

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139853.10 + phase: 0
(342 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
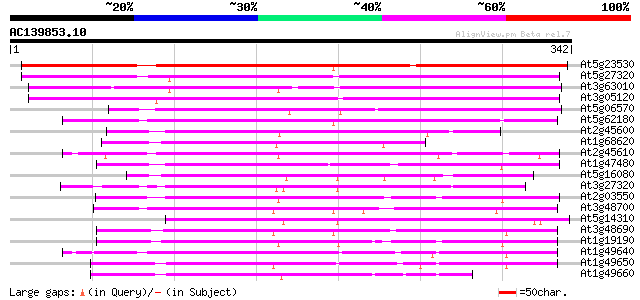
Score E
Sequences producing significant alignments: (bits) Value
At5g23530 unknown protein 381 e-106
At5g27320 Unknown protein 245 3e-65
At3g63010 unknown protein 229 2e-60
At3g05120 unknown protein 228 4e-60
At5g06570 unknown protein 176 1e-44
At5g62180 putative protein 164 5e-41
At2g45600 unknown protein 145 3e-35
At1g68620 putative carboxylesterase 142 3e-34
At2g45610 unknown protein 139 2e-33
At1g47480 unknown protein 139 2e-33
At5g16080 unknown protein 129 3e-30
At3g27320 esterase, putative 122 3e-28
At2g03550 putative esterase 121 6e-28
At3g48700 unknown protein 120 1e-27
At5g14310 unknown protein 116 2e-26
At3g48690 unknown protein 116 2e-26
At1g19190 unknown protein 114 6e-26
At1g49640 hypothetical protein 105 4e-23
At1g49650 unknown protein 100 1e-21
At1g49660 unknown protein 96 4e-20
>At5g23530 unknown protein
Length = 335
Score = 381 bits (979), Expect = e-106
Identities = 182/337 (54%), Positives = 237/337 (70%), Gaps = 18/337 (5%)
Query: 8 KLPFPLKTRLSISLIMTLTDAACRSNGSVNRRLLNFLDNKTSAKATPINGVSTKDITVDA 67
KL PLKTR+++++I T+TD A R +G++NRR L D + P+N VST D VD
Sbjct: 10 KLTLPLKTRIALTVISTMTDNAQRPDGTINRRFLRLFDFRAPPNPKPVNIVSTSDFVVDQ 69
Query: 68 ESKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTFMSPASLSYDTICRRFSRE 127
+WFRL+TP + +PVV+FFHGGGF F+SP + YD +CRRF+R+
Sbjct: 70 SRDLWFRLYTP-----------HVSGDKIPVVVFFHGGGFAFLSPNAYPYDNVCRRFARK 118
Query: 128 LNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDENK-SVLPENVDVSKCFLAGDSAGANL 186
L V+SVNYR PE+RYP QY+DG ALK+++EN S+LP N D+S+CF AGDSAG N+
Sbjct: 119 LPAYVISVNYRLAPEHRYPAQYDDGFDALKYIEENHGSILPANADLSRCFFAGDSAGGNI 178
Query: 187 AHHVAVRACK---AGLQRIRVAGLISMQPFFGGEERTEAEIRLEGSLMISMARTDWMWKV 243
AH+VA+R C+ + +++ GLIS+QPFFGGEERTEAE +L G+ ++S RTDW WK
Sbjct: 179 AHNVAIRICREPRSSFTAVKLIGLISIQPFFGGEERTEAEKQLVGAPLVSPDRTDWCWKA 238
Query: 244 FLPEGSNRDHNAANVSGPNAEDLSRLDYPDTLVFVGGLDGLYDWQKRYYEWLKISGKKAQ 303
G NRDH A NV GPNA D+S LDYP+T+V V G D L DWQ+ YYEWLK+ GKKA
Sbjct: 239 M---GLNRDHEAVNVGGPNAVDISGLDYPETMVVVAGFDPLKDWQRSYYEWLKLCGKKAT 295
Query: 304 LIEYPNMMHGFYAFPNVPEASQLILQIKDFINNRVSN 340
LIEYPNM H FY FP +PEA QLI++IKDF++ RV++
Sbjct: 296 LIEYPNMFHAFYIFPELPEAGQLIMRIKDFVDERVAS 332
>At5g27320 Unknown protein
Length = 344
Score = 245 bits (625), Expect = 3e-65
Identities = 134/337 (39%), Positives = 190/337 (55%), Gaps = 18/337 (5%)
Query: 8 KLPFPLKTRLSISLIMTLTDAACRSNGSVNRRLLNFLDNKTSAKATPINGVSTKDITVDA 67
K PL T + IS + R +G+ NR L FLD K A A P+NGV + D+ +D
Sbjct: 13 KTVVPLNTWVLISNFKLAYNLLRRPDGTFNRHLAEFLDRKVPANANPVNGVFSFDVIIDR 72
Query: 68 ESKIWFRLFTPTGINASAGGGSNTETTSL---------PVVIFFHGGGFTFMSPASLSYD 118
++ + R++ P A G++ T L PV++FFHGG F S S YD
Sbjct: 73 QTNLLSRVYRP------ADAGTSPSITDLQNPVDGEIVPVIVFFHGGSFAHSSANSAIYD 126
Query: 119 TICRRFSRELNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDENKSVLPENVDVSKCFLA 178
T+CRR VVVSVNYRR PE RYP Y+DG LK+++ + + + + FLA
Sbjct: 127 TLCRRLVGLCGAVVVSVNYRRAPENRYPCAYDDGWAVLKWVNSSSWLRSKKDSKVRIFLA 186
