

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139841.5 + phase: 0
(252 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
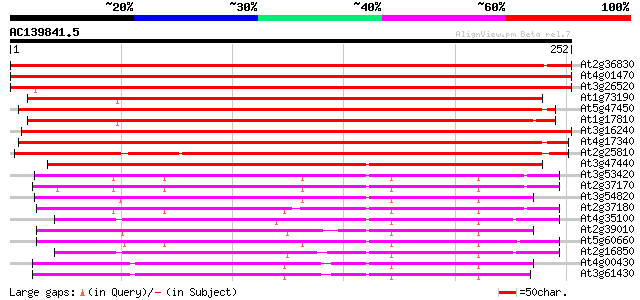
Sequences producing significant alignments: (bits) Value
At2g36830 putative aquaporin (tonoplast intrinsic protein gamma) 410 e-115
At4g01470 putative water channel protein 404 e-113
At3g26520 salt-stress induced tonoplast intrinsic protein 388 e-108
At1g73190 tonoplast intrinsic protein, alpha (alpha-TIP) 298 2e-81
At5g47450 membrane channel protein-like; aquaporin (tonoplast in... 296 7e-81
At1g17810 putative protein 295 2e-80
At3g16240 delta tonoplast integral protein (delta-TIP) 295 2e-80
At4g17340 membrane channel like protein 294 4e-80
At2g25810 putative aquaporin (tonoplast intrinsic protein) 265 1e-71
At3g47440 aquaporin-like protein 184 3e-47
At3g53420 plasma membrane intrinsic protein 2a 140 5e-34
At2g37170 aquaporin (plasma membrane intrinsic protein 2B) 140 5e-34
At3g54820 aquaporin/MIP - like protein 140 8e-34
At2g37180 aquaporin (plasma membrane intrinsic protein 2C) 137 5e-33
At4g35100 plasma membrane intrinsic protein PIP3 137 7e-33
At2g39010 putative aquaporin (water channel protein) 136 1e-32
At5g60660 mipC protein - like (aquaporin) 134 4e-32
At2g16850 putative aquaporin (plasma membrane intrinsic protein) 134 4e-32
At4g00430 transmembrane protein 125 2e-29
At3g61430 plasma membrane intrinsic protein 1a 124 6e-29
>At2g36830 putative aquaporin (tonoplast intrinsic protein gamma)
Length = 251
Score = 410 bits (1054), Expect = e-115
Identities = 202/252 (80%), Positives = 221/252 (87%), Gaps = 1/252 (0%)
Query: 1 MPISRIAIGNPSEFGKADALKAALAEFISMLIFVFAGEGSGMAYNKLTNNGAATPAGLVA 60
MPI IAIG P E + DALKAALAEFIS LIFV AG GSGMA+NKLT NGA TP+GLVA
Sbjct: 1 MPIRNIAIGRPDEATRPDALKAALAEFISTLIFVVAGSGSGMAFNKLTENGATTPSGLVA 60
Query: 61 ASLSHAFALFVAVSVGANISGGHVNPAVTFGAFIGGHITLIRGLLYWIAQLLGSVVACLL 120
A+++HAF LFVAVSVGANISGGHVNPAVTFGAFIGG+ITL+RG+LYWIAQLLGSVVACL+
Sbjct: 61 AAVAHAFGLFVAVSVGANISGGHVNPAVTFGAFIGGNITLLRGILYWIAQLLGSVVACLI 120
Query: 121 LKIATGGLETSAFSLSSGVGATNALVFEIVMTFGLVYTVYATAVDPKNGSLGTIAPIAIG 180
LK ATGGL AF LS+GVG NA VFEIVMTFGLVYTVYATA+DPKNGSLGTIAPIAIG
Sbjct: 121 LKFATGGLAVPAFGLSAGVGVLNAFVFEIVMTFGLVYTVYATAIDPKNGSLGTIAPIAIG 180
Query: 181 FIVGANILAGGAFDGASMNPAVSFGPAVVSWTWANHWVYWVGPLIGSAIAAVVYETFFIT 240
FIVGANILAGGAF GASMNPAV+FGPAVVSWTW NHWVYW GPL+G IA ++YE FFI
Sbjct: 181 FIVGANILAGGAFSGASMNPAVAFGPAVVSWTWTNHWVYWAGPLVGGGIAGLIYEVFFIN 240
Query: 241 PSSYEQLPVTDY 252
+++EQLP TDY
Sbjct: 241 -TTHEQLPTTDY 251
>At4g01470 putative water channel protein
Length = 252
Score = 404 bits (1039), Expect = e-113
Identities = 197/252 (78%), Positives = 222/252 (87%)
Query: 1 MPISRIAIGNPSEFGKADALKAALAEFISMLIFVFAGEGSGMAYNKLTNNGAATPAGLVA 60
MPI+RIAIG P E + DA++AA AEF SM+IFVFAG+GSGMAY KLT +G ATPAGLVA
