVPVGSGRRKNKGWPSSNHYLQVTSED 200
Query: 62 AKLNSPNGIHSLGNGAAVLTFDSNPPLCNTMASVLNIAERTQNCVSNGFHHASSYGGEND 121
N+ I S G+ + +T G H +
Sbjct: 201 CDNNNSGTILSFGSSESSVT-------------------------ETGKHQSGD------ 229
Query: 122 HSIGVSVTASNSSERKSHTSTNGLVDKGVEGFPPQLQHIPSPFLPYTWNSAMPPPTFCPP 181
TA S++ S +K +GF P P LP N++ P P P
Sbjct: 230 -------TAKISADSVSQE------NKSYQGFLP-----PQVMLP---NNSSPWPYQWSP 268
Query: 182 NYPLAFYTPVTPPAYWGCMPPPWNIPCISPGSASVNESDSAHSSVPTLGKHSRDGNIITS 241
P A + PV P YWGC P I P S E+ S LGK SRD
Sbjct: 269 TGPNASFYPV--PFYWGCTVP------IYPTS----ETSSC------LGKRSRD------ 304
Query: 242 VNSPKEKPETEHNSTESSV------LIPKTLRFDDPSEAAKSSLWSKLGINNDKANSLNG 295
+ E N T +++ L+ ++LR + EA+KS++WSKL +K G
Sbjct: 305 ------QTEGRINDTNTTITTTRARLVSESLRMN--IEASKSAVWSKLPTKPEK--KTQG 354
Query: 296 GGMFNGFQSKGKDMNHSV--GTSPLMYANPAALSRSRTFHE 334
+FNGF +KG S+ TS + ANPAA+SR+ F E
Sbjct: 355 FSLFNGFDTKGNSNRSSLVSETSHSLQANPAAMSRAMNFRE 395
>At1g26790 H-protein promoter binding factor-2b, putative
Length = 396
Score = 129 bits (325), Expect = 2e-30
Identities = 103/334 (30%), Positives = 149/334 (43%), Gaps = 84/334 (25%)
Query: 2 DTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKN-LSTSHYRHLIIPE 60
DTKFCY+NNYNVNQPRHFC+KCQRYWTAGG+MR VPVG+GRRKNK +S+ Y H+ +
Sbjct: 144 DTKFCYYNNYNVNQPRHFCRKCQRYWTAGGSMRIVPVGSGRRKNKGWVSSDQYLHITSED 203
Query: 61 GAKLNSPNGIHSLGNGAAVLTFDSNPPLCNTMASVLNIAERTQNCVSNGFHHASSYGGEN 120
NS + +L+F+S+ L + ER ++
Sbjct: 204 TDNYNS--------SSTKILSFESSDSL---------VTERPKH---------------- 230
Query: 121 DHSIGVSVTASNSSERKSHTSTNGLVDKGVEGFPPQLQHIPSPFLPYTWNSAMPPPTFCP 180
S V + A S+ + + GL+ PPQ + P+ PY + P
Sbjct: 231 -QSNEVKINAEPVSQEPN--NFQGLL-------PPQASPVSPPW-PYQY----------P 269
Query: 181 PNYPLAFYTPVTPPAYWGCMPPPWNIPCISPGSASVNESDSAHSSVPTLGKHSRDGNIIT 240
PN P ++ PV YWGC P W S + LGK +RD
Sbjct: 270 PN-PSFYHMPV----YWGCAIPVW----------------STLDTSTCLGKRTRDETSHE 308
Query: 241 SVNSPKEKPETEHNSTESSVLIPKTLRFDDPSEAAKSSLWSKLGINNDKANSLNGGGMFN 300
+V K E +S+L+ ++ S A + +W + + +K + N
Sbjct: 309 TVKESKNAFE------RTSLLLESQSIKNETSMATNNHVWYPVPMTREKTQEFS--FFSN 360
Query: 301 GFQSKGKDMNHSVGTSPLMYANPAALSRSRTFHE 334
G ++K + T + ANPAA++RS F E
Sbjct: 361 GAETKSSNNRFVPETYLNLQANPAAMARSMNFRE 394
>At1g29160 ascorbate oxidase promoter-binding protein, putative
Length = 175
Score = 99.4 bits (246), Expect = 2e-21
Identities = 41/46 (89%), Positives = 44/46 (95%)
Query: 1 MDTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNK 46
M+TKFCYFNNYNVNQPRHFCK CQRYWTAGG +RNVPVGAGRRK+K
Sbjct: 70 METKFCYFNNYNVNQPRHFCKGCQRYWTAGGALRNVPVGAGRRKSK 115