Query: 179 GDSAGANLAHHVAVRACKAGLQRIRVAGLISMQPFFGGEERTEAEIRLEGSLMISMARTD 238
GDS+G N+ H+VAVRA ++ RI V G I + P FGG ERTE+E RL+G +++ D
Sbjct: 187 GDSSGGNIVHNVAVRAVES---RIDVLGNILLNPMFGGTERTESEKRLDGKYFVTVRDRD 243
Query: 239 WMWKVFLPEGSNRDHNAANVSGPNAEDLSRLDYPDTLVFVGGLDGLYDWQKRYYEWLKIS 298
W W+ FLPEG +R+H A + GP ++ L L +P +LV V GLD + DWQ +Y E LK +
Sbjct: 244 WYWRAFLPEGEDREHPACSPFGPRSKSLEGLSFPKSLVVVAGLDLIQDWQLKYAEGLKKA 303
Query: 299 GKKAQLIEYPNMMHGFYAFPNVPEASQLILQIKDFIN 335
G++ +L+ GFY PN ++ +I F+N
Sbjct: 304 GQEVKLLYLEQATIGFYLLPNNNHFHTVMDEIAAFVN 340
>At3g63010 unknown protein
Length = 358
Score = 229 bits (583), Expect = 2e-60
Identities = 133/334 (39%), Positives = 187/334 (55%), Gaps = 16/334 (4%)
Query: 12 PLKTRLSISLIMTLTDAACRSNGSVNRRLLNFLDNKTSAKATPINGVSTKDITVDAESKI 71
PL T + IS R +GS NR L FLD K A + P++GV + D VD+ + +
Sbjct: 17 PLNTWVLISNFKLAYKVLRRPDGSFNRDLAEFLDRKVPANSFPLDGVFSFD-HVDSTTNL 75
Query: 72 WFRLFTPTGINASAGGGSNTETTSL------PVVIFFHGGGFTFMSPASLSYDTICRRFS 125
R++ P + G+ T L PV+IFFHGG FT S S YDT CRR
Sbjct: 76 LTRIYQPASLLHQTRHGTLELTKPLSTTEIVPVLIFFHGGSFTHSSANSAIYDTFCRRLV 135
Query: 126 RELNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDEN---KSVLPENVDVSKCFLAGDSA 182
VVVVSV+YRR+PE+RYP Y+DG AL ++ +S NV V +LAGDS+
Sbjct: 136 TICGVVVVSVDYRRSPEHRYPCAYDDGWNALNWVKSRVWLQSGKDSNVYV---YLAGDSS 192
Query: 183 GANLAHHVAVRACKAGLQRIRVAGLISMQPFFGGEERTEAEIRLEGSLMISMARTDWMWK 242
G N+AH+VAVRA G ++V G I + P FGG+ERT++E L+G +++ DW W+
Sbjct: 193 GGNIAHNVAVRATNEG---VKVLGNILLHPMFGGQERTQSEKTLDGKYFVTIQDRDWYWR 249
Query: 243 VFLPEGSNRDHNAANVSGPNAEDLSRLDYPDTLVFVGGLDGLYDWQKRYYEWLKISGKKA 302
+LPEG +RDH A N GP + L +++P +LV V GLD + DWQ Y + LK +G +
Sbjct: 250 AYLPEGEDRDHPACNPFGPRGQSLKGVNFPKSLVVVAGLDLVQDWQLAYVDGLKKTGLEV 309
Query: 303 QLIEYPNMMHGFYAFPNVPEASQLILQIKDFINN 336
L+ GFY PN L+ ++ F+++
Sbjct: 310 NLLYLKQATIGFYFLPNNDHFHCLMEELNKFVHS 343
>At3g05120 unknown protein
Length = 345
Score = 228 bits (581), Expect = 4e-60
Identities = 124/329 (37%), Positives = 179/329 (53%), Gaps = 8/329 (2%)
Query: 12 PLKTRLSISLIMTLTDAACRSNGSVNRRLLNFLDNKTSAKATPINGVSTKDITVDAESKI 71
PL T + IS + R +G+ NR L +LD K +A A P++GV + D+ +D +
Sbjct: 17 PLNTWVLISNFKVAYNILRRPDGTFNRHLAEYLDRKVTANANPVDGVFSFDVLIDRRINL 76
Query: 72 WFRLFTPTGINASAGGG-----SNTETTSLPVVIFFHGGGFTFMSPASLSYDTICRRFSR 126
R++ P + + +PV++FFHGG F S S YDT+CRR
Sbjct: 77 LSRVYRPAYADQEQPPSILDLEKPVDGDIVPVILFFHGGSFAHSSANSAIYDTLCRRLVG 136
Query: 127 ELNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDENKSVLPENVDVSKCFLAGDSAGANL 186
VVVSVNYRR PE YP Y+DG AL +++ + + FLAGDS+G N+
Sbjct: 137 LCKCVVVSVNYRRAPENPYPCAYDDGWIALNWVNSRSWLKSKKDSKVHIFLAGDSSGGNI 196
Query: 187 AHHVAVRACKAGLQRIRVAGLISMQPFFGGEERTEAEIRLEGSLMISMARTDWMWKVFLP 246
AH+VA+RA ++G+ V G I + P FGG ERTE+E L+G +++ DW WK FLP
Sbjct: 197 AHNVALRAGESGID---VLGNILLNPMFGGNERTESEKSLDGKYFVTVRDRDWYWKAFLP 253
Query: 247 EGSNRDHNAANVSGPNAEDLSRLDYPDTLVFVGGLDGLYDWQKRYYEWLKISGKKAQLIE 306
EG +R+H A N P + L + +P +LV V GLD + DWQ Y E LK +G++ +L+
Sbjct: 254 EGEDREHPACNPFSPRGKSLEGVSFPKSLVVVAGLDLIRDWQLAYAEGLKKAGQEVKLMH 313
Query: 307 YPNMMHGFYAFPNVPEASQLILQIKDFIN 335
GFY PN ++ +I F+N
Sbjct: 314 LEKATVGFYLLPNNNHFHNVMDEISAFVN 342
>At5g06570 unknown protein
Length = 329