Sbjct: 1 MPINRIAIGTPGEASRPDAIRAAFAEFFSMVIFVFAGQGSGMAYGKLTGDGPATPAGLVA 60
Query: 61 ASLSHAFALFVAVSVGANISGGHVNPAVTFGAFIGGHITLIRGLLYWIAQLLGSVVACLL 120
ASLSHAFALFVAVSVGAN+SGGHVNPAVTFGAFIGG+ITL+R +LYWIAQLLG+VVACLL
Sbjct: 61 ASLSHAFALFVAVSVGANVSGGHVNPAVTFGAFIGGNITLLRAILYWIAQLLGAVVACLL 120
Query: 121 LKIATGGLETSAFSLSSGVGATNALVFEIVMTFGLVYTVYATAVDPKNGSLGTIAPIAIG 180
LK++TGG+ET+AFSLS GV NA+VFEIVMTFGLVYTVYATAVDPK G +G IAP+AIG
Sbjct: 121 LKVSTGGMETAAFSLSYGVTPWNAVVFEIVMTFGLVYTVYATAVDPKKGDIGIIAPLAIG 180
Query: 181 FIVGANILAGGAFDGASMNPAVSFGPAVVSWTWANHWVYWVGPLIGSAIAAVVYETFFIT 240
IVGANIL GGAFDGASMNPAVSFGPAVVSW W NHWVYWVGP IG+AIAA+VY+T FI
Sbjct: 181 LIVGANILVGGAFDGASMNPAVSFGPAVVSWIWTNHWVYWVGPFIGAAIAAIVYDTIFIG 240
Query: 241 PSSYEQLPVTDY 252
+ +E LP D+
Sbjct: 241 SNGHEPLPSNDF 252
>At3g26520 salt-stress induced tonoplast intrinsic protein
Length = 253
Score = 388 bits (997), Expect = e-108
Identities = 192/253 (75%), Positives = 215/253 (84%), Gaps = 1/253 (0%)
Query: 1 MPISRIAIGN-PSEFGKADALKAALAEFISMLIFVFAGEGSGMAYNKLTNNGAATPAGLV 59
MP IAIG E +AL+AALAEFIS LIFVFAG GSG+A+NK+T+NGA TP+GLV
Sbjct: 1 MPTRNIAIGGVQEEVYHPNALRAALAEFISTLIFVFAGSGSGIAFNKITDNGATTPSGLV 60
Query: 60 AASLSHAFALFVAVSVGANISGGHVNPAVTFGAFIGGHITLIRGLLYWIAQLLGSVVACL 119
AA+L+HAF LFVAVSVGANISGGHVNPAVTFG +GG+ITL+RG+LYWIAQLLGSV AC
Sbjct: 61 AAALAHAFGLFVAVSVGANISGGHVNPAVTFGVLLGGNITLLRGILYWIAQLLGSVAACF 120
Query: 120 LLKIATGGLETSAFSLSSGVGATNALVFEIVMTFGLVYTVYATAVDPKNGSLGTIAPIAI 179
LL ATGG AF LS+GVG+ NALVFEIVMTFGLVYTVYATAVDPKNGSLGTIAPIAI
Sbjct: 121 LLSFATGGEPIPAFGLSAGVGSLNALVFEIVMTFGLVYTVYATAVDPKNGSLGTIAPIAI 180
Query: 180 GFIVGANILAGGAFDGASMNPAVSFGPAVVSWTWANHWVYWVGPLIGSAIAAVVYETFFI 239
GFIVGANILAGGAF GASMNPAV+FGPAVVSWTW NHWVYW GPLIG +A ++Y+ FI
Sbjct: 181 GFIVGANILAGGAFSGASMNPAVAFGPAVVSWTWTNHWVYWAGPLIGGGLAGIIYDFVFI 240
Query: 240 TPSSYEQLPVTDY 252
+++EQLP TDY
Sbjct: 241 DENAHEQLPTTDY 253
>At1g73190 tonoplast intrinsic protein, alpha (alpha-TIP)
Length = 268
Score = 298 bits (763), Expect = 2e-81
Identities = 141/236 (59%), Positives = 176/236 (73%), Gaps = 5/236 (2%)
Query: 9 GNPSEFGKADALKAALAEFISMLIFVFAGEGSGMAYNKL-----TNNGAATPAGLVAASL 63
G E D+++A LAEF+S +FVFA EGS ++ +KL + G TP GL+ +L
Sbjct: 12 GRADEATHPDSIRATLAEFLSTFVFVFAAEGSILSLDKLYWEHAAHAGTNTPGGLILVAL 71
Query: 64 SHAFALFVAVSVGANISGGHVNPAVTFGAFIGGHITLIRGLLYWIAQLLGSVVACLLLKI 123
+HAFALF AVS N+SGGHVNPAVTFGA +GG +T IR + YWIAQLLG+++ACLLL++
Sbjct: 72 AHAFALFAAVSAAINVSGGHVNPAVTFGALVGGRVTAIRAIYYWIAQLLGAILACLLLRL 131
Query: 124 ATGGLETSAFSLSSGVGATNALVFEIVMTFGLVYTVYATAVDPKNGSLGTIAPIAIGFIV 183
T G+ F L+SGVGA N LV EI++TFGLVY VY+T +DPK GSLG IAP+AIG IV
Sbjct: 132 TTNGMRPVGFRLASGVGAVNGLVLEIILTFGLVYVVYSTLIDPKRGSLGIIAPLAIGLIV 191
Query: 184 GANILAGGAFDGASMNPAVSFGPAVVSWTWANHWVYWVGPLIGSAIAAVVYETFFI 239
GANIL GG F GASMNPA +FGPA+V W W +HW+YWVGP IGSA+AA++YE I