>At2g34140 putative DOF zinc finger protein
Length = 170
Score = 97.4 bits (241), Expect = 9e-21
Identities = 40/46 (86%), Positives = 43/46 (92%)
Query: 1 MDTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNK 46
M+TKFCYFNNYNVNQPRHFCK C RYWTAGG +RNVPVGAGRRK+K
Sbjct: 66 METKFCYFNNYNVNQPRHFCKGCHRYWTAGGALRNVPVGAGRRKSK 111
>At5g60200 zinc finger protein - like
Length = 257
Score = 91.3 bits (225), Expect = 7e-19
Identities = 56/160 (35%), Positives = 86/160 (53%), Gaps = 9/160 (5%)
Query: 2 DTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTSHYRHL-IIPE 60
+TKFCY+NNY++ QPR+FCK C+RYWT GGT+RN+PVG G RKNK ++S R L PE
Sbjct: 64 NTKFCYYNNYSLTQPRYFCKSCRRYWTKGGTLRNIPVGGGCRKNKRSTSSAARSLRTTPE 123
Query: 61 GAKLNSPNGIHSLGNGAAVLTFDSNPPLCNTMASVLNIAERTQNCVSNGFHH--ASSYGG 118
A + + + A + +N + ++A L + + GFH SS+
Sbjct: 124 PASHDG-----KVFSAAGFNGYSNNEHIDLSLAFALLNKQHPGSSSQLGFHSELGSSHQS 178
Query: 119 ENDHSIGVSVTASNSSERKSHTSTN-GLVDKGVEGFPPQL 157
+ + G S N++ + S+ G + + GFP Q+
Sbjct: 179 DMEGMFGTSQQKENATYAFGNGSSGLGDPSRVLWGFPWQM 218
>At1g28310 zinc finger like protein OBP3
Length = 311
Score = 90.5 bits (223), Expect = 1e-18
Identities = 60/186 (32%), Positives = 89/186 (47%), Gaps = 24/186 (12%)
Query: 2 DTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNL---STSHYRHLII 58
+TKFCY+NNY+++QPRHFCK C+RYWT GGT+RNVPVG RKNK + ST+
Sbjct: 36 NTKFCYYNNYSLSQPRHFCKACKRYWTRGGTLRNVPVGGSYRKNKRVKRPSTATTTTAST 95
Query: 59 PEGAKLNSPNGIHSLGNGAAVLTFDSNPPLCNTMASVLN--------IAERTQNC----- 105
+SPN H + + +++ L + M+S N ++ ++ C
Sbjct: 96 VSTTNSSSPNNPHQISHFSSMNHHPLFYGLSDHMSSCNNNLPMIPSRFSDSSKTCSSSGL 155
Query: 106 ----VSNGFHHASSYGGENDHSIGVSVTASNSSERKSHTS----TNGLVDKGVEGFPPQL 157
+S+GF S+ G H + T + S S T+ +GL + L
Sbjct: 156 ESEFLSSGFSSLSALGLGLPHQMSHDHTINGSFINNSTTNKPFLLSGLFGSSMSSSSTLL 215
Query: 158 QHIPSP 163
QH P
Sbjct: 216 QHPHKP 221
>At3g55370 zinc finger protein OBP3
Length = 323
Score = 87.8 bits (216), Expect = 8e-18
Identities = 42/105 (40%), Positives = 61/105 (58%), Gaps = 14/105 (13%)
Query: 2 DTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTSHYRHLII--- 58
+TKFCYFNNY++ QPRHFCK C+RYWT GG++RNVPVG G R+NK + +++
Sbjct: 85 NTKFCYFNNYSLTQPRHFCKTCRRYWTRGGSLRNVPVGGGFRRNKRSKSRSKSTVVVSTD 144
Query: 59 --------PEGAKLNSPNGIHSLGNGAAVLTFDSNPPLCNTMASV 95
++P+ HS G + F+SN P+ + S+
Sbjct: 145 NTTSTSSLTSRPSYSNPSKFHSYGQ---IPEFNSNLPILPPLQSL 186
>At2g28810 putative DOF zinc finger protein
Length = 320
Score = 87.8 bits (216), Expect = 8e-18
Identities = 52/139 (37%), Positives = 75/139 (53%), Gaps = 12/139 (8%)