Score = 176 bits (447), Expect = 1e-44
Identities = 107/288 (37%), Positives = 152/288 (52%), Gaps = 22/288 (7%)
Query: 61 KDITVDAESKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTFMSPASLSYDTI 120
KD + + RL+ P S + T+LPVV+FFHGGGF F S + +
Sbjct: 50 KDSIYHKPNNLHLRLYKPI---------SASNRTALPVVVFFHGGGFCFGSRSWPHFHNF 100
Query: 121 CRRFSRELNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDENKSVLPEN--------VDV 172
C + LN +VVS +YR PE+R P +ED E L +L + N VD
Sbjct: 101 CLTLASSLNALVVSPDYRLAPEHRLPAAFEDAEAVLTWLWDQAVSDGVNHWFEDGTDVDF 160
Query: 173 SKCFLAGDSAGANLAHHVAVRACKAGLQ--RIRVAGLISMQPFFGGEERTEAEIRLEGSL 230
+ F+ GDS+G N+AH +AVR ++ +RV G + M PFFGGEERT +E
Sbjct: 161 DRVFVVGDSSGGNIAHQLAVRFGSGSIELTPVRVRGYVLMGPFFGGEERTNSE-NGPSEA 219
Query: 231 MISMARTDWMWKVFLPEGSNRDHNAANVSGPNAEDLSRLDYPDTLVFVGGLDGLYDWQKR 290
++S+ D W++ LP G+ RDH+ AN GP + L + LV VGG + L D K
Sbjct: 220 LLSLDLLDKFWRLSLPNGATRDHHMANPFGPTSPTLESISLEPMLVIVGGSELLRDRAKE 279
Query: 291 Y-YEWLKISGKKAQLIEYPNMMHGFYA-FPNVPEASQLILQIKDFINN 336
Y Y+ K+ GK+ IE+ N HGFY+ +P+ A Q++ I DF+NN
Sbjct: 280 YAYKLKKMGGKRVDYIEFENKEHGFYSNYPSSEAAEQVLRIIGDFMNN 327
>At5g62180 putative protein
Length = 327
Score = 164 bits (416), Expect = 5e-41
Identities = 107/309 (34%), Positives = 164/309 (52%), Gaps = 13/309 (4%)
Query: 33 NGSVNRRLLNFLDNKTSAKATPINGVSTKDITVDAESKIWFRLFTPTGINASAGGGSNTE 92
+GS+ R L NF + +P+N +KD+ V+ W RL+ P+ SA N
Sbjct: 21 DGSITRDLSNFPCTAATPDPSPLNPAVSKDLPVNQLKSTWLRLYLPS----SAVNEGNVS 76
Query: 93 TTSLPVVIFFHGGGFTFMSPASLSYDTICRRFSRELNVVVVSVNYRRTPEYRYPTQYEDG 152
+ LP+V+++HGGGF S + C +R+LN +VVS +YR PE+R P Y+DG
Sbjct: 77 SQKLPIVVYYHGGGFILCSVDMQLFHDFCSEVARDLNAIVVSPSYRLAPEHRLPAAYDDG 136
Query: 153 ETALKFL-DENKSVLPENVDVSKCFLAGDSAGANLAHHVAVRACK--AGLQRIRVAGLIS 209
AL ++ + + + D S FL G SAG NLA++V +R+ + L +++ GLI
Sbjct: 137 VEALDWIKTSDDEWIKSHADFSNVFLMGTSAGGNLAYNVGLRSVDSVSDLSPLQIRGLIL 196
Query: 210 MQPFFGGEERTEAEIRLEGSLMISMARTDWMWKVFLPEGSNRDHNAANVS-GPNAEDLSR 268
PFFGGEER+E+EIRL + TD MW + LP G +RDH +N + G +E L +
Sbjct: 197 HHPFFGGEERSESEIRLMNDQVCPPIVTDVMWDLSLPVGVDRDHEYSNPTVGDGSEKLEK 256
Query: 269 LD-YPDTLVFVGGLDG-LYDWQKRYYEWLKISGKKAQLIEYPNMMHGFYAFPNVP-EASQ 325
+ ++ +GG D + D QK + +K G +++E+ H A P +
Sbjct: 257 IGRLRWKVMMIGGEDDPMIDLQKDVAKLMKKKG--VEVVEHYTGGHVHGAEIRDPSKRKT 314
Query: 326 LILQIKDFI 334
L L IK+FI
Sbjct: 315 LFLSIKNFI 323
>At2g45600 unknown protein
Length = 329
Score = 145 bits (366), Expect = 3e-35
Identities = 85/254 (33%), Positives = 133/254 (51%), Gaps = 25/254 (9%)
Query: 60 TKDITVDAESKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTFMSPASLSYDT 119
+KDI ++ + + R+F P I + LP++++FHGGGF S AS +
Sbjct: 39 SKDIPLNQTNNTFIRIFKPRNIPPES---------KLPILVYFHGGGFILYSAASAPFHE 89
Query: 120 ICRRFSRELNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDENK----------SVLPEN 169
C + + L +++SV YR PE+R P YED A+ +L + + L +
Sbjct: 90 SCTKMADRLQTIILSVEYRLAPEHRLPAAYEDAVEAILWLRDQARGPINGGDCDTWLKDG 149
Query: 170 VDVSKCFLAGDSAGANLAHHVAVRACKAGLQRIRVAGLISMQPFFGGEERTEAEIRLEGS 229
VD SKC++ G S+G N+ ++VA+R L +++ GLI Q FFGG E +++E RL+
Sbjct: 150 VDFSKCYVMGSSSGGNIVYNVALRVVDTDLSPVKIQGLIMNQAFFGGVEPSDSESRLKDD 209
Query: 230 LMISMARTDWMWKVFLPEGSNRDH---NAANVSGPNAED-LSRLDYPDTLVFVGGLDGLY 285