Sbjct: 192 GANILVGGPFSGASMNPARAFGPALVGWRWHDHWIYWVGPFIGSALAALIYEYMVI 247
>At5g47450 membrane channel protein-like; aquaporin (tonoplast
intrinsic protein)-like
Length = 250
Score = 296 bits (758), Expect = 7e-81
Identities = 150/241 (62%), Positives = 180/241 (74%), Gaps = 2/241 (0%)
Query: 5 RIAIGNPSEFGKADALKAALAEFISMLIFVFAGEGSGMAYNKLTNNGAATPAGLVAASLS 64
+I +G+ + +LKA L+EFI+ L+FVFAG GS +A+ KLT++GA PAGLVA +++
Sbjct: 3 KIEVGSVGDSFSVSSLKAYLSEFIATLLFVFAGVGSAVAFAKLTSDGALDPAGLVAIAIA 62
Query: 65 HAFALFVAVSVGANISGGHVNPAVTFGAFIGGHITLIRGLLYWIAQLLGSVVACLLLKIA 124
HAFALFV VS+ ANISGGH+NPAVT G IGG+ITLI G YWIAQ LGS+VACLLL
Sbjct: 63 HAFALFVGVSIAANISGGHLNPAVTLGLAIGGNITLITGFFYWIAQCLGSIVACLLLVFV 122
Query: 125 TGGLETSAFSLSSGVGATNALVFEIVMTFGLVYTVYATAVDPKNGSLGTIAPIAIGFIVG 184
T G +S+G+GA +V EIV+TF LVYTVYATA DPK GSLGTIAPIAIGFIVG
Sbjct: 123 TNGKSVPTHGVSAGLGAVEGVVMEIVVTFALVYTVYATAADPKKGSLGTIAPIAIGFIVG 182
Query: 185 ANILAGGAFDGASMNPAVSFGPAVVSWTWANHWVYWVGPLIGSAIAAVVYETFFITPSSY 244
ANILA G F G SMNPA SFGPAVVS + W+YWVGPL+G A+A ++Y FI SY
Sbjct: 183 ANILAAGPFSGGSMNPARSFGPAVVSGDLSQIWIYWVGPLVGGALAGLIYGDVFI--GSY 240
Query: 245 E 245
E
Sbjct: 241 E 241
>At1g17810 putative protein
Length = 267
Score = 295 bits (755), Expect = 2e-80
Identities = 141/242 (58%), Positives = 180/242 (74%), Gaps = 6/242 (2%)
Query: 9 GNPSEFGKADALKAALAEFISMLIFVFAGEGSGMAYNKL-----TNNGAATPAGLVAASL 63
G E D+++A LAEF+S +FVFAGEGS +A +KL + G TP GLV +L
Sbjct: 12 GRADEATHPDSIRATLAEFLSTFVFVFAGEGSILALDKLYWDTAAHTGTNTPGGLVLVAL 71
Query: 64 SHAFALFVAVSVGANISGGHVNPAVTFGAFIGGHITLIRGLLYWIAQLLGSVVACLLLKI 123
+HA ALF AVS N+SGGHVNPAVTF A IGG I++IR + YW+AQL+G+++ACLLL++
Sbjct: 72 AHALALFAAVSAAINVSGGHVNPAVTFAALIGGRISVIRAIYYWVAQLIGAILACLLLRL 131
Query: 124 ATGGLETSAFSLSSGVGATNALVFEIVMTFGLVYTVYATAVDPKNGSLGTIAPIAIGFIV 183
AT GL F ++SGV + L+ EI++TF LVY VY+TA+DPK GS+G IAP+AIG IV
Sbjct: 132 ATNGLRPVGFHVASGVSELHGLLMEIILTFALVYVVYSTAIDPKRGSIGIIAPLAIGLIV 191
Query: 184 GANILAGGAFDGASMNPAVSFGPAVVSWTWANHWVYWVGPLIGSAIAAVVYETFFITPSS 243
GANIL GG FDGASMNPA +FGPA+V W W+NHW+YWVGP IG A+AA++YE + I PS
Sbjct: 192 GANILVGGPFDGASMNPARAFGPALVGWRWSNHWIYWVGPFIGGALAALIYE-YMIIPSV 250
Query: 244 YE 245
E
Sbjct: 251 NE 252
>At3g16240 delta tonoplast integral protein (delta-TIP)
Length = 250
Score = 295 bits (754), Expect = 2e-80
Identities = 146/247 (59%), Positives = 180/247 (72%)
Query: 6 IAIGNPSEFGKADALKAALAEFISMLIFVFAGEGSGMAYNKLTNNGAATPAGLVAASLSH 65
+A G+ + +L+A LAEFIS L+FVFAG GS +AY KLT++ A GLVA ++ H
Sbjct: 4 VAFGSFDDSFSLASLRAYLAEFISTLLFVFAGVGSAIAYAKLTSDAALDTPGLVAIAVCH 63
Query: 66 AFALFVAVSVGANISGGHVNPAVTFGAFIGGHITLIRGLLYWIAQLLGSVVACLLLKIAT 125
FALFVAV++GANISGGHVNPAVTFG +GG IT+I G+ YWIAQLLGS AC LLK T
Sbjct: 64 GFALFVAVAIGANISGGHVNPAVTFGLAVGGQITVITGVFYWIAQLLGSTAACFLLKYVT 123
Query: 126 GGLETSAFSLSSGVGATNALVFEIVMTFGLVYTVYATAVDPKNGSLGTIAPIAIGFIVGA 185
GGL S+++G+G+ +V EI++TF LVYTVYATA DPK GSLGTIAP+AIG IVGA