Query: 2 DTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTSHYRHLIIPEG 61
+TKFCYFNNYN+ QPRHFCK C+RYWT GG +RNVPVG G R+NK + + + +
Sbjct: 103 NTKFCYFNNYNLTQPRHFCKACRRYWTRGGALRNVPVGGGCRRNKKGKSGNSKSSSSSQN 162
Query: 62 AKLNSPNGIHSLGNGAAV-LTFDSNPPLCNTMASV-------LNIAERTQNCVSNGFHHA 113
+ S S N + V L +S P T+ ++ LN+A N NG
Sbjct: 163 KQSTSMVNATSPTNTSNVQLQTNSQFPFLPTLQNLTQLGGIGLNLAAINGNNGGNG---- 218
Query: 114 SSYGGENDHSIGVSVTASN 132
G N++++ S+ +S+
Sbjct: 219 PVMGNNNENNLMTSLGSSS 237
>At1g21340 DNA-binding protein, putative
Length = 260
Score = 87.4 bits (215), Expect = 1e-17
Identities = 40/87 (45%), Positives = 60/87 (67%), Gaps = 11/87 (12%)
Query: 2 DTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAG-RRKNKNLSTSHYRHLIIPE 60
+TKFCY+NNY+++QPR+FCK C+RYWT GG++RN+PVG G R+++++ SH R
Sbjct: 47 NTKFCYYNNYSLSQPRYFCKGCRRYWTKGGSLRNIPVGGGCRKRSRSRQNSHKRF----- 101
Query: 61 GAKLNSPNGIHSLGNGAAVLTFDSNPP 87
G N P+G+ + +G F S+PP
Sbjct: 102 GRNENRPDGLINQDDG-----FQSSPP 123
>At5g60850 zinc finger protein OBP4 - like
Length = 307
Score = 87.0 bits (214), Expect = 1e-17
Identities = 36/67 (53%), Positives = 46/67 (67%), Gaps = 5/67 (7%)
Query: 1 MDTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTSHYRHLIIPE 60
++TKFCY+NNYN++QPRHFCK C+RYWT GG +RNVPVG G RK K T +P
Sbjct: 59 LNTKFCYYNNYNLSQPRHFCKNCRRYWTKGGVLRNVPVGGGCRKAKRSKTKQ-----VPS 113
Query: 61 GAKLNSP 67
+ + P
Sbjct: 114 SSSADKP 120
>At3g50410 DNA binding protein
Length = 253
Score = 87.0 bits (214), Expect = 1e-17
Identities = 54/145 (37%), Positives = 77/145 (52%), Gaps = 24/145 (16%)
Query: 2 DTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTSHYRHLIIPEG 61
+TKFCY+NNYN +QPRHFCK C+RYWT GGT+R+VPVG G RK+ S +
Sbjct: 39 NTKFCYYNNYNFSQPRHFCKACRRYWTHGGTLRDVPVGGGTRKSAKRSRT-------CSN 91
Query: 62 AKLNSPNGIHSLGNGAAVLTFDSNPPLCNTMASVLNIAERTQNCVSNGFHH--ASSYGGE 119
+ +S +G+ S NG + T P L Q+ +SNG H S G
Sbjct: 92 SSSSSVSGVVSNSNGVPLQT---TPVLF------------PQSSISNGVTHTVTESDGKG 136
Query: 120 NDHSIGVSVTASNSSERKSHTSTNG 144
+ S+ S T++ + + T+T+G
Sbjct: 137 SALSLCGSFTSTLLNHNAAATATHG 161
>At2g46590 DOF zinc finger like protein
Length = 357
Score = 87.0 bits (214), Expect = 1e-17
Identities = 42/84 (50%), Positives = 57/84 (67%), Gaps = 7/84 (8%)
Query: 2 DTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTSHYRHLI--IP 59
+TKFCY+NNY++ QPR+FCK C+RYWT GG++RNVPVG RKNK S+S +++ IP
Sbjct: 77 NTKFCYYNNYSLTQPRYFCKGCRRYWTEGGSLRNVPVGGSSRKNKRSSSSSSSNILQTIP 136
Query: 60 EG-AKLNSP----NGIHSLGNGAA 78
LN P N IH+ G++
Sbjct: 137 SSLPDLNPPILFSNQIHNKSKGSS 160
>At2g28510 DOF zinc finger like protein