+ + T +W + LP+G +RDH N SGP +D + R +P TL+ G D L
Sbjct: 210 KICPLPATHLLWSLCLPDGVDRDHVYSNPIKSSGPQEKDKMGR--FPSTLINGYGGDPLV 267
Query: 286 DWQKRYYEWLKISG 299
D Q+ E LK G
Sbjct: 268 DRQRHVAEMLKGRG 281
>At1g68620 putative carboxylesterase
Length = 336
Score = 142 bits (357), Expect = 3e-34
Identities = 74/201 (36%), Positives = 116/201 (56%), Gaps = 11/201 (5%)
Query: 57 GVSTKDITVDAESKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTFMSPASLS 116
GV+ D+ +D + +W RL+ P S+ + LP++++FHGGGF S + L
Sbjct: 57 GVTCSDVVIDKLTNVWARLYVPMTTTKSS-------VSKLPLIVYFHGGGFCVGSASWLC 109
Query: 117 YDTICRRFSRELNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDE--NKSVLPENVDVSK 174
Y R S +V+SVNYR PE P YEDG A+ +L++ N ++ + D +
Sbjct: 110 YHEFLARLSARSRCLVMSVNYRLAPENPLPAAYEDGVNAILWLNKARNDNLWAKQCDFGR 169
Query: 175 CFLAGDSAGANLAHHVAVRACKAGLQRIRVAGLISMQPFFGGEERTEAEIRL--EGSLMI 232
FLAGDSAG N+A VA R +++ G I +QPF+ GEERTE+E R+ + + ++
Sbjct: 170 IFLAGDSAGGNIAQQVAARLASPEDLALKIEGTILIQPFYSGEERTESERRVGNDKTAVL 229
Query: 233 SMARTDWMWKVFLPEGSNRDH 253
++A +D W++ LP G+NR+H
Sbjct: 230 TLASSDAWWRMSLPRGANREH 250
>At2g45610 unknown protein
Length = 324
Score = 139 bits (350), Expect = 2e-33
Identities = 97/316 (30%), Positives = 160/316 (49%), Gaps = 26/316 (8%)
Query: 33 NGSVNRRLLNFLDNKTSAKATPING--VSTKDITVDAESKIWFRLFTPTGINASAGGGSN 90
NGS R +F+ + P G ++KD+T++ E+ + R+F PT + ++ +
Sbjct: 22 NGSCTR---HFVWPRVEPDPDPCPGKLAASKDVTINHETGVSVRIFRPTNLPSN-----D 73
Query: 91 TETTSLPVVIFFHGGGFTFMSPASLSYDTICRRFSRELNVVVVSVNYRRTPEYRYPTQYE 150
LP++I HG G+ S + D C + + EL V+VVSV+YR PE+R P QY+
Sbjct: 74 NAVARLPIIIHLHGSGWILYPANSAANDRCCSQMASELTVIVVSVHYRLPPEHRLPAQYD 133
Query: 151 DGETALKFLDEN-------KSVLPENVDVSKCFLAGDSAGANLAHHVAVRACKAGLQRIR 203
D AL ++ + + L + D S+C++ G S GAN+A +A+R+ L ++
Sbjct: 134 DALDALLWVKQQVVDSTNGEPWLKDYADFSRCYICGSSNGANIAFQLALRSLDHDLTPLQ 193
Query: 204 VAGLISMQPFFGGEERTEAEIRLEGSLMISMARTDWMWKVFLPEGSNRDHNAANVSG--P 261
+ G + QP FGG+ RT++E++ ++ + D MW++ LP G +RDH N G P
Sbjct: 194 IDGCVFYQPLFGGKTRTKSELKNFADPVMPVPAVDAMWELSLPVGVDRDHRYCNPLGYLP 253
Query: 262 NAEDLSRLDYPDTLVFVGGLDGLYDWQKRYYEWLKISGKKAQLIEYPNMMHGFYAFPNVP 321
E + RL LV G D D Q+ + L +G + +E GF++ V
Sbjct: 254 QKEKVGRLG--RCLVIGYGGDTSLDRQQDFVNLLVAAGVR---VEARFDDAGFHSIELVD 308
Query: 322 --EASQLILQIKDFIN 335
A L+ I+DFI+
Sbjct: 309 PRRAVALLNMIRDFIS 324
>At1g47480 unknown protein
Length = 314
Score = 139 bits (350), Expect = 2e-33
Identities = 93/286 (32%), Positives = 142/286 (49%), Gaps = 18/286 (6%)
Query: 54 PINGVSTKDITVDAESKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTFMSPA 113
PI GV +KDI ++ ++ + R++ P I +P++++FHGG F S +
Sbjct: 39 PITGVFSKDIIIEPKTGLSARIYRPFSIQPGQ---------KIPLMLYFHGGAFLISSTS 89
Query: 114 SLSYDTICRRFSRELNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDE-NKSVLPENVDV 172
SY T + + NV+ VSVNYR PE+ PT YED TALK + N+ + + D+
Sbjct: 90 FPSYHTSLNKIVNQANVIAVSVNYRLAPEHPLPTAYEDSWTALKNIQAINEPWINDYADL 149
Query: 173 SKCFLAGDSAGANLAHHVAVRACKAGLQRIRVAGLISMQPFFGGEERTEAEIRLEGSLMI 232
FL GDSAGAN++HH+A RA K Q +++ G+ + P+F G + AEI+ E +
Sbjct: 150 DSLFLVGDSAGANISHHLAFRA-KQSDQTLKIKGIGMIHPYFWGTQPIGAEIKDEARKQM 208