Sbjct: 124 GGLAVPTHSVAAGLGSIEGVVMEIIITFALVYTVYATAADPKKGSLGTIAPLAIGLIVGA 183
Query: 186 NILAGGAFDGASMNPAVSFGPAVVSWTWANHWVYWVGPLIGSAIAAVVYETFFITPSSYE 245
NILA G F G SMNPA SFGPAV + ++ HWVYWVGPLIG +A ++Y F+ S +
Sbjct: 184 NILAAGPFSGGSMNPARSFGPAVAAGDFSGHWVYWVGPLIGGGLAGLIYGNVFMGSSEHV 243
Query: 246 QLPVTDY 252
L D+
Sbjct: 244 PLASADF 250
>At4g17340 membrane channel like protein
Length = 250
Score = 294 bits (752), Expect = 4e-80
Identities = 148/247 (59%), Positives = 182/247 (72%), Gaps = 2/247 (0%)
Query: 5 RIAIGNPSEFGKADALKAALAEFISMLIFVFAGEGSGMAYNKLTNNGAATPAGLVAASLS 64
+I IG+ + +LKA L+EFI+ L+FVFAG GS +A+ KLT++ A PAGLVA +++
Sbjct: 3 KIEIGSVGDSFSVASLKAYLSEFIATLLFVFAGVGSALAFAKLTSDAALDPAGLVAVAVA 62
Query: 65 HAFALFVAVSVGANISGGHVNPAVTFGAFIGGHITLIRGLLYWIAQLLGSVVACLLLKIA 124
HAFALFV VS+ ANISGGH+NPAVT G +GG+IT+I G YWIAQ LGS+VACLLL
Sbjct: 63 HAFALFVGVSIAANISGGHLNPAVTLGLAVGGNITVITGFFYWIAQCLGSIVACLLLVFV 122
Query: 125 TGGLETSAFSLSSGVGATNALVFEIVMTFGLVYTVYATAVDPKNGSLGTIAPIAIGFIVG 184
T G +++G+GA +V EIV+TF LVYTVYATA DPK GSLGTIAPIAIGFIVG
Sbjct: 123 TNGESVPTHGVAAGLGAIEGVVMEIVVTFALVYTVYATAADPKKGSLGTIAPIAIGFIVG 182
Query: 185 ANILAGGAFDGASMNPAVSFGPAVVSWTWANHWVYWVGPLIGSAIAAVVYETFFITPSSY 244
ANILA G F G SMNPA SFGPAVVS ++ W+YWVGPL+G A+A ++Y FI SY
Sbjct: 183 ANILAAGPFSGGSMNPARSFGPAVVSGDFSQIWIYWVGPLVGGALAGLIYGDVFI--GSY 240
Query: 245 EQLPVTD 251
P T+
Sbjct: 241 APAPTTE 247
>At2g25810 putative aquaporin (tonoplast intrinsic protein)
Length = 249
Score = 265 bits (678), Expect = 1e-71
Identities = 133/249 (53%), Positives = 169/249 (67%), Gaps = 7/249 (2%)
Query: 3 ISRIAIGNPSEFGKADALKAALAEFISMLIFVFAGEGSGMAYNKLTNNGAATPAGLVAAS 62
+ +I +G+ SE K D +KA + EFI+ +FVFAG GS MA + L N T GL A +
Sbjct: 1 MKKIELGHHSEAAKPDCIKALIVEFITTFLFVFAGVGSAMATDSLVGN---TLVGLFAVA 57
Query: 63 LSHAFALFVAVSVGANISGGHVNPAVTFGAFIGGHITLIRGLLYWIAQLLGSVVACLLLK 122
++HAF + V +S G +ISGGH+NPAVT G +GGHI++ R LYWI QLL S AC LL
Sbjct: 58 VAHAFVVAVMISAG-HISGGHLNPAVTLGLLLGGHISVFRAFLYWIDQLLASSAACFLLS 116
Query: 123 IATGGLETSAFSLSSGVGATNALVFEIVMTFGLVYTVYATAVDPKNGSLGTIAPIAIGFI 182
TGG+ T +L+SGV T +++EI++TF L++TVYAT VDPK GSL P+ GF+
Sbjct: 117 YLTGGMGTPVHTLASGVSYTQGIIWEIILTFSLLFTVYATIVDPKKGSLDGFGPLLTGFV 176
Query: 183 VGANILAGGAFDGASMNPAVSFGPAVVSWTWANHWVYWVGPLIGSAIAAVVYETFFITPS 242
VGANILAGGAF GASMNPA SFGPA+VS W +HWVYWVGPLIG +A +YE I
Sbjct: 177 VGANILAGGAFSGASMNPARSFGPALVSGNWTDHWVYWVGPLIGGGLAGFIYENVLI--- 233
Query: 243 SYEQLPVTD 251
+PV D
Sbjct: 234 DRPHVPVAD 242
>At3g47440 aquaporin-like protein
Length = 256
Score = 184 bits (468), Expect = 3e-47
Identities = 95/222 (42%), Positives = 138/222 (61%), Gaps = 1/222 (0%)
Query: 18 DALKAALAEFISMLIFVFAGEGSGMAYNKLTNNGAATPAGLVAASLSHAFALFVAVSVGA 77
+AL+ ++EFIS FV A GS M+ KL + P G++ ++++A AL +V +
Sbjct: 20 NALRCYVSEFISTFFFVLAAVGSVMSSRKLMAGDVSGPFGVLIPAIANALALSSSVYISW 79