Length = 288
Score = 87.0 bits (214), Expect = 1e-17
Identities = 34/52 (65%), Positives = 45/52 (86%)
Query: 2 DTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTSHY 53
+TKFCY+NNY+++QPR+FCK C+RYWT GGT+RNVPVG G R+NK S+S +
Sbjct: 57 NTKFCYYNNYSLSQPRYFCKSCRRYWTKGGTLRNVPVGGGCRRNKRSSSSAF 108
>At5g62940 Dof zinc finger protein - like
Length = 372
Score = 86.7 bits (213), Expect = 2e-17
Identities = 34/50 (68%), Positives = 44/50 (88%)
Query: 3 TKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTSH 52
TKFCY+NNY+++QPR+FCK C+RYWT GGT+RN+PVG G RKNK S+S+
Sbjct: 83 TKFCYYNNYSLSQPRYFCKTCRRYWTKGGTLRNIPVGGGCRKNKKPSSSN 132
>At2g37590 putative DOF zinc finger protein
Length = 330
Score = 86.7 bits (213), Expect = 2e-17
Identities = 39/84 (46%), Positives = 53/84 (62%), Gaps = 4/84 (4%)
Query: 2 DTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTSHYRHLIIPEG 61
+TKFCYFNNY++ QPRHFCK C+RYWT GG +RNVPVG G R+N+ ++ +
Sbjct: 98 NTKFCYFNNYSLTQPRHFCKTCRRYWTRGGALRNVPVGGGCRRNRRTKSNSNNN----NN 153
Query: 62 AKLNSPNGIHSLGNGAAVLTFDSN 85
+ S N S GN + + T S+
Sbjct: 154 STATSNNTSFSSGNASTISTILSS 177
>At1g64620 zinc finger protein, putative
Length = 352
Score = 86.7 bits (213), Expect = 2e-17
Identities = 33/51 (64%), Positives = 45/51 (87%)
Query: 1 MDTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTS 51
++TKFCY+NNY++ QPR+FCK C+RYWTAGG++RN+PVG G RKNK S++
Sbjct: 57 LNTKFCYYNNYSLTQPRYFCKDCRRYWTAGGSLRNIPVGGGVRKNKRSSSN 107
>At3g21270 Dof zinc finger protein
Length = 204
Score = 86.3 bits (212), Expect = 2e-17
Identities = 33/44 (75%), Positives = 39/44 (88%)
Query: 2 DTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKN 45
+TKFCY+NNYN++QPRHFCK C+RYWT GG +RNVPVG G RKN
Sbjct: 38 NTKFCYYNNYNLSQPRHFCKSCRRYWTKGGALRNVPVGGGSRKN 81
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.313 0.130 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,192,680
Number of Sequences: 26719
Number of extensions: 463978
Number of successful extensions: 2501
Number of sequences better than 10.0: 159
Number of HSP's better than 10.0 without gapping: 69
Number of HSP's successfully gapped in prelim test: 91
Number of HSP's that attempted gapping in prelim test: 1188
Number of HSP's gapped (non-prelim): 758
length of query: 336
length of database: 11,318,596
effective HSP length: 100
effective length of query: 236
effective length of database: 8,646,696
effective search space: 2040620256
effective search space used: 2040620256
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC140030.1