Query: 233 SMARTDWMWKVFLPEGSNRDHNAANVSGPNAEDLSRLDYPDTLVFVGGLDGLYDWQKRYY 292
D W+ P D N + DL L ++ V D L + K YY
Sbjct: 209 ----VDGWWEFVCPSEKGSDDPWINPFADGSPDLGGLGCERVMITVAEKDILNERGKMYY 264
Query: 293 EWLKIS--GKKAQLIEYPNMMHGFYAF-PNVPEASQLILQIKDFIN 335
E L S K +++E H F+ F P+ EA +++ + FIN
Sbjct: 265 ERLVKSEWKGKVEIMETKEKDHVFHIFEPDCDEAMEMVRCLALFIN 310
>At5g16080 unknown protein
Length = 344
Score = 129 bits (323), Expect = 3e-30
Identities = 83/261 (31%), Positives = 133/261 (50%), Gaps = 26/261 (9%)
Query: 72 WFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTFMSPASLSYDTICRRFSRELNVV 131
W R++ P AS + +LP++++FHGGGF S A Y + + V
Sbjct: 75 WTRVYIPDAAAASP-------SVTLPLLVYFHGGGFCVGSAAWSCYHDFLTSLAVKARCV 127
Query: 132 VVSVNYRRTPEYRYPTQYEDGETALKFLDENKSVLP-------ENVDVSKCFLAGDSAGA 184
+VSVNYR PE+R P Y+DG + +L + + ++S FLAGDSAGA
Sbjct: 128 IVSVNYRLAPEHRLPAAYDDGVNVVSWLVKQQISTGGGYPSWLSKCNLSNVFLAGDSAGA 187
Query: 185 NLAHHVAVRACKAG--LQRIRVAGLISMQPFFGGEERTEAEIRLE--GSLMISMARTDWM 240
N+A+ VAVR +G + + G+I + PFFGGE RT +E + S ++++ +D
Sbjct: 188 NIAYQVAVRIMASGKYANTLHLKGIILIHPFFGGESRTSSEKQQHHTKSSALTLSASDAY 247
Query: 241 WKVFLPEGSNRDHNAAN--VSGPNAEDLSRLDYPDTLVFVGGLDGLYDWQKRYYEWLKIS 298
W++ LP G++RDH N +S A+ P T+VF+ D L + + ++
Sbjct: 248 WRLALPRGASRDHPWCNPLMSSAGAK------LPTTMVFMAEFDILKERNLEMCKVMRSH 301
Query: 299 GKKAQLIEYPNMMHGFYAFPN 319
GK+ + I + + H F+ N
Sbjct: 302 GKRVEGIVHGGVGHAFHILDN 322
>At3g27320 esterase, putative
Length = 460
Score = 122 bits (306), Expect = 3e-28
Identities = 99/324 (30%), Positives = 149/324 (45%), Gaps = 55/324 (16%)
Query: 32 SNGSVNRRLLNFLDNKTSAKATPINGVSTKDITVDAESKIWFRLFTPTGINASAGGGSNT 91
S GS N L + ++++ ++ G +T + +A S +R + P+ S+GG S
Sbjct: 115 SLGSSNSLLSHKVESRRNSY-----GYTTGSSSPEAGSSDVYRGYAPS----SSGGNSR- 164
Query: 92 ETTSLPVVIFFHGGGFTFMSPASLSYDTICRRFSRELNVVVVSVNYRRTPEYRYPTQYED 151
LPV++ FHGGG+ S S++ D CRR ++ +++V++V YR PE RYP ED
Sbjct: 165 ---KLPVMLQFHGGGWVSGSNDSVANDFFCRRMAKHCDIIVLAVGYRLAPENRYPAACED 221
Query: 152 GETALKFLDE-------NKSV-------------------------------LPENVDVS 173
G LK+L + NKS+ L + D S
Sbjct: 222 GFKVLKWLGKQANLAECNKSMGNSRRPGGEVKKSEVNKHIVDAFGASLVEPWLANHADPS 281
Query: 174 KCFLAGDSAGANLAHHVAVRACKAG--LQRIRVAGLISMQPFFGGEERTEAEIRLEGSLM 231
+C L G S GAN+A +VA +A + G L ++V + M PFF G T++EI+ S
Sbjct: 282 RCVLLGVSCGANIADYVARKAIEVGQNLDPVKVVAQVLMYPFFIGSVPTQSEIKQANSYF 341
Query: 232 ISMARTDWMWKVFLPEGS-NRDHNAANVSGPNAEDLSRLDYPDTLVFVGGLDGLYDWQKR 290
WK+FLPE + DH AAN P + P TL V D + D
Sbjct: 342 YDKPMCILAWKLFLPEEEFSLDHQAANPLVPGRSPPLKF-MPPTLTIVAEHDWMRDRAIA 400
Query: 291 YYEWLKISGKKAQLIEYPNMMHGF 314
Y E L+ A ++EY + +H F
Sbjct: 401 YSEELRKVNVDAPVLEYKDAVHEF 424
>At2g03550 putative esterase
Length = 312
Score = 121 bits (303), Expect = 6e-28
Identities = 95/294 (32%), Positives = 140/294 (47%), Gaps = 28/294 (9%)
Query: 53 TPINGVSTKDITVDAESKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTFMSP 112
TP NGV +KDI E + R++ P + LP++I+FHGGGF +
Sbjct: 35 TPQNGVVSKDIIHSPEKNLSLRIYLPEKVTVK----------KLPILIYFHGGGFIIETA 84
Query: 113 ASLSYDTICRRFSRELNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDEN------KSVL 166
S Y T N + +SVNYRR PE+ P YED +LK++ + ++ +
Sbjct: 85 FSPPYHTFLTSAVAAANCLAISVNYRRAPEFPVPIPYEDSWDSLKWVLTHITGTGPETWI 144