Query: 78 NISGGHVNPAVTFGAFIGGHITLIRGLLYWIAQLLGSVVACLLLKIATGGLETSAFSLSS 137
N+SGGHVNPAVTF + G I++ + YW +Q++ SV+ACL+LK+ + ++
Sbjct: 80 NVSGGHVNPAVTFAMAVAGRISVPTAMFYWTSQMIASVMACLVLKVTVMEQHVPIYKIAG 139
Query: 138 GVGATNALVFEIVMTFGLVYTVYATAVDPKNGSLGTIAPIAIGFIVGANILAGGAFDGAS 197
+ A V E V+ F LVYTV+ TA DP+ G + PI IGF+ GAN+LA G F G S
Sbjct: 140 EMTGFGASVLEGVLAFVLVYTVF-TASDPRRGLPLAVGPIFIGFVAGANVLAAGPFSGGS 198
Query: 198 MNPAVSFGPAVVSWTWANHWVYWVGPLIGSAIAAVVYETFFI 239
MNPA +FG A+V ++ N VYWVGPL+G A AA+VY+ +
Sbjct: 199 MNPACAFGSAMVYGSFKNQAVYWVGPLLGGATAALVYDNVVV 240
>At3g53420 plasma membrane intrinsic protein 2a
Length = 287
Score = 140 bits (354), Expect = 5e-34
Identities = 95/251 (37%), Positives = 134/251 (52%), Gaps = 17/251 (6%)
Query: 12 SEFGKADALKAALAEFISMLIFVFAGEGSGMAYN--KLTNNGAATPAGLVAASLSHAFA- 68
+E K +A +AEF++ L+F++ + + Y T+ G G+ ++ AF
Sbjct: 30 AELKKWSFYRAVIAEFVATLLFLYITVLTVIGYKIQSDTDAGGVDCGGVGILGIAWAFGG 89
Query: 69 -LFVAVSVGANISGGHVNPAVTFGAFIGGHITLIRGLLYWIAQLLGSVVACLLLKIATGG 127
+F+ V A ISGGH+NPAVTFG F+ ++L R LLY IAQ LG++ +K
Sbjct: 90 MIFILVYCTAGISGGHINPAVTFGLFLARKVSLPRALLYIIAQCLGAICGVGFVKAFQSS 149
Query: 128 LET----SAFSLSSGVGATNALVFEIVMTFGLVYTVYATAVDPKNGS----LGTIAPIAI 179
T A SL+ G L EI+ TF LVYTV+ +A DPK + + +AP+ I
Sbjct: 150 YYTRYGGGANSLADGYSTGTGLAAEIIGTFVLVYTVF-SATDPKRSARDSHVPVLAPLPI 208
Query: 180 GFIVGANILAGGAFDGASMNPAVSFGPAVV---SWTWANHWVYWVGPLIGSAIAAVVYET 236
GF V LA G +NPA SFG AV+ S W +HW++WVGP IG+AIAA Y
Sbjct: 209 GFAVFMVHLATIPITGTGINPARSFGAAVIYNKSKPWDDHWIFWVGPFIGAAIAA-FYHQ 267
Query: 237 FFITPSSYEQL 247
F + S + L
Sbjct: 268 FVLRASGSKSL 278
>At2g37170 aquaporin (plasma membrane intrinsic protein 2B)
Length = 285
Score = 140 bits (354), Expect = 5e-34
Identities = 96/258 (37%), Positives = 136/258 (52%), Gaps = 23/258 (8%)
Query: 11 PSEFGKADAL------KAALAEFISMLIFVFAGEGSGMAYN--KLTNNGAATPAGLVAAS 62
P+ F AD L +A +AEF++ L+F++ + + Y T G G+
Sbjct: 21 PTPFFDADELTKWSLYRAVIAEFVATLLFLYITVLTVIGYKIQSDTKAGGVDCGGVGILG 80
Query: 63 LSHAFA--LFVAVSVGANISGGHVNPAVTFGAFIGGHITLIRGLLYWIAQLLGSVVACLL 120
++ AF +F+ V A ISGGH+NPAVTFG F+ ++LIR +LY +AQ LG++
Sbjct: 81 IAWAFGGMIFILVYCTAGISGGHINPAVTFGLFLARKVSLIRAVLYMVAQCLGAICGVGF 140
Query: 121 LKIATGGLET----SAFSLSSGVGATNALVFEIVMTFGLVYTVYATAVDPKNGS----LG 172
+K A SL+ G L EI+ TF LVYTV+ +A DPK + +
Sbjct: 141 VKAFQSSYYDRYGGGANSLADGYNTGTGLAAEIIGTFVLVYTVF-SATDPKRNARDSHVP 199
Query: 173 TIAPIAIGFIVGANILAGGAFDGASMNPAVSFGPAVV---SWTWANHWVYWVGPLIGSAI 229
+AP+ IGF V LA G +NPA SFG AV+ S W +HW++WVGP IG+AI
Sbjct: 200 VLAPLPIGFAVFMVHLATIPITGTGINPARSFGAAVIYNKSKPWDDHWIFWVGPFIGAAI 259
Query: 230 AAVVYETFFITPSSYEQL 247
AA Y F + S + L
Sbjct: 260 AA-FYHQFVLRASGSKSL 276
>At3g54820 aquaporin/MIP - like protein
Length = 286
Score = 140 bits (352), Expect = 8e-34
Identities = 85/239 (35%), Positives = 127/239 (52%), Gaps = 16/239 (6%)
Query: 12 SEFGKADALKAALAEFISMLIFVFAGEGSGMAYNKLT----NNGAATPAGLVAASLSHAF 67
+E GK +A +AEFI+ L+F++ + + Y T N T G++ + +
Sbjct: 29 TELGKWSFYRALIAEFIATLLFLYVTIMTVIGYKSQTDPALNPDQCTGVGVLGIAWAFGG 88
Query: 68 ALFVAVSVGANISGGHVNPAVTFGAFIGGHITLIRGLLYWIAQLLGSVVACLLLKIATGG 127
+F+ V A ISGGH+NPAVTFG + +TL+R ++Y +AQ LG++ L+K
Sbjct: 89 MIFILVYCTAGISGGHINPAVTFGLLLARKVTLVRAVMYMVAQCLGAICGVALVKAFQSA 148
Query: 128 LET----SAFSLSSGVGATNALVFEIVMTFGLVYTVYATAVDPKNGS----LGTIAPIAI 179
T A LS G + EI+ TF LVYTV+ +A DPK + + +AP+ I
Sbjct: 149 YFTRYGGGANGLSDGYSIGTGVAAEIIGTFVLVYTVF-SATDPKRSARDSHVPVLAPLPI 207
Query: 180 GFIVGANILAGGAFDGASMNPAVSFGPAVV---SWTWANHWVYWVGPLIGSAIAAVVYE 235
GF V LA G +NPA S G A++ W +HW++WVGP G+AIAA ++
Sbjct: 208 GFAVFIVHLATIPITGTGINPARSLGAAIIYNKDKAWDHHWIFWVGPFAGAAIAAFYHQ 266
>At2g37180 aquaporin (plasma membrane intrinsic protein 2C)
Length = 285
Score = 137 bits (345), Expect = 5e-33
Identities = 93/253 (36%), Positives = 133/253 (51%), Gaps = 23/253 (9%)
Query: 13 EFGKADALKAALAEFISMLIFVFAGEGSGMAYN--KLTNNGAATPAGLVAASLSHAFA-- 68
E K +A +AEF++ L+F++ + + Y T G G+ ++ AF
Sbjct: 29 ELTKWSLYRAVIAEFVATLLFLYVTVLTVIGYKIQSDTKAGGVDCGGVGILGIAWAFGGM 88
Query: 69 LFVAVSVGANISGGHVNPAVTFGAFIGGHITLIRGLLYWIAQLLGSVVACLLLK------ 122
+F+ V A ISGGH+NPAVTFG F+ ++LIR +LY +AQ LG++ +K
Sbjct: 89 IFILVYCTAGISGGHINPAVTFGLFLARKVSLIRAVLYMVAQCLGAICGVGFVKAFQSSH 148
Query: 123 -IATGGLETSAFSLSSGVGATNALVFEIVMTFGLVYTVYATAVDPKNGS----LGTIAPI 177
+ GG A L+ G L EI+ TF LVYTV+ +A DPK + + +AP+
Sbjct: 149 YVNYGG---GANFLADGYNTGTGLAAEIIGTFVLVYTVF-SATDPKRNARDSHVPVLAPL 204
Query: 178 AIGFIVGANILAGGAFDGASMNPAVSFGPAVV---SWTWANHWVYWVGPLIGSAIAAVVY 234
IGF V LA G +NPA SFG AV+ S W +HW++WVGP IG+ IAA Y
Sbjct: 205 PIGFAVFMVHLATIPITGTGINPARSFGAAVIFNKSKPWDDHWIFWVGPFIGATIAA-FY 263
Query: 235 ETFFITPSSYEQL 247
F + S + L
Sbjct: 264 HQFVLRASGSKSL 276
>At4g35100 plasma membrane intrinsic protein PIP3
Length = 280
Score = 137 bits (344), Expect = 7e-33
Identities = 87/238 (36%), Positives = 129/238 (53%), Gaps = 15/238 (6%)
Query: 21 KAALAEFISMLIFVFAGEGSGMAYNKLTNNGAATPAGLVAASLSHAFALFVAVSVGANIS 80
+A +AEFI+ L+F++ + + + K T G GL+ + + +FV V A IS
Sbjct: 38 RALIAEFIATLLFLYVTVATVIGHKKQT--GPCDGVGLLGIAWAFGGMIFVLVYCTAGIS 95
Query: 81 GGHVNPAVTFGAFIGGHITLIRGLLYWIAQLLGSVVAC----LLLKIATGGLETSAFSLS 136
GGH+NPAVTFG F+ ++L+R L Y IAQ LG++ +K L A +++
Sbjct: 96 GGHINPAVTFGLFLARKVSLVRALGYMIAQCLGAICGVGFVKAFMKTPYNTLGGGANTVA 155
Query: 137 SGVGATNALVFEIVMTFGLVYTVYATAVDPKNGS----LGTIAPIAIGFIVGANILAGGA 192
G AL EI+ TF LVYTV+ +A DPK + + +AP+ IGF V LA
Sbjct: 156 DGYSKGTALGAEIIGTFVLVYTVF-SATDPKRSARDSHIPVLAPLPIGFAVFMVHLATIP 214
Query: 193 FDGASMNPAVSFGPAVV---SWTWANHWVYWVGPLIGSAIAAVVYETFFITPSSYEQL 247
G +NPA SFG AV+ W + W++WVGP +G A+AA Y + + S+ + L
Sbjct: 215 ITGTGINPARSFGAAVIYNNEKAWDDQWIFWVGPFLG-ALAAAAYHQYILRASAIKAL 271
>At2g39010 putative aquaporin (water channel protein)