Query: 167 PENVDVSKCFLAGDSAGANLAHHVAVRACKAGLQRIRVAGLISMQPFFGGEER-TEAEIR 225
++ D K FLAGDSAG N++HH+ +RA K L ++G+I + P+F + E E+R
Sbjct: 145 NKHGDFGKVFLAGDSAGGNISHHLTMRAKKEKLCDSLISGIILIHPYFWSKTPIDEFEVR 204
Query: 226 LEGSLMISMARTDWMWKVFLPEGSNR-DHNAANVSGPNAEDLSRLDYPDTLVFVGGLDGL 284
G + W+V P D NV G D S L LV V G D
Sbjct: 205 DVG----KTKGVEGSWRVASPNSKQGVDDPWLNVVG---SDPSGLGCGRVLVMVAGDDLF 257
Query: 285 YDWQKRYYEWLKISG--KKAQLIEYPNMMHGFY-AFPNVPEASQLILQIKDFIN 335
Y E LK SG + +++E N H F+ PN A Q++ ++++FIN
Sbjct: 258 VRQGWCYAEKLKKSGWEGEVEVMETKNEGHVFHLKNPNSDNARQVVKKLEEFIN 311
>At3g48700 unknown protein
Length = 329
Score = 120 bits (300), Expect = 1e-27
Identities = 91/305 (29%), Positives = 143/305 (46%), Gaps = 35/305 (11%)
Query: 52 ATPINGVSTKDITVDAESKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTFMS 111
+ P NGV +KD+ ++ + R++ P A + LP++++FHGGGF +
Sbjct: 34 SNPQNGVVSKDVVYSPDNNLSLRIYLPE----KAATAETEASVKLPLLVYFHGGGFLVET 89
Query: 112 PASLSYDTICRRFSRELNVVVVSVNYRRTPEYRYPTQYEDGETALKFL------DENKSV 165
S +Y T + V VSV+YRR PE+ PT Y+D TALK++ ++
Sbjct: 90 AFSPTYHTFLTAAVSASDCVAVSVDYRRAPEHPIPTSYDDSWTALKWVFSHIAGSGSEDW 149
Query: 166 LPENVDVSKCFLAGDSAGANLAHHVAVRACK-----AGLQRIRVAGLISMQPFF------ 214
L ++ D SK FLAGDSAGAN+ HH+ ++A K L ++G+I + P+F
Sbjct: 150 LNKHADFSKVFLAGDSAGANITHHMTMKAAKDKLSPESLNESGISGIILVHPYFWSKTPV 209
Query: 215 GGEERTEAEIRLEGSLMISMARTDWMWKVFLPEGSN-RDHNAANVSGPNAEDLSRLDYPD 273
+E T+ IR + +W + P + D NV + DLS L
Sbjct: 210 DDKETTDVAIR---------TWIESVWTLASPNSKDGSDDPFINVVQSESVDLSGLGCGK 260
Query: 274 TLVFVGGLDGLYDWQKRYYEWL---KISGKKAQLIEYPNMMHGFY-AFPNVPEASQLILQ 329
LV V D L Y+E L + +G+ ++E H F+ PN +A +L+ +
Sbjct: 261 VLVMVAEKDALVRQGWGYWEKLGKSRWNGEVLDVVETKGEGHVFHLRDPNSEKAHELVHR 320
Query: 330 IKDFI 334
FI
Sbjct: 321 FAGFI 325
>At5g14310 unknown protein
Length = 446
Score = 116 bits (291), Expect = 2e-26
Identities = 87/294 (29%), Positives = 131/294 (43%), Gaps = 48/294 (16%)
Query: 96 LPVVIFFHGGGFTFMSPASLSYDTICRRFSRELNVVVVSVNYRRTPEYRYPTQYEDGETA 155
LPV++ FHGGG+ S S + D CRR ++ +V+V++V YR PE RYP +EDG
Sbjct: 151 LPVMLQFHGGGWVSGSSDSAANDFFCRRIAKVCDVIVLAVGYRLAPENRYPAAFEDGVKV 210
Query: 156 LKFLDENKSV--------------------------------------LPENVDVSKCFL 177
L +L + ++ L + D S+C L
Sbjct: 211 LHWLGKQANLADCCKSLGNRRVNGVEVKKLNVQGQIVDAFGASMVEPWLAAHADPSRCVL 270
Query: 178 AGDSAGANLAHHVAVRACKAG--LQRIRVAGLISMQPFFGGEERTEAEIRLEGSLMISMA 235
G S G N+A +VA +A +AG L+ ++V + M PFF G T++EI+L S
Sbjct: 271 LGVSCGGNIADYVARKAVEAGKLLEPVKVVAQVLMYPFFIGNNPTQSEIKLANSYFYDKP 330
Query: 236 RTDWMWKVFLPEGS-NRDHNAANVSGPNAEDLSRLDYPDTLVFVGGLDGLYDWQKRYYEW 294
+ WK+FLPE + DH AAN N P TL V D + D Y E
Sbjct: 331 VSVLAWKLFLPEKEFDFDHPAANPLAHNRSGPPLKLMPPTLTVVAEHDWMRDRAIAYSEE 390
Query: 295 LKISGKKAQLIEYPNMMHGFYAFP---NVPE----ASQLILQIKDFINNRVSNF 341
L+ + ++EY + +H F P+ A + + +K +I+ R F
Sbjct: 391 LRKVNVDSPVLEYKDAVHEFATLDMLLKTPQAQACAEDIAIWVKKYISLRGHEF 444
Score = 28.1 bits (61), Expect = 7.2
Identities = 21/68 (30%), Positives = 33/68 (47%), Gaps = 8/68 (11%)
Query: 13 LKTRLSISLIMTLTDAACRSNGSVNRRLLNFLDNKTSAKATP--INGVSTKDITVDAESK 70
LK RL + ++ D S G R +++ A A P +GV+TKDI +D +
Sbjct: 17 LKHRLQNLISISAADGLSDSFGVSTR------SDESVAAANPSFTDGVATKDIHIDPMTS 70