Length = 289
Score = 136 bits (342), Expect = 1e-32
Identities = 85/244 (34%), Positives = 130/244 (52%), Gaps = 28/244 (11%)
Query: 13 EFGKADALKAALAEFISMLIFVFAGEGSGMAYNKLTN----NGAATPAGLVAASLSHAFA 68
E K +A +AEFI+ L+F++ + + + T+ GA GL+ S +
Sbjct: 30 ELKKWSFYRAVIAEFIATLLFLYVTVLTVIGFKSQTDINAGGGACASVGLLGISWAFGGM 89
Query: 69 LFVAVSVGANISGGHVNPAVTFGAFIGGHITLIRGLLYWIAQLLGSVVACLLLKI----- 123
+F+ V A ISGGH+NPAVTFG F+ ++L+R + Y +AQ LG+ L+K+
Sbjct: 90 IFILVYCTAGISGGHINPAVTFGLFLASKVSLVRAVSYMVAQCLGATCGVGLVKVFQSTY 149
Query: 124 -----ATGGLETSAFSLSSGVGATNALVFEIVMTFGLVYTVYATAVDPKNGS----LGTI 174
+ + +++ GVGA EI+ TF LVYTV+ +A DPK + + +
Sbjct: 150 YNRYGGGANMLSDGYNVGVGVGA------EIIGTFVLVYTVF-SATDPKRNARDSHIPVL 202
Query: 175 APIAIGFIVGANILAGGAFDGASMNPAVSFGPAVV---SWTWANHWVYWVGPLIGSAIAA 231
AP+ IGF V LA G +NPA SFG AV+ W + W++WVGP +G+AIAA
Sbjct: 203 APLPIGFSVFMVHLATIPITGTGINPARSFGAAVIYNNQKAWDDQWIFWVGPFVGAAIAA 262
Query: 232 VVYE 235
++
Sbjct: 263 FYHQ 266
>At5g60660 mipC protein - like (aquaporin)
Length = 291
Score = 134 bits (338), Expect = 4e-32
Identities = 89/250 (35%), Positives = 132/250 (52%), Gaps = 17/250 (6%)
Query: 13 EFGKADALKAALAEFISMLIFVFAGEGSGMAYNKLTNN--GAATPAGLVAASLSHAFA-- 68
E K +A +AEF++ L+F++ + + Y T+ G G+ ++ AF
Sbjct: 31 ELRKWPLYRAVIAEFVATLLFLYVSILTVIGYKAQTDATAGGVDCGGVGILGIAWAFGGM 90
Query: 69 LFVAVSVGANISGGHVNPAVTFGAFIGGHITLIRGLLYWIAQLLGSVVACLLLKIATGGL 128
+FV V A ISGGH+NPAVT G F+ ++L+R +LY +AQ LG++ C +K
Sbjct: 91 IFVLVYCTAGISGGHINPAVTVGLFLARKVSLVRTVLYIVAQCLGAICGCGFVKAFQSSY 150
Query: 129 ET----SAFSLSSGVGATNALVFEIVMTFGLVYTVYATAVDPKNGS----LGTIAPIAIG 180
T A L+ G L EI+ TF LVYTV+ +A DPK + + +AP+ IG
Sbjct: 151 YTRYGGGANELADGYNKGTGLGAEIIGTFVLVYTVF-SATDPKRNARDSHVPVLAPLPIG 209
Query: 181 FIVGANILAGGAFDGASMNPAVSFGPAVV---SWTWANHWVYWVGPLIGSAIAAVVYETF 237
F V LA G +NPA SFG AV+ W + W++WVGP+IG+A AA Y F
Sbjct: 210 FAVFMVHLATIPITGTGINPARSFGAAVIYNNEKAWDDQWIFWVGPMIGAA-AAAFYHQF 268
Query: 238 FITPSSYEQL 247
+ ++ + L
Sbjct: 269 ILRAAAIKAL 278
>At2g16850 putative aquaporin (plasma membrane intrinsic protein)
Length = 278
Score = 134 bits (338), Expect = 4e-32
Identities = 88/242 (36%), Positives = 130/242 (53%), Gaps = 23/242 (9%)
Query: 21 KAALAEFISMLIFVFAGEGSGMAYNKLTNNGAATPAGLVAASLSHAFALFVAVSVGANIS 80
+A +AEFI+ L+F++ + + + T G GL+ + + +FV V A IS
Sbjct: 36 RAIIAEFIATLLFLYVTVATVIGHKNQT--GPCGGVGLLGIAWAFGGMIFVLVYCTAGIS 93
Query: 81 GGHVNPAVTFGAFIGGHITLIRGLLYWIAQLLGSVVACLLLKI--------ATGGLETSA 132
GGH+NPAVTFG F+ ++L R + Y +AQ LG++ L+K GG T A
Sbjct: 94 GGHINPAVTFGLFLARKVSLPRAVAYMVAQCLGAICGVGLVKAFMMTPYKRLGGGANTVA 153
Query: 133 FSLSSGVGATNALVFEIVMTFGLVYTVYATAVDPKNGS----LGTIAPIAIGFIVGANIL 188
S+G AL EI+ TF LVYTV+ +A DPK + + +AP+ IGF V L
Sbjct: 154 DGYSTG----TALGAEIIGTFVLVYTVF-SATDPKRSARDSHVPVLAPLPIGFAVFMVHL 208
Query: 189 AGGAFDGASMNPAVSFGPAVV---SWTWANHWVYWVGPLIGSAIAAVVYETFFITPSSYE 245
A G +NPA SFG AV+ W +HW++WVGP +G A+AA Y + + ++ +
Sbjct: 209 ATIPITGTGINPARSFGAAVIYNNEKAWDDHWIFWVGPFVG-ALAAAAYHQYILRAAAIK 267
Query: 246 QL 247
L
Sbjct: 268 AL 269
>At4g00430 transmembrane protein
Length = 287
Score = 125 bits (314), Expect = 2e-29
Identities = 82/240 (34%), Positives = 124/240 (51%), Gaps = 22/240 (9%)
Query: 11 PSEFGKADALKAALAEFISMLIFVFAGEGSGMAYNKLTNNGAATPAGLVAASLSHAFALF 70
P E +A +AEFI+ +F++ + M + N A+ G+ + + +F
Sbjct: 43 PGELSSWSFYRAGIAEFIATFLFLYITVLTVMGVKRAPNMCASV--GIQGIAWAFGGMIF 100
Query: 71 VAVSVGANISGGHVNPAVTFGAFIGGHITLIRGLLYWIAQLLGSVVACLLLK-------- 122
V A ISGGH+NPAVTFG F+ ++L R + Y I Q LG++ ++K
Sbjct: 101 ALVYCTAGISGGHINPAVTFGLFLARKLSLTRAVFYMIMQCLGAICGAGVVKGFQPTPYQ 160
Query: 123 IATGGLETSAFSLSSGVGATNALVFEIVMTFGLVYTVYATAVDPKNGS----LGTIAPIA 178
GG T A + G G L EI+ TF LVYTV+ +A D K + + +AP+
Sbjct: 161 TLGGGANTVAHGYTKGSG----LGAEIIGTFVLVYTVF-SATDAKRSARDSHVPILAPLP 215
Query: 179 IGFIVGANILAGGAFDGASMNPAVSFGPAVV---SWTWANHWVYWVGPLIGSAIAAVVYE 235
IGF V LA G +NPA S G A++ +W +HW++WVGP IG+A+AA+ ++
Sbjct: 216 IGFAVFLVHLATIPITGTGINPARSLGAAIIYNKDHSWDDHWIFWVGPFIGAALAALYHQ 275
>At3g61430 plasma membrane intrinsic protein 1a
Length = 286
Score = 124 bits (310), Expect = 6e-29
Identities = 83/239 (34%), Positives = 123/239 (50%), Gaps = 22/239 (9%)
Query: 11 PSEFGKADALKAALAEFISMLIFVFAGEGSGMAYNKLTNNGAATPAGLVAASLSHAFALF 70
P E +A +AEFI+ +F++ + M + N A+ G+ + + +F
Sbjct: 42 PGELSSWSFWRAGIAEFIATFLFLYITVLTVMGVKRSPNMCASV--GIQGIAWAFGGMIF 99
Query: 71 VAVSVGANISGGHVNPAVTFGAFIGGHITLIRGLLYWIAQLLGSVVACLLLK-------- 122
V A ISGGH+NPAVTFG F+ ++L R L Y + Q LG++ ++K
Sbjct: 100 ALVYCTAGISGGHINPAVTFGLFLARKLSLTRALYYIVMQCLGAICGAGVVKGFQPKQYQ 159
Query: 123 IATGGLETSAFSLSSGVGATNALVFEIVMTFGLVYTVYATAVDPKNGS----LGTIAPIA 178
GG T A + G G L EI+ TF LVYTV+ +A D K + + +AP+
Sbjct: 160 ALGGGANTVAHGYTKGSG----LGAEIIGTFVLVYTVF-SATDAKRNARDSHVPILAPLP 214
Query: 179 IGFIVGANILAGGAFDGASMNPAVSFGPAVV---SWTWANHWVYWVGPLIGSAIAAVVY 234
IGF V LA G +NPA S G A++ +W +HWV+WVGP IG+A+AA+ +
Sbjct: 215 IGFAVFLVHLATIPITGTGINPARSLGAAIIYNKDHSWDDHWVFWVGPFIGAALAALYH 273
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.137 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,510,431
Number of Sequences: 26719
Number of extensions: 223673
Number of successful extensions: 693
Number of sequences better than 10.0: 53
Number of HSP's better than 10.0 without gapping: 39
Number of HSP's successfully gapped in prelim test: 14
Number of HSP's that attempted gapping in prelim test: 513
Number of HSP's gapped (non-prelim): 61
length of query: 252
length of database: 11,318,596
effective HSP length: 97
effective length of query: 155
effective length of database: 8,726,853
effective search space: 1352662215
effective search space used: 1352662215
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC139841.5