Query: 71 IWFRLFTP 78
+ R+F P
Sbjct: 71 LTVRIFLP 78
>At3g48690 unknown protein
Length = 324
Score = 116 bits (290), Expect = 2e-26
Identities = 87/295 (29%), Positives = 140/295 (46%), Gaps = 24/295 (8%)
Query: 54 PINGVSTKDITVDAESKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTFMSPA 113
P NGV +KD+ A++ + R++ P A + LP++++FHGGGF +
Sbjct: 36 PQNGVVSKDVVYSADNNLSVRIYLPEKAAAETD-------SKLPLLVYFHGGGFIIETAF 88
Query: 114 SLSYDTICRRFSRELNVVVVSVNYRRTPEYRYPTQYEDGETALKFL------DENKSVLP 167
S +Y T N V VSV+YRR PE+ ++D TALK++ + L
Sbjct: 89 SPTYHTFLTTSVSASNCVAVSVDYRRAPEHPISVPFDDSWTALKWVFTHITGSGQEDWLN 148
Query: 168 ENVDVSKCFLAGDSAGANLAHHVAVRACK----AGLQRIRVAGLISMQPFFGGEERTEAE 223
++ D S+ FL+GDSAGAN+ HH+A+RA K GL ++G+I + P+F + + +
Sbjct: 149 KHADFSRVFLSGDSAGANIVHHMAMRAAKEKLSPGLNDTGISGIILLHPYFWSKTPIDEK 208
Query: 224 IRLEGSLMISMARTDWMWKVFLPEGSN-RDHNAANVSGPNAEDLSRLDYPDTLVFVGGLD 282
+ +L + + + W + P + D NV + DLS L LV V D
Sbjct: 209 DTKDETLRM---KIEAFWMMASPNSKDGTDDPLLNVVQSESVDLSGLGCGKVLVMVAEKD 265
Query: 283 GLYDWQKRYYEWLKISGKK--AQLIEYPNMMHGFYAF-PNVPEASQLILQIKDFI 334
L Y L+ SG K +++E H F+ P A +++ + FI
Sbjct: 266 ALVRQGWGYAAKLEKSGWKGEVEVVESEGEDHVFHLLKPECDNAIEVMHKFSGFI 320
>At1g19190 unknown protein
Length = 318
Score = 114 bits (286), Expect = 6e-26
Identities = 89/293 (30%), Positives = 141/293 (47%), Gaps = 27/293 (9%)
Query: 54 PINGVSTKDITVDAESKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTFMSPA 113
P NGV +KD E + R++ P G +P++++FHGGGF +
Sbjct: 36 PENGVVSKDAVYSPEKNLSLRIYLPQNSVYETG------EKKIPLLVYFHGGGFIMETAF 89
Query: 114 SLSYDTICRRFSRELNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDEN------KSVLP 167
S Y T + + VSV YRR PE+ PT YED A++++ + + L
Sbjct: 90 SPIYHTFLTSAVSATDCIAVSVEYRRAPEHPIPTLYEDSWDAIQWIFTHITRSGPEDWLN 149
Query: 168 ENVDVSKCFLAGDSAGANLAHHVAVRACKAGL--QRIRVAGLISMQPFFGGEERTEAEIR 225
++ D SK FLAGDSAGAN+AHH+A+R K L + +++G+I P+F + E E+
Sbjct: 150 KHADFSKVFLAGDSAGANIAHHMAIRVDKEKLPPENFKISGMILFHPYFLSKALIE-EME 208
Query: 226 LEGSLMISMARTDWMWKVFLPEGSNRDHNA-ANVSGPNAEDLSRLDYPDTLVFVGGLDGL 284
+E +M + +W++ P+ N + NV G DL+ L LV V G D L
Sbjct: 209 VE-----AMRYYERLWRIASPDSGNGVEDPWINVVG---SDLTGLGCRRVLVMVAGNDVL 260
Query: 285 YDWQKRYYEWLKISG--KKAQLIEYPNMMHGFY-AFPNVPEASQLILQIKDFI 334
Y L+ SG K +++E H F+ P+ A +++ +F+
Sbjct: 261 ARGGWSYVAELEKSGWIGKVKVMETKEEGHVFHLRDPDSENARRVLRNFAEFL 313
>At1g49640 hypothetical protein
Length = 315
Score = 105 bits (262), Expect = 4e-23
Identities = 89/308 (28%), Positives = 146/308 (46%), Gaps = 19/308 (6%)
Query: 33 NGSVNRRLLNFLDNKTSAKATPINGVSTKDITVDAESKIWFRLFTPTGINASAGGGSNTE 92
NG V R L+ D K ++ P N V +KD+ ++ + R+F P + +T
Sbjct: 19 NGRVER--LSGNDIKPTS-LNPQNDVVSKDVMYSSDHNLSVRMFLP-----NKSRKLDTA 70
Query: 93 TTSLPVVIFFHGGGFTFMSPASLSYDTICRRFSRELNVVVVSVNYRRTPEYRYPTQYEDG 152
+P++I+FHGG + SP S Y N + VSV YR PE+ P Y+D
Sbjct: 71 GNKIPLLIYFHGGAYIIQSPFSPVYHNYLTEVVITANCLAVSVQYRLAPEHPVPAAYDDS 130
Query: 153 ETALKFL-DENKSVLPENVDVSKCFLAGDSAGANLAHHVAVRACKAGLQRIRVAGLISMQ 211
+A++++ + + E D + F+AGDSAGAN++HH+ +RA K L + G++ +
Sbjct: 131 WSAIQWIFSHSDDWINEYADFDRVFIAGDSAGANISHHMGIRAGKEKLSP-TIKGIVMVH 189
Query: 212 PFFGGEERTEAEIRLEGSLMISMARTDWMWKVFLPEGSNRDHNAA--NVSGPNAEDLSRL 269
P F G+E + +G + +A ++W+ + S N NV G + D+S +
Sbjct: 190 PGFWGKEPIDEHDVQDGEVRNKIA---YIWENIVSPNSVDGVNDPWFNVVG-SGSDVSEM 245
Query: 270 DYPDTLVFVGGLDGLYDWQKRYYEWLKISGKK--AQLIEYPNMMHGFYAF-PNVPEASQL 326
LV V G D + Y L+ S K ++IE H F+ N AS+L
Sbjct: 246 GCEKVLVAVAGKDVFWRQGLAYAAKLEKSQWKGSVEVIEEEEEGHCFHLHNHNSQNASKL 305
Query: 327 ILQIKDFI 334
+ + +FI
Sbjct: 306 MQKFLEFI 313
>At1g49650 unknown protein
Length = 374
Score = 100 bits (249), Expect = 1e-21
Identities = 85/296 (28%), Positives = 132/296 (43%), Gaps = 22/296 (7%)
Query: 50 AKATPINGVSTKDITVDAESKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTF 109
A P N V +KD+ + RLF P A G LP++I+FHGG +
Sbjct: 88 ASLNPRNDVVSKDVVYSPGHNLSVRLFLPHKSTQLAAGNK------LPLLIYFHGGAWIN 141
Query: 110 MSPASLSYDTICRRFSRELNVVVVSVNYRRTPEYRYPTQYEDGETALKFL------DENK 163
SP S Y + N + VSV YRR PE P YED +A++++ +
Sbjct: 142 ESPFSPIYHNFLTEVVKSANCLAVSVQYRRAPEDPVPAAYEDTWSAIQWIFSHSCGSGEE 201
Query: 164 SVLPENVDVSKCFLAGDSAGANLAHHVAVRACKAGLQRIRVAGLISMQPFFGGEERTEAE 223
+ + D + FLAGDSAG N++HH+A+RA K L + R+ G + + P G++ +
Sbjct: 202 DWINKYADFERVFLAGDSAGGNISHHMAMRAGKEKL-KPRIKGTVIVHPAIWGKDPVDEH 260
Query: 224 IRLEGSLMISMARTDWMWKVFLPEGS--NRDHNAANVSGPNAEDLSRLDYPDTLVFVGGL 281
+ + +A +W+ + S D NV G + + S + LV V G
Sbjct: 261 DVQDREIRDGVAE---VWEKIVSPNSVDGADDPWFNVVG-SGSNFSGMGCDKVLVEVAGK 316
Query: 282 DGLYDWQKRYYEWLKISGKK--AQLIEYPNMMHGFYAF-PNVPEASQLILQIKDFI 334
D + Y LK SG K ++IE + H F+ P+ A + + +FI
Sbjct: 317 DVFWRQGLAYAAKLKKSGWKGEVEVIEEEDEEHCFHLLNPSSENAPSFMKRFVEFI 372
>At1g49660 unknown protein
Length = 319
Score = 95.5 bits (236), Expect = 4e-20
Identities = 72/240 (30%), Positives = 109/240 (45%), Gaps = 16/240 (6%)
Query: 50 AKATPINGVSTKDITVDAESKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTF 109
A P V +KD+ E+ + RLF P G LP++I+ HGG +
Sbjct: 32 ASLDPTYDVVSKDVIYSPENNLSVRLFLPHKSTKLTAGNK------LPLLIYIHGGAWII 85
Query: 110 MSPASLSYDTICRRFSRELNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDENKS----- 164
SP S Y + N + VSV YRR PE P YED +A++++ + +
Sbjct: 86 ESPFSPLYHNYLTEVVKSANCLAVSVQYRRAPEDPVPAAYEDVWSAIQWIFAHSNGSGPV 145
Query: 165 -VLPENVDVSKCFLAGDSAGANLAHHVAVRACKAGLQRIRVAGLISMQPFFGGEERTEAE 223
+ ++ D K FL GDSAG N++HH+A++A K +++ G+ + P F G + + E
Sbjct: 146 DWINKHADFGKVFLGGDSAGGNISHHMAMKAGKEKKLDLKIKGIAVVHPAFWGTDPVD-E 204
Query: 224 IRLEGSLMISMARTDWMWKVFLPEGSN-RDHNAANVSGPNAEDLSRLDYPDTLVFVGGLD 282
++ S W K+ P N D NV+G + D S L LV V G D
Sbjct: 205 YDVQDKETRSGIAEIWE-KIASPNSVNGTDDPLFNVNG-SGSDFSGLGCDKVLVAVAGKD 262
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.136 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,798,495
Number of Sequences: 26719
Number of extensions: 331940
Number of successful extensions: 892
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 26
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 810
Number of HSP's gapped (non-prelim): 34
length of query: 342
length of database: 11,318,596
effective HSP length: 100
effective length of query: 242
effective length of database: 8,646,696
effective search space: 2092500432
effective search space used: 2092500432
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC139853.10


